Project
averaged GNF SymAtlas + NCI-60 cancer cell lines, human (Su, 2004; Ross, 2000)
Navigation
Downloads
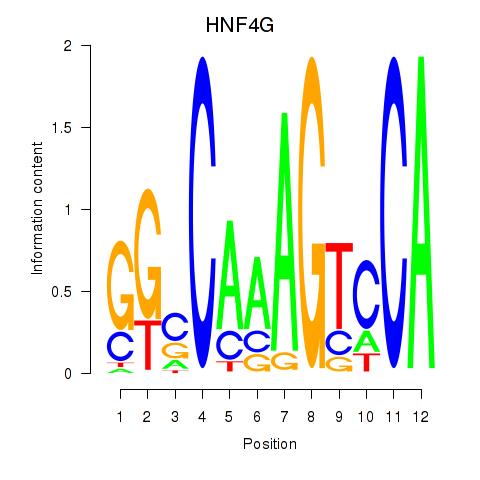
Results for HNF4G
Z-value: 1.61
Transcription factors associated with HNF4G
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HNF4G
|
ENSG00000164749.7 | hepatocyte nuclear factor 4 gamma |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HNF4G | hg19_v2_chr8_+_76452097_76452126 | -0.23 | 6.1e-04 | Click! |
Activity profile of HNF4G motif
Sorted Z-values of HNF4G motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_+_6309963 | 17.57 |
ENST00000382515.2
|
CD9
|
CD9 molecule |
| chr2_+_201170770 | 17.42 |
ENST00000409988.3
ENST00000409385.1 |
SPATS2L
|
spermatogenesis associated, serine-rich 2-like |
| chr2_+_201170596 | 17.29 |
ENST00000439084.1
ENST00000409718.1 |
SPATS2L
|
spermatogenesis associated, serine-rich 2-like |
| chr1_-_24126892 | 17.03 |
ENST00000374497.3
ENST00000425913.1 |
GALE
|
UDP-galactose-4-epimerase |
| chr8_-_145641864 | 16.45 |
ENST00000276833.5
|
SLC39A4
|
solute carrier family 39 (zinc transporter), member 4 |
| chr1_-_211752073 | 15.33 |
ENST00000367001.4
|
SLC30A1
|
solute carrier family 30 (zinc transporter), member 1 |
| chr8_-_145642267 | 14.81 |
ENST00000301305.3
|
SLC39A4
|
solute carrier family 39 (zinc transporter), member 4 |
| chr11_+_32112431 | 14.55 |
ENST00000054950.3
|
RCN1
|
reticulocalbin 1, EF-hand calcium binding domain |
| chr22_+_38071615 | 14.45 |
ENST00000215909.5
|
LGALS1
|
lectin, galactoside-binding, soluble, 1 |
| chr16_+_29690358 | 14.10 |
ENST00000395384.4
ENST00000562473.1 |
QPRT
|
quinolinate phosphoribosyltransferase |
| chr16_+_56659687 | 13.88 |
ENST00000568293.1
ENST00000330439.6 |
MT1E
|
metallothionein 1E |
| chr18_+_3412005 | 13.85 |
ENST00000401449.1
|
TGIF1
|
TGFB-induced factor homeobox 1 |
| chr5_+_137514834 | 13.63 |
ENST00000508792.1
ENST00000504621.1 |
KIF20A
|
kinesin family member 20A |
| chr17_-_38545799 | 13.57 |
ENST00000577541.1
|
TOP2A
|
topoisomerase (DNA) II alpha 170kDa |
| chr16_-_56701933 | 13.41 |
ENST00000568675.1
ENST00000569500.1 ENST00000444837.2 ENST00000379811.3 |
MT1G
|
metallothionein 1G |
| chr19_+_18496957 | 12.20 |
ENST00000252809.3
|
GDF15
|
growth differentiation factor 15 |
| chr20_+_19867150 | 12.15 |
ENST00000255006.6
|
RIN2
|
Ras and Rab interactor 2 |
| chr5_+_137514687 | 12.07 |
ENST00000394894.3
|
KIF20A
|
kinesin family member 20A |
| chr22_+_37415700 | 11.98 |
ENST00000397129.1
|
MPST
|
mercaptopyruvate sulfurtransferase |
| chr14_+_55595762 | 11.63 |
ENST00000254301.9
|
LGALS3
|
lectin, galactoside-binding, soluble, 3 |
| chr5_+_34757309 | 11.58 |
ENST00000397449.1
|
RAI14
|
retinoic acid induced 14 |
| chr22_+_37415776 | 11.37 |
ENST00000341116.3
ENST00000429360.2 ENST00000404393.1 |
MPST
|
mercaptopyruvate sulfurtransferase |
| chr3_-_123339418 | 11.27 |
ENST00000583087.1
|
MYLK
|
myosin light chain kinase |
| chr17_+_1944790 | 11.18 |
ENST00000575162.1
|
DPH1
|
diphthamide biosynthesis 1 |
| chr3_-_185542817 | 11.06 |
ENST00000382199.2
|
IGF2BP2
|
insulin-like growth factor 2 mRNA binding protein 2 |
| chr3_-_185542761 | 10.98 |
ENST00000457616.2
ENST00000346192.3 |
IGF2BP2
|
insulin-like growth factor 2 mRNA binding protein 2 |
| chr3_-_123339343 | 10.87 |
ENST00000578202.1
|
MYLK
|
myosin light chain kinase |
| chr1_-_151965048 | 10.21 |
ENST00000368809.1
|
S100A10
|
S100 calcium binding protein A10 |
| chr22_+_39378346 | 10.06 |
ENST00000407298.3
|
APOBEC3B
|
apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3B |
| chr3_+_122785895 | 9.94 |
ENST00000316218.7
|
PDIA5
|
protein disulfide isomerase family A, member 5 |
| chr1_+_94883991 | 9.93 |
ENST00000370214.4
|
ABCD3
|
ATP-binding cassette, sub-family D (ALD), member 3 |
| chr1_+_180165672 | 9.75 |
ENST00000443059.1
|
QSOX1
|
quiescin Q6 sulfhydryl oxidase 1 |
| chr2_-_238499303 | 9.65 |
ENST00000409576.1
|
RAB17
|
RAB17, member RAS oncogene family |
| chr22_-_37415475 | 9.56 |
ENST00000403892.3
ENST00000249042.3 ENST00000438203.1 |
TST
|
thiosulfate sulfurtransferase (rhodanese) |
| chr14_+_55595960 | 9.30 |
ENST00000554715.1
|
LGALS3
|
lectin, galactoside-binding, soluble, 3 |
| chr22_+_37415676 | 9.23 |
ENST00000401419.3
|
MPST
|
mercaptopyruvate sulfurtransferase |
| chr7_-_94285472 | 9.18 |
ENST00000437425.2
ENST00000447873.1 ENST00000415788.2 |
SGCE
|
sarcoglycan, epsilon |
| chr22_+_37415728 | 9.16 |
ENST00000404802.3
|
MPST
|
mercaptopyruvate sulfurtransferase |
| chr7_-_94285511 | 9.12 |
ENST00000265735.7
|
SGCE
|
sarcoglycan, epsilon |
| chr1_-_153508460 | 9.07 |
ENST00000462776.2
|
S100A6
|
S100 calcium binding protein A6 |
| chr1_+_94883931 | 8.95 |
ENST00000394233.2
ENST00000454898.2 ENST00000536817.1 |
ABCD3
|
ATP-binding cassette, sub-family D (ALD), member 3 |
| chr10_+_17272608 | 8.94 |
ENST00000421459.2
|
VIM
|
vimentin |
| chr22_-_19165917 | 8.84 |
ENST00000451283.1
|
SLC25A1
|
solute carrier family 25 (mitochondrial carrier; citrate transporter), member 1 |
| chr12_+_54422142 | 8.82 |
ENST00000243108.4
|
HOXC6
|
homeobox C6 |
| chr7_-_94285402 | 8.68 |
ENST00000428696.2
ENST00000445866.2 |
SGCE
|
sarcoglycan, epsilon |
| chr1_+_236686717 | 8.62 |
ENST00000341872.6
ENST00000450372.2 |
LGALS8
|
lectin, galactoside-binding, soluble, 8 |
| chr16_+_56642041 | 8.56 |
ENST00000245185.5
|
MT2A
|
metallothionein 2A |
| chr12_-_50419177 | 8.54 |
ENST00000454520.2
ENST00000546595.1 ENST00000548824.1 ENST00000549777.1 ENST00000546723.1 ENST00000427314.2 ENST00000552157.1 ENST00000552310.1 ENST00000548644.1 ENST00000312377.5 ENST00000546786.1 ENST00000550149.1 ENST00000546764.1 ENST00000552004.1 ENST00000548320.1 ENST00000547905.1 ENST00000550651.1 ENST00000551145.1 ENST00000434422.1 ENST00000552921.1 |
RACGAP1
|
Rac GTPase activating protein 1 |
| chr10_+_95256356 | 8.40 |
ENST00000371485.3
|
CEP55
|
centrosomal protein 55kDa |
| chr6_-_31926629 | 8.34 |
ENST00000375425.5
ENST00000426722.1 ENST00000441998.1 ENST00000444811.2 ENST00000375429.3 |
NELFE
|
negative elongation factor complex member E |
| chr2_-_150444116 | 8.16 |
ENST00000428879.1
ENST00000422782.2 |
MMADHC
|
methylmalonic aciduria (cobalamin deficiency) cblD type, with homocystinuria |
| chr11_+_12399071 | 8.12 |
ENST00000539723.1
ENST00000550549.1 |
PARVA
|
parvin, alpha |
| chrX_-_153599578 | 8.10 |
ENST00000360319.4
ENST00000344736.4 |
FLNA
|
filamin A, alpha |
| chrX_-_99891796 | 8.03 |
ENST00000373020.4
|
TSPAN6
|
tetraspanin 6 |
| chr19_-_6720686 | 8.02 |
ENST00000245907.6
|
C3
|
complement component 3 |
| chr2_-_20251744 | 7.92 |
ENST00000175091.4
|
LAPTM4A
|
lysosomal protein transmembrane 4 alpha |
| chr2_-_47168906 | 7.91 |
ENST00000444761.2
ENST00000409147.1 |
MCFD2
|
multiple coagulation factor deficiency 2 |
| chr12_-_56709674 | 7.85 |
ENST00000551286.1
ENST00000549318.1 |
CNPY2
RP11-977G19.10
|
canopy FGF signaling regulator 2 Uncharacterized protein |
| chr9_-_117853297 | 7.83 |
ENST00000542877.1
ENST00000537320.1 ENST00000341037.4 |
TNC
|
tenascin C |
| chr6_+_24667257 | 7.80 |
ENST00000537591.1
ENST00000230048.4 |
ACOT13
|
acyl-CoA thioesterase 13 |
| chr11_+_842808 | 7.77 |
ENST00000397397.2
ENST00000397411.2 ENST00000397396.1 |
TSPAN4
|
tetraspanin 4 |
| chr2_-_161350305 | 7.72 |
ENST00000348849.3
|
RBMS1
|
RNA binding motif, single stranded interacting protein 1 |
| chr16_+_55512742 | 7.66 |
ENST00000568715.1
ENST00000219070.4 |
MMP2
|
matrix metallopeptidase 2 (gelatinase A, 72kDa gelatinase, 72kDa type IV collagenase) |
| chr2_+_39005325 | 7.66 |
ENST00000281950.3
|
GEMIN6
|
gem (nuclear organelle) associated protein 6 |
| chr14_+_24458093 | 7.65 |
ENST00000558753.1
ENST00000537912.1 |
DHRS4L2
|
dehydrogenase/reductase (SDR family) member 4 like 2 |
| chr14_+_53173910 | 7.60 |
ENST00000606149.1
ENST00000555339.1 ENST00000556813.1 |
PSMC6
|
proteasome (prosome, macropain) 26S subunit, ATPase, 6 |
| chr2_-_47168850 | 7.56 |
ENST00000409207.1
|
MCFD2
|
multiple coagulation factor deficiency 2 |
| chr17_-_73149921 | 7.49 |
ENST00000481647.1
ENST00000470924.1 |
HN1
|
hematological and neurological expressed 1 |
| chr9_+_133320339 | 7.47 |
ENST00000372394.1
ENST00000372393.3 ENST00000422569.1 |
ASS1
|
argininosuccinate synthase 1 |
| chr3_-_52488048 | 7.43 |
ENST00000232975.3
|
TNNC1
|
troponin C type 1 (slow) |
| chr14_+_53173890 | 7.42 |
ENST00000445930.2
|
PSMC6
|
proteasome (prosome, macropain) 26S subunit, ATPase, 6 |
| chr17_-_17494972 | 7.23 |
ENST00000435340.2
ENST00000255389.5 ENST00000395781.2 |
PEMT
|
phosphatidylethanolamine N-methyltransferase |
| chr20_+_48807351 | 7.13 |
ENST00000303004.3
|
CEBPB
|
CCAAT/enhancer binding protein (C/EBP), beta |
| chr12_-_56709786 | 6.96 |
ENST00000547423.1
ENST00000548360.1 ENST00000551475.1 |
RP11-977G19.10
CNPY2
|
Uncharacterized protein canopy FGF signaling regulator 2 |
| chr15_+_89010923 | 6.92 |
ENST00000353598.6
|
MRPS11
|
mitochondrial ribosomal protein S11 |
| chr2_+_39005336 | 6.82 |
ENST00000409566.1
|
GEMIN6
|
gem (nuclear organelle) associated protein 6 |
| chr1_-_209824643 | 6.76 |
ENST00000391911.1
ENST00000415782.1 |
LAMB3
|
laminin, beta 3 |
| chr20_-_33872518 | 6.73 |
ENST00000374436.3
|
EIF6
|
eukaryotic translation initiation factor 6 |
| chr17_-_39677971 | 6.69 |
ENST00000393976.2
|
KRT15
|
keratin 15 |
| chr17_-_40575535 | 6.60 |
ENST00000357037.5
|
PTRF
|
polymerase I and transcript release factor |
| chr22_-_22901477 | 6.45 |
ENST00000420709.1
ENST00000398741.1 ENST00000405655.3 |
PRAME
|
preferentially expressed antigen in melanoma |
| chr1_-_153935938 | 6.42 |
ENST00000368621.1
ENST00000368623.3 |
SLC39A1
|
solute carrier family 39 (zinc transporter), member 1 |
| chr12_+_52463751 | 6.38 |
ENST00000336854.4
ENST00000550604.1 ENST00000553049.1 ENST00000548915.1 |
C12orf44
|
chromosome 12 open reading frame 44 |
| chr22_-_22901636 | 6.35 |
ENST00000406503.1
ENST00000439106.1 ENST00000402697.1 ENST00000543184.1 ENST00000398743.2 |
PRAME
|
preferentially expressed antigen in melanoma |
| chr10_+_89419370 | 6.31 |
ENST00000361175.4
ENST00000456849.1 |
PAPSS2
|
3'-phosphoadenosine 5'-phosphosulfate synthase 2 |
| chr5_-_133968529 | 6.27 |
ENST00000402673.2
|
SAR1B
|
SAR1 homolog B (S. cerevisiae) |
| chr7_-_100239132 | 6.22 |
ENST00000223051.3
ENST00000431692.1 |
TFR2
|
transferrin receptor 2 |
| chr19_-_14628645 | 6.22 |
ENST00000598235.1
|
DNAJB1
|
DnaJ (Hsp40) homolog, subfamily B, member 1 |
| chr19_-_11308190 | 6.18 |
ENST00000586659.1
ENST00000592903.1 ENST00000589359.1 ENST00000588724.1 ENST00000432929.2 |
KANK2
|
KN motif and ankyrin repeat domains 2 |
| chr17_+_72983674 | 6.07 |
ENST00000337231.5
|
CDR2L
|
cerebellar degeneration-related protein 2-like |
| chr15_-_89010607 | 6.06 |
ENST00000312475.4
|
MRPL46
|
mitochondrial ribosomal protein L46 |
| chr11_+_4116005 | 6.05 |
ENST00000300738.5
|
RRM1
|
ribonucleotide reductase M1 |
| chr2_+_217524323 | 5.83 |
ENST00000456764.1
|
IGFBP2
|
insulin-like growth factor binding protein 2, 36kDa |
| chr11_+_116700614 | 5.81 |
ENST00000375345.1
|
APOC3
|
apolipoprotein C-III |
| chr2_-_238499131 | 5.79 |
ENST00000538644.1
|
RAB17
|
RAB17, member RAS oncogene family |
| chr22_+_38093005 | 5.77 |
ENST00000406386.3
|
TRIOBP
|
TRIO and F-actin binding protein |
| chr3_-_158390282 | 5.72 |
ENST00000264265.3
|
LXN
|
latexin |
| chr11_+_66247880 | 5.65 |
ENST00000360510.2
ENST00000453114.1 ENST00000541961.1 ENST00000532019.1 ENST00000526515.1 ENST00000530165.1 ENST00000533725.1 |
DPP3
|
dipeptidyl-peptidase 3 |
| chr11_-_2158507 | 5.55 |
ENST00000381392.1
ENST00000381395.1 ENST00000418738.2 |
IGF2
|
insulin-like growth factor 2 (somatomedin A) |
| chr2_-_238499725 | 5.52 |
ENST00000264601.3
|
RAB17
|
RAB17, member RAS oncogene family |
| chrX_-_151999269 | 5.49 |
ENST00000370277.3
|
CETN2
|
centrin, EF-hand protein, 2 |
| chr2_-_238499337 | 5.44 |
ENST00000411462.1
ENST00000409822.1 |
RAB17
|
RAB17, member RAS oncogene family |
| chr14_-_24664540 | 5.33 |
ENST00000530563.1
ENST00000528895.1 ENST00000528669.1 ENST00000532632.1 |
TM9SF1
|
transmembrane 9 superfamily member 1 |
| chr2_-_150444300 | 5.32 |
ENST00000303319.5
|
MMADHC
|
methylmalonic aciduria (cobalamin deficiency) cblD type, with homocystinuria |
| chr5_+_179125907 | 5.31 |
ENST00000247461.4
ENST00000452673.2 ENST00000502498.1 ENST00000507307.1 ENST00000513246.1 ENST00000502673.1 ENST00000506654.1 ENST00000512607.2 ENST00000510810.1 |
CANX
|
calnexin |
| chr9_+_114659046 | 5.30 |
ENST00000374279.3
|
UGCG
|
UDP-glucose ceramide glucosyltransferase |
| chr18_+_7754957 | 5.29 |
ENST00000400053.4
|
PTPRM
|
protein tyrosine phosphatase, receptor type, M |
| chr11_+_116700600 | 5.20 |
ENST00000227667.3
|
APOC3
|
apolipoprotein C-III |
| chr1_-_67896009 | 5.20 |
ENST00000370990.5
|
SERBP1
|
SERPINE1 mRNA binding protein 1 |
| chr20_+_1115821 | 5.19 |
ENST00000435720.1
|
PSMF1
|
proteasome (prosome, macropain) inhibitor subunit 1 (PI31) |
| chr19_+_45449301 | 5.19 |
ENST00000591597.1
|
APOC2
|
apolipoprotein C-II |
| chr3_+_184080387 | 5.17 |
ENST00000455712.1
|
POLR2H
|
polymerase (RNA) II (DNA directed) polypeptide H |
| chr10_+_88854926 | 5.16 |
ENST00000298784.1
ENST00000298786.4 |
FAM35A
|
family with sequence similarity 35, member A |
| chr7_-_2281802 | 5.08 |
ENST00000242257.8
ENST00000440306.2 |
FTSJ2
|
FtsJ RNA methyltransferase homolog 2 (E. coli) |
| chr7_+_6048856 | 5.03 |
ENST00000223029.3
ENST00000400479.2 ENST00000395236.2 |
AIMP2
|
aminoacyl tRNA synthetase complex-interacting multifunctional protein 2 |
| chr8_+_22436635 | 5.03 |
ENST00000452226.1
ENST00000397760.4 ENST00000339162.7 ENST00000397761.2 |
PDLIM2
|
PDZ and LIM domain 2 (mystique) |
| chr3_+_52812523 | 5.00 |
ENST00000540715.1
|
ITIH1
|
inter-alpha-trypsin inhibitor heavy chain 1 |
| chr7_+_2281882 | 4.97 |
ENST00000397046.1
ENST00000397048.1 ENST00000454650.1 |
NUDT1
|
nudix (nucleoside diphosphate linked moiety X)-type motif 1 |
| chr7_+_150759634 | 4.97 |
ENST00000392826.2
ENST00000461735.1 |
SLC4A2
|
solute carrier family 4 (anion exchanger), member 2 |
| chr10_+_105726862 | 4.96 |
ENST00000335753.4
ENST00000369755.3 |
SLK
|
STE20-like kinase |
| chr10_-_96829246 | 4.87 |
ENST00000371270.3
ENST00000535898.1 ENST00000539050.1 |
CYP2C8
|
cytochrome P450, family 2, subfamily C, polypeptide 8 |
| chr1_-_19638566 | 4.86 |
ENST00000330072.5
ENST00000235835.3 |
AKR7A2
|
aldo-keto reductase family 7, member A2 (aflatoxin aldehyde reductase) |
| chr8_-_124428569 | 4.80 |
ENST00000521903.1
|
ATAD2
|
ATPase family, AAA domain containing 2 |
| chr1_+_236687232 | 4.79 |
ENST00000416919.2
ENST00000323938.6 |
LGALS8
|
lectin, galactoside-binding, soluble, 8 |
| chr12_+_56477093 | 4.77 |
ENST00000549672.1
ENST00000415288.2 |
ERBB3
|
v-erb-b2 avian erythroblastic leukemia viral oncogene homolog 3 |
| chr2_+_241807870 | 4.77 |
ENST00000307503.3
|
AGXT
|
alanine-glyoxylate aminotransferase |
| chr17_-_40288449 | 4.76 |
ENST00000552162.1
ENST00000550504.1 |
RAB5C
|
RAB5C, member RAS oncogene family |
| chr19_-_42806919 | 4.72 |
ENST00000595530.1
ENST00000538771.1 ENST00000601865.1 |
PAFAH1B3
|
platelet-activating factor acetylhydrolase 1b, catalytic subunit 3 (29kDa) |
| chr1_+_155099927 | 4.72 |
ENST00000368407.3
|
EFNA1
|
ephrin-A1 |
| chr13_+_37574678 | 4.70 |
ENST00000389704.3
|
EXOSC8
|
exosome component 8 |
| chr17_-_7082668 | 4.70 |
ENST00000573083.1
ENST00000574388.1 |
ASGR1
|
asialoglycoprotein receptor 1 |
| chr7_+_77166592 | 4.69 |
ENST00000248594.6
|
PTPN12
|
protein tyrosine phosphatase, non-receptor type 12 |
| chr12_+_3069037 | 4.67 |
ENST00000397122.2
|
TEAD4
|
TEA domain family member 4 |
| chr17_-_46178527 | 4.67 |
ENST00000393408.3
|
CBX1
|
chromobox homolog 1 |
| chr1_+_155583012 | 4.57 |
ENST00000462250.2
|
MSTO1
|
misato 1, mitochondrial distribution and morphology regulator |
| chr17_+_48624450 | 4.55 |
ENST00000006658.6
ENST00000356488.4 ENST00000393244.3 |
SPATA20
|
spermatogenesis associated 20 |
| chr7_+_2281843 | 4.47 |
ENST00000356714.1
ENST00000397049.1 |
NUDT1
|
nudix (nucleoside diphosphate linked moiety X)-type motif 1 |
| chr1_+_164528866 | 4.38 |
ENST00000420696.2
|
PBX1
|
pre-B-cell leukemia homeobox 1 |
| chr11_-_65325430 | 4.36 |
ENST00000322147.4
|
LTBP3
|
latent transforming growth factor beta binding protein 3 |
| chr6_-_144329531 | 4.36 |
ENST00000429150.1
ENST00000392309.1 ENST00000416623.1 ENST00000392307.1 |
PLAGL1
|
pleiomorphic adenoma gene-like 1 |
| chr3_+_157823609 | 4.32 |
ENST00000480820.1
|
RSRC1
|
arginine/serine-rich coiled-coil 1 |
| chr20_-_43883197 | 4.30 |
ENST00000338380.2
|
SLPI
|
secretory leukocyte peptidase inhibitor |
| chr3_+_37284668 | 4.25 |
ENST00000361924.2
ENST00000444882.1 ENST00000356847.4 ENST00000450863.2 ENST00000429018.1 |
GOLGA4
|
golgin A4 |
| chr1_+_24120143 | 4.21 |
ENST00000374501.1
|
LYPLA2
|
lysophospholipase II |
| chr1_+_220267429 | 4.21 |
ENST00000366922.1
ENST00000302637.5 |
IARS2
|
isoleucyl-tRNA synthetase 2, mitochondrial |
| chr1_-_155270770 | 4.19 |
ENST00000392414.3
|
PKLR
|
pyruvate kinase, liver and RBC |
| chr18_+_19749386 | 4.18 |
ENST00000269216.3
|
GATA6
|
GATA binding protein 6 |
| chr1_+_145727681 | 4.15 |
ENST00000417171.1
ENST00000451928.2 |
PDZK1
|
PDZ domain containing 1 |
| chr20_-_35274548 | 4.13 |
ENST00000262866.4
|
SLA2
|
Src-like-adaptor 2 |
| chr1_+_236687173 | 4.11 |
ENST00000238181.7
|
LGALS8
|
lectin, galactoside-binding, soluble, 8 |
| chr22_-_43045574 | 4.09 |
ENST00000352397.5
|
CYB5R3
|
cytochrome b5 reductase 3 |
| chr16_+_88872176 | 4.08 |
ENST00000569140.1
|
CDT1
|
chromatin licensing and DNA replication factor 1 |
| chr17_-_46178741 | 4.07 |
ENST00000581003.1
ENST00000225603.4 |
CBX1
|
chromobox homolog 1 |
| chr12_+_20963647 | 4.07 |
ENST00000381545.3
|
SLCO1B3
|
solute carrier organic anion transporter family, member 1B3 |
| chrX_+_138612889 | 4.00 |
ENST00000218099.2
ENST00000394090.2 |
F9
|
coagulation factor IX |
| chr19_+_45449266 | 3.99 |
ENST00000592257.1
|
APOC2
|
apolipoprotein C-II |
| chr9_-_179018 | 3.97 |
ENST00000431099.2
ENST00000382447.4 ENST00000382389.1 ENST00000377447.3 ENST00000314367.10 ENST00000356521.4 ENST00000382393.1 ENST00000377400.4 |
CBWD1
|
COBW domain containing 1 |
| chr11_+_18417813 | 3.97 |
ENST00000540430.1
ENST00000379412.5 |
LDHA
|
lactate dehydrogenase A |
| chrX_+_70503037 | 3.97 |
ENST00000535149.1
|
NONO
|
non-POU domain containing, octamer-binding |
| chr1_-_24194771 | 3.96 |
ENST00000374479.3
|
FUCA1
|
fucosidase, alpha-L- 1, tissue |
| chr12_+_20963632 | 3.93 |
ENST00000540853.1
ENST00000261196.2 |
SLCO1B3
|
solute carrier organic anion transporter family, member 1B3 |
| chr3_-_46000064 | 3.91 |
ENST00000433878.1
|
FYCO1
|
FYVE and coiled-coil domain containing 1 |
| chr3_-_52860850 | 3.88 |
ENST00000441637.2
|
ITIH4
|
inter-alpha-trypsin inhibitor heavy chain family, member 4 |
| chr4_-_40517984 | 3.82 |
ENST00000381795.6
|
RBM47
|
RNA binding motif protein 47 |
| chr8_-_11710979 | 3.77 |
ENST00000415599.2
|
CTSB
|
cathepsin B |
| chr16_-_57831676 | 3.76 |
ENST00000465878.2
ENST00000539578.1 ENST00000561524.1 |
KIFC3
|
kinesin family member C3 |
| chr13_+_43597269 | 3.69 |
ENST00000379221.2
|
DNAJC15
|
DnaJ (Hsp40) homolog, subfamily C, member 15 |
| chr3_-_141747439 | 3.69 |
ENST00000467667.1
|
TFDP2
|
transcription factor Dp-2 (E2F dimerization partner 2) |
| chr16_-_58163299 | 3.68 |
ENST00000262498.3
|
C16orf80
|
chromosome 16 open reading frame 80 |
| chr1_-_153518270 | 3.68 |
ENST00000354332.4
ENST00000368716.4 |
S100A4
|
S100 calcium binding protein A4 |
| chr9_+_37486005 | 3.66 |
ENST00000377792.3
|
POLR1E
|
polymerase (RNA) I polypeptide E, 53kDa |
| chr11_-_118550346 | 3.65 |
ENST00000530256.1
|
TREH
|
trehalase (brush-border membrane glycoprotein) |
| chr9_-_14314518 | 3.64 |
ENST00000397581.2
|
NFIB
|
nuclear factor I/B |
| chr19_-_14629224 | 3.61 |
ENST00000254322.2
|
DNAJB1
|
DnaJ (Hsp40) homolog, subfamily B, member 1 |
| chr14_+_24458021 | 3.60 |
ENST00000397071.1
ENST00000559411.1 ENST00000335125.6 |
DHRS4L2
|
dehydrogenase/reductase (SDR family) member 4 like 2 |
| chr17_+_80186908 | 3.58 |
ENST00000582743.1
ENST00000578684.1 ENST00000577650.1 ENST00000582715.1 |
SLC16A3
|
solute carrier family 16 (monocarboxylate transporter), member 3 |
| chr22_+_30163340 | 3.57 |
ENST00000330029.6
ENST00000401406.3 |
UQCR10
|
ubiquinol-cytochrome c reductase, complex III subunit X |
| chr3_+_184081175 | 3.56 |
ENST00000452961.1
ENST00000296223.3 |
POLR2H
|
polymerase (RNA) II (DNA directed) polypeptide H |
| chr2_-_27545431 | 3.56 |
ENST00000233545.2
|
MPV17
|
MpV17 mitochondrial inner membrane protein |
| chr9_-_14314566 | 3.55 |
ENST00000397579.2
|
NFIB
|
nuclear factor I/B |
| chr5_-_89705537 | 3.52 |
ENST00000522864.1
ENST00000522083.1 ENST00000522565.1 ENST00000522842.1 ENST00000283122.3 |
CETN3
|
centrin, EF-hand protein, 3 |
| chrX_-_48827976 | 3.49 |
ENST00000218176.3
|
KCND1
|
potassium voltage-gated channel, Shal-related subfamily, member 1 |
| chr1_-_154909329 | 3.49 |
ENST00000368467.3
|
PMVK
|
phosphomevalonate kinase |
| chr2_+_191208196 | 3.48 |
ENST00000392329.2
ENST00000322522.4 ENST00000430311.1 ENST00000541441.1 |
INPP1
|
inositol polyphosphate-1-phosphatase |
| chr12_+_118454500 | 3.48 |
ENST00000537315.1
ENST00000229043.3 ENST00000484086.2 ENST00000420967.1 ENST00000454402.2 ENST00000392542.2 ENST00000535092.1 |
RFC5
|
replication factor C (activator 1) 5, 36.5kDa |
| chr6_+_146864829 | 3.47 |
ENST00000367495.3
|
RAB32
|
RAB32, member RAS oncogene family |
| chr14_-_53331239 | 3.47 |
ENST00000553663.1
|
FERMT2
|
fermitin family member 2 |
| chr19_+_45449228 | 3.44 |
ENST00000252490.4
|
APOC2
|
apolipoprotein C-II |
| chr8_-_27469196 | 3.43 |
ENST00000546343.1
ENST00000560566.1 |
CLU
|
clusterin |
| chr3_+_184081213 | 3.43 |
ENST00000429568.1
|
POLR2H
|
polymerase (RNA) II (DNA directed) polypeptide H |
| chr1_+_207262578 | 3.39 |
ENST00000243611.5
ENST00000367076.3 |
C4BPB
|
complement component 4 binding protein, beta |
| chr1_-_22263790 | 3.39 |
ENST00000374695.3
|
HSPG2
|
heparan sulfate proteoglycan 2 |
| chr21_-_38445443 | 3.36 |
ENST00000360525.4
|
PIGP
|
phosphatidylinositol glycan anchor biosynthesis, class P |
| chr8_-_105601134 | 3.35 |
ENST00000276654.5
ENST00000424843.2 |
LRP12
|
low density lipoprotein receptor-related protein 12 |
| chr11_+_67071050 | 3.30 |
ENST00000376757.5
|
SSH3
|
slingshot protein phosphatase 3 |
| chr11_-_118550375 | 3.29 |
ENST00000525958.1
ENST00000264029.4 ENST00000397925.1 ENST00000529101.1 |
TREH
|
trehalase (brush-border membrane glycoprotein) |
| chr11_+_64009072 | 3.27 |
ENST00000535135.1
ENST00000394540.3 |
FKBP2
|
FK506 binding protein 2, 13kDa |
| chr14_+_77790901 | 3.25 |
ENST00000553586.1
ENST00000555583.1 |
GSTZ1
|
glutathione S-transferase zeta 1 |
| chr1_+_201924619 | 3.24 |
ENST00000367287.4
|
TIMM17A
|
translocase of inner mitochondrial membrane 17 homolog A (yeast) |
| chr9_+_70856397 | 3.22 |
ENST00000360171.6
|
CBWD3
|
COBW domain containing 3 |
Network of associatons between targets according to the STRING database.
First level regulatory network of HNF4G
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.3 | 51.3 | GO:0009439 | cyanate metabolic process(GO:0009439) cyanate catabolic process(GO:0009440) |
| 7.0 | 20.9 | GO:2000521 | negative regulation of immunological synapse formation(GO:2000521) |
| 5.3 | 26.4 | GO:0002415 | immunoglobulin transcytosis in epithelial cells mediated by polymeric immunoglobulin receptor(GO:0002415) |
| 4.7 | 46.6 | GO:0071578 | zinc II ion transmembrane import(GO:0071578) |
| 4.2 | 12.6 | GO:0010902 | positive regulation of very-low-density lipoprotein particle remodeling(GO:0010902) |
| 4.2 | 16.7 | GO:0015910 | peroxisomal long-chain fatty acid import(GO:0015910) |
| 3.7 | 18.4 | GO:1904977 | lymphatic endothelial cell migration(GO:1904977) |
| 3.7 | 11.0 | GO:2000909 | regulation of cholesterol import(GO:0060620) negative regulation of cholesterol import(GO:0060621) regulation of sterol import(GO:2000909) negative regulation of sterol import(GO:2000910) |
| 3.6 | 14.4 | GO:0002317 | plasma cell differentiation(GO:0002317) |
| 2.8 | 14.1 | GO:0072526 | pyridine-containing compound catabolic process(GO:0072526) |
| 2.8 | 22.1 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 2.7 | 8.0 | GO:0002894 | positive regulation of type IIa hypersensitivity(GO:0001798) positive regulation of type II hypersensitivity(GO:0002894) |
| 2.6 | 7.8 | GO:0060447 | bud outgrowth involved in lung branching(GO:0060447) |
| 2.4 | 9.4 | GO:0006203 | dGTP catabolic process(GO:0006203) |
| 2.2 | 8.8 | GO:0006842 | tricarboxylic acid transport(GO:0006842) citrate transport(GO:0015746) |
| 2.1 | 8.5 | GO:0044837 | assembly of actomyosin apparatus involved in cytokinesis(GO:0000912) actomyosin contractile ring assembly(GO:0000915) actomyosin contractile ring organization(GO:0044837) |
| 2.1 | 36.2 | GO:0000920 | cell separation after cytokinesis(GO:0000920) |
| 2.1 | 6.2 | GO:0070563 | negative regulation of vitamin D receptor signaling pathway(GO:0070563) |
| 2.0 | 12.1 | GO:0042866 | pyruvate biosynthetic process(GO:0042866) |
| 1.9 | 1.9 | GO:0009822 | alkaloid catabolic process(GO:0009822) |
| 1.9 | 7.4 | GO:0032972 | diaphragm contraction(GO:0002086) regulation of muscle filament sliding speed(GO:0032972) |
| 1.8 | 16.4 | GO:0060510 | Type II pneumocyte differentiation(GO:0060510) |
| 1.8 | 5.3 | GO:0006679 | glucosylceramide biosynthetic process(GO:0006679) |
| 1.7 | 8.6 | GO:0010273 | detoxification of copper ion(GO:0010273) stress response to copper ion(GO:1990169) |
| 1.7 | 6.8 | GO:1903377 | negative regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway(GO:1903377) |
| 1.7 | 6.7 | GO:1902626 | assembly of large subunit precursor of preribosome(GO:1902626) |
| 1.6 | 14.8 | GO:0045716 | positive regulation of low-density lipoprotein particle receptor biosynthetic process(GO:0045716) |
| 1.6 | 6.4 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 1.6 | 17.6 | GO:0009414 | response to water deprivation(GO:0009414) |
| 1.6 | 12.8 | GO:0017182 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 1.6 | 6.3 | GO:0000103 | sulfate assimilation(GO:0000103) |
| 1.6 | 4.7 | GO:0090675 | intermicrovillar adhesion(GO:0090675) |
| 1.5 | 7.7 | GO:0001957 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 1.5 | 3.0 | GO:0036500 | ATF6-mediated unfolded protein response(GO:0036500) |
| 1.5 | 6.0 | GO:0001189 | RNA polymerase I transcriptional preinitiation complex assembly(GO:0001188) RNA polymerase I transcriptional preinitiation complex assembly at the promoter for the nuclear large rRNA transcript(GO:0001189) |
| 1.4 | 4.2 | GO:0006428 | isoleucyl-tRNA aminoacylation(GO:0006428) |
| 1.4 | 13.9 | GO:0042117 | monocyte activation(GO:0042117) |
| 1.4 | 4.1 | GO:1902595 | regulation of DNA replication origin binding(GO:1902595) |
| 1.4 | 13.6 | GO:0030263 | apoptotic chromosome condensation(GO:0030263) |
| 1.3 | 8.1 | GO:0016479 | negative regulation of transcription from RNA polymerase I promoter(GO:0016479) |
| 1.3 | 1.3 | GO:0009441 | glycolate metabolic process(GO:0009441) |
| 1.3 | 17.0 | GO:0019388 | galactose catabolic process(GO:0019388) |
| 1.2 | 3.6 | GO:1901253 | negative regulation of intracellular transport of viral material(GO:1901253) |
| 1.2 | 4.7 | GO:0014028 | notochord formation(GO:0014028) |
| 1.2 | 4.7 | GO:0034473 | U1 snRNA 3'-end processing(GO:0034473) U5 snRNA 3'-end processing(GO:0034476) |
| 1.2 | 3.5 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 1.2 | 5.8 | GO:0030047 | actin modification(GO:0030047) |
| 1.1 | 4.4 | GO:1902460 | regulation of mesenchymal stem cell proliferation(GO:1902460) positive regulation of mesenchymal stem cell proliferation(GO:1902462) |
| 1.1 | 3.2 | GO:0046098 | guanine metabolic process(GO:0046098) |
| 1.1 | 6.4 | GO:1902847 | macrophage proliferation(GO:0061517) microglial cell proliferation(GO:0061518) regulation of neuronal signal transduction(GO:1902847) positive regulation of neurofibrillary tangle assembly(GO:1902998) |
| 1.0 | 6.2 | GO:0098707 | ferrous iron import into cell(GO:0097460) ferrous iron import across plasma membrane(GO:0098707) |
| 1.0 | 3.0 | GO:0042822 | pyridoxal phosphate metabolic process(GO:0042822) pyridoxal phosphate biosynthetic process(GO:0042823) |
| 1.0 | 7.8 | GO:0007598 | blood coagulation, extrinsic pathway(GO:0007598) |
| 0.9 | 7.5 | GO:0006526 | arginine biosynthetic process(GO:0006526) |
| 0.9 | 7.2 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.9 | 3.6 | GO:0035879 | plasma membrane lactate transport(GO:0035879) |
| 0.9 | 8.9 | GO:0060020 | Bergmann glial cell differentiation(GO:0060020) |
| 0.9 | 5.3 | GO:2000825 | positive regulation of androgen receptor activity(GO:2000825) |
| 0.9 | 2.6 | GO:1901526 | positive regulation of macromitophagy(GO:1901526) positive regulation of mitophagy in response to mitochondrial depolarization(GO:1904925) |
| 0.8 | 4.2 | GO:0002084 | protein depalmitoylation(GO:0002084) |
| 0.8 | 0.8 | GO:1902908 | regulation of melanosome transport(GO:1902908) |
| 0.8 | 8.3 | GO:0034244 | negative regulation of transcription elongation from RNA polymerase II promoter(GO:0034244) |
| 0.8 | 11.3 | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 0.7 | 6.6 | GO:0030450 | regulation of complement activation, classical pathway(GO:0030450) negative regulation of complement activation, classical pathway(GO:0045959) regulation of opsonization(GO:1903027) |
| 0.7 | 13.9 | GO:0071294 | cellular response to zinc ion(GO:0071294) |
| 0.7 | 10.1 | GO:0010528 | regulation of transposition(GO:0010528) negative regulation of transposition(GO:0010529) |
| 0.7 | 2.8 | GO:0006447 | regulation of translational initiation by iron(GO:0006447) |
| 0.7 | 9.8 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
| 0.7 | 3.5 | GO:1900264 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 0.7 | 4.9 | GO:0002933 | lipid hydroxylation(GO:0002933) |
| 0.7 | 2.7 | GO:0019322 | xylulose metabolic process(GO:0005997) pentose biosynthetic process(GO:0019322) |
| 0.7 | 3.9 | GO:1901096 | regulation of autophagosome maturation(GO:1901096) positive regulation of autophagosome maturation(GO:1901098) |
| 0.6 | 10.2 | GO:0001765 | membrane raft assembly(GO:0001765) |
| 0.6 | 3.8 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.6 | 5.5 | GO:0038028 | insulin receptor signaling pathway via phosphatidylinositol 3-kinase(GO:0038028) |
| 0.6 | 3.7 | GO:1902956 | regulation of mitochondrial electron transport, NADH to ubiquinone(GO:1902956) negative regulation of mitochondrial electron transport, NADH to ubiquinone(GO:1902957) |
| 0.6 | 4.3 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.6 | 2.4 | GO:0006458 | 'de novo' protein folding(GO:0006458) |
| 0.6 | 8.5 | GO:0034975 | protein folding in endoplasmic reticulum(GO:0034975) |
| 0.6 | 15.0 | GO:0045899 | positive regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045899) |
| 0.6 | 1.8 | GO:0014707 | branchiomeric skeletal muscle development(GO:0014707) |
| 0.6 | 7.1 | GO:1990440 | positive regulation of transcription from RNA polymerase II promoter in response to endoplasmic reticulum stress(GO:1990440) |
| 0.6 | 1.8 | GO:0060769 | lateral sprouting involved in mammary gland duct morphogenesis(GO:0060599) positive regulation of epithelial cell proliferation involved in prostate gland development(GO:0060769) |
| 0.6 | 1.7 | GO:0051710 | regulation of cytolysis in other organism(GO:0051710) |
| 0.6 | 2.8 | GO:0001306 | age-dependent response to oxidative stress(GO:0001306) age-dependent response to reactive oxygen species(GO:0001315) regulation of systemic arterial blood pressure by acetylcholine(GO:0003068) vasodilation by acetylcholine involved in regulation of systemic arterial blood pressure(GO:0003069) regulation of systemic arterial blood pressure by neurotransmitter(GO:0003070) age-dependent general metabolic decline(GO:0007571) |
| 0.6 | 2.2 | GO:0048205 | COPI-coated vesicle budding(GO:0035964) Golgi transport vesicle coating(GO:0048200) COPI coating of Golgi vesicle(GO:0048205) |
| 0.5 | 6.5 | GO:1901223 | negative regulation of NIK/NF-kappaB signaling(GO:1901223) |
| 0.5 | 6.5 | GO:0005984 | disaccharide metabolic process(GO:0005984) |
| 0.5 | 4.8 | GO:0031936 | negative regulation of chromatin silencing(GO:0031936) |
| 0.5 | 12.2 | GO:1901741 | positive regulation of myoblast fusion(GO:1901741) |
| 0.5 | 2.1 | GO:0046271 | phenylpropanoid catabolic process(GO:0046271) |
| 0.5 | 1.5 | GO:2000830 | vacuolar phosphate transport(GO:0007037) positive regulation of mitotic cell cycle DNA replication(GO:1903465) positive regulation of parathyroid hormone secretion(GO:2000830) |
| 0.5 | 3.5 | GO:1903232 | melanosome assembly(GO:1903232) |
| 0.5 | 4.0 | GO:0016139 | glycoside catabolic process(GO:0016139) |
| 0.5 | 3.0 | GO:0030948 | negative regulation of vascular endothelial growth factor receptor signaling pathway(GO:0030948) |
| 0.5 | 4.9 | GO:0044598 | polyketide metabolic process(GO:0030638) daunorubicin metabolic process(GO:0044597) doxorubicin metabolic process(GO:0044598) |
| 0.5 | 2.4 | GO:1902896 | terminal web assembly(GO:1902896) |
| 0.5 | 2.8 | GO:0051012 | microtubule sliding(GO:0051012) |
| 0.5 | 5.6 | GO:0071577 | zinc II ion transmembrane transport(GO:0071577) |
| 0.5 | 9.8 | GO:0016242 | negative regulation of macroautophagy(GO:0016242) |
| 0.5 | 3.3 | GO:0006572 | tyrosine catabolic process(GO:0006572) |
| 0.5 | 5.5 | GO:0006089 | lactate metabolic process(GO:0006089) |
| 0.4 | 3.6 | GO:0017062 | respiratory chain complex III assembly(GO:0017062) mitochondrial respiratory chain complex III assembly(GO:0034551) mitochondrial respiratory chain complex III biogenesis(GO:0097033) |
| 0.4 | 6.7 | GO:0032836 | glomerular basement membrane development(GO:0032836) |
| 0.4 | 2.1 | GO:0032910 | transforming growth factor beta3 production(GO:0032907) regulation of transforming growth factor beta3 production(GO:0032910) positive regulation of transforming growth factor beta3 production(GO:0032916) negative regulation of lung blood pressure(GO:0061767) |
| 0.4 | 6.1 | GO:0071670 | smooth muscle cell chemotaxis(GO:0071670) |
| 0.4 | 3.9 | GO:0038203 | TORC2 signaling(GO:0038203) |
| 0.4 | 4.7 | GO:2000587 | negative regulation of platelet-derived growth factor receptor-beta signaling pathway(GO:2000587) |
| 0.4 | 22.7 | GO:0018279 | protein N-linked glycosylation via asparagine(GO:0018279) |
| 0.4 | 4.8 | GO:0048227 | plasma membrane to endosome transport(GO:0048227) |
| 0.4 | 15.3 | GO:0006363 | transcription elongation from RNA polymerase I promoter(GO:0006362) termination of RNA polymerase I transcription(GO:0006363) |
| 0.3 | 2.0 | GO:0007386 | compartment pattern specification(GO:0007386) |
| 0.3 | 1.6 | GO:0046874 | quinolinate metabolic process(GO:0046874) |
| 0.3 | 14.5 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.3 | 0.9 | GO:0019477 | L-lysine catabolic process to acetyl-CoA(GO:0019474) L-lysine catabolic process(GO:0019477) L-lysine metabolic process(GO:0046440) |
| 0.3 | 3.5 | GO:0046834 | lipid phosphorylation(GO:0046834) phosphatidylinositol phosphorylation(GO:0046854) |
| 0.3 | 0.9 | GO:1904430 | negative regulation of t-circle formation(GO:1904430) |
| 0.3 | 10.0 | GO:0043567 | regulation of insulin-like growth factor receptor signaling pathway(GO:0043567) |
| 0.3 | 6.9 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.3 | 5.8 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.3 | 6.0 | GO:0006206 | pyrimidine nucleobase metabolic process(GO:0006206) |
| 0.3 | 4.6 | GO:0048311 | mitochondrion distribution(GO:0048311) |
| 0.3 | 3.7 | GO:0048762 | mesenchymal cell differentiation(GO:0048762) |
| 0.2 | 1.0 | GO:1905225 | response to thyrotropin-releasing hormone(GO:1905225) |
| 0.2 | 6.2 | GO:0050965 | detection of temperature stimulus involved in sensory perception(GO:0050961) detection of temperature stimulus involved in sensory perception of pain(GO:0050965) |
| 0.2 | 3.2 | GO:1902166 | negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator(GO:1902166) |
| 0.2 | 4.2 | GO:0047497 | establishment of mitochondrion localization, microtubule-mediated(GO:0034643) mitochondrion transport along microtubule(GO:0047497) establishment of mitochondrion localization(GO:0051654) |
| 0.2 | 2.4 | GO:1902455 | negative regulation of stem cell population maintenance(GO:1902455) |
| 0.2 | 4.8 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.2 | 7.7 | GO:0032008 | positive regulation of TOR signaling(GO:0032008) |
| 0.2 | 5.5 | GO:0000717 | nucleotide-excision repair, DNA duplex unwinding(GO:0000717) |
| 0.2 | 2.9 | GO:0034138 | toll-like receptor 3 signaling pathway(GO:0034138) |
| 0.2 | 4.7 | GO:0016254 | preassembly of GPI anchor in ER membrane(GO:0016254) |
| 0.2 | 6.8 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.2 | 5.8 | GO:0010591 | regulation of lamellipodium assembly(GO:0010591) |
| 0.2 | 1.6 | GO:1900045 | negative regulation of histone ubiquitination(GO:0033183) histone H2A K63-linked ubiquitination(GO:0070535) negative regulation of protein K63-linked ubiquitination(GO:1900045) regulation of histone H2A K63-linked ubiquitination(GO:1901314) negative regulation of histone H2A K63-linked ubiquitination(GO:1901315) negative regulation of protein polyubiquitination(GO:1902915) |
| 0.2 | 0.5 | GO:0071109 | superior temporal gyrus development(GO:0071109) |
| 0.2 | 1.4 | GO:0007256 | activation of JNKK activity(GO:0007256) |
| 0.2 | 3.4 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.2 | 2.0 | GO:0016191 | synaptic vesicle uncoating(GO:0016191) |
| 0.2 | 1.1 | GO:0030200 | heparan sulfate proteoglycan catabolic process(GO:0030200) |
| 0.2 | 1.1 | GO:0098706 | ferric iron import(GO:0033216) ferric iron import into cell(GO:0097461) ferric iron import across plasma membrane(GO:0098706) |
| 0.2 | 1.8 | GO:0045542 | positive regulation of cholesterol biosynthetic process(GO:0045542) |
| 0.2 | 1.1 | GO:1904338 | regulation of dopaminergic neuron differentiation(GO:1904338) |
| 0.2 | 0.6 | GO:2001046 | positive regulation of integrin-mediated signaling pathway(GO:2001046) |
| 0.2 | 3.4 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.2 | 4.1 | GO:0050849 | negative regulation of calcium-mediated signaling(GO:0050849) |
| 0.2 | 10.7 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.2 | 1.9 | GO:0061314 | Notch signaling involved in heart development(GO:0061314) |
| 0.2 | 0.2 | GO:0036115 | fatty-acyl-CoA catabolic process(GO:0036115) malonyl-CoA metabolic process(GO:2001293) |
| 0.2 | 5.3 | GO:0000042 | protein targeting to Golgi(GO:0000042) |
| 0.2 | 4.7 | GO:0009235 | cobalamin metabolic process(GO:0009235) |
| 0.2 | 4.4 | GO:0010971 | positive regulation of G2/M transition of mitotic cell cycle(GO:0010971) |
| 0.2 | 2.2 | GO:0030202 | heparin metabolic process(GO:0030202) heparin biosynthetic process(GO:0030210) |
| 0.2 | 2.5 | GO:0019852 | L-ascorbic acid metabolic process(GO:0019852) |
| 0.2 | 4.1 | GO:0015879 | carnitine transport(GO:0015879) |
| 0.2 | 4.7 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.2 | 3.8 | GO:0046697 | decidualization(GO:0046697) |
| 0.2 | 7.2 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.2 | 1.0 | GO:1903433 | regulation of constitutive secretory pathway(GO:1903433) positive regulation of constitutive secretory pathway(GO:1903435) |
| 0.2 | 1.6 | GO:0046784 | viral mRNA export from host cell nucleus(GO:0046784) |
| 0.2 | 0.6 | GO:0019323 | pentose catabolic process(GO:0019323) |
| 0.1 | 2.7 | GO:0046341 | CDP-diacylglycerol metabolic process(GO:0046341) |
| 0.1 | 1.0 | GO:0009732 | detection of carbohydrate stimulus(GO:0009730) detection of hexose stimulus(GO:0009732) detection of monosaccharide stimulus(GO:0034287) detection of glucose(GO:0051594) |
| 0.1 | 0.9 | GO:0051547 | regulation of keratinocyte migration(GO:0051547) |
| 0.1 | 1.3 | GO:0010891 | negative regulation of sequestering of triglyceride(GO:0010891) |
| 0.1 | 5.6 | GO:0015721 | bile acid and bile salt transport(GO:0015721) |
| 0.1 | 0.6 | GO:0007068 | negative regulation of transcription during mitosis(GO:0007068) negative regulation of transcription from RNA polymerase II promoter during mitosis(GO:0007070) |
| 0.1 | 0.4 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.1 | 0.8 | GO:0007044 | cell-substrate junction assembly(GO:0007044) |
| 0.1 | 0.5 | GO:0046340 | diacylglycerol catabolic process(GO:0046340) |
| 0.1 | 2.9 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.1 | 0.7 | GO:0001992 | regulation of systemic arterial blood pressure by vasopressin(GO:0001992) negative regulation of urine volume(GO:0035811) |
| 0.1 | 5.0 | GO:0031122 | cytoplasmic microtubule organization(GO:0031122) |
| 0.1 | 0.7 | GO:0061484 | hematopoietic stem cell homeostasis(GO:0061484) |
| 0.1 | 0.7 | GO:0006065 | UDP-glucuronate biosynthetic process(GO:0006065) UDP-glucuronate metabolic process(GO:0046398) |
| 0.1 | 2.6 | GO:1904707 | positive regulation of vascular smooth muscle cell proliferation(GO:1904707) |
| 0.1 | 1.8 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.1 | 24.2 | GO:0043488 | regulation of mRNA stability(GO:0043488) |
| 0.1 | 4.5 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.1 | 2.2 | GO:0051292 | nuclear pore complex assembly(GO:0051292) |
| 0.1 | 8.6 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.1 | 2.6 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.1 | 0.8 | GO:0007506 | gonadal mesoderm development(GO:0007506) |
| 0.1 | 6.1 | GO:0002474 | antigen processing and presentation of peptide antigen via MHC class I(GO:0002474) |
| 0.1 | 4.7 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.1 | 1.6 | GO:0032802 | low-density lipoprotein particle receptor catabolic process(GO:0032802) |
| 0.1 | 0.3 | GO:0048250 | mitochondrial iron ion transport(GO:0048250) |
| 0.1 | 28.1 | GO:0007160 | cell-matrix adhesion(GO:0007160) |
| 0.1 | 0.1 | GO:0032571 | response to vitamin K(GO:0032571) |
| 0.1 | 1.1 | GO:0070070 | proton-transporting V-type ATPase complex assembly(GO:0070070) vacuolar proton-transporting V-type ATPase complex assembly(GO:0070072) |
| 0.1 | 1.6 | GO:0006465 | signal peptide processing(GO:0006465) |
| 0.1 | 1.4 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.1 | 6.4 | GO:0070125 | mitochondrial translational elongation(GO:0070125) mitochondrial translational termination(GO:0070126) |
| 0.1 | 0.4 | GO:0045144 | meiosis II(GO:0007135) meiotic sister chromatid segregation(GO:0045144) |
| 0.1 | 0.8 | GO:0006228 | UTP biosynthetic process(GO:0006228) UTP metabolic process(GO:0046051) |
| 0.1 | 1.6 | GO:0042481 | regulation of odontogenesis(GO:0042481) |
| 0.1 | 1.2 | GO:1900017 | positive regulation of cytokine production involved in inflammatory response(GO:1900017) |
| 0.1 | 0.3 | GO:0043335 | protein unfolding(GO:0043335) |
| 0.1 | 2.3 | GO:0031440 | regulation of mRNA 3'-end processing(GO:0031440) |
| 0.1 | 1.9 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.1 | 2.0 | GO:0051642 | centrosome localization(GO:0051642) |
| 0.1 | 3.1 | GO:0048146 | positive regulation of fibroblast proliferation(GO:0048146) |
| 0.1 | 1.0 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.1 | 0.4 | GO:0071763 | nuclear membrane organization(GO:0071763) |
| 0.1 | 1.4 | GO:0007398 | ectoderm development(GO:0007398) |
| 0.1 | 1.1 | GO:1900078 | positive regulation of cellular response to insulin stimulus(GO:1900078) |
| 0.1 | 3.4 | GO:0006027 | glycosaminoglycan catabolic process(GO:0006027) |
| 0.1 | 4.3 | GO:0000380 | alternative mRNA splicing, via spliceosome(GO:0000380) |
| 0.1 | 0.2 | GO:0060370 | susceptibility to T cell mediated cytotoxicity(GO:0060370) |
| 0.1 | 0.6 | GO:0051601 | exocyst localization(GO:0051601) |
| 0.1 | 0.2 | GO:0031443 | fast-twitch skeletal muscle fiber contraction(GO:0031443) |
| 0.1 | 4.3 | GO:0032091 | negative regulation of protein binding(GO:0032091) |
| 0.1 | 7.1 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.1 | 0.4 | GO:2000143 | negative regulation of transcription initiation from RNA polymerase II promoter(GO:0060633) negative regulation of DNA-templated transcription, initiation(GO:2000143) |
| 0.0 | 1.4 | GO:0044247 | glycogen catabolic process(GO:0005980) glucan catabolic process(GO:0009251) cellular polysaccharide catabolic process(GO:0044247) |
| 0.0 | 0.1 | GO:0009698 | phenylpropanoid metabolic process(GO:0009698) |
| 0.0 | 0.3 | GO:1990822 | L-cystine transport(GO:0015811) basic amino acid transmembrane transport(GO:1990822) |
| 0.0 | 0.4 | GO:0071372 | cellular response to follicle-stimulating hormone stimulus(GO:0071372) |
| 0.0 | 2.8 | GO:0030574 | collagen catabolic process(GO:0030574) |
| 0.0 | 0.2 | GO:1903361 | protein localization to basolateral plasma membrane(GO:1903361) |
| 0.0 | 15.6 | GO:0043687 | post-translational protein modification(GO:0043687) |
| 0.0 | 1.9 | GO:0034260 | negative regulation of GTPase activity(GO:0034260) |
| 0.0 | 2.2 | GO:0000245 | spliceosomal complex assembly(GO:0000245) |
| 0.0 | 0.4 | GO:0060836 | lymphatic endothelial cell differentiation(GO:0060836) |
| 0.0 | 0.5 | GO:0043085 | positive regulation of catalytic activity(GO:0043085) |
| 0.0 | 1.1 | GO:0015949 | nucleobase-containing small molecule interconversion(GO:0015949) |
| 0.0 | 4.3 | GO:0048706 | embryonic skeletal system development(GO:0048706) |
| 0.0 | 0.4 | GO:0033299 | secretion of lysosomal enzymes(GO:0033299) |
| 0.0 | 4.0 | GO:0070268 | cornification(GO:0070268) |
| 0.0 | 2.6 | GO:0044275 | cellular carbohydrate catabolic process(GO:0044275) |
| 0.0 | 0.8 | GO:0006884 | cell volume homeostasis(GO:0006884) |
| 0.0 | 0.1 | GO:1900738 | psychomotor behavior(GO:0036343) positive regulation of phospholipase C-activating G-protein coupled receptor signaling pathway(GO:1900738) |
| 0.0 | 1.2 | GO:0010862 | positive regulation of pathway-restricted SMAD protein phosphorylation(GO:0010862) |
| 0.0 | 0.7 | GO:0008608 | attachment of spindle microtubules to kinetochore(GO:0008608) |
| 0.0 | 0.6 | GO:0050927 | positive regulation of positive chemotaxis(GO:0050927) |
| 0.0 | 2.1 | GO:0048565 | digestive tract development(GO:0048565) |
| 0.0 | 0.5 | GO:0006590 | thyroid hormone generation(GO:0006590) |
| 0.0 | 0.7 | GO:0080171 | lysosome organization(GO:0007040) lytic vacuole organization(GO:0080171) |
| 0.0 | 0.3 | GO:0032411 | positive regulation of transporter activity(GO:0032411) |
| 0.0 | 0.2 | GO:0019530 | taurine metabolic process(GO:0019530) |
| 0.0 | 0.6 | GO:0015695 | organic cation transport(GO:0015695) |
| 0.0 | 0.3 | GO:0002566 | somatic diversification of immune receptors via somatic mutation(GO:0002566) somatic hypermutation of immunoglobulin genes(GO:0016446) |
| 0.0 | 0.1 | GO:0035814 | negative regulation of renal sodium excretion(GO:0035814) |
| 0.0 | 0.6 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.0 | 0.0 | GO:0015808 | L-alanine transport(GO:0015808) |
| 0.0 | 0.2 | GO:0007080 | mitotic metaphase plate congression(GO:0007080) |
| 0.0 | 1.3 | GO:0007098 | centrosome cycle(GO:0007098) |
| 0.0 | 0.7 | GO:0045444 | fat cell differentiation(GO:0045444) |
| 0.0 | 0.5 | GO:0030855 | epithelial cell differentiation(GO:0030855) |
| 0.0 | 0.7 | GO:0050817 | blood coagulation(GO:0007596) coagulation(GO:0050817) |
| 0.0 | 1.6 | GO:0006364 | rRNA processing(GO:0006364) |
| 0.0 | 1.6 | GO:0090305 | nucleic acid phosphodiester bond hydrolysis(GO:0090305) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.5 | 13.6 | GO:0009330 | DNA topoisomerase complex (ATP-hydrolyzing)(GO:0009330) |
| 3.4 | 27.0 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 3.0 | 23.6 | GO:0034363 | intermediate-density lipoprotein particle(GO:0034363) |
| 2.7 | 8.1 | GO:0031523 | Myb complex(GO:0031523) |
| 1.8 | 14.5 | GO:0097504 | Gemini of coiled bodies(GO:0097504) |
| 1.7 | 8.7 | GO:0001940 | male pronucleus(GO:0001940) |
| 1.7 | 8.5 | GO:0097149 | centralspindlin complex(GO:0097149) |
| 1.7 | 8.3 | GO:0032021 | NELF complex(GO:0032021) |
| 1.5 | 6.0 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 1.3 | 15.0 | GO:0031595 | nuclear proteasome complex(GO:0031595) |
| 1.2 | 7.4 | GO:1990584 | cardiac Troponin complex(GO:1990584) |
| 1.2 | 21.3 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 1.1 | 12.0 | GO:0044233 | ER-mitochondrion membrane contact site(GO:0044233) |
| 0.9 | 4.7 | GO:0044214 | spanning component of plasma membrane(GO:0044214) spanning component of membrane(GO:0089717) |
| 0.8 | 6.8 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.7 | 7.8 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.7 | 5.3 | GO:0000421 | autophagosome membrane(GO:0000421) |
| 0.6 | 48.5 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.6 | 3.1 | GO:0071986 | Ragulator complex(GO:0071986) |
| 0.6 | 7.2 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 0.6 | 8.3 | GO:0090543 | Flemming body(GO:0090543) |
| 0.6 | 3.5 | GO:0031390 | Ctf18 RFC-like complex(GO:0031390) |
| 0.6 | 32.3 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.6 | 17.6 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.6 | 6.2 | GO:0034366 | spherical high-density lipoprotein particle(GO:0034366) |
| 0.5 | 3.8 | GO:0036021 | endolysosome lumen(GO:0036021) |
| 0.5 | 4.8 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.5 | 6.2 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
| 0.5 | 4.0 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.4 | 3.1 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.4 | 4.7 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.4 | 2.1 | GO:0034457 | Mpp10 complex(GO:0034457) |
| 0.4 | 23.4 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.4 | 2.7 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 0.4 | 23.4 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.4 | 11.7 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.3 | 3.4 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.3 | 21.2 | GO:0012507 | ER to Golgi transport vesicle membrane(GO:0012507) |
| 0.3 | 3.6 | GO:0045275 | mitochondrial respiratory chain complex III(GO:0005750) respiratory chain complex III(GO:0045275) |
| 0.3 | 5.5 | GO:0000109 | nucleotide-excision repair complex(GO:0000109) |
| 0.3 | 4.3 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 0.3 | 5.4 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.3 | 15.8 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.3 | 0.8 | GO:0071062 | alphav-beta3 integrin-vitronectin complex(GO:0071062) |
| 0.3 | 3.2 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 0.3 | 1.3 | GO:0031501 | mannosyltransferase complex(GO:0031501) |
| 0.3 | 6.6 | GO:0044215 | other organism(GO:0044215) other organism cell(GO:0044216) other organism part(GO:0044217) |
| 0.3 | 2.1 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.3 | 6.9 | GO:0000314 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.2 | 4.5 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 0.2 | 10.9 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.2 | 6.7 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.2 | 4.8 | GO:0017101 | aminoacyl-tRNA synthetase multienzyme complex(GO:0017101) |
| 0.2 | 1.6 | GO:0000347 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.2 | 2.9 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.2 | 0.2 | GO:0060053 | neurofilament cytoskeleton(GO:0060053) |
| 0.2 | 3.2 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.2 | 0.6 | GO:0000145 | exocyst(GO:0000145) |
| 0.2 | 3.9 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.2 | 67.5 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.2 | 2.1 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.2 | 0.9 | GO:0043564 | Ku70:Ku80 complex(GO:0043564) |
| 0.2 | 6.4 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.2 | 3.1 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.2 | 3.5 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.2 | 2.2 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.2 | 5.2 | GO:0005839 | proteasome core complex(GO:0005839) |
| 0.2 | 4.2 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.2 | 4.7 | GO:0101002 | ficolin-1-rich granule(GO:0101002) ficolin-1-rich granule lumen(GO:1904813) |
| 0.1 | 0.4 | GO:0000818 | nuclear MIS12/MIND complex(GO:0000818) condensed chromosome inner kinetochore(GO:0000939) |
| 0.1 | 18.1 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.1 | 2.4 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.1 | 2.4 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.1 | 1.1 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.1 | 11.5 | GO:0000779 | condensed chromosome, centromeric region(GO:0000779) |
| 0.1 | 5.5 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.1 | 5.9 | GO:0002102 | podosome(GO:0002102) |
| 0.1 | 1.8 | GO:0033162 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.1 | 9.6 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.1 | 3.7 | GO:1990391 | DNA repair complex(GO:1990391) |
| 0.1 | 3.5 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.1 | 2.9 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.1 | 3.0 | GO:0099738 | cell cortex region(GO:0099738) |
| 0.1 | 3.3 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.1 | 6.0 | GO:0005604 | basement membrane(GO:0005604) |
| 0.1 | 2.5 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 0.1 | 2.0 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.1 | 1.6 | GO:0036020 | endolysosome membrane(GO:0036020) |
| 0.1 | 16.0 | GO:0019897 | extrinsic component of plasma membrane(GO:0019897) |
| 0.1 | 2.6 | GO:0005753 | mitochondrial proton-transporting ATP synthase complex(GO:0005753) |
| 0.1 | 2.5 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 6.8 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.1 | 28.4 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.1 | 24.5 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.1 | 2.7 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.1 | 32.6 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.1 | 0.5 | GO:0033010 | paranodal junction(GO:0033010) |
| 0.1 | 4.2 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.1 | 167.6 | GO:0070062 | extracellular exosome(GO:0070062) |
| 0.1 | 7.6 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.1 | 2.0 | GO:0001725 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.0 | 0.7 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.0 | 0.7 | GO:0005922 | connexon complex(GO:0005922) |
| 0.0 | 1.0 | GO:0030673 | axolemma(GO:0030673) |
| 0.0 | 5.3 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 2.9 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 1.1 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.0 | 2.2 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 5.7 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 16.1 | GO:0048471 | perinuclear region of cytoplasm(GO:0048471) |
| 0.0 | 3.4 | GO:0005681 | spliceosomal complex(GO:0005681) |
| 0.0 | 0.5 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.0 | 0.5 | GO:0005861 | troponin complex(GO:0005861) |
| 0.0 | 12.8 | GO:0005730 | nucleolus(GO:0005730) |
| 0.0 | 0.2 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.0 | 0.4 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 0.7 | GO:0030665 | clathrin-coated vesicle membrane(GO:0030665) |
| 0.0 | 0.9 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) potassium channel complex(GO:0034705) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.4 | 41.7 | GO:0016784 | 3-mercaptopyruvate sulfurtransferase activity(GO:0016784) |
| 5.7 | 17.0 | GO:0003978 | UDP-N-acetylglucosamine 4-epimerase activity(GO:0003974) UDP-glucose 4-epimerase activity(GO:0003978) |
| 4.8 | 14.4 | GO:0048030 | disaccharide binding(GO:0048030) |
| 4.8 | 9.6 | GO:0004792 | thiosulfate sulfurtransferase activity(GO:0004792) |
| 4.7 | 14.1 | GO:0004514 | nicotinate-nucleotide diphosphorylase (carboxylating) activity(GO:0004514) |
| 4.5 | 13.6 | GO:0003918 | DNA topoisomerase type II (ATP-hydrolyzing) activity(GO:0003918) DNA topoisomerase II activity(GO:0061505) |
| 4.4 | 22.1 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 3.1 | 9.4 | GO:0008413 | 8-oxo-7,8-dihydroguanosine triphosphate pyrophosphatase activity(GO:0008413) 8-oxo-7,8-dihydrodeoxyguanosine triphosphate pyrophosphatase activity(GO:0035539) |
| 2.8 | 11.0 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) thiol oxidase activity(GO:0016972) |
| 2.8 | 11.0 | GO:0070653 | high-density lipoprotein particle receptor binding(GO:0070653) |
| 2.7 | 52.2 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 2.6 | 20.9 | GO:0019863 | IgE binding(GO:0019863) |
| 2.5 | 12.6 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 2.4 | 7.2 | GO:0000773 | phosphatidyl-N-methylethanolamine N-methyltransferase activity(GO:0000773) phosphatidylethanolamine N-methyltransferase activity(GO:0004608) phosphatidyl-N-dimethylethanolamine N-methyltransferase activity(GO:0080101) |
| 2.4 | 16.7 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 2.2 | 8.8 | GO:0015142 | citrate transmembrane transporter activity(GO:0015137) tricarboxylic acid transmembrane transporter activity(GO:0015142) |
| 2.1 | 6.3 | GO:0004781 | adenylylsulfate kinase activity(GO:0004020) sulfate adenylyltransferase activity(GO:0004779) sulfate adenylyltransferase (ATP) activity(GO:0004781) |
| 2.0 | 8.1 | GO:0034988 | Fc-gamma receptor I complex binding(GO:0034988) |
| 2.0 | 10.1 | GO:0047844 | deoxycytidine deaminase activity(GO:0047844) |
| 1.8 | 7.4 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 1.6 | 6.2 | GO:0004998 | transferrin receptor activity(GO:0004998) |
| 1.5 | 6.0 | GO:0004748 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 1.5 | 8.9 | GO:1990254 | keratin filament binding(GO:1990254) |
| 1.4 | 4.2 | GO:0004822 | isoleucine-tRNA ligase activity(GO:0004822) |
| 1.4 | 15.0 | GO:0036402 | proteasome-activating ATPase activity(GO:0036402) |
| 1.3 | 6.7 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 1.2 | 21.3 | GO:0001054 | RNA polymerase I activity(GO:0001054) |
| 1.1 | 5.5 | GO:0032795 | heterotrimeric G-protein binding(GO:0032795) |
| 1.1 | 3.2 | GO:0061752 | telomeric repeat-containing RNA binding(GO:0061752) |
| 1.0 | 11.4 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 1.0 | 4.1 | GO:0005124 | scavenger receptor binding(GO:0005124) |
| 1.0 | 4.9 | GO:0033695 | oxidoreductase activity, acting on CH or CH2 groups, quinone or similar compound as acceptor(GO:0033695) caffeine oxidase activity(GO:0034875) |
| 1.0 | 8.7 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 1.0 | 2.9 | GO:0000253 | 3-keto sterol reductase activity(GO:0000253) |
| 0.9 | 4.7 | GO:0047179 | platelet-activating factor acetyltransferase activity(GO:0047179) |
| 0.9 | 8.9 | GO:0045545 | syndecan binding(GO:0045545) |
| 0.9 | 3.5 | GO:0035651 | AP-1 adaptor complex binding(GO:0035650) AP-3 adaptor complex binding(GO:0035651) |
| 0.9 | 6.9 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.9 | 2.6 | GO:0030272 | 5-formyltetrahydrofolate cyclo-ligase activity(GO:0030272) |
| 0.8 | 6.7 | GO:0004873 | asialoglycoprotein receptor activity(GO:0004873) |
| 0.8 | 5.8 | GO:0004301 | epoxide hydrolase activity(GO:0004301) |
| 0.8 | 2.5 | GO:0070260 | tyrosyl-RNA phosphodiesterase activity(GO:0036317) 5'-tyrosyl-DNA phosphodiesterase activity(GO:0070260) |
| 0.7 | 22.0 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.7 | 2.2 | GO:0030158 | protein xylosyltransferase activity(GO:0030158) |
| 0.7 | 2.1 | GO:0042134 | rRNA primary transcript binding(GO:0042134) |
| 0.7 | 7.4 | GO:0031013 | troponin I binding(GO:0031013) |
| 0.7 | 2.7 | GO:0016744 | transferase activity, transferring aldehyde or ketonic groups(GO:0016744) |
| 0.7 | 8.6 | GO:0046870 | cadmium ion binding(GO:0046870) |
| 0.7 | 2.6 | GO:0001512 | dihydronicotinamide riboside quinone reductase activity(GO:0001512) melatonin binding(GO:1904408) |
| 0.6 | 7.1 | GO:0035251 | UDP-glucosyltransferase activity(GO:0035251) |
| 0.6 | 1.7 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 0.6 | 4.0 | GO:0004459 | lactate dehydrogenase activity(GO:0004457) L-lactate dehydrogenase activity(GO:0004459) |
| 0.5 | 10.3 | GO:0003756 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.5 | 4.2 | GO:0008474 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.5 | 15.0 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.5 | 3.6 | GO:0016681 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.5 | 3.5 | GO:0003689 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.5 | 7.6 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.5 | 2.8 | GO:0016647 | oxidoreductase activity, acting on the CH-NH group of donors, oxygen as acceptor(GO:0016647) |
| 0.4 | 1.3 | GO:0016899 | oxidoreductase activity, acting on the CH-OH group of donors, oxygen as acceptor(GO:0016899) |
| 0.4 | 1.8 | GO:0032810 | sterol response element binding(GO:0032810) |
| 0.4 | 3.5 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.4 | 5.0 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.4 | 9.1 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.4 | 3.2 | GO:0015450 | P-P-bond-hydrolysis-driven protein transmembrane transporter activity(GO:0015450) |
| 0.4 | 8.5 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 0.4 | 3.6 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.4 | 4.8 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.4 | 34.3 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.4 | 11.3 | GO:0050750 | low-density lipoprotein particle receptor binding(GO:0050750) |
| 0.4 | 7.1 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.4 | 27.4 | GO:0019003 | GDP binding(GO:0019003) |
| 0.4 | 2.8 | GO:0004784 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.4 | 1.8 | GO:0033857 | diphosphoinositol-pentakisphosphate kinase activity(GO:0033857) |
| 0.3 | 3.5 | GO:0016312 | inositol bisphosphate phosphatase activity(GO:0016312) |
| 0.3 | 14.3 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.3 | 4.1 | GO:0004128 | cytochrome-b5 reductase activity, acting on NAD(P)H(GO:0004128) |
| 0.3 | 1.4 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.3 | 3.4 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.3 | 2.9 | GO:0004305 | ethanolamine kinase activity(GO:0004305) |
| 0.3 | 1.6 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.3 | 1.9 | GO:0038049 | transcription factor activity, ligand-activated RNA polymerase II transcription factor binding(GO:0038049) |
| 0.3 | 3.3 | GO:0005041 | low-density lipoprotein receptor activity(GO:0005041) |
| 0.3 | 6.7 | GO:0015125 | bile acid transmembrane transporter activity(GO:0015125) |
| 0.3 | 4.8 | GO:0019841 | retinol binding(GO:0019841) |
| 0.3 | 1.8 | GO:0004882 | androgen receptor activity(GO:0004882) |
| 0.3 | 12.8 | GO:0042974 | retinoic acid receptor binding(GO:0042974) |
| 0.3 | 2.3 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.3 | 4.1 | GO:0004726 | non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 0.3 | 1.1 | GO:0050692 | 9-cis retinoic acid receptor activity(GO:0004886) DBD domain binding(GO:0050692) LBD domain binding(GO:0050693) vitamin D response element binding(GO:0070644) |
| 0.3 | 5.0 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.3 | 8.5 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.3 | 1.1 | GO:0052851 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 0.3 | 5.7 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.3 | 0.8 | GO:0003881 | CDP-diacylglycerol-inositol 3-phosphatidyltransferase activity(GO:0003881) |
| 0.3 | 1.1 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.3 | 5.6 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.3 | 7.2 | GO:0005527 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.3 | 3.0 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.2 | 5.2 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.2 | 12.8 | GO:0005160 | transforming growth factor beta receptor binding(GO:0005160) |
| 0.2 | 40.3 | GO:0005178 | integrin binding(GO:0005178) |
| 0.2 | 1.9 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.2 | 1.4 | GO:0031014 | troponin T binding(GO:0031014) |
| 0.2 | 3.2 | GO:0001075 | transcription factor activity, RNA polymerase II core promoter sequence-specific binding involved in preinitiation complex assembly(GO:0001075) |
| 0.2 | 4.4 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.2 | 4.8 | GO:0008483 | transaminase activity(GO:0008483) |
| 0.2 | 1.9 | GO:0030976 | thiamine pyrophosphate binding(GO:0030976) |
| 0.2 | 4.7 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.2 | 10.7 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.2 | 5.3 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.2 | 3.7 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.2 | 0.5 | GO:0033878 | hormone-sensitive lipase activity(GO:0033878) |
| 0.2 | 2.4 | GO:0019870 | potassium channel inhibitor activity(GO:0019870) |
| 0.2 | 3.0 | GO:0043495 | protein anchor(GO:0043495) |
| 0.2 | 6.8 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.1 | 1.5 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 0.1 | 0.6 | GO:0047288 | monosialoganglioside sialyltransferase activity(GO:0047288) |
| 0.1 | 3.2 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.1 | 1.0 | GO:0004996 | thyroid-stimulating hormone receptor activity(GO:0004996) |
| 0.1 | 2.7 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.1 | 2.7 | GO:0017070 | U6 snRNA binding(GO:0017070) |
| 0.1 | 4.8 | GO:0043394 | proteoglycan binding(GO:0043394) |
| 0.1 | 1.5 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.1 | 5.8 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.1 | 0.7 | GO:0005000 | vasopressin receptor activity(GO:0005000) |
| 0.1 | 3.3 | GO:0016859 | cis-trans isomerase activity(GO:0016859) |
| 0.1 | 1.0 | GO:0004340 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.1 | 0.5 | GO:0005166 | neurotrophin p75 receptor binding(GO:0005166) |
| 0.1 | 1.5 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.1 | 4.7 | GO:0004532 | exoribonuclease activity(GO:0004532) |
| 0.1 | 3.5 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.1 | 0.4 | GO:0017002 | activin-activated receptor activity(GO:0017002) |
| 0.1 | 0.7 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.1 | 1.9 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.1 | 3.8 | GO:0030546 | receptor activator activity(GO:0030546) |
| 0.1 | 3.5 | GO:0016776 | phosphotransferase activity, phosphate group as acceptor(GO:0016776) |
| 0.1 | 3.0 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.1 | 4.1 | GO:0005547 | phosphatidylinositol-3,4,5-trisphosphate binding(GO:0005547) |
| 0.1 | 1.9 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.1 | 0.7 | GO:0015165 | pyrimidine nucleotide-sugar transmembrane transporter activity(GO:0015165) |
| 0.1 | 6.7 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.1 | 8.1 | GO:0032947 | protein complex scaffold(GO:0032947) |
| 0.1 | 3.0 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.1 | 4.3 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.1 | 0.5 | GO:0004447 | iodide peroxidase activity(GO:0004447) |
| 0.1 | 8.5 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 0.9 | GO:0051575 | 5'-deoxyribose-5-phosphate lyase activity(GO:0051575) |
| 0.1 | 15.9 | GO:0004866 | endopeptidase inhibitor activity(GO:0004866) |
| 0.1 | 1.6 | GO:0046966 | thyroid hormone receptor binding(GO:0046966) |
| 0.1 | 0.3 | GO:0016979 | lipoate-protein ligase activity(GO:0016979) |
| 0.1 | 2.9 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.1 | 0.5 | GO:0005157 | macrophage colony-stimulating factor receptor binding(GO:0005157) |
| 0.1 | 0.6 | GO:0016857 | racemase and epimerase activity, acting on carbohydrates and derivatives(GO:0016857) |
| 0.1 | 0.4 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.1 | 1.6 | GO:0008198 | ferrous iron binding(GO:0008198) |
| 0.1 | 1.3 | GO:0016712 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) |
| 0.1 | 0.8 | GO:0035240 | dopamine binding(GO:0035240) |
| 0.1 | 13.2 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 0.8 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.1 | 1.3 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.1 | 3.0 | GO:0030170 | pyridoxal phosphate binding(GO:0030170) |
| 0.1 | 2.3 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.1 | 2.9 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.1 | 0.6 | GO:0022820 | potassium:chloride symporter activity(GO:0015379) potassium ion symporter activity(GO:0022820) |
| 0.1 | 1.5 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.1 | 1.3 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.1 | 0.7 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.1 | 1.0 | GO:0004862 | cAMP-dependent protein kinase inhibitor activity(GO:0004862) |
| 0.1 | 2.6 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.1 | 0.6 | GO:0019534 | toxin transporter activity(GO:0019534) |
| 0.0 | 0.3 | GO:0015184 | L-cystine transmembrane transporter activity(GO:0015184) |
| 0.0 | 2.0 | GO:0008237 | metallopeptidase activity(GO:0008237) |
| 0.0 | 0.3 | GO:0005381 | iron ion transmembrane transporter activity(GO:0005381) |
| 0.0 | 2.6 | GO:0016879 | ligase activity, forming carbon-nitrogen bonds(GO:0016879) |
| 0.0 | 2.0 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.0 | 0.2 | GO:0097016 | L27 domain binding(GO:0097016) |
| 0.0 | 4.8 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.0 | 7.2 | GO:0005125 | cytokine activity(GO:0005125) |
| 0.0 | 1.1 | GO:0046961 | proton-transporting ATPase activity, rotational mechanism(GO:0046961) |
| 0.0 | 0.3 | GO:0032052 | bile acid binding(GO:0032052) |
| 0.0 | 3.2 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.0 | 26.8 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 0.4 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.0 | 0.1 | GO:0022850 | serotonin-gated cation channel activity(GO:0022850) |
| 0.0 | 0.6 | GO:0016493 | C-C chemokine receptor activity(GO:0016493) |
| 0.0 | 10.8 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.0 | 2.5 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.0 | 1.1 | GO:0003707 | steroid hormone receptor activity(GO:0003707) |
| 0.0 | 0.6 | GO:0001164 | RNA polymerase I regulatory region DNA binding(GO:0001013) RNA polymerase I regulatory region sequence-specific DNA binding(GO:0001163) RNA polymerase I CORE element sequence-specific DNA binding(GO:0001164) |
| 0.0 | 0.2 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 0.0 | 0.4 | GO:0005537 | mannose binding(GO:0005537) |
| 0.0 | 3.1 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 0.1 | GO:0015266 | protein channel activity(GO:0015266) |
| 0.0 | 0.0 | GO:0022858 | L-alanine transmembrane transporter activity(GO:0015180) alanine transmembrane transporter activity(GO:0022858) |
| 0.0 | 0.3 | GO:0017081 | chloride channel regulator activity(GO:0017081) |
| 0.0 | 3.2 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 4.4 | GO:0051020 | GTPase binding(GO:0051020) |
| 0.0 | 1.4 | GO:0047485 | protein N-terminus binding(GO:0047485) |
| 0.0 | 0.1 | GO:0016206 | catechol O-methyltransferase activity(GO:0016206) |
| 0.0 | 0.3 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.0 | 0.6 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.0 | 0.2 | GO:0019825 | oxygen binding(GO:0019825) |
| 0.0 | 0.6 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 67.8 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.7 | 33.4 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.4 | 8.6 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.3 | 30.2 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.3 | 20.1 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.3 | 4.7 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.2 | 5.5 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.2 | 10.1 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.2 | 10.4 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.2 | 13.0 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.2 | 7.7 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.2 | 8.1 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 0.2 | 12.6 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.2 | 14.6 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.2 | 5.3 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.2 | 3.8 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.1 | 8.8 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.1 | 13.1 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.1 | 6.4 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.1 | 4.7 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.1 | 11.6 | PID E2F PATHWAY | E2F transcription factor network |
| 0.1 | 3.4 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.1 | 6.4 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.1 | 7.8 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.1 | 27.5 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.1 | 1.4 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.1 | 2.8 | PID FOXO PATHWAY | FoxO family signaling |
| 0.1 | 2.0 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 4.8 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.1 | 2.2 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.1 | 3.5 | PID ATR PATHWAY | ATR signaling pathway |
| 0.1 | 12.1 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.1 | 9.5 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 1.2 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.0 | 0.5 | SA TRKA RECEPTOR | The TrkA receptor binds nerve growth factor to activate MAP kinase pathways and promote cell growth. |
| 0.0 | 15.9 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 4.0 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 1.0 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 1.9 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 0.4 | PID ALK1 PATHWAY | ALK1 signaling events |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.3 | 52.2 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 1.5 | 33.3 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 1.3 | 34.5 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.9 | 24.6 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.7 | 22.5 | REACTOME RNA POL III TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase III Transcription Termination |
| 0.7 | 20.9 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.7 | 9.4 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.6 | 6.9 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.6 | 8.5 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.5 | 14.6 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.5 | 8.5 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.5 | 4.7 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.5 | 9.8 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.4 | 11.6 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.4 | 2.4 | REACTOME DOWNSTREAM SIGNAL TRANSDUCTION | Genes involved in Downstream signal transduction |
| 0.4 | 6.3 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.3 | 37.0 | REACTOME MUSCLE CONTRACTION | Genes involved in Muscle contraction |
| 0.3 | 12.1 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.3 | 18.6 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 0.3 | 21.7 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.3 | 14.5 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.3 | 24.3 | REACTOME CDT1 ASSOCIATION WITH THE CDC6 ORC ORIGIN COMPLEX | Genes involved in CDT1 association with the CDC6:ORC:origin complex |
| 0.3 | 7.6 | REACTOME PYRUVATE METABOLISM | Genes involved in Pyruvate metabolism |
| 0.3 | 3.8 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.3 | 7.0 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.3 | 1.9 | REACTOME RECEPTOR LIGAND BINDING INITIATES THE SECOND PROTEOLYTIC CLEAVAGE OF NOTCH RECEPTOR | Genes involved in Receptor-ligand binding initiates the second proteolytic cleavage of Notch receptor |
| 0.3 | 7.2 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.3 | 2.9 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.2 | 2.2 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.2 | 6.6 | REACTOME RNA POL I TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase I Transcription Termination |
| 0.2 | 4.4 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 0.2 | 3.5 | REACTOME POL SWITCHING | Genes involved in Polymerase switching |
| 0.2 | 5.8 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.2 | 32.0 | REACTOME DIABETES PATHWAYS | Genes involved in Diabetes pathways |
| 0.2 | 14.8 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.2 | 4.6 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.2 | 7.1 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.2 | 4.7 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.2 | 6.4 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.2 | 4.2 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.2 | 2.7 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.2 | 23.9 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.2 | 2.6 | REACTOME SIGNALING BY ACTIVATED POINT MUTANTS OF FGFR1 | Genes involved in Signaling by activated point mutants of FGFR1 |
| 0.2 | 2.3 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.2 | 5.8 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.2 | 4.1 | REACTOME FATTY ACYL COA BIOSYNTHESIS | Genes involved in Fatty Acyl-CoA Biosynthesis |
| 0.1 | 4.8 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.1 | 2.9 | REACTOME OXYGEN DEPENDENT PROLINE HYDROXYLATION OF HYPOXIA INDUCIBLE FACTOR ALPHA | Genes involved in Oxygen-dependent Proline Hydroxylation of Hypoxia-inducible Factor Alpha |
| 0.1 | 6.5 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.1 | 2.0 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |
| 0.1 | 8.9 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.1 | 3.9 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 1.6 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.1 | 0.7 | REACTOME REGULATION OF WATER BALANCE BY RENAL AQUAPORINS | Genes involved in Regulation of Water Balance by Renal Aquaporins |
| 0.1 | 1.9 | REACTOME TRIGLYCERIDE BIOSYNTHESIS | Genes involved in Triglyceride Biosynthesis |
| 0.1 | 1.8 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.1 | 2.1 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.1 | 0.4 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 0.1 | 2.0 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.1 | 1.0 | REACTOME HORMONE LIGAND BINDING RECEPTORS | Genes involved in Hormone ligand-binding receptors |
| 0.1 | 6.4 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.1 | 4.6 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.1 | 18.1 | REACTOME METABOLISM OF CARBOHYDRATES | Genes involved in Metabolism of carbohydrates |
| 0.1 | 3.5 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 1.5 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.1 | 6.4 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.1 | 4.8 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.1 | 2.0 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.1 | 5.7 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.1 | 0.9 | REACTOME INTEGRATION OF PROVIRUS | Genes involved in Integration of provirus |
| 0.0 | 1.0 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.0 | 7.2 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.0 | 1.9 | REACTOME TRAFFICKING OF AMPA RECEPTORS | Genes involved in Trafficking of AMPA receptors |
| 0.0 | 3.4 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.0 | 1.1 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.0 | 0.7 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.0 | 0.7 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.0 | 0.1 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GLP1 | Genes involved in Synthesis, Secretion, and Inactivation of Glucagon-like Peptide-1 (GLP-1) |
| 0.0 | 4.7 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.0 | 1.1 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 2.6 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 3.8 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 0.1 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 0.0 | 0.6 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.0 | 0.4 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.0 | 0.9 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |