Project
GNF SymAtlas + NCI-60 cancer cell lines, comparison of cancers vs non-cancers, human (Su, 2004; Ross, 2000)
Navigation
Downloads
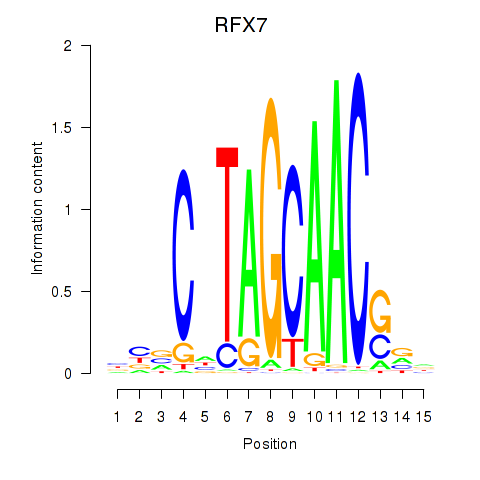
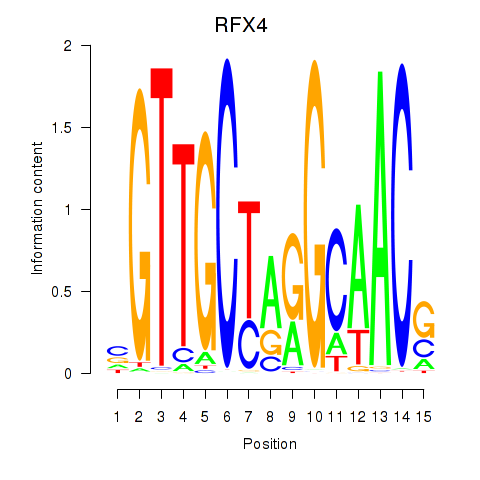
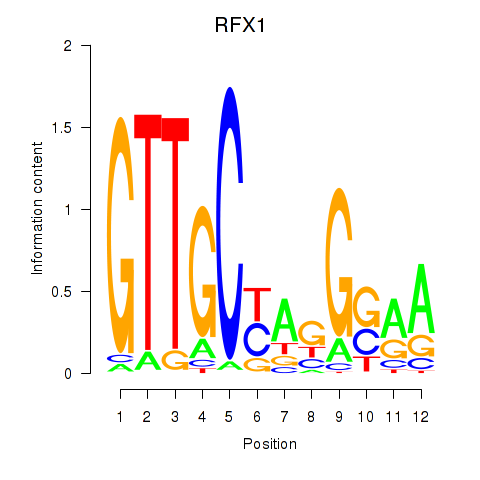
Results for RFX7_RFX4_RFX1
Z-value: 1.28
Transcription factors associated with RFX7_RFX4_RFX1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
RFX7
|
ENSG00000181827.10 | regulatory factor X7 |
|
RFX4
|
ENSG00000111783.8 | regulatory factor X4 |
|
RFX1
|
ENSG00000132005.4 | regulatory factor X1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| RFX1 | hg19_v2_chr19_-_14117074_14117141 | 0.35 | 1.0e-07 | Click! |
| RFX7 | hg19_v2_chr15_-_56535464_56535521 | 0.25 | 2.3e-04 | Click! |
Activity profile of RFX7_RFX4_RFX1 motif
Sorted Z-values of RFX7_RFX4_RFX1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of RFX7_RFX4_RFX1
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 50.8 | 203.2 | GO:0090299 | fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:0060825) regulation of neural crest formation(GO:0090299) negative regulation of neural crest formation(GO:0090301) regulation of fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:2000313) negative regulation of fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:2000314) |
| 12.0 | 84.2 | GO:0060294 | cilium movement involved in cell motility(GO:0060294) |
| 10.4 | 31.3 | GO:2000777 | positive regulation of oocyte maturation(GO:1900195) negative regulation of NLRP3 inflammasome complex assembly(GO:1900226) positive regulation of proteasomal ubiquitin-dependent protein catabolic process involved in cellular response to hypoxia(GO:2000777) |
| 9.8 | 48.8 | GO:0070829 | response to vitamin B2(GO:0033274) heterochromatin maintenance(GO:0070829) |
| 9.3 | 27.9 | GO:0090521 | glomerular visceral epithelial cell migration(GO:0090521) |
| 8.9 | 26.8 | GO:0018307 | enzyme active site formation(GO:0018307) tRNA thio-modification(GO:0034227) |
| 8.6 | 25.8 | GO:1990168 | protein K33-linked deubiquitination(GO:1990168) |
| 8.4 | 25.3 | GO:0071988 | regulation of spindle elongation(GO:0032887) regulation of mitotic spindle elongation(GO:0032888) anastral spindle assembly(GO:0055048) protein localization to spindle pole body(GO:0071988) regulation of protein localization to spindle pole body(GO:1902363) positive regulation of protein localization to spindle pole body(GO:1902365) positive regulation of mitotic spindle elongation(GO:1902846) |
| 8.1 | 40.4 | GO:0035617 | stress granule disassembly(GO:0035617) |
| 7.2 | 50.7 | GO:0001539 | cilium or flagellum-dependent cell motility(GO:0001539) |
| 6.1 | 24.3 | GO:0060613 | fat pad development(GO:0060613) |
| 6.1 | 18.2 | GO:0050760 | negative regulation of thymidylate synthase biosynthetic process(GO:0050760) |
| 5.6 | 39.4 | GO:0034316 | negative regulation of Arp2/3 complex-mediated actin nucleation(GO:0034316) |
| 5.6 | 22.4 | GO:1904491 | protein localization to ciliary transition zone(GO:1904491) |
| 5.4 | 32.7 | GO:2001181 | positive regulation of interleukin-10 secretion(GO:2001181) |
| 4.5 | 22.4 | GO:2000638 | regulation of SREBP signaling pathway(GO:2000638) negative regulation of SREBP signaling pathway(GO:2000639) |
| 4.2 | 12.7 | GO:0098937 | dendritic transport(GO:0098935) anterograde dendritic transport(GO:0098937) |
| 4.1 | 49.0 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 3.6 | 21.4 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 3.5 | 17.4 | GO:2001106 | regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001106) |
| 3.3 | 10.0 | GO:0006106 | fumarate metabolic process(GO:0006106) aspartate catabolic process(GO:0006533) |
| 3.3 | 19.9 | GO:0051013 | microtubule severing(GO:0051013) |
| 3.3 | 9.9 | GO:0035610 | protein side chain deglutamylation(GO:0035610) |
| 3.3 | 19.8 | GO:0033133 | fructose 2,6-bisphosphate metabolic process(GO:0006003) positive regulation of glucokinase activity(GO:0033133) positive regulation of hexokinase activity(GO:1903301) |
| 3.1 | 55.6 | GO:0019614 | catechol-containing compound catabolic process(GO:0019614) catecholamine catabolic process(GO:0042424) |
| 3.0 | 8.9 | GO:0001694 | histamine biosynthetic process(GO:0001694) |
| 3.0 | 8.9 | GO:1903031 | regulation of microtubule plus-end binding(GO:1903031) positive regulation of microtubule plus-end binding(GO:1903033) |
| 2.8 | 24.9 | GO:0003351 | epithelial cilium movement(GO:0003351) |
| 2.7 | 8.0 | GO:1905154 | negative regulation of membrane invagination(GO:1905154) |
| 2.4 | 109.7 | GO:0035735 | intraciliary transport involved in cilium morphogenesis(GO:0035735) |
| 2.4 | 7.3 | GO:1904395 | retinal rod cell differentiation(GO:0060221) positive regulation of skeletal muscle acetylcholine-gated channel clustering(GO:1904395) negative regulation of neuromuscular junction development(GO:1904397) |
| 2.4 | 4.7 | GO:0070286 | axonemal dynein complex assembly(GO:0070286) |
| 2.3 | 16.4 | GO:2000286 | receptor internalization involved in canonical Wnt signaling pathway(GO:2000286) |
| 2.3 | 4.6 | GO:1900425 | negative regulation of defense response to bacterium(GO:1900425) |
| 2.3 | 6.8 | GO:0050902 | leukocyte adhesive activation(GO:0050902) |
| 2.3 | 11.3 | GO:0061088 | regulation of sequestering of zinc ion(GO:0061088) |
| 2.2 | 8.7 | GO:0051754 | meiotic sister chromatid cohesion, centromeric(GO:0051754) |
| 2.1 | 14.9 | GO:0042791 | 5S class rRNA transcription from RNA polymerase III type 1 promoter(GO:0042791) tRNA transcription from RNA polymerase III promoter(GO:0042797) |
| 2.1 | 19.2 | GO:1902412 | regulation of mitotic cytokinesis(GO:1902412) |
| 2.1 | 10.4 | GO:1903564 | regulation of protein localization to cilium(GO:1903564) |
| 1.9 | 24.6 | GO:1904896 | ESCRT complex disassembly(GO:1904896) ESCRT III complex disassembly(GO:1904903) |
| 1.9 | 9.4 | GO:0035845 | photoreceptor cell outer segment organization(GO:0035845) |
| 1.9 | 5.6 | GO:1905224 | clathrin-coated pit assembly(GO:1905224) |
| 1.7 | 18.8 | GO:0048102 | autophagic cell death(GO:0048102) |
| 1.7 | 48.9 | GO:0006884 | cell volume homeostasis(GO:0006884) |
| 1.6 | 9.6 | GO:1903750 | regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:1903750) negative regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:1903751) |
| 1.6 | 4.8 | GO:0060370 | susceptibility to T cell mediated cytotoxicity(GO:0060370) |
| 1.6 | 4.7 | GO:0035359 | negative regulation of peroxisome proliferator activated receptor signaling pathway(GO:0035359) |
| 1.5 | 15.3 | GO:0008343 | adult feeding behavior(GO:0008343) |
| 1.5 | 11.9 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 1.5 | 14.8 | GO:0051612 | negative regulation of neurotransmitter uptake(GO:0051581) serotonin uptake(GO:0051610) regulation of serotonin uptake(GO:0051611) negative regulation of serotonin uptake(GO:0051612) |
| 1.4 | 8.6 | GO:0098535 | de novo centriole assembly(GO:0098535) |
| 1.4 | 4.3 | GO:0002581 | negative regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002581) |
| 1.4 | 16.8 | GO:0030321 | transepithelial chloride transport(GO:0030321) |
| 1.4 | 12.6 | GO:0002318 | myeloid progenitor cell differentiation(GO:0002318) |
| 1.3 | 6.7 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) |
| 1.3 | 14.6 | GO:0022008 | neurogenesis(GO:0022008) |
| 1.3 | 16.9 | GO:1904714 | regulation of chaperone-mediated autophagy(GO:1904714) |
| 1.3 | 30.6 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 1.3 | 25.2 | GO:0032012 | regulation of ARF protein signal transduction(GO:0032012) |
| 1.2 | 9.7 | GO:0090043 | regulation of tubulin deacetylation(GO:0090043) |
| 1.2 | 2.4 | GO:0061055 | myotome development(GO:0061055) |
| 1.2 | 2.4 | GO:2001190 | positive regulation of T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:2001190) |
| 1.2 | 2.4 | GO:0010637 | negative regulation of mitochondrial fusion(GO:0010637) |
| 1.2 | 4.7 | GO:0035625 | regulation of epinephrine secretion(GO:0014060) negative regulation of epinephrine secretion(GO:0032811) epidermal growth factor-activated receptor transactivation by G-protein coupled receptor signaling pathway(GO:0035625) epinephrine secretion(GO:0048242) |
| 1.1 | 17.1 | GO:0099514 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 1.1 | 4.5 | GO:1904450 | negative regulation of gamma-aminobutyric acid secretion(GO:0014053) aspartate secretion(GO:0061528) regulation of aspartate secretion(GO:1904448) positive regulation of aspartate secretion(GO:1904450) |
| 1.1 | 10.1 | GO:0010499 | proteasomal ubiquitin-independent protein catabolic process(GO:0010499) |
| 1.1 | 8.9 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 1.1 | 52.5 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 1.1 | 6.5 | GO:0010615 | positive regulation of cardiac muscle adaptation(GO:0010615) positive regulation of cardiac muscle hypertrophy in response to stress(GO:1903244) |
| 1.1 | 3.2 | GO:0061074 | regulation of neural retina development(GO:0061074) regulation of retina development in camera-type eye(GO:1902866) |
| 1.1 | 6.4 | GO:0034983 | peptidyl-lysine deacetylation(GO:0034983) |
| 1.0 | 2.1 | GO:0061511 | centriole elongation(GO:0061511) |
| 1.0 | 8.3 | GO:0009957 | epidermal cell fate specification(GO:0009957) |
| 1.0 | 8.2 | GO:0006659 | phosphatidylserine biosynthetic process(GO:0006659) |
| 1.0 | 15.3 | GO:0034315 | regulation of Arp2/3 complex-mediated actin nucleation(GO:0034315) |
| 0.9 | 8.5 | GO:0030916 | otic vesicle formation(GO:0030916) |
| 0.9 | 13.8 | GO:0002244 | hematopoietic progenitor cell differentiation(GO:0002244) |
| 0.9 | 3.6 | GO:0048511 | rhythmic process(GO:0048511) |
| 0.9 | 7.2 | GO:0007506 | gonadal mesoderm development(GO:0007506) |
| 0.9 | 15.2 | GO:0019374 | galactosylceramide metabolic process(GO:0006681) galactolipid metabolic process(GO:0019374) |
| 0.9 | 4.4 | GO:0051533 | positive regulation of NFAT protein import into nucleus(GO:0051533) |
| 0.9 | 12.9 | GO:0010839 | negative regulation of keratinocyte proliferation(GO:0010839) |
| 0.9 | 8.5 | GO:0016127 | cholesterol catabolic process(GO:0006707) sterol catabolic process(GO:0016127) |
| 0.8 | 5.8 | GO:1900113 | negative regulation of histone H3-K9 trimethylation(GO:1900113) |
| 0.8 | 2.4 | GO:0097359 | UDP-glucosylation(GO:0097359) |
| 0.8 | 5.5 | GO:0016080 | synaptic vesicle targeting(GO:0016080) |
| 0.8 | 8.6 | GO:0098789 | pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.8 | 8.4 | GO:0015886 | heme transport(GO:0015886) |
| 0.8 | 1.5 | GO:2000969 | positive regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000969) |
| 0.8 | 4.5 | GO:0032510 | endosome to lysosome transport via multivesicular body sorting pathway(GO:0032510) |
| 0.7 | 8.9 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.7 | 2.2 | GO:1900220 | semaphorin-plexin signaling pathway involved in bone trabecula morphogenesis(GO:1900220) |
| 0.7 | 4.3 | GO:0035093 | spermatogenesis, exchange of chromosomal proteins(GO:0035093) |
| 0.7 | 12.8 | GO:2001241 | positive regulation of extrinsic apoptotic signaling pathway in absence of ligand(GO:2001241) |
| 0.7 | 25.9 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 0.7 | 2.7 | GO:1903644 | regulation of chaperone-mediated protein folding(GO:1903644) |
| 0.7 | 8.1 | GO:0008037 | binding of sperm to zona pellucida(GO:0007339) cell recognition(GO:0008037) cell-cell recognition(GO:0009988) sperm-egg recognition(GO:0035036) |
| 0.7 | 21.7 | GO:0008045 | motor neuron axon guidance(GO:0008045) |
| 0.7 | 9.1 | GO:0002021 | response to dietary excess(GO:0002021) |
| 0.6 | 3.9 | GO:0006287 | base-excision repair, gap-filling(GO:0006287) |
| 0.6 | 6.4 | GO:1900227 | positive regulation of NLRP3 inflammasome complex assembly(GO:1900227) |
| 0.6 | 1.3 | GO:0045144 | meiotic sister chromatid segregation(GO:0045144) |
| 0.6 | 1.9 | GO:0007288 | sperm axoneme assembly(GO:0007288) |
| 0.6 | 3.1 | GO:2001288 | positive regulation of caveolin-mediated endocytosis(GO:2001288) |
| 0.6 | 27.4 | GO:0060612 | adipose tissue development(GO:0060612) |
| 0.6 | 7.4 | GO:0043653 | mitochondrial fragmentation involved in apoptotic process(GO:0043653) |
| 0.6 | 21.3 | GO:2000300 | regulation of synaptic vesicle exocytosis(GO:2000300) |
| 0.6 | 2.4 | GO:0050882 | voluntary musculoskeletal movement(GO:0050882) |
| 0.6 | 8.7 | GO:0007196 | adenylate cyclase-inhibiting G-protein coupled glutamate receptor signaling pathway(GO:0007196) |
| 0.6 | 4.5 | GO:1903546 | protein localization to photoreceptor outer segment(GO:1903546) |
| 0.6 | 8.8 | GO:0045475 | locomotor rhythm(GO:0045475) |
| 0.5 | 3.3 | GO:0061737 | leukotriene signaling pathway(GO:0061737) |
| 0.5 | 6.5 | GO:0070070 | proton-transporting V-type ATPase complex assembly(GO:0070070) vacuolar proton-transporting V-type ATPase complex assembly(GO:0070072) |
| 0.5 | 2.6 | GO:1905232 | cellular response to L-glutamate(GO:1905232) |
| 0.5 | 15.1 | GO:0097120 | receptor localization to synapse(GO:0097120) |
| 0.5 | 4.6 | GO:0051026 | chiasma assembly(GO:0051026) |
| 0.5 | 2.5 | GO:0060075 | regulation of resting membrane potential(GO:0060075) |
| 0.5 | 22.1 | GO:0016579 | protein deubiquitination(GO:0016579) |
| 0.5 | 9.2 | GO:1901663 | ubiquinone biosynthetic process(GO:0006744) quinone biosynthetic process(GO:1901663) |
| 0.5 | 1.9 | GO:0043376 | regulation of CD8-positive, alpha-beta T cell differentiation(GO:0043376) |
| 0.5 | 11.2 | GO:0045662 | negative regulation of myoblast differentiation(GO:0045662) |
| 0.5 | 1.9 | GO:0042357 | thiamine diphosphate metabolic process(GO:0042357) |
| 0.5 | 21.9 | GO:0043268 | positive regulation of potassium ion transport(GO:0043268) |
| 0.4 | 17.0 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.4 | 1.3 | GO:0090187 | positive regulation of pancreatic juice secretion(GO:0090187) |
| 0.4 | 1.3 | GO:0044314 | protein K27-linked ubiquitination(GO:0044314) |
| 0.4 | 2.1 | GO:0022605 | ovarian cumulus expansion(GO:0001550) oogenesis stage(GO:0022605) fused antrum stage(GO:0048165) |
| 0.4 | 3.2 | GO:0070986 | left/right axis specification(GO:0070986) |
| 0.4 | 4.0 | GO:0015871 | choline transport(GO:0015871) |
| 0.4 | 7.7 | GO:0071420 | cellular response to histamine(GO:0071420) |
| 0.4 | 8.5 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.4 | 7.3 | GO:0042340 | keratan sulfate catabolic process(GO:0042340) |
| 0.4 | 0.8 | GO:0006288 | base-excision repair, DNA ligation(GO:0006288) |
| 0.3 | 1.0 | GO:0044805 | late nucleophagy(GO:0044805) |
| 0.3 | 32.3 | GO:0007286 | spermatid development(GO:0007286) |
| 0.3 | 16.6 | GO:0071377 | cellular response to glucagon stimulus(GO:0071377) |
| 0.3 | 2.7 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.3 | 6.3 | GO:0097484 | dendrite extension(GO:0097484) |
| 0.3 | 9.3 | GO:0015695 | organic cation transport(GO:0015695) |
| 0.3 | 1.2 | GO:0008626 | granzyme-mediated apoptotic signaling pathway(GO:0008626) |
| 0.3 | 1.7 | GO:0031022 | nuclear migration along microfilament(GO:0031022) |
| 0.3 | 1.9 | GO:0034287 | detection of carbohydrate stimulus(GO:0009730) detection of hexose stimulus(GO:0009732) detection of monosaccharide stimulus(GO:0034287) detection of glucose(GO:0051594) |
| 0.3 | 11.4 | GO:0042059 | negative regulation of epidermal growth factor receptor signaling pathway(GO:0042059) |
| 0.2 | 7.4 | GO:0008214 | protein demethylation(GO:0006482) protein dealkylation(GO:0008214) |
| 0.2 | 3.8 | GO:0006004 | fucose metabolic process(GO:0006004) |
| 0.2 | 14.6 | GO:0048384 | retinoic acid receptor signaling pathway(GO:0048384) |
| 0.2 | 1.1 | GO:1901525 | negative regulation of macromitophagy(GO:1901525) |
| 0.2 | 3.3 | GO:0002638 | negative regulation of immunoglobulin production(GO:0002638) |
| 0.2 | 4.4 | GO:0034724 | DNA replication-independent nucleosome assembly(GO:0006336) DNA replication-independent nucleosome organization(GO:0034724) |
| 0.2 | 8.3 | GO:0035418 | protein localization to synapse(GO:0035418) |
| 0.2 | 4.1 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.2 | 2.1 | GO:0015889 | cobalamin transport(GO:0015889) |
| 0.2 | 3.7 | GO:0071625 | vocalization behavior(GO:0071625) |
| 0.2 | 12.5 | GO:0045839 | negative regulation of mitotic nuclear division(GO:0045839) |
| 0.2 | 8.4 | GO:0046949 | fatty-acyl-CoA biosynthetic process(GO:0046949) |
| 0.2 | 5.4 | GO:0030207 | chondroitin sulfate catabolic process(GO:0030207) |
| 0.2 | 3.2 | GO:0007501 | mesodermal cell fate specification(GO:0007501) |
| 0.2 | 6.3 | GO:0030317 | sperm motility(GO:0030317) |
| 0.2 | 11.0 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.2 | 11.1 | GO:0019933 | cAMP-mediated signaling(GO:0019933) |
| 0.2 | 7.6 | GO:1901379 | regulation of potassium ion transmembrane transport(GO:1901379) |
| 0.2 | 4.0 | GO:0036498 | IRE1-mediated unfolded protein response(GO:0036498) |
| 0.2 | 5.8 | GO:0045898 | regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045898) |
| 0.2 | 8.4 | GO:0031338 | regulation of vesicle fusion(GO:0031338) |
| 0.2 | 31.5 | GO:0060271 | cilium morphogenesis(GO:0060271) |
| 0.2 | 1.8 | GO:0021692 | cerebellar Purkinje cell layer morphogenesis(GO:0021692) |
| 0.2 | 6.3 | GO:0043666 | regulation of phosphoprotein phosphatase activity(GO:0043666) |
| 0.2 | 0.7 | GO:1902715 | positive regulation of interferon-gamma secretion(GO:1902715) |
| 0.2 | 4.1 | GO:0036152 | phosphatidylethanolamine acyl-chain remodeling(GO:0036152) |
| 0.2 | 0.8 | GO:0030205 | dermatan sulfate metabolic process(GO:0030205) dermatan sulfate biosynthetic process(GO:0030208) |
| 0.2 | 6.1 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.2 | 3.7 | GO:0010669 | epithelial structure maintenance(GO:0010669) |
| 0.2 | 6.3 | GO:0086010 | membrane depolarization during action potential(GO:0086010) |
| 0.2 | 5.4 | GO:0050855 | regulation of B cell receptor signaling pathway(GO:0050855) |
| 0.2 | 8.3 | GO:2000649 | regulation of sodium ion transmembrane transporter activity(GO:2000649) |
| 0.2 | 1.2 | GO:0030968 | endoplasmic reticulum unfolded protein response(GO:0030968) |
| 0.1 | 9.9 | GO:0006501 | C-terminal protein lipidation(GO:0006501) |
| 0.1 | 15.5 | GO:0051897 | positive regulation of protein kinase B signaling(GO:0051897) |
| 0.1 | 5.9 | GO:0030513 | positive regulation of BMP signaling pathway(GO:0030513) |
| 0.1 | 12.2 | GO:0038083 | peptidyl-tyrosine autophosphorylation(GO:0038083) |
| 0.1 | 5.3 | GO:1904031 | positive regulation of cyclin-dependent protein kinase activity(GO:1904031) |
| 0.1 | 8.4 | GO:0009060 | aerobic respiration(GO:0009060) |
| 0.1 | 1.1 | GO:1904263 | positive regulation of TORC1 signaling(GO:1904263) |
| 0.1 | 0.9 | GO:0071372 | response to light intensity(GO:0009642) cellular response to follicle-stimulating hormone stimulus(GO:0071372) |
| 0.1 | 1.2 | GO:0015074 | DNA integration(GO:0015074) |
| 0.1 | 5.1 | GO:0016266 | O-glycan processing(GO:0016266) |
| 0.1 | 8.1 | GO:1903955 | positive regulation of protein targeting to mitochondrion(GO:1903955) |
| 0.1 | 3.3 | GO:0002437 | inflammatory response to antigenic stimulus(GO:0002437) |
| 0.1 | 0.6 | GO:0060154 | cellular process regulating host cell cycle in response to virus(GO:0060154) |
| 0.1 | 2.9 | GO:0060996 | dendritic spine development(GO:0060996) |
| 0.1 | 0.4 | GO:0010793 | regulation of mRNA export from nucleus(GO:0010793) |
| 0.1 | 3.7 | GO:0031572 | G2 DNA damage checkpoint(GO:0031572) |
| 0.1 | 1.0 | GO:0061051 | positive regulation of cell growth involved in cardiac muscle cell development(GO:0061051) |
| 0.1 | 4.2 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.1 | 0.6 | GO:0023021 | termination of signal transduction(GO:0023021) |
| 0.1 | 1.7 | GO:0031468 | nuclear envelope reassembly(GO:0031468) |
| 0.1 | 2.8 | GO:0090630 | activation of GTPase activity(GO:0090630) |
| 0.1 | 1.1 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 0.1 | 0.7 | GO:0010172 | embryonic body morphogenesis(GO:0010172) |
| 0.1 | 0.9 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 0.1 | 0.2 | GO:0090219 | negative regulation of lipid kinase activity(GO:0090219) |
| 0.1 | 2.1 | GO:0009083 | branched-chain amino acid catabolic process(GO:0009083) |
| 0.1 | 1.8 | GO:0045026 | plasma membrane fusion(GO:0045026) |
| 0.1 | 1.1 | GO:0007635 | chemosensory behavior(GO:0007635) |
| 0.1 | 4.5 | GO:0000245 | spliceosomal complex assembly(GO:0000245) |
| 0.1 | 1.1 | GO:0001573 | ganglioside metabolic process(GO:0001573) |
| 0.1 | 1.5 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.1 | 0.5 | GO:0010727 | negative regulation of hydrogen peroxide metabolic process(GO:0010727) |
| 0.1 | 1.2 | GO:0006359 | regulation of transcription from RNA polymerase III promoter(GO:0006359) |
| 0.1 | 2.2 | GO:0001961 | positive regulation of cytokine-mediated signaling pathway(GO:0001961) |
| 0.1 | 0.2 | GO:0016999 | antibiotic metabolic process(GO:0016999) cellular amide catabolic process(GO:0043605) |
| 0.1 | 7.2 | GO:0008277 | regulation of G-protein coupled receptor protein signaling pathway(GO:0008277) |
| 0.1 | 1.9 | GO:0007274 | neuromuscular synaptic transmission(GO:0007274) |
| 0.0 | 15.6 | GO:0098916 | chemical synaptic transmission(GO:0007268) anterograde trans-synaptic signaling(GO:0098916) |
| 0.0 | 0.8 | GO:0021522 | spinal cord motor neuron differentiation(GO:0021522) |
| 0.0 | 0.3 | GO:1990592 | protein ufmylation(GO:0071569) protein polyufmylation(GO:1990564) protein K69-linked ufmylation(GO:1990592) |
| 0.0 | 0.9 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 0.0 | 4.6 | GO:0021915 | neural tube development(GO:0021915) |
| 0.0 | 0.7 | GO:0003094 | glomerular filtration(GO:0003094) |
| 0.0 | 5.4 | GO:0006937 | regulation of muscle contraction(GO:0006937) |
| 0.0 | 2.4 | GO:1900181 | negative regulation of protein localization to nucleus(GO:1900181) |
| 0.0 | 0.7 | GO:0043488 | regulation of mRNA stability(GO:0043488) |
| 0.0 | 0.1 | GO:0051958 | methotrexate transport(GO:0051958) reduced folate transmembrane transport(GO:0098838) |
| 0.0 | 0.9 | GO:0032620 | interleukin-17 production(GO:0032620) |
| 0.0 | 2.0 | GO:0071349 | interleukin-12-mediated signaling pathway(GO:0035722) response to interleukin-12(GO:0070671) cellular response to interleukin-12(GO:0071349) |
| 0.0 | 0.1 | GO:0035814 | negative regulation of urine volume(GO:0035811) negative regulation of renal sodium excretion(GO:0035814) |
| 0.0 | 1.5 | GO:1904837 | beta-catenin-TCF complex assembly(GO:1904837) |
| 0.0 | 3.9 | GO:0051092 | positive regulation of NF-kappaB transcription factor activity(GO:0051092) |
| 0.0 | 1.2 | GO:0014047 | glutamate secretion(GO:0014047) |
| 0.0 | 21.6 | GO:0007417 | central nervous system development(GO:0007417) |
| 0.0 | 1.0 | GO:0042552 | myelination(GO:0042552) |
| 0.0 | 1.6 | GO:0060041 | retina development in camera-type eye(GO:0060041) |
| 0.0 | 5.7 | GO:0045860 | positive regulation of protein kinase activity(GO:0045860) |
| 0.0 | 0.1 | GO:0042953 | lipoprotein transport(GO:0042953) lipoprotein localization(GO:0044872) |
| 0.0 | 0.2 | GO:1901661 | quinone metabolic process(GO:1901661) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 8.4 | 25.3 | GO:0061673 | cortical microtubule(GO:0055028) mitotic spindle astral microtubule(GO:0061673) |
| 7.1 | 99.7 | GO:0005858 | axonemal dynein complex(GO:0005858) |
| 4.9 | 29.2 | GO:0044447 | axoneme part(GO:0044447) |
| 4.1 | 32.7 | GO:0045179 | apical cortex(GO:0045179) |
| 3.9 | 15.4 | GO:0016939 | kinesin II complex(GO:0016939) |
| 3.8 | 19.2 | GO:0005818 | astral microtubule(GO:0000235) aster(GO:0005818) |
| 3.8 | 45.3 | GO:0033010 | paranodal junction(GO:0033010) |
| 3.2 | 22.4 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
| 3.1 | 18.8 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 3.1 | 18.5 | GO:0002177 | manchette(GO:0002177) |
| 2.9 | 63.5 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 2.9 | 8.6 | GO:0098536 | deuterosome(GO:0098536) |
| 2.5 | 7.4 | GO:0044609 | DBIRD complex(GO:0044609) |
| 2.3 | 32.9 | GO:0036038 | MKS complex(GO:0036038) |
| 2.2 | 4.4 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 2.2 | 6.5 | GO:0002199 | zona pellucida receptor complex(GO:0002199) |
| 2.1 | 14.9 | GO:0000127 | transcription factor TFIIIC complex(GO:0000127) |
| 2.1 | 14.5 | GO:0001520 | outer dense fiber(GO:0001520) |
| 2.1 | 39.1 | GO:0031045 | dense core granule(GO:0031045) |
| 2.0 | 18.2 | GO:1990812 | growth cone lamellipodium(GO:1990761) growth cone filopodium(GO:1990812) |
| 1.9 | 23.1 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 1.8 | 140.5 | GO:0036126 | sperm flagellum(GO:0036126) |
| 1.5 | 36.1 | GO:0034451 | centriolar satellite(GO:0034451) |
| 1.5 | 37.9 | GO:0032391 | photoreceptor connecting cilium(GO:0032391) |
| 1.2 | 65.5 | GO:0016235 | aggresome(GO:0016235) |
| 1.1 | 4.5 | GO:0032144 | 4-aminobutyrate transaminase complex(GO:0032144) |
| 1.0 | 5.1 | GO:0070847 | core mediator complex(GO:0070847) |
| 1.0 | 11.9 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 1.0 | 14.4 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.9 | 6.4 | GO:0097427 | microtubule bundle(GO:0097427) |
| 0.8 | 5.8 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 0.8 | 9.0 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.7 | 6.5 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.7 | 9.8 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.7 | 8.2 | GO:0043190 | ATP-binding cassette (ABC) transporter complex(GO:0043190) |
| 0.6 | 18.6 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.6 | 24.9 | GO:0036064 | ciliary basal body(GO:0036064) |
| 0.6 | 7.9 | GO:0005798 | Golgi-associated vesicle(GO:0005798) |
| 0.5 | 12.8 | GO:0071565 | nBAF complex(GO:0071565) |
| 0.5 | 2.5 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.5 | 32.5 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.5 | 6.1 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.5 | 1.8 | GO:0008091 | spectrin(GO:0008091) |
| 0.4 | 13.0 | GO:0097546 | ciliary base(GO:0097546) |
| 0.4 | 14.7 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.4 | 11.3 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.4 | 1.3 | GO:0000818 | nuclear MIS12/MIND complex(GO:0000818) condensed chromosome inner kinetochore(GO:0000939) |
| 0.4 | 4.6 | GO:0032300 | mismatch repair complex(GO:0032300) |
| 0.4 | 69.6 | GO:0030426 | growth cone(GO:0030426) |
| 0.4 | 19.6 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.4 | 2.2 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.4 | 5.1 | GO:0001673 | male germ cell nucleus(GO:0001673) |
| 0.4 | 8.3 | GO:0099738 | cell cortex region(GO:0099738) |
| 0.4 | 1.1 | GO:0034686 | integrin alphav-beta8 complex(GO:0034686) |
| 0.3 | 9.6 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.3 | 12.2 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.3 | 11.0 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.3 | 0.9 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.3 | 24.5 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.3 | 3.7 | GO:0031252 | cell leading edge(GO:0031252) |
| 0.3 | 27.9 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.2 | 1.7 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.2 | 16.5 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.2 | 3.9 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.2 | 6.2 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.2 | 102.4 | GO:0010008 | endosome membrane(GO:0010008) |
| 0.2 | 1.1 | GO:1990130 | EGO complex(GO:0034448) Iml1 complex(GO:1990130) Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.2 | 77.8 | GO:0005815 | microtubule organizing center(GO:0005815) |
| 0.2 | 57.6 | GO:0045202 | synapse(GO:0045202) |
| 0.2 | 5.0 | GO:0016592 | mediator complex(GO:0016592) |
| 0.1 | 16.4 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 0.1 | 6.6 | GO:0000786 | nucleosome(GO:0000786) |
| 0.1 | 4.6 | GO:0005859 | muscle myosin complex(GO:0005859) |
| 0.1 | 3.1 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.1 | 9.0 | GO:0005902 | microvillus(GO:0005902) |
| 0.1 | 11.4 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) potassium channel complex(GO:0034705) |
| 0.1 | 14.1 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.1 | 191.8 | GO:0042995 | cell projection(GO:0042995) |
| 0.1 | 17.7 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.1 | 7.0 | GO:0036477 | somatodendritic compartment(GO:0036477) |
| 0.1 | 6.3 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.1 | 14.3 | GO:0031461 | cullin-RING ubiquitin ligase complex(GO:0031461) |
| 0.1 | 13.0 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.1 | 4.1 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.1 | 11.5 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.1 | 2.1 | GO:0045334 | clathrin-coated endocytic vesicle(GO:0045334) |
| 0.1 | 1.1 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) |
| 0.1 | 0.5 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.1 | 3.0 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.1 | 4.7 | GO:0005811 | lipid particle(GO:0005811) |
| 0.1 | 16.3 | GO:0005773 | vacuole(GO:0005773) |
| 0.1 | 10.9 | GO:0005874 | microtubule(GO:0005874) |
| 0.1 | 4.3 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.1 | 8.2 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.0 | 4.7 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 2.6 | GO:0005769 | early endosome(GO:0005769) |
| 0.0 | 0.9 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 0.0 | GO:1903439 | G-protein coupled receptor dimeric complex(GO:0038037) G-protein coupled receptor complex(GO:0097648) calcitonin family receptor complex(GO:1903439) amylin receptor complex(GO:1903440) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.4 | 31.3 | GO:0034739 | histone deacetylase activity (H4-K16 specific)(GO:0034739) |
| 9.8 | 48.8 | GO:0004489 | methylenetetrahydrofolate reductase (NAD(P)H) activity(GO:0004489) |
| 7.9 | 55.6 | GO:0016206 | catechol O-methyltransferase activity(GO:0016206) |
| 7.7 | 61.3 | GO:0004565 | beta-galactosidase activity(GO:0004565) |
| 6.2 | 18.6 | GO:1904455 | ubiquitin-specific protease activity involved in negative regulation of ERAD pathway(GO:1904455) |
| 5.2 | 15.5 | GO:0019912 | cyclin-dependent protein kinase activating kinase activity(GO:0019912) |
| 4.6 | 36.8 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 4.5 | 26.8 | GO:0004792 | thiosulfate sulfurtransferase activity(GO:0004792) |
| 4.3 | 21.4 | GO:0070740 | tubulin-glutamic acid ligase activity(GO:0070740) |
| 3.7 | 91.9 | GO:0045503 | dynein light chain binding(GO:0045503) |
| 3.3 | 10.0 | GO:0080130 | phosphatidylserine decarboxylase activity(GO:0004609) L-phenylalanine aminotransferase activity(GO:0070546) L-phenylalanine:2-oxoglutarate aminotransferase activity(GO:0080130) |
| 3.3 | 19.9 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 3.3 | 19.8 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) fructose-2,6-bisphosphate 2-phosphatase activity(GO:0004331) |
| 2.9 | 23.4 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 2.8 | 8.4 | GO:0015439 | heme-transporting ATPase activity(GO:0015439) |
| 2.6 | 33.8 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 2.4 | 11.8 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 2.2 | 48.9 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 2.1 | 6.4 | GO:0004618 | phosphoglycerate kinase activity(GO:0004618) |
| 2.1 | 8.2 | GO:0017169 | CDP-alcohol phosphatidyltransferase activity(GO:0017169) |
| 2.1 | 16.4 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 2.0 | 6.1 | GO:0045322 | unmethylated CpG binding(GO:0045322) |
| 2.0 | 30.5 | GO:0070679 | inositol 1,4,5 trisphosphate binding(GO:0070679) |
| 1.9 | 5.7 | GO:0008321 | Ral guanyl-nucleotide exchange factor activity(GO:0008321) |
| 1.8 | 7.3 | GO:0035373 | chondroitin sulfate proteoglycan binding(GO:0035373) |
| 1.8 | 8.9 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 1.7 | 8.7 | GO:0001641 | group II metabotropic glutamate receptor activity(GO:0001641) |
| 1.7 | 8.3 | GO:0097108 | hedgehog family protein binding(GO:0097108) |
| 1.6 | 4.9 | GO:0004616 | phosphogluconate dehydrogenase (decarboxylating) activity(GO:0004616) |
| 1.6 | 32.7 | GO:0043560 | insulin receptor substrate binding(GO:0043560) |
| 1.6 | 4.7 | GO:0004938 | alpha2-adrenergic receptor activity(GO:0004938) |
| 1.4 | 7.2 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) |
| 1.4 | 16.8 | GO:0005229 | intracellular calcium activated chloride channel activity(GO:0005229) |
| 1.4 | 16.4 | GO:0048156 | tau protein binding(GO:0048156) |
| 1.3 | 6.7 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 1.3 | 6.4 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 1.2 | 3.7 | GO:0016215 | stearoyl-CoA 9-desaturase activity(GO:0004768) acyl-CoA desaturase activity(GO:0016215) |
| 1.2 | 11.9 | GO:0004579 | dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 1.2 | 18.8 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 1.2 | 12.8 | GO:0004645 | phosphorylase activity(GO:0004645) |
| 1.1 | 34.3 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 1.1 | 4.5 | GO:0032145 | 4-aminobutyrate transaminase activity(GO:0003867) succinate-semialdehyde dehydrogenase binding(GO:0032145) (S)-3-amino-2-methylpropionate transaminase activity(GO:0047298) |
| 1.1 | 80.5 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.9 | 21.3 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 0.9 | 19.1 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.9 | 3.6 | GO:0005222 | intracellular cAMP activated cation channel activity(GO:0005222) |
| 0.9 | 16.3 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.8 | 2.4 | GO:0003980 | UDP-glucose:glycoprotein glucosyltransferase activity(GO:0003980) |
| 0.8 | 3.8 | GO:0070568 | guanylyltransferase activity(GO:0070568) |
| 0.7 | 5.8 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.7 | 2.1 | GO:0015235 | cobalamin transporter activity(GO:0015235) |
| 0.7 | 2.7 | GO:0070137 | ubiquitin-like protein-specific endopeptidase activity(GO:0070137) SUMO-specific endopeptidase activity(GO:0070139) |
| 0.7 | 4.0 | GO:0039552 | RIG-I binding(GO:0039552) |
| 0.7 | 3.3 | GO:0004974 | leukotriene receptor activity(GO:0004974) |
| 0.6 | 16.8 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.6 | 7.7 | GO:0022851 | GABA-gated chloride ion channel activity(GO:0022851) |
| 0.6 | 1.7 | GO:0071566 | UFM1 activating enzyme activity(GO:0071566) |
| 0.5 | 4.3 | GO:0047144 | 2-acylglycerol-3-phosphate O-acyltransferase activity(GO:0047144) |
| 0.5 | 3.7 | GO:0030160 | GKAP/Homer scaffold activity(GO:0030160) |
| 0.5 | 3.1 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.5 | 5.4 | GO:0004415 | hyalurononglucosaminidase activity(GO:0004415) |
| 0.5 | 9.9 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.5 | 16.3 | GO:0030332 | cyclin binding(GO:0030332) |
| 0.5 | 19.8 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.5 | 6.4 | GO:0017136 | NAD-dependent histone deacetylase activity(GO:0017136) |
| 0.5 | 12.4 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.4 | 19.3 | GO:0030552 | cAMP binding(GO:0030552) |
| 0.4 | 11.4 | GO:0048487 | beta-tubulin binding(GO:0048487) |
| 0.4 | 7.8 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.4 | 0.8 | GO:0032356 | oxidized DNA binding(GO:0032356) |
| 0.3 | 7.3 | GO:0008484 | sulfuric ester hydrolase activity(GO:0008484) |
| 0.3 | 8.9 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.3 | 3.7 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.3 | 9.3 | GO:0008519 | ammonium transmembrane transporter activity(GO:0008519) |
| 0.3 | 5.1 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.3 | 1.2 | GO:0035473 | lipase binding(GO:0035473) |
| 0.3 | 20.8 | GO:0030276 | clathrin binding(GO:0030276) |
| 0.3 | 6.1 | GO:0019789 | SUMO transferase activity(GO:0019789) |
| 0.3 | 11.8 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.3 | 3.0 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.3 | 19.6 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.3 | 4.2 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.3 | 0.8 | GO:0047757 | chondroitin-glucuronate 5-epimerase activity(GO:0047757) |
| 0.3 | 5.4 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 0.3 | 64.9 | GO:0008017 | microtubule binding(GO:0008017) |
| 0.3 | 4.6 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.3 | 6.5 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.3 | 4.0 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
| 0.3 | 5.6 | GO:0015278 | calcium-release channel activity(GO:0015278) ligand-gated calcium channel activity(GO:0099604) |
| 0.3 | 4.7 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.3 | 2.5 | GO:0022841 | potassium ion leak channel activity(GO:0022841) |
| 0.3 | 8.5 | GO:0008395 | steroid hydroxylase activity(GO:0008395) |
| 0.2 | 7.4 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.2 | 1.9 | GO:0004844 | uracil DNA N-glycosylase activity(GO:0004844) deaminated base DNA N-glycosylase activity(GO:0097506) |
| 0.2 | 14.8 | GO:0048365 | Rac GTPase binding(GO:0048365) |
| 0.2 | 4.0 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.2 | 1.9 | GO:0016595 | glutamate binding(GO:0016595) |
| 0.2 | 5.6 | GO:0071949 | FAD binding(GO:0071949) |
| 0.2 | 6.5 | GO:0046961 | proton-transporting ATPase activity, rotational mechanism(GO:0046961) |
| 0.2 | 10.9 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.2 | 2.1 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.2 | 9.5 | GO:0015631 | tubulin binding(GO:0015631) |
| 0.2 | 1.3 | GO:0034191 | apolipoprotein A-I receptor binding(GO:0034191) |
| 0.2 | 1.1 | GO:0051575 | 5'-deoxyribose-5-phosphate lyase activity(GO:0051575) |
| 0.2 | 2.2 | GO:0031433 | telethonin binding(GO:0031433) |
| 0.2 | 2.4 | GO:0046790 | virion binding(GO:0046790) |
| 0.2 | 8.0 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.2 | 1.1 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.2 | 2.7 | GO:0003691 | double-stranded telomeric DNA binding(GO:0003691) |
| 0.2 | 2.9 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.2 | 8.6 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.2 | 1.2 | GO:0003960 | NADPH:quinone reductase activity(GO:0003960) |
| 0.2 | 3.7 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.2 | 12.9 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.2 | 8.9 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.2 | 0.9 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.2 | 2.2 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.1 | 0.9 | GO:0032393 | MHC class I receptor activity(GO:0032393) |
| 0.1 | 0.4 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.1 | 2.9 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.1 | 1.1 | GO:0031386 | protein tag(GO:0031386) |
| 0.1 | 2.0 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.1 | 8.7 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.1 | 1.9 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.1 | 10.5 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 1.1 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.1 | 2.4 | GO:0001223 | transcription coactivator binding(GO:0001223) |
| 0.1 | 0.8 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.1 | 18.4 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.1 | 7.3 | GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds(GO:0004553) |
| 0.1 | 1.5 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.1 | 1.1 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.1 | 1.0 | GO:0003836 | beta-galactoside (CMP) alpha-2,3-sialyltransferase activity(GO:0003836) |
| 0.1 | 1.2 | GO:1990381 | ubiquitin-specific protease binding(GO:1990381) |
| 0.1 | 4.5 | GO:0001618 | virus receptor activity(GO:0001618) |
| 0.1 | 8.4 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.1 | 1.7 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.1 | 2.2 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.1 | 8.4 | GO:0005178 | integrin binding(GO:0005178) |
| 0.1 | 0.6 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 0.0 | 0.1 | GO:0042954 | lipoprotein transporter activity(GO:0042954) |
| 0.0 | 2.3 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.0 | 4.4 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 6.6 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.0 | 0.8 | GO:0051018 | protein kinase A binding(GO:0051018) |
| 0.0 | 0.1 | GO:0015350 | reduced folate carrier activity(GO:0008518) methotrexate transporter activity(GO:0015350) |
| 0.0 | 10.0 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.0 | 2.6 | GO:0004497 | monooxygenase activity(GO:0004497) |
| 0.0 | 1.3 | GO:0050681 | androgen receptor binding(GO:0050681) |
| 0.0 | 0.5 | GO:0004551 | nucleotide diphosphatase activity(GO:0004551) |
| 0.0 | 4.2 | GO:0035091 | phosphatidylinositol binding(GO:0035091) |
| 0.0 | 0.8 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 1.9 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 0.6 | GO:0016798 | hydrolase activity, acting on glycosyl bonds(GO:0016798) |
| 0.0 | 1.2 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.9 | GO:0008536 | Ran GTPase binding(GO:0008536) |
| 0.0 | 10.1 | GO:0008047 | enzyme activator activity(GO:0008047) |
| 0.0 | 0.7 | GO:0008143 | poly(A) binding(GO:0008143) |
| 0.0 | 1.6 | GO:0005249 | voltage-gated potassium channel activity(GO:0005249) |
| 0.0 | 1.8 | GO:0004714 | transmembrane receptor protein tyrosine kinase activity(GO:0004714) |
| 0.0 | 0.2 | GO:0043548 | phosphatidylinositol 3-kinase binding(GO:0043548) |
| 0.0 | 4.2 | GO:0003779 | actin binding(GO:0003779) |
| 0.0 | 0.0 | GO:0031716 | calcitonin receptor activity(GO:0004948) calcitonin receptor binding(GO:0031716) |
| 0.0 | 0.2 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.9 | 2.9 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 1.0 | 25.2 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.8 | 32.7 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.7 | 25.0 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.6 | 17.2 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.4 | 40.2 | ST T CELL SIGNAL TRANSDUCTION | T Cell Signal Transduction |
| 0.4 | 23.2 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.3 | 19.0 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.3 | 21.4 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.3 | 3.3 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
| 0.3 | 20.2 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.3 | 7.3 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.3 | 14.0 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.3 | 19.8 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.3 | 12.8 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.2 | 8.9 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.2 | 11.7 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.2 | 4.9 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.2 | 19.9 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.2 | 3.5 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.2 | 4.8 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.2 | 4.9 | ST PHOSPHOINOSITIDE 3 KINASE PATHWAY | PI3K Pathway |
| 0.1 | 3.7 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 6.9 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.1 | 3.7 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.1 | 3.6 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.1 | 2.0 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.1 | 6.2 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.1 | 1.1 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.1 | 1.6 | PID FGF PATHWAY | FGF signaling pathway |
| 0.1 | 3.3 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.1 | 1.9 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
| 0.1 | 1.5 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.1 | 0.6 | PID PI3KCI AKT PATHWAY | Class I PI3K signaling events mediated by Akt |
| 0.1 | 1.1 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.0 | 2.0 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.0 | 1.5 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.0 | 2.1 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 0.6 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.0 | 0.7 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.0 | 0.5 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.0 | 0.3 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.0 | 0.9 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 2.2 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 3.3 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 25.3 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 1.4 | 39.4 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 1.4 | 32.7 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 1.2 | 24.9 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 1.0 | 70.1 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.9 | 8.5 | REACTOME REGULATION OF INSULIN SECRETION | Genes involved in Regulation of Insulin Secretion |
| 0.9 | 8.5 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 0.7 | 16.6 | REACTOME G BETA GAMMA SIGNALLING THROUGH PLC BETA | Genes involved in G beta:gamma signalling through PLC beta |
| 0.6 | 14.8 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.5 | 8.7 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.5 | 8.7 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.4 | 5.3 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.4 | 11.0 | REACTOME EGFR DOWNREGULATION | Genes involved in EGFR downregulation |
| 0.4 | 11.3 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.4 | 29.7 | REACTOME NUCLEAR SIGNALING BY ERBB4 | Genes involved in Nuclear signaling by ERBB4 |
| 0.4 | 12.7 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 2 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 2 Promoter |
| 0.4 | 23.9 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.4 | 10.0 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.4 | 2.5 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 0.3 | 4.7 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |
| 0.3 | 2.2 | REACTOME G ALPHA1213 SIGNALLING EVENTS | Genes involved in G alpha (12/13) signalling events |
| 0.3 | 2.4 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.3 | 11.1 | REACTOME REGULATION OF INSULIN SECRETION BY GLUCAGON LIKE PEPTIDE1 | Genes involved in Regulation of Insulin Secretion by Glucagon-like Peptide-1 |
| 0.2 | 7.7 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.2 | 9.3 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.2 | 4.7 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.2 | 4.9 | REACTOME ERK MAPK TARGETS | Genes involved in ERK/MAPK targets |
| 0.2 | 8.4 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.2 | 4.6 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.2 | 1.7 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.2 | 3.3 | REACTOME EICOSANOID LIGAND BINDING RECEPTORS | Genes involved in Eicosanoid ligand-binding receptors |
| 0.2 | 4.1 | REACTOME ACYL CHAIN REMODELLING OF PE | Genes involved in Acyl chain remodelling of PE |
| 0.2 | 6.7 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.1 | 4.8 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 4.5 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.1 | 2.7 | REACTOME RESOLUTION OF AP SITES VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Resolution of AP sites via the single-nucleotide replacement pathway |
| 0.1 | 7.5 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.1 | 3.4 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.1 | 3.9 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.1 | 4.8 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.1 | 2.9 | REACTOME RIP MEDIATED NFKB ACTIVATION VIA DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 0.1 | 8.3 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.1 | 1.5 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.1 | 5.5 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.1 | 5.9 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.1 | 1.9 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.1 | 2.1 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.1 | 1.1 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.1 | 8.9 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.1 | 2.1 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.1 | 1.6 | REACTOME SIGNAL TRANSDUCTION BY L1 | Genes involved in Signal transduction by L1 |
| 0.1 | 1.0 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.0 | 15.6 | REACTOME GENERIC TRANSCRIPTION PATHWAY | Genes involved in Generic Transcription Pathway |
| 0.0 | 0.9 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 1.6 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.0 | 1.7 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 2.2 | REACTOME SIGNALING BY ERBB2 | Genes involved in Signaling by ERBB2 |
| 0.0 | 0.6 | REACTOME INFLAMMASOMES | Genes involved in Inflammasomes |
| 0.0 | 1.6 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 1.1 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.0 | 1.0 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |