Project
GNF SymAtlas + NCI-60 cancer cell lines, comparison of cancers vs non-cancers, human (Su, 2004; Ross, 2000)
Navigation
Downloads
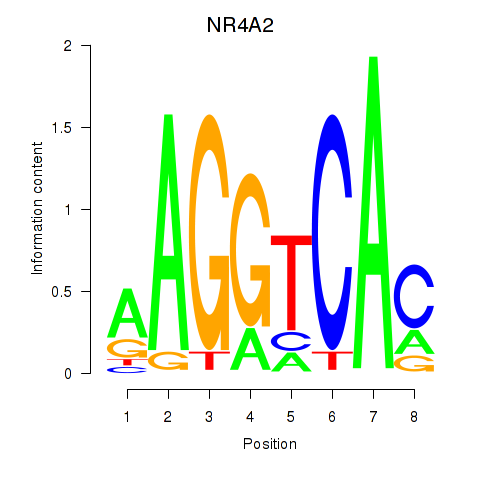
Results for NR4A2
Z-value: 1.21
Transcription factors associated with NR4A2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NR4A2
|
ENSG00000153234.9 | nuclear receptor subfamily 4 group A member 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NR4A2 | hg19_v2_chr2_-_157198860_157198978 | -0.39 | 3.2e-09 | Click! |
Activity profile of NR4A2 motif
Sorted Z-values of NR4A2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_-_72936531 | 31.21 |
ENST00000339594.4
|
BAZ1B
|
bromodomain adjacent to zinc finger domain, 1B |
| chr2_-_176046391 | 25.24 |
ENST00000392541.3
ENST00000409194.1 |
ATP5G3
|
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit C3 (subunit 9) |
| chr10_+_81107271 | 24.98 |
ENST00000448165.1
|
PPIF
|
peptidylprolyl isomerase F |
| chr11_-_64014379 | 22.55 |
ENST00000309318.3
|
PPP1R14B
|
protein phosphatase 1, regulatory (inhibitor) subunit 14B |
| chr11_-_64013663 | 20.87 |
ENST00000392210.2
|
PPP1R14B
|
protein phosphatase 1, regulatory (inhibitor) subunit 14B |
| chr7_+_128399002 | 19.09 |
ENST00000493278.1
|
CALU
|
calumenin |
| chr10_+_51371390 | 17.94 |
ENST00000478381.1
ENST00000451577.2 ENST00000374098.2 ENST00000374097.2 |
TIMM23B
|
translocase of inner mitochondrial membrane 23 homolog B (yeast) |
| chr20_+_3776936 | 17.92 |
ENST00000439880.2
|
CDC25B
|
cell division cycle 25B |
| chr20_+_3776371 | 17.80 |
ENST00000245960.5
|
CDC25B
|
cell division cycle 25B |
| chr4_-_71705060 | 16.86 |
ENST00000514161.1
|
GRSF1
|
G-rich RNA sequence binding factor 1 |
| chr4_-_71705027 | 16.79 |
ENST00000545193.1
|
GRSF1
|
G-rich RNA sequence binding factor 1 |
| chr10_+_81107216 | 16.39 |
ENST00000394579.3
ENST00000225174.3 |
PPIF
|
peptidylprolyl isomerase F |
| chr8_+_145149930 | 16.12 |
ENST00000318911.4
|
CYC1
|
cytochrome c-1 |
| chr14_-_104387888 | 15.69 |
ENST00000286953.3
|
C14orf2
|
chromosome 14 open reading frame 2 |
| chr3_-_167452262 | 14.17 |
ENST00000487947.2
|
PDCD10
|
programmed cell death 10 |
| chr20_-_60718430 | 13.89 |
ENST00000370873.4
ENST00000370858.3 |
PSMA7
|
proteasome (prosome, macropain) subunit, alpha type, 7 |
| chrX_-_109683446 | 13.80 |
ENST00000372057.1
|
AMMECR1
|
Alport syndrome, mental retardation, midface hypoplasia and elliptocytosis chromosomal region gene 1 |
| chr8_-_131028869 | 13.39 |
ENST00000518283.1
ENST00000519110.1 |
FAM49B
|
family with sequence similarity 49, member B |
| chr1_-_17380630 | 13.36 |
ENST00000375499.3
|
SDHB
|
succinate dehydrogenase complex, subunit B, iron sulfur (Ip) |
| chr14_-_104387831 | 13.28 |
ENST00000557040.1
ENST00000414262.2 ENST00000555030.1 ENST00000554713.1 ENST00000553430.1 |
C14orf2
|
chromosome 14 open reading frame 2 |
| chr2_-_207023918 | 12.91 |
ENST00000455934.2
ENST00000449699.1 ENST00000454195.1 |
NDUFS1
|
NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) |
| chr2_-_207024134 | 12.66 |
ENST00000457011.1
ENST00000440274.1 ENST00000432169.1 |
NDUFS1
|
NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) |
| chr1_-_241683001 | 12.26 |
ENST00000366560.3
|
FH
|
fumarate hydratase |
| chr1_+_32757668 | 11.75 |
ENST00000373548.3
|
HDAC1
|
histone deacetylase 1 |
| chr5_-_133340326 | 11.52 |
ENST00000425992.1
ENST00000395044.3 ENST00000395047.2 |
VDAC1
|
voltage-dependent anion channel 1 |
| chr3_-_167452298 | 11.44 |
ENST00000475915.2
ENST00000462725.2 ENST00000461494.1 |
PDCD10
|
programmed cell death 10 |
| chr11_+_34938119 | 11.34 |
ENST00000227868.4
ENST00000430469.2 ENST00000533262.1 |
PDHX
|
pyruvate dehydrogenase complex, component X |
| chr10_-_51623203 | 10.92 |
ENST00000444743.1
ENST00000374065.3 ENST00000374064.3 ENST00000260867.4 |
TIMM23
|
translocase of inner mitochondrial membrane 23 homolog (yeast) |
| chr16_-_69368774 | 10.86 |
ENST00000562949.1
|
RP11-343C2.12
|
Conserved oligomeric Golgi complex subunit 8 |
| chr4_-_74088800 | 10.84 |
ENST00000509867.2
|
ANKRD17
|
ankyrin repeat domain 17 |
| chr10_-_120938303 | 10.78 |
ENST00000356951.3
ENST00000298510.2 |
PRDX3
|
peroxiredoxin 3 |
| chr19_-_2328572 | 10.71 |
ENST00000252622.10
|
LSM7
|
LSM7 homolog, U6 small nuclear RNA associated (S. cerevisiae) |
| chr10_+_14880157 | 10.47 |
ENST00000378372.3
|
HSPA14
|
heat shock 70kDa protein 14 |
| chr1_+_155583012 | 10.45 |
ENST00000462250.2
|
MSTO1
|
misato 1, mitochondrial distribution and morphology regulator |
| chr3_+_138340067 | 10.43 |
ENST00000479848.1
|
FAIM
|
Fas apoptotic inhibitory molecule |
| chr17_+_79670386 | 10.39 |
ENST00000333676.3
ENST00000571730.1 ENST00000541223.1 |
MRPL12
SLC25A10
SLC25A10
|
mitochondrial ribosomal protein L12 Mitochondrial dicarboxylate carrier; Uncharacterized protein; cDNA FLJ60124, highly similar to Mitochondrial dicarboxylate carrier solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 |
| chr1_+_68150744 | 10.37 |
ENST00000370986.4
ENST00000370985.3 |
GADD45A
|
growth arrest and DNA-damage-inducible, alpha |
| chr16_-_85833109 | 10.33 |
ENST00000253457.3
|
EMC8
|
ER membrane protein complex subunit 8 |
| chr3_-_113465065 | 9.84 |
ENST00000497255.1
ENST00000478020.1 ENST00000240922.3 ENST00000493900.1 |
NAA50
|
N(alpha)-acetyltransferase 50, NatE catalytic subunit |
| chr11_+_86748863 | 9.57 |
ENST00000340353.7
|
TMEM135
|
transmembrane protein 135 |
| chr8_+_145582633 | 9.40 |
ENST00000540505.1
|
SLC52A2
|
solute carrier family 52 (riboflavin transporter), member 2 |
| chr22_-_43036607 | 9.19 |
ENST00000505920.1
|
ATP5L2
|
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit G2 |
| chr15_+_75335604 | 9.11 |
ENST00000563393.1
|
PPCDC
|
phosphopantothenoylcysteine decarboxylase |
| chr1_+_155179012 | 8.95 |
ENST00000609421.1
|
MTX1
|
metaxin 1 |
| chr11_+_66624527 | 8.86 |
ENST00000393952.3
|
LRFN4
|
leucine rich repeat and fibronectin type III domain containing 4 |
| chr15_-_55562479 | 8.72 |
ENST00000564609.1
|
RAB27A
|
RAB27A, member RAS oncogene family |
| chr8_-_131028641 | 8.27 |
ENST00000523509.1
|
FAM49B
|
family with sequence similarity 49, member B |
| chr19_+_8509842 | 8.27 |
ENST00000325495.4
ENST00000600092.1 ENST00000594907.1 ENST00000596984.1 ENST00000601645.1 |
HNRNPM
|
heterogeneous nuclear ribonucleoprotein M |
| chr2_+_198365122 | 7.98 |
ENST00000604458.1
|
HSPE1-MOB4
|
HSPE1-MOB4 readthrough |
| chr2_+_85804614 | 7.81 |
ENST00000263864.5
ENST00000409760.1 |
VAMP8
|
vesicle-associated membrane protein 8 |
| chr3_-_10028366 | 7.75 |
ENST00000429759.1
|
EMC3
|
ER membrane protein complex subunit 3 |
| chr4_+_41614720 | 7.61 |
ENST00000509277.1
|
LIMCH1
|
LIM and calponin homology domains 1 |
| chr1_+_201924619 | 7.49 |
ENST00000367287.4
|
TIMM17A
|
translocase of inner mitochondrial membrane 17 homolog A (yeast) |
| chr4_-_71705082 | 7.23 |
ENST00000439371.1
ENST00000499044.2 |
GRSF1
|
G-rich RNA sequence binding factor 1 |
| chr6_-_24667180 | 7.16 |
ENST00000545995.1
|
TDP2
|
tyrosyl-DNA phosphodiesterase 2 |
| chr5_-_137878887 | 7.16 |
ENST00000507939.1
ENST00000572514.1 ENST00000499810.2 ENST00000360541.5 |
ETF1
|
eukaryotic translation termination factor 1 |
| chr4_-_139163491 | 7.13 |
ENST00000280612.5
|
SLC7A11
|
solute carrier family 7 (anionic amino acid transporter light chain, xc- system), member 11 |
| chr19_-_8008533 | 7.12 |
ENST00000597926.1
|
TIMM44
|
translocase of inner mitochondrial membrane 44 homolog (yeast) |
| chr19_-_10446449 | 7.02 |
ENST00000592439.1
|
ICAM3
|
intercellular adhesion molecule 3 |
| chr6_-_24666819 | 6.78 |
ENST00000341060.3
|
TDP2
|
tyrosyl-DNA phosphodiesterase 2 |
| chr8_-_100905925 | 6.75 |
ENST00000518171.1
|
COX6C
|
cytochrome c oxidase subunit VIc |
| chr2_+_234621551 | 6.61 |
ENST00000608381.1
ENST00000373414.3 |
UGT1A1
UGT1A5
|
UDP glucuronosyltransferase 1 family, polypeptide A8 UDP glucuronosyltransferase 1 family, polypeptide A5 |
| chr14_+_23791159 | 6.57 |
ENST00000557702.1
|
PABPN1
|
poly(A) binding protein, nuclear 1 |
| chr3_+_138340049 | 6.53 |
ENST00000464668.1
|
FAIM
|
Fas apoptotic inhibitory molecule |
| chr3_+_179322573 | 6.43 |
ENST00000493866.1
ENST00000472629.1 ENST00000482604.1 |
NDUFB5
|
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 5, 16kDa |
| chr7_-_107642348 | 6.42 |
ENST00000393561.1
|
LAMB1
|
laminin, beta 1 |
| chr16_-_30457048 | 6.41 |
ENST00000500504.2
ENST00000542752.1 |
SEPHS2
|
selenophosphate synthetase 2 |
| chr12_+_120875910 | 6.41 |
ENST00000551806.1
|
AL021546.6
|
Glutamyl-tRNA(Gln) amidotransferase subunit C, mitochondrial |
| chr10_+_22605374 | 6.40 |
ENST00000448361.1
|
COMMD3
|
COMM domain containing 3 |
| chr22_+_30163340 | 6.35 |
ENST00000330029.6
ENST00000401406.3 |
UQCR10
|
ubiquinol-cytochrome c reductase, complex III subunit X |
| chr19_+_45394477 | 6.23 |
ENST00000252487.5
ENST00000405636.2 ENST00000592434.1 ENST00000426677.2 ENST00000589649.1 |
TOMM40
|
translocase of outer mitochondrial membrane 40 homolog (yeast) |
| chr1_-_1711508 | 6.22 |
ENST00000378625.1
|
NADK
|
NAD kinase |
| chr8_-_53626974 | 5.95 |
ENST00000435644.2
ENST00000518710.1 ENST00000025008.5 ENST00000517963.1 |
RB1CC1
|
RB1-inducible coiled-coil 1 |
| chr1_-_205719295 | 5.91 |
ENST00000367142.4
|
NUCKS1
|
nuclear casein kinase and cyclin-dependent kinase substrate 1 |
| chr19_+_1407517 | 5.90 |
ENST00000336761.6
ENST00000233078.4 |
DAZAP1
|
DAZ associated protein 1 |
| chr14_-_55369525 | 5.88 |
ENST00000543643.2
ENST00000536224.2 ENST00000395514.1 ENST00000491895.2 |
GCH1
|
GTP cyclohydrolase 1 |
| chr3_+_113465866 | 5.72 |
ENST00000273398.3
ENST00000538620.1 ENST00000496747.1 ENST00000475322.1 |
ATP6V1A
|
ATPase, H+ transporting, lysosomal 70kDa, V1 subunit A |
| chr1_-_27816641 | 5.66 |
ENST00000430629.2
|
WASF2
|
WAS protein family, member 2 |
| chr2_-_26467557 | 5.63 |
ENST00000380649.3
|
HADHA
|
hydroxyacyl-CoA dehydrogenase/3-ketoacyl-CoA thiolase/enoyl-CoA hydratase (trifunctional protein), alpha subunit |
| chr2_+_198365095 | 5.63 |
ENST00000409468.1
|
HSPE1
|
heat shock 10kDa protein 1 |
| chr9_+_115983808 | 5.41 |
ENST00000374210.6
ENST00000374212.4 |
SLC31A1
|
solute carrier family 31 (copper transporter), member 1 |
| chr2_+_109237717 | 5.41 |
ENST00000409441.1
|
LIMS1
|
LIM and senescent cell antigen-like domains 1 |
| chr4_-_140216948 | 5.33 |
ENST00000265500.4
|
NDUFC1
|
NADH dehydrogenase (ubiquinone) 1, subcomplex unknown, 1, 6kDa |
| chr9_-_34048873 | 4.96 |
ENST00000449054.1
ENST00000379239.4 ENST00000539807.1 ENST00000379238.1 ENST00000418786.2 ENST00000360802.1 ENST00000412543.1 |
UBAP2
|
ubiquitin associated protein 2 |
| chr5_-_137911049 | 4.84 |
ENST00000297185.3
|
HSPA9
|
heat shock 70kDa protein 9 (mortalin) |
| chr16_+_4674787 | 4.82 |
ENST00000262370.7
|
MGRN1
|
mahogunin ring finger 1, E3 ubiquitin protein ligase |
| chr18_-_12377283 | 4.70 |
ENST00000269143.3
|
AFG3L2
|
AFG3-like AAA ATPase 2 |
| chr19_+_46850251 | 4.63 |
ENST00000012443.4
|
PPP5C
|
protein phosphatase 5, catalytic subunit |
| chr3_+_159570722 | 4.59 |
ENST00000482804.1
|
SCHIP1
|
schwannomin interacting protein 1 |
| chr3_-_197025447 | 4.57 |
ENST00000346964.2
ENST00000357674.4 ENST00000314062.3 ENST00000448528.2 ENST00000419553.1 |
DLG1
|
discs, large homolog 1 (Drosophila) |
| chr10_+_22605304 | 4.49 |
ENST00000475460.2
ENST00000602390.1 ENST00000489125.2 ENST00000456711.1 ENST00000444869.1 |
COMMD3-BMI1
COMMD3
|
COMMD3-BMI1 readthrough COMM domain containing 3 |
| chr16_-_70729496 | 4.48 |
ENST00000567648.1
|
VAC14
|
Vac14 homolog (S. cerevisiae) |
| chr8_-_145018905 | 4.45 |
ENST00000398774.2
|
PLEC
|
plectin |
| chr2_-_207024233 | 4.44 |
ENST00000423725.1
ENST00000233190.6 |
NDUFS1
|
NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) |
| chr3_+_179322481 | 4.42 |
ENST00000259037.3
|
NDUFB5
|
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 5, 16kDa |
| chr15_-_55562582 | 4.40 |
ENST00000396307.2
|
RAB27A
|
RAB27A, member RAS oncogene family |
| chr8_-_100905850 | 4.30 |
ENST00000520271.1
ENST00000522940.1 ENST00000523016.1 ENST00000517682.2 ENST00000297564.2 |
COX6C
|
cytochrome c oxidase subunit VIc |
| chr18_-_19284724 | 4.29 |
ENST00000580981.1
ENST00000289119.2 |
ABHD3
|
abhydrolase domain containing 3 |
| chr20_-_62130474 | 4.09 |
ENST00000217182.3
|
EEF1A2
|
eukaryotic translation elongation factor 1 alpha 2 |
| chr7_+_39663485 | 4.05 |
ENST00000436179.1
|
RALA
|
v-ral simian leukemia viral oncogene homolog A (ras related) |
| chr4_+_81118647 | 3.97 |
ENST00000415738.2
|
PRDM8
|
PR domain containing 8 |
| chr15_+_78441663 | 3.89 |
ENST00000299518.2
ENST00000558554.1 ENST00000557826.1 ENST00000561279.1 ENST00000559186.1 ENST00000560770.1 ENST00000559881.1 ENST00000559205.1 ENST00000441490.2 |
IDH3A
|
isocitrate dehydrogenase 3 (NAD+) alpha |
| chr22_+_31518938 | 3.83 |
ENST00000412985.1
ENST00000331075.5 ENST00000412277.2 ENST00000420017.1 ENST00000400294.2 ENST00000405300.1 ENST00000404390.3 |
INPP5J
|
inositol polyphosphate-5-phosphatase J |
| chr2_+_228678550 | 3.70 |
ENST00000409189.3
ENST00000358813.4 |
CCL20
|
chemokine (C-C motif) ligand 20 |
| chr1_-_20987851 | 3.59 |
ENST00000464364.1
ENST00000602624.2 |
DDOST
|
dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit (non-catalytic) |
| chr20_-_13765526 | 3.54 |
ENST00000202816.1
|
ESF1
|
ESF1, nucleolar pre-rRNA processing protein, homolog (S. cerevisiae) |
| chr19_+_6739662 | 3.52 |
ENST00000313285.8
ENST00000313244.9 ENST00000596758.1 |
TRIP10
|
thyroid hormone receptor interactor 10 |
| chr2_-_26467465 | 3.52 |
ENST00000457468.2
|
HADHA
|
hydroxyacyl-CoA dehydrogenase/3-ketoacyl-CoA thiolase/enoyl-CoA hydratase (trifunctional protein), alpha subunit |
| chr7_-_140179276 | 3.51 |
ENST00000443720.2
ENST00000255977.2 |
MKRN1
|
makorin ring finger protein 1 |
| chr11_-_9336234 | 3.47 |
ENST00000528080.1
|
TMEM41B
|
transmembrane protein 41B |
| chr17_-_56358287 | 3.37 |
ENST00000225275.3
ENST00000340482.3 |
MPO
|
myeloperoxidase |
| chr17_-_56350797 | 3.35 |
ENST00000577220.1
|
MPO
|
myeloperoxidase |
| chr11_+_67798363 | 3.27 |
ENST00000525628.1
|
NDUFS8
|
NADH dehydrogenase (ubiquinone) Fe-S protein 8, 23kDa (NADH-coenzyme Q reductase) |
| chr1_-_20987889 | 3.27 |
ENST00000415136.2
|
DDOST
|
dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit (non-catalytic) |
| chr16_-_30537839 | 3.22 |
ENST00000380412.5
|
ZNF768
|
zinc finger protein 768 |
| chr8_-_100905363 | 3.02 |
ENST00000524245.1
|
COX6C
|
cytochrome c oxidase subunit VIc |
| chr3_-_151034734 | 2.97 |
ENST00000260843.4
|
GPR87
|
G protein-coupled receptor 87 |
| chr5_+_218356 | 2.96 |
ENST00000264932.6
ENST00000504309.1 ENST00000510361.1 |
SDHA
|
succinate dehydrogenase complex, subunit A, flavoprotein (Fp) |
| chr20_-_33999766 | 2.95 |
ENST00000349714.5
ENST00000438533.1 ENST00000359226.2 ENST00000374384.2 ENST00000374377.5 ENST00000407996.2 ENST00000424405.1 ENST00000542501.1 ENST00000397554.1 ENST00000540457.1 ENST00000374380.2 ENST00000374385.5 |
UQCC1
|
ubiquinol-cytochrome c reductase complex assembly factor 1 |
| chr17_-_79167828 | 2.94 |
ENST00000570817.1
|
AZI1
|
5-azacytidine induced 1 |
| chr3_+_159557637 | 2.93 |
ENST00000445224.2
|
SCHIP1
|
schwannomin interacting protein 1 |
| chr4_-_40517984 | 2.92 |
ENST00000381795.6
|
RBM47
|
RNA binding motif protein 47 |
| chr20_-_1373606 | 2.91 |
ENST00000381715.1
ENST00000439640.2 ENST00000381719.3 |
FKBP1A
|
FK506 binding protein 1A, 12kDa |
| chr2_+_234627424 | 2.80 |
ENST00000373409.3
|
UGT1A4
|
UDP glucuronosyltransferase 1 family, polypeptide A4 |
| chr2_+_119981384 | 2.67 |
ENST00000393108.2
ENST00000354888.5 ENST00000450943.2 ENST00000393110.2 ENST00000393106.2 ENST00000409811.1 ENST00000393107.2 |
STEAP3
|
STEAP family member 3, metalloreductase |
| chr12_-_57039739 | 2.67 |
ENST00000552959.1
ENST00000551020.1 ENST00000553007.2 ENST00000552919.1 ENST00000552104.1 ENST00000262030.3 |
ATP5B
|
ATP synthase, H+ transporting, mitochondrial F1 complex, beta polypeptide |
| chr4_-_151936416 | 2.55 |
ENST00000510413.1
ENST00000507224.1 |
LRBA
|
LPS-responsive vesicle trafficking, beach and anchor containing |
| chrX_+_106045891 | 2.50 |
ENST00000357242.5
ENST00000310452.2 ENST00000481617.2 ENST00000276175.3 |
TBC1D8B
|
TBC1 domain family, member 8B (with GRAM domain) |
| chr15_+_43885252 | 2.49 |
ENST00000453782.1
ENST00000300283.6 ENST00000437924.1 ENST00000450086.2 |
CKMT1B
|
creatine kinase, mitochondrial 1B |
| chr4_-_151936865 | 2.47 |
ENST00000535741.1
|
LRBA
|
LPS-responsive vesicle trafficking, beach and anchor containing |
| chr4_+_41614909 | 2.44 |
ENST00000509454.1
ENST00000396595.3 ENST00000381753.4 |
LIMCH1
|
LIM and calponin homology domains 1 |
| chr15_+_43985725 | 2.36 |
ENST00000413453.2
|
CKMT1A
|
creatine kinase, mitochondrial 1A |
| chr7_-_150777874 | 2.35 |
ENST00000540185.1
|
FASTK
|
Fas-activated serine/threonine kinase |
| chr19_-_46476791 | 2.33 |
ENST00000263257.5
|
NOVA2
|
neuro-oncological ventral antigen 2 |
| chr11_+_67798114 | 2.25 |
ENST00000453471.2
ENST00000528492.1 ENST00000526339.1 ENST00000525419.1 |
NDUFS8
|
NADH dehydrogenase (ubiquinone) Fe-S protein 8, 23kDa (NADH-coenzyme Q reductase) |
| chr1_+_206858328 | 2.17 |
ENST00000367103.3
|
MAPKAPK2
|
mitogen-activated protein kinase-activated protein kinase 2 |
| chr7_-_150777949 | 2.12 |
ENST00000482571.1
|
FASTK
|
Fas-activated serine/threonine kinase |
| chr15_+_43985084 | 2.06 |
ENST00000434505.1
ENST00000411750.1 |
CKMT1A
|
creatine kinase, mitochondrial 1A |
| chr14_+_22320634 | 2.01 |
ENST00000390435.1
|
TRAV8-3
|
T cell receptor alpha variable 8-3 |
| chr20_+_47538357 | 1.99 |
ENST00000371917.4
|
ARFGEF2
|
ADP-ribosylation factor guanine nucleotide-exchange factor 2 (brefeldin A-inhibited) |
| chr9_+_113431029 | 1.97 |
ENST00000189978.5
ENST00000374448.4 ENST00000374440.3 |
MUSK
|
muscle, skeletal, receptor tyrosine kinase |
| chr21_-_35014027 | 1.93 |
ENST00000399442.1
ENST00000413017.2 ENST00000445393.1 ENST00000417979.1 ENST00000426935.1 ENST00000381540.3 ENST00000361534.2 ENST00000381554.3 |
CRYZL1
|
crystallin, zeta (quinone reductase)-like 1 |
| chr11_+_22696314 | 1.91 |
ENST00000532398.1
ENST00000433790.1 |
GAS2
|
growth arrest-specific 2 |
| chr7_-_150777920 | 1.90 |
ENST00000353841.2
ENST00000297532.6 |
FASTK
|
Fas-activated serine/threonine kinase |
| chr11_-_66139199 | 1.81 |
ENST00000357440.2
|
SLC29A2
|
solute carrier family 29 (equilibrative nucleoside transporter), member 2 |
| chr9_+_113431059 | 1.78 |
ENST00000416899.2
|
MUSK
|
muscle, skeletal, receptor tyrosine kinase |
| chr17_-_9940058 | 1.72 |
ENST00000585266.1
|
GAS7
|
growth arrest-specific 7 |
| chr22_+_40573921 | 1.71 |
ENST00000454349.2
ENST00000335727.9 |
TNRC6B
|
trinucleotide repeat containing 6B |
| chr11_+_122709200 | 1.63 |
ENST00000227348.4
|
CRTAM
|
cytotoxic and regulatory T cell molecule |
| chr19_-_49552006 | 1.58 |
ENST00000391869.3
|
CGB1
|
chorionic gonadotropin, beta polypeptide 1 |
| chr10_-_33625154 | 1.55 |
ENST00000265371.4
|
NRP1
|
neuropilin 1 |
| chr7_-_91875109 | 1.51 |
ENST00000412043.2
ENST00000430102.1 ENST00000425073.1 ENST00000394503.2 ENST00000454017.1 ENST00000440209.1 ENST00000413688.1 ENST00000452773.1 ENST00000433016.1 ENST00000394505.2 ENST00000422347.1 ENST00000458493.1 ENST00000425919.1 |
KRIT1
|
KRIT1, ankyrin repeat containing |
| chr1_-_154164534 | 1.50 |
ENST00000271850.7
ENST00000368530.2 |
TPM3
|
tropomyosin 3 |
| chr7_-_151433342 | 1.45 |
ENST00000433631.2
|
PRKAG2
|
protein kinase, AMP-activated, gamma 2 non-catalytic subunit |
| chr12_+_58166431 | 1.41 |
ENST00000333012.5
|
METTL21B
|
methyltransferase like 21B |
| chr7_+_73245193 | 1.40 |
ENST00000340958.2
|
CLDN4
|
claudin 4 |
| chr22_-_37880543 | 1.37 |
ENST00000442496.1
|
MFNG
|
MFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| chr20_+_30865429 | 1.36 |
ENST00000375712.3
|
KIF3B
|
kinesin family member 3B |
| chr5_-_149829314 | 1.35 |
ENST00000407193.1
|
RPS14
|
ribosomal protein S14 |
| chr7_+_112090483 | 1.34 |
ENST00000403825.3
ENST00000429071.1 |
IFRD1
|
interferon-related developmental regulator 1 |
| chr7_-_151433393 | 1.33 |
ENST00000492843.1
|
PRKAG2
|
protein kinase, AMP-activated, gamma 2 non-catalytic subunit |
| chr22_-_30960876 | 1.30 |
ENST00000401975.1
ENST00000428682.1 ENST00000423299.1 |
GAL3ST1
|
galactose-3-O-sulfotransferase 1 |
| chr3_-_197024965 | 1.29 |
ENST00000392382.2
|
DLG1
|
discs, large homolog 1 (Drosophila) |
| chr12_+_6309963 | 1.26 |
ENST00000382515.2
|
CD9
|
CD9 molecule |
| chr12_-_108954933 | 1.25 |
ENST00000431469.2
ENST00000546815.1 |
SART3
|
squamous cell carcinoma antigen recognized by T cells 3 |
| chr17_-_10276319 | 1.24 |
ENST00000252172.4
ENST00000418404.3 |
MYH13
|
myosin, heavy chain 13, skeletal muscle |
| chr3_-_15540055 | 1.21 |
ENST00000605797.1
ENST00000435459.2 |
COLQ
|
collagen-like tail subunit (single strand of homotrimer) of asymmetric acetylcholinesterase |
| chr7_-_91875358 | 1.21 |
ENST00000458177.1
ENST00000394507.1 ENST00000340022.2 ENST00000444960.1 |
KRIT1
|
KRIT1, ankyrin repeat containing |
| chr2_+_191334212 | 1.18 |
ENST00000444317.1
ENST00000535751.1 |
MFSD6
|
major facilitator superfamily domain containing 6 |
| chrX_-_63005405 | 1.16 |
ENST00000374878.1
ENST00000437457.2 |
ARHGEF9
|
Cdc42 guanine nucleotide exchange factor (GEF) 9 |
| chr13_+_102142296 | 1.14 |
ENST00000376162.3
|
ITGBL1
|
integrin, beta-like 1 (with EGF-like repeat domains) |
| chr12_+_120875887 | 1.03 |
ENST00000229379.2
|
COX6A1
|
cytochrome c oxidase subunit VIa polypeptide 1 |
| chr5_-_149829294 | 1.01 |
ENST00000401695.3
|
RPS14
|
ribosomal protein S14 |
| chr1_-_29508499 | 0.91 |
ENST00000373795.4
|
SRSF4
|
serine/arginine-rich splicing factor 4 |
| chr3_-_46000064 | 0.85 |
ENST00000433878.1
|
FYCO1
|
FYVE and coiled-coil domain containing 1 |
| chr19_+_45251804 | 0.73 |
ENST00000164227.5
|
BCL3
|
B-cell CLL/lymphoma 3 |
| chr7_-_81399355 | 0.62 |
ENST00000457544.2
|
HGF
|
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr19_-_49552363 | 0.59 |
ENST00000448456.3
ENST00000355414.2 |
CGB8
|
chorionic gonadotropin, beta polypeptide 8 |
| chr17_+_79679299 | 0.48 |
ENST00000331531.5
|
SLC25A10
|
solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 |
| chr16_+_3704822 | 0.44 |
ENST00000414110.2
|
DNASE1
|
deoxyribonuclease I |
| chr12_-_108955070 | 0.43 |
ENST00000228284.3
ENST00000546611.1 |
SART3
|
squamous cell carcinoma antigen recognized by T cells 3 |
| chr9_+_33629119 | 0.38 |
ENST00000331828.4
|
TRBV21OR9-2
|
T cell receptor beta variable 21/OR9-2 (pseudogene) |
| chr17_-_27503770 | 0.36 |
ENST00000533112.1
|
MYO18A
|
myosin XVIIIA |
| chrX_-_153640420 | 0.34 |
ENST00000451865.1
ENST00000432135.1 ENST00000369809.1 ENST00000393638.1 ENST00000424626.1 ENST00000309585.5 |
DNASE1L1
|
deoxyribonuclease I-like 1 |
| chr5_-_148758839 | 0.29 |
ENST00000261796.3
|
IL17B
|
interleukin 17B |
| chr6_-_31628512 | 0.26 |
ENST00000375911.1
|
C6orf47
|
chromosome 6 open reading frame 47 |
| chr11_-_19223523 | 0.21 |
ENST00000265968.3
|
CSRP3
|
cysteine and glycine-rich protein 3 (cardiac LIM protein) |
| chr6_+_107077435 | 0.17 |
ENST00000369046.4
|
QRSL1
|
glutaminyl-tRNA synthase (glutamine-hydrolyzing)-like 1 |
| chr1_+_212738676 | 0.01 |
ENST00000366981.4
ENST00000366987.2 |
ATF3
|
activating transcription factor 3 |
Network of associatons between targets according to the STRING database.
First level regulatory network of NR4A2
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 13.8 | 41.4 | GO:2000276 | regulation of mitochondrial membrane permeability involved in programmed necrotic cell death(GO:1902445) negative regulation of oxidative phosphorylation uncoupler activity(GO:2000276) |
| 5.1 | 25.6 | GO:1903588 | negative regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903588) |
| 4.1 | 12.3 | GO:0006106 | fumarate metabolic process(GO:0006106) |
| 3.6 | 10.8 | GO:0071423 | thiosulfate transport(GO:0015709) oxaloacetate transport(GO:0015729) malate transport(GO:0015743) malate transmembrane transport(GO:0071423) oxaloacetate(2-) transmembrane transport(GO:1902356) |
| 3.6 | 21.7 | GO:2000567 | memory T cell activation(GO:0035709) regulation of memory T cell activation(GO:2000567) positive regulation of memory T cell activation(GO:2000568) |
| 3.6 | 10.8 | GO:0018171 | peptidyl-cysteine oxidation(GO:0018171) |
| 3.6 | 35.7 | GO:0007144 | female meiosis I(GO:0007144) |
| 3.5 | 10.5 | GO:0051083 | 'de novo' cotranslational protein folding(GO:0051083) |
| 3.3 | 49.7 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 2.7 | 10.8 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 2.6 | 7.8 | GO:1903595 | positive regulation of histamine secretion by mast cell(GO:1903595) |
| 2.3 | 11.7 | GO:0061198 | fungiform papilla formation(GO:0061198) |
| 2.3 | 9.4 | GO:0032218 | riboflavin transport(GO:0032218) |
| 2.2 | 6.6 | GO:1904247 | positive regulation of polynucleotide adenylyltransferase activity(GO:1904247) |
| 2.2 | 13.1 | GO:1903435 | positive regulation of constitutive secretory pathway(GO:1903435) |
| 2.1 | 25.8 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 2.1 | 6.4 | GO:0016260 | selenocysteine biosynthetic process(GO:0016260) |
| 2.0 | 5.9 | GO:0051066 | dihydrobiopterin metabolic process(GO:0051066) |
| 1.6 | 9.8 | GO:0070601 | centromeric sister chromatid cohesion(GO:0070601) |
| 1.6 | 22.5 | GO:0006122 | mitochondrial electron transport, ubiquinol to cytochrome c(GO:0006122) |
| 1.6 | 6.4 | GO:0021812 | neuronal-glial interaction involved in cerebral cortex radial glia guided migration(GO:0021812) |
| 1.6 | 9.4 | GO:0006789 | bilirubin conjugation(GO:0006789) |
| 1.5 | 5.9 | GO:0030242 | pexophagy(GO:0030242) |
| 1.5 | 5.9 | GO:0019046 | release from viral latency(GO:0019046) |
| 1.3 | 11.3 | GO:0061732 | mitochondrial acetyl-CoA biosynthetic process from pyruvate(GO:0061732) |
| 1.2 | 3.7 | GO:0045360 | regulation of interleukin-1 biosynthetic process(GO:0045360) positive regulation of interleukin-1 biosynthetic process(GO:0045362) |
| 1.2 | 16.3 | GO:0006105 | succinate metabolic process(GO:0006105) |
| 1.2 | 8.1 | GO:0015677 | copper ion import(GO:0015677) |
| 1.1 | 6.4 | GO:0035965 | cardiolipin acyl-chain remodeling(GO:0035965) |
| 1.0 | 6.2 | GO:0006741 | NADP biosynthetic process(GO:0006741) |
| 0.9 | 36.3 | GO:0015985 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.8 | 15.1 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.8 | 5.9 | GO:1903760 | regulation of voltage-gated potassium channel activity involved in ventricular cardiac muscle cell action potential repolarization(GO:1903760) |
| 0.8 | 16.2 | GO:0006851 | mitochondrial calcium ion transport(GO:0006851) |
| 0.8 | 49.2 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.7 | 6.7 | GO:0001878 | response to yeast(GO:0001878) |
| 0.7 | 5.0 | GO:0008582 | regulation of synaptic growth at neuromuscular junction(GO:0008582) |
| 0.6 | 40.9 | GO:0006378 | mRNA polyadenylation(GO:0006378) |
| 0.6 | 4.8 | GO:1902037 | negative regulation of hematopoietic stem cell differentiation(GO:1902037) |
| 0.6 | 2.9 | GO:0070131 | positive regulation of mitochondrial translation(GO:0070131) |
| 0.6 | 10.4 | GO:0071850 | mitotic cell cycle arrest(GO:0071850) |
| 0.6 | 4.4 | GO:0006600 | creatine metabolic process(GO:0006600) |
| 0.5 | 2.9 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.5 | 7.8 | GO:0034975 | protein folding in endoplasmic reticulum(GO:0034975) |
| 0.5 | 7.2 | GO:0006449 | regulation of translational termination(GO:0006449) |
| 0.4 | 1.3 | GO:0007518 | myoblast fate determination(GO:0007518) |
| 0.4 | 9.1 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.4 | 4.8 | GO:0043951 | negative regulation of cAMP-mediated signaling(GO:0043951) |
| 0.4 | 7.1 | GO:0089711 | L-glutamate transmembrane transport(GO:0089711) |
| 0.3 | 4.1 | GO:0051665 | membrane raft localization(GO:0051665) |
| 0.3 | 1.5 | GO:0021649 | vestibulocochlear nerve structural organization(GO:0021649) positive regulation of cytokine activity(GO:0060301) ganglion morphogenesis(GO:0061552) endothelial tip cell fate specification(GO:0097102) VEGF-activated neuropilin signaling pathway involved in axon guidance(GO:1902378) dorsal root ganglion morphogenesis(GO:1904835) otic placode development(GO:1905040) |
| 0.3 | 4.6 | GO:0016576 | histone dephosphorylation(GO:0016576) |
| 0.3 | 4.1 | GO:1904714 | regulation of chaperone-mediated autophagy(GO:1904714) |
| 0.3 | 2.9 | GO:0022417 | protein maturation by protein folding(GO:0022417) |
| 0.3 | 10.7 | GO:0043928 | exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 0.3 | 5.7 | GO:0036295 | cellular response to increased oxygen levels(GO:0036295) |
| 0.3 | 5.7 | GO:0035855 | megakaryocyte development(GO:0035855) |
| 0.2 | 0.7 | GO:0045082 | positive regulation of interleukin-10 biosynthetic process(GO:0045082) |
| 0.2 | 3.8 | GO:0031115 | negative regulation of microtubule polymerization(GO:0031115) |
| 0.2 | 2.2 | GO:0072383 | plus-end-directed vesicle transport along microtubule(GO:0072383) |
| 0.2 | 1.6 | GO:0002860 | positive regulation of natural killer cell mediated immune response to tumor cell(GO:0002857) positive regulation of natural killer cell mediated cytotoxicity directed against tumor cell target(GO:0002860) |
| 0.2 | 1.7 | GO:0060213 | regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060211) positive regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060213) |
| 0.2 | 7.2 | GO:0090140 | regulation of mitochondrial fission(GO:0090140) |
| 0.2 | 2.8 | GO:0006853 | carnitine shuttle(GO:0006853) |
| 0.2 | 5.9 | GO:0001893 | maternal placenta development(GO:0001893) |
| 0.2 | 1.1 | GO:0044351 | macropinocytosis(GO:0044351) |
| 0.2 | 4.4 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.1 | 3.0 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.1 | 1.3 | GO:0006682 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 0.1 | 1.4 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 0.1 | 2.7 | GO:2000114 | regulation of establishment of cell polarity(GO:2000114) |
| 0.1 | 5.4 | GO:0051894 | positive regulation of focal adhesion assembly(GO:0051894) |
| 0.1 | 13.9 | GO:0002479 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-dependent(GO:0002479) |
| 0.1 | 0.4 | GO:1903028 | positive regulation of opsonization(GO:1903028) |
| 0.1 | 1.8 | GO:1901642 | nucleoside transmembrane transport(GO:1901642) |
| 0.1 | 6.9 | GO:0018279 | protein N-linked glycosylation via asparagine(GO:0018279) |
| 0.1 | 8.0 | GO:0006457 | protein folding(GO:0006457) |
| 0.1 | 2.0 | GO:0032012 | regulation of ARF protein signal transduction(GO:0032012) |
| 0.1 | 1.3 | GO:0030913 | paranodal junction assembly(GO:0030913) |
| 0.1 | 2.3 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.1 | 4.0 | GO:0014003 | oligodendrocyte development(GO:0014003) |
| 0.1 | 3.6 | GO:0006656 | phosphatidylcholine biosynthetic process(GO:0006656) |
| 0.1 | 1.4 | GO:0061436 | establishment of skin barrier(GO:0061436) |
| 0.1 | 4.5 | GO:0043550 | regulation of lipid kinase activity(GO:0043550) |
| 0.1 | 7.0 | GO:0002223 | stimulatory C-type lectin receptor signaling pathway(GO:0002223) |
| 0.1 | 1.7 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 0.1 | 8.9 | GO:0006626 | protein targeting to mitochondrion(GO:0006626) |
| 0.1 | 1.6 | GO:0030728 | ovulation(GO:0030728) |
| 0.1 | 0.2 | GO:0035995 | detection of muscle stretch(GO:0035995) |
| 0.1 | 10.7 | GO:0006302 | double-strand break repair(GO:0006302) |
| 0.0 | 1.9 | GO:1901661 | quinone metabolic process(GO:1901661) |
| 0.0 | 15.7 | GO:0043687 | post-translational protein modification(GO:0043687) |
| 0.0 | 2.5 | GO:0031338 | regulation of vesicle fusion(GO:0031338) |
| 0.0 | 0.9 | GO:0048025 | negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.0 | 3.8 | GO:0008543 | fibroblast growth factor receptor signaling pathway(GO:0008543) |
| 0.0 | 5.4 | GO:0006986 | response to unfolded protein(GO:0006986) |
| 0.0 | 0.3 | GO:1900017 | positive regulation of cytokine production involved in inflammatory response(GO:1900017) |
| 0.0 | 2.0 | GO:0043066 | negative regulation of apoptotic process(GO:0043066) negative regulation of programmed cell death(GO:0043069) |
| 0.0 | 0.8 | GO:0006308 | DNA catabolic process(GO:0006308) |
| 0.0 | 14.3 | GO:0045087 | innate immune response(GO:0045087) |
| 0.0 | 3.3 | GO:0007050 | cell cycle arrest(GO:0007050) |
| 0.0 | 5.5 | GO:0002283 | neutrophil activation involved in immune response(GO:0002283) neutrophil degranulation(GO:0043312) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.0 | 30.0 | GO:0045272 | plasma membrane respiratory chain complex I(GO:0045272) |
| 4.1 | 16.3 | GO:0045283 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 3.0 | 36.3 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 2.4 | 77.6 | GO:0005753 | mitochondrial proton-transporting ATP synthase complex(GO:0005753) |
| 2.1 | 6.4 | GO:0005607 | laminin-2 complex(GO:0005607) |
| 2.0 | 22.5 | GO:0005750 | mitochondrial respiratory chain complex III(GO:0005750) respiratory chain complex III(GO:0045275) |
| 2.0 | 18.1 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 1.5 | 10.7 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 1.4 | 9.8 | GO:0031415 | NatA complex(GO:0031415) |
| 1.3 | 66.4 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 1.2 | 13.4 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 1.2 | 13.9 | GO:0019773 | proteasome core complex, alpha-subunit complex(GO:0019773) |
| 1.1 | 11.3 | GO:0045254 | pyruvate dehydrogenase complex(GO:0045254) |
| 1.1 | 13.1 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 1.0 | 8.3 | GO:0042382 | paraspeckles(GO:0042382) |
| 1.0 | 5.9 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 1.0 | 25.8 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.9 | 6.2 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.8 | 10.8 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.7 | 2.0 | GO:0005879 | axonemal microtubule(GO:0005879) |
| 0.7 | 4.6 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.6 | 4.5 | GO:0035032 | phosphatidylinositol 3-kinase complex, class III(GO:0035032) |
| 0.6 | 2.9 | GO:1990425 | ryanodine receptor complex(GO:1990425) |
| 0.6 | 6.9 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.5 | 11.7 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.5 | 5.9 | GO:0097025 | lateral loop(GO:0043219) MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.5 | 1.4 | GO:0005873 | plus-end kinesin complex(GO:0005873) |
| 0.4 | 5.7 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.4 | 5.2 | GO:0045239 | tricarboxylic acid cycle enzyme complex(GO:0045239) |
| 0.4 | 1.7 | GO:0071001 | U4/U6 snRNP(GO:0071001) |
| 0.4 | 4.1 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.4 | 5.7 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.3 | 6.6 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.3 | 19.2 | GO:0030964 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.3 | 4.0 | GO:0042405 | nuclear inclusion body(GO:0042405) |
| 0.3 | 10.4 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.2 | 13.9 | GO:0016235 | aggresome(GO:0016235) |
| 0.2 | 5.7 | GO:0098800 | inner mitochondrial membrane protein complex(GO:0098800) |
| 0.2 | 35.7 | GO:0000922 | spindle pole(GO:0000922) |
| 0.2 | 4.4 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.2 | 0.7 | GO:0033257 | Bcl3/NF-kappaB2 complex(GO:0033257) |
| 0.2 | 3.5 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.1 | 1.5 | GO:0002116 | semaphorin receptor complex(GO:0002116) sorting endosome(GO:0097443) |
| 0.1 | 16.9 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.1 | 2.8 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.1 | 7.8 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.1 | 3.2 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.1 | 2.5 | GO:0012505 | endomembrane system(GO:0012505) |
| 0.1 | 1.2 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.1 | 7.1 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.1 | 1.5 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.1 | 3.6 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.1 | 10.9 | GO:1904813 | ficolin-1-rich granule(GO:0101002) ficolin-1-rich granule lumen(GO:1904813) |
| 0.1 | 6.7 | GO:0035578 | azurophil granule lumen(GO:0035578) |
| 0.1 | 19.3 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.1 | 12.8 | GO:0005840 | ribosome(GO:0005840) |
| 0.1 | 0.9 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.1 | 4.1 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.1 | 1.2 | GO:0032982 | myosin filament(GO:0032982) |
| 0.1 | 17.9 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.0 | 3.3 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 1.3 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.0 | 1.4 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.0 | 6.0 | GO:0005741 | mitochondrial outer membrane(GO:0005741) organelle outer membrane(GO:0031968) |
| 0.0 | 3.7 | GO:0043235 | receptor complex(GO:0043235) |
| 0.0 | 3.9 | GO:0005770 | late endosome(GO:0005770) |
| 0.0 | 1.7 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 1.6 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.0 | 5.9 | GO:0016607 | nuclear speck(GO:0016607) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.4 | 16.3 | GO:0008177 | succinate dehydrogenase (ubiquinone) activity(GO:0008177) |
| 5.4 | 16.1 | GO:0045155 | electron transporter, transferring electrons from CoQH2-cytochrome c reductase complex and cytochrome c oxidase complex activity(GO:0045155) |
| 4.6 | 13.9 | GO:0036317 | tyrosyl-RNA phosphodiesterase activity(GO:0036317) 5'-tyrosyl-DNA phosphodiesterase activity(GO:0070260) |
| 3.9 | 42.6 | GO:0015266 | protein channel activity(GO:0015266) |
| 3.6 | 10.8 | GO:0015117 | thiosulfate transmembrane transporter activity(GO:0015117) oxaloacetate transmembrane transporter activity(GO:0015131) |
| 3.6 | 21.7 | GO:0023025 | MHC class Ib protein complex binding(GO:0023025) MHC class Ib protein binding, via antigen binding groove(GO:0023030) |
| 3.0 | 41.4 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 2.3 | 9.4 | GO:0032217 | riboflavin transporter activity(GO:0032217) |
| 2.3 | 9.2 | GO:0003985 | acetyl-CoA C-acetyltransferase activity(GO:0003985) |
| 2.1 | 6.4 | GO:0004756 | selenide, water dikinase activity(GO:0004756) phosphotransferase activity, paired acceptors(GO:0016781) |
| 2.1 | 6.4 | GO:0033867 | Fas-activated serine/threonine kinase activity(GO:0033867) |
| 1.5 | 30.0 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 1.4 | 11.5 | GO:0015288 | porin activity(GO:0015288) |
| 1.4 | 7.1 | GO:0005294 | neutral L-amino acid secondary active transmembrane transporter activity(GO:0005294) |
| 1.4 | 11.3 | GO:0004738 | pyruvate dehydrogenase activity(GO:0004738) pyruvate dehydrogenase [NAD(P)+] activity(GO:0034603) pyruvate dehydrogenase (NAD+) activity(GO:0034604) |
| 1.3 | 10.8 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 1.3 | 36.3 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 1.2 | 3.7 | GO:0031731 | CCR6 chemokine receptor binding(GO:0031731) |
| 1.2 | 7.2 | GO:0008079 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 1.1 | 6.3 | GO:0016681 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 1.0 | 5.9 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 1.0 | 3.9 | GO:0004449 | isocitrate dehydrogenase (NAD+) activity(GO:0004449) |
| 0.9 | 40.7 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.9 | 25.8 | GO:0035173 | histone kinase activity(GO:0035173) |
| 0.9 | 4.3 | GO:0052740 | 1-acyl-2-lysophosphatidylserine acylhydrolase activity(GO:0052740) |
| 0.8 | 11.7 | GO:0032041 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.8 | 9.8 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.7 | 7.8 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) |
| 0.7 | 6.9 | GO:0004579 | dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.7 | 2.7 | GO:0052851 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 0.7 | 12.4 | GO:0017070 | U6 snRNA binding(GO:0017070) |
| 0.6 | 4.4 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.5 | 5.4 | GO:0005375 | copper ion transmembrane transporter activity(GO:0005375) |
| 0.5 | 5.9 | GO:0097016 | L27 domain binding(GO:0097016) |
| 0.5 | 15.1 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.5 | 13.1 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.5 | 13.9 | GO:0004298 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.4 | 19.2 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.4 | 3.8 | GO:0004445 | inositol-polyphosphate 5-phosphatase activity(GO:0004445) phosphatidylinositol-3,4,5-trisphosphate 5-phosphatase activity(GO:0034485) |
| 0.3 | 1.4 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.3 | 5.9 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.3 | 1.9 | GO:0003960 | NADPH:quinone reductase activity(GO:0003960) |
| 0.3 | 2.3 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.3 | 2.8 | GO:0008607 | phosphorylase kinase regulator activity(GO:0008607) |
| 0.3 | 35.7 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.3 | 5.7 | GO:0044769 | ATPase activity, coupled to transmembrane movement of ions, rotational mechanism(GO:0044769) |
| 0.2 | 9.4 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.2 | 1.5 | GO:0038085 | vascular endothelial growth factor binding(GO:0038085) |
| 0.2 | 6.2 | GO:0003951 | NAD+ kinase activity(GO:0003951) |
| 0.2 | 2.9 | GO:0019855 | calcium channel inhibitor activity(GO:0019855) type I transforming growth factor beta receptor binding(GO:0034713) |
| 0.2 | 4.1 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.2 | 12.3 | GO:0016836 | hydro-lyase activity(GO:0016836) |
| 0.2 | 30.7 | GO:0047485 | protein N-terminus binding(GO:0047485) |
| 0.2 | 1.3 | GO:0001733 | galactosylceramide sulfotransferase activity(GO:0001733) galactose 3-O-sulfotransferase activity(GO:0050694) |
| 0.2 | 9.1 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.2 | 4.4 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.1 | 4.6 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.1 | 3.0 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.1 | 6.7 | GO:0004601 | peroxidase activity(GO:0004601) |
| 0.1 | 12.7 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.1 | 0.4 | GO:0004530 | deoxyribonuclease I activity(GO:0004530) |
| 0.1 | 6.6 | GO:0070063 | RNA polymerase binding(GO:0070063) |
| 0.1 | 0.9 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.1 | 1.8 | GO:0005337 | nucleoside transmembrane transporter activity(GO:0005337) |
| 0.1 | 2.0 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.1 | 0.9 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.1 | 19.2 | GO:0044822 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.1 | 1.1 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.1 | 6.4 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 4.4 | GO:0032947 | protein complex scaffold(GO:0032947) |
| 0.0 | 2.7 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 3.7 | GO:0004714 | transmembrane receptor protein tyrosine kinase activity(GO:0004714) |
| 0.0 | 5.7 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.0 | 1.4 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.0 | 4.7 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 2.1 | GO:0017022 | myosin binding(GO:0017022) |
| 0.0 | 1.2 | GO:0000146 | microfilament motor activity(GO:0000146) |
| 0.0 | 1.7 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 4.9 | GO:0061630 | ubiquitin protein ligase activity(GO:0061630) |
| 0.0 | 2.5 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.0 | 23.2 | GO:0003723 | RNA binding(GO:0003723) |
| 0.0 | 1.2 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.0 | 0.2 | GO:0016884 | carbon-nitrogen ligase activity, with glutamine as amido-N-donor(GO:0016884) |
| 0.0 | 1.5 | GO:0005178 | integrin binding(GO:0005178) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.6 | 35.7 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.8 | 11.7 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.6 | 48.8 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.3 | 10.4 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.3 | 13.9 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.3 | 36.4 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.3 | 6.4 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.2 | 11.5 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.2 | 2.9 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.2 | 6.7 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 0.2 | 14.3 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.1 | 7.0 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.1 | 3.5 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.1 | 4.1 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.1 | 1.5 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.1 | 0.7 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.1 | 4.6 | PID ERBB1 DOWNSTREAM PATHWAY | ErbB1 downstream signaling |
| 0.0 | 1.1 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.0 | 1.3 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.0 | 1.2 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 1.9 | PID MTOR 4PATHWAY | mTOR signaling pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 79.2 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 1.4 | 32.5 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 1.3 | 35.7 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.8 | 64.3 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.7 | 11.3 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.7 | 9.1 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.5 | 11.7 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.5 | 9.2 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.5 | 13.1 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.4 | 4.5 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.4 | 6.6 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.4 | 9.4 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.3 | 5.9 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.3 | 10.8 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.3 | 8.4 | REACTOME TRANSFERRIN ENDOCYTOSIS AND RECYCLING | Genes involved in Transferrin endocytosis and recycling |
| 0.3 | 5.9 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.2 | 5.4 | REACTOME METAL ION SLC TRANSPORTERS | Genes involved in Metal ion SLC transporters |
| 0.2 | 43.2 | REACTOME INFLUENZA VIRAL RNA TRANSCRIPTION AND REPLICATION | Genes involved in Influenza Viral RNA Transcription and Replication |
| 0.2 | 13.9 | REACTOME CDK MEDIATED PHOSPHORYLATION AND REMOVAL OF CDC6 | Genes involved in CDK-mediated phosphorylation and removal of Cdc6 |
| 0.2 | 5.4 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.2 | 6.9 | REACTOME SRP DEPENDENT COTRANSLATIONAL PROTEIN TARGETING TO MEMBRANE | Genes involved in SRP-dependent cotranslational protein targeting to membrane |
| 0.2 | 6.4 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.2 | 5.1 | REACTOME P38MAPK EVENTS | Genes involved in p38MAPK events |
| 0.2 | 7.1 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.2 | 1.7 | REACTOME REGULATORY RNA PATHWAYS | Genes involved in Regulatory RNA pathways |
| 0.2 | 6.2 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 0.1 | 2.9 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.1 | 6.1 | REACTOME NONSENSE MEDIATED DECAY ENHANCED BY THE EXON JUNCTION COMPLEX | Genes involved in Nonsense Mediated Decay Enhanced by the Exon Junction Complex |
| 0.1 | 1.5 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.1 | 2.8 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.1 | 15.6 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.1 | 8.7 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.1 | 3.8 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.1 | 9.2 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.1 | 2.0 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.1 | 1.4 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.1 | 3.7 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.1 | 2.5 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.1 | 1.4 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.0 | 1.8 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 4.7 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |
| 0.0 | 1.4 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.0 | 1.2 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.0 | 2.9 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.0 | 1.3 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.0 | 1.5 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.4 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.0 | 3.5 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 3.1 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |