Project
GNF SymAtlas + NCI-60 cancer cell lines, human (Su, 2004; Ross, 2000)
Navigation
Downloads
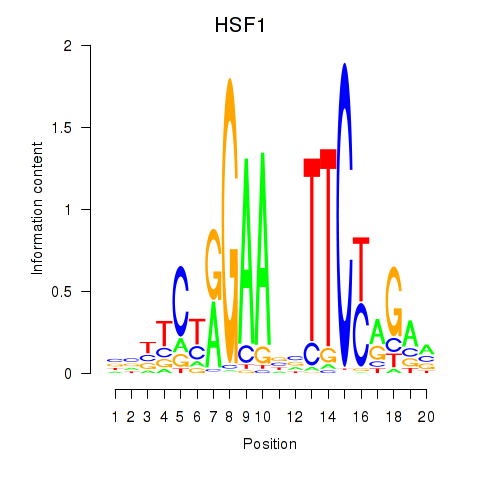
Results for HSF1
Z-value: 1.22
Transcription factors associated with HSF1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HSF1
|
ENSG00000185122.6 | heat shock transcription factor 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HSF1 | hg19_v2_chr8_+_145515263_145515299 | -0.12 | 7.8e-02 | Click! |
Activity profile of HSF1 motif
Sorted Z-values of HSF1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_-_33041378 | 41.32 |
ENST00000428995.1
|
HLA-DPA1
|
major histocompatibility complex, class II, DP alpha 1 |
| chr19_+_1065922 | 26.17 |
ENST00000539243.2
|
HMHA1
|
histocompatibility (minor) HA-1 |
| chr1_+_111415757 | 22.73 |
ENST00000429072.2
ENST00000271324.5 |
CD53
|
CD53 molecule |
| chr4_-_84030996 | 21.37 |
ENST00000411416.2
|
PLAC8
|
placenta-specific 8 |
| chr2_-_89513402 | 18.91 |
ENST00000498435.1
|
IGKV1-27
|
immunoglobulin kappa variable 1-27 |
| chr6_-_32557610 | 18.30 |
ENST00000360004.5
|
HLA-DRB1
|
major histocompatibility complex, class II, DR beta 1 |
| chr1_-_153363452 | 18.04 |
ENST00000368732.1
ENST00000368733.3 |
S100A8
|
S100 calcium binding protein A8 |
| chr6_-_32498046 | 17.51 |
ENST00000374975.3
|
HLA-DRB5
|
major histocompatibility complex, class II, DR beta 5 |
| chr6_+_32605195 | 17.31 |
ENST00000374949.2
|
HLA-DQA1
|
major histocompatibility complex, class II, DQ alpha 1 |
| chr14_-_106092403 | 17.14 |
ENST00000390543.2
|
IGHG4
|
immunoglobulin heavy constant gamma 4 (G4m marker) |
| chr5_+_118690466 | 13.11 |
ENST00000503646.1
|
TNFAIP8
|
tumor necrosis factor, alpha-induced protein 8 |
| chr7_+_150264365 | 13.05 |
ENST00000255945.2
ENST00000461940.1 |
GIMAP4
|
GTPase, IMAP family member 4 |
| chr16_+_85942594 | 12.20 |
ENST00000566369.1
|
IRF8
|
interferon regulatory factor 8 |
| chr14_-_106926724 | 11.80 |
ENST00000434710.1
|
IGHV3-43
|
immunoglobulin heavy variable 3-43 |
| chr6_+_32811885 | 11.76 |
ENST00000458296.1
ENST00000413039.1 ENST00000429600.1 ENST00000412095.1 ENST00000415067.1 ENST00000395330.1 |
TAPSAR1
PSMB9
|
TAP1 and PSMB8 antisense RNA 1 proteasome (prosome, macropain) subunit, beta type, 9 |
| chr3_-_46506563 | 11.73 |
ENST00000231751.4
|
LTF
|
lactotransferrin |
| chr6_-_41909561 | 11.65 |
ENST00000372991.4
|
CCND3
|
cyclin D3 |
| chr3_-_46506358 | 11.52 |
ENST00000417439.1
ENST00000431944.1 |
LTF
|
lactotransferrin |
| chr6_-_41909466 | 11.45 |
ENST00000414200.2
|
CCND3
|
cyclin D3 |
| chr6_+_32605134 | 11.11 |
ENST00000343139.5
ENST00000395363.1 ENST00000496318.1 |
HLA-DQA1
|
major histocompatibility complex, class II, DQ alpha 1 |
| chr22_-_17680472 | 10.95 |
ENST00000330232.4
|
CECR1
|
cat eye syndrome chromosome region, candidate 1 |
| chr1_+_161494036 | 10.89 |
ENST00000309758.4
|
HSPA6
|
heat shock 70kDa protein 6 (HSP70B') |
| chr14_-_106573756 | 10.80 |
ENST00000390601.2
|
IGHV3-11
|
immunoglobulin heavy variable 3-11 (gene/pseudogene) |
| chr2_-_201729284 | 10.69 |
ENST00000434813.2
|
CLK1
|
CDC-like kinase 1 |
| chr6_-_41909191 | 10.21 |
ENST00000512426.1
ENST00000372987.4 |
CCND3
|
cyclin D3 |
| chr19_-_54876558 | 10.21 |
ENST00000391742.2
ENST00000434277.2 |
LAIR1
|
leukocyte-associated immunoglobulin-like receptor 1 |
| chr6_-_32812420 | 10.16 |
ENST00000374881.2
|
PSMB8
|
proteasome (prosome, macropain) subunit, beta type, 8 |
| chr2_+_85922491 | 9.86 |
ENST00000526018.1
|
GNLY
|
granulysin |
| chr6_+_29910301 | 9.77 |
ENST00000376809.5
ENST00000376802.2 |
HLA-A
|
major histocompatibility complex, class I, A |
| chr19_-_2051223 | 9.34 |
ENST00000309340.7
ENST00000589534.1 ENST00000250896.3 ENST00000589509.1 |
MKNK2
|
MAP kinase interacting serine/threonine kinase 2 |
| chr6_-_32811771 | 9.08 |
ENST00000395339.3
ENST00000374882.3 |
PSMB8
|
proteasome (prosome, macropain) subunit, beta type, 8 |
| chr6_+_35265586 | 8.63 |
ENST00000542066.1
ENST00000316637.5 |
DEF6
|
differentially expressed in FDCP 6 homolog (mouse) |
| chr22_-_26961328 | 8.30 |
ENST00000398110.2
|
TPST2
|
tyrosylprotein sulfotransferase 2 |
| chr7_+_150434430 | 8.07 |
ENST00000358647.3
|
GIMAP5
|
GTPase, IMAP family member 5 |
| chr7_-_139763521 | 8.03 |
ENST00000263549.3
|
PARP12
|
poly (ADP-ribose) polymerase family, member 12 |
| chr15_-_55581954 | 7.88 |
ENST00000336787.1
|
RAB27A
|
RAB27A, member RAS oncogene family |
| chr5_-_171615315 | 7.28 |
ENST00000176763.5
|
STK10
|
serine/threonine kinase 10 |
| chr14_-_106552755 | 7.23 |
ENST00000390600.2
|
IGHV3-9
|
immunoglobulin heavy variable 3-9 |
| chr17_-_39661849 | 7.14 |
ENST00000246635.3
ENST00000336861.3 ENST00000587544.1 ENST00000587435.1 |
KRT13
|
keratin 13 |
| chr11_+_809961 | 7.13 |
ENST00000530797.1
|
RPLP2
|
ribosomal protein, large, P2 |
| chr1_+_212782012 | 7.03 |
ENST00000341491.4
ENST00000366985.1 |
ATF3
|
activating transcription factor 3 |
| chr2_+_61404624 | 7.00 |
ENST00000394457.3
|
AHSA2
|
AHA1, activator of heat shock 90kDa protein ATPase homolog 2 (yeast) |
| chr14_+_23012122 | 6.98 |
ENST00000390534.1
|
TRAJ3
|
T cell receptor alpha joining 3 |
| chr11_-_102323489 | 6.96 |
ENST00000361236.3
|
TMEM123
|
transmembrane protein 123 |
| chr1_-_17215868 | 6.81 |
ENST00000422124.1
|
RP11-108M9.4
|
RP11-108M9.4 |
| chr5_-_169694286 | 6.78 |
ENST00000521416.1
ENST00000520344.1 |
LCP2
|
lymphocyte cytosolic protein 2 (SH2 domain containing leukocyte protein of 76kDa) |
| chr1_-_17216109 | 6.72 |
ENST00000416869.1
|
RP11-108M9.4
|
RP11-108M9.4 |
| chr19_-_54876414 | 6.60 |
ENST00000474878.1
ENST00000348231.4 |
LAIR1
|
leukocyte-associated immunoglobulin-like receptor 1 |
| chr22_-_26986045 | 6.58 |
ENST00000442495.1
ENST00000440953.1 ENST00000450022.1 ENST00000338754.4 |
TPST2
|
tyrosylprotein sulfotransferase 2 |
| chr12_+_10460549 | 6.58 |
ENST00000543420.1
ENST00000543777.1 |
KLRD1
|
killer cell lectin-like receptor subfamily D, member 1 |
| chr12_-_57505121 | 6.51 |
ENST00000538913.2
ENST00000537215.2 ENST00000454075.3 ENST00000554825.1 ENST00000553275.1 ENST00000300134.3 |
STAT6
|
signal transducer and activator of transcription 6, interleukin-4 induced |
| chr6_+_89674246 | 6.49 |
ENST00000369474.1
|
AL079342.1
|
Uncharacterized protein; cDNA FLJ27030 fis, clone SLV07741 |
| chr6_-_167369612 | 6.49 |
ENST00000507747.1
|
RP11-514O12.4
|
RP11-514O12.4 |
| chr5_-_175964366 | 6.43 |
ENST00000274811.4
|
RNF44
|
ring finger protein 44 |
| chr6_+_31783291 | 6.33 |
ENST00000375651.5
ENST00000608703.1 ENST00000458062.2 |
HSPA1A
|
heat shock 70kDa protein 1A |
| chr18_-_19284724 | 6.33 |
ENST00000580981.1
ENST00000289119.2 |
ABHD3
|
abhydrolase domain containing 3 |
| chr11_-_104840093 | 6.32 |
ENST00000417440.2
ENST00000444739.2 |
CASP4
|
caspase 4, apoptosis-related cysteine peptidase |
| chr16_+_68119764 | 6.27 |
ENST00000570212.1
ENST00000562926.1 |
NFATC3
|
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 3 |
| chr11_-_64512273 | 6.24 |
ENST00000377497.3
ENST00000377487.1 ENST00000430645.1 |
RASGRP2
|
RAS guanyl releasing protein 2 (calcium and DAG-regulated) |
| chr1_+_6105974 | 6.06 |
ENST00000378083.3
|
KCNAB2
|
potassium voltage-gated channel, shaker-related subfamily, beta member 2 |
| chr1_-_20306909 | 6.00 |
ENST00000375111.3
ENST00000400520.3 |
PLA2G2A
|
phospholipase A2, group IIA (platelets, synovial fluid) |
| chr22_-_36556821 | 5.91 |
ENST00000531095.1
ENST00000397293.2 ENST00000349314.2 |
APOL3
|
apolipoprotein L, 3 |
| chr5_+_156696362 | 5.89 |
ENST00000377576.3
|
CYFIP2
|
cytoplasmic FMR1 interacting protein 2 |
| chr19_+_47813110 | 5.85 |
ENST00000355085.3
|
C5AR1
|
complement component 5a receptor 1 |
| chr14_-_25045446 | 5.82 |
ENST00000216336.2
|
CTSG
|
cathepsin G |
| chr22_-_29196546 | 5.81 |
ENST00000403532.3
ENST00000216037.6 |
XBP1
|
X-box binding protein 1 |
| chr17_-_76123101 | 5.69 |
ENST00000392467.3
|
TMC6
|
transmembrane channel-like 6 |
| chr11_-_104905840 | 5.67 |
ENST00000526568.1
ENST00000393136.4 ENST00000531166.1 ENST00000534497.1 ENST00000527979.1 ENST00000446369.1 ENST00000353247.5 ENST00000528974.1 ENST00000533400.1 ENST00000525825.1 ENST00000436863.3 |
CASP1
|
caspase 1, apoptosis-related cysteine peptidase |
| chr16_+_222846 | 5.63 |
ENST00000251595.6
ENST00000397806.1 |
HBA2
|
hemoglobin, alpha 2 |
| chr19_+_13842559 | 5.53 |
ENST00000586600.1
|
CCDC130
|
coiled-coil domain containing 130 |
| chr12_+_6898638 | 5.45 |
ENST00000011653.4
|
CD4
|
CD4 molecule |
| chr21_+_42733870 | 5.43 |
ENST00000330714.3
ENST00000436410.1 ENST00000435611.1 |
MX2
|
myxovirus (influenza virus) resistance 2 (mouse) |
| chr14_-_106725723 | 5.37 |
ENST00000390609.2
|
IGHV3-23
|
immunoglobulin heavy variable 3-23 |
| chr18_+_57567180 | 5.26 |
ENST00000316660.6
ENST00000269518.9 |
PMAIP1
|
phorbol-12-myristate-13-acetate-induced protein 1 |
| chr15_+_81589254 | 5.26 |
ENST00000394652.2
|
IL16
|
interleukin 16 |
| chr14_-_106668095 | 5.22 |
ENST00000390606.2
|
IGHV3-20
|
immunoglobulin heavy variable 3-20 |
| chr14_-_107049312 | 5.16 |
ENST00000390627.2
|
IGHV3-53
|
immunoglobulin heavy variable 3-53 |
| chr4_-_25865159 | 5.01 |
ENST00000502949.1
ENST00000264868.5 ENST00000513691.1 ENST00000514872.1 |
SEL1L3
|
sel-1 suppressor of lin-12-like 3 (C. elegans) |
| chr13_-_49018789 | 4.98 |
ENST00000378434.4
|
LPAR6
|
lysophosphatidic acid receptor 6 |
| chr11_+_117063295 | 4.96 |
ENST00000525478.1
ENST00000532062.1 |
SIDT2
|
SID1 transmembrane family, member 2 |
| chr12_+_51632508 | 4.95 |
ENST00000449723.3
|
DAZAP2
|
DAZ associated protein 2 |
| chr12_+_10460417 | 4.89 |
ENST00000381908.3
ENST00000336164.4 ENST00000350274.5 |
KLRD1
|
killer cell lectin-like receptor subfamily D, member 1 |
| chr1_+_212738676 | 4.70 |
ENST00000366981.4
ENST00000366987.2 |
ATF3
|
activating transcription factor 3 |
| chr7_+_130020932 | 4.67 |
ENST00000484324.1
|
CPA1
|
carboxypeptidase A1 (pancreatic) |
| chr1_-_236030216 | 4.62 |
ENST00000389794.3
ENST00000389793.2 |
LYST
|
lysosomal trafficking regulator |
| chr20_-_4795747 | 4.61 |
ENST00000379376.2
|
RASSF2
|
Ras association (RalGDS/AF-6) domain family member 2 |
| chr22_+_37309662 | 4.58 |
ENST00000403662.3
ENST00000262825.5 |
CSF2RB
|
colony stimulating factor 2 receptor, beta, low-affinity (granulocyte-macrophage) |
| chr7_+_150065278 | 4.56 |
ENST00000519397.1
ENST00000479668.1 ENST00000540729.1 |
REPIN1
|
replication initiator 1 |
| chr1_-_225615599 | 4.51 |
ENST00000421383.1
ENST00000272163.4 |
LBR
|
lamin B receptor |
| chr3_+_42947600 | 4.49 |
ENST00000328199.6
ENST00000541208.1 |
ZNF662
|
zinc finger protein 662 |
| chr11_+_10471836 | 4.47 |
ENST00000444303.2
|
AMPD3
|
adenosine monophosphate deaminase 3 |
| chrX_-_73072534 | 4.45 |
ENST00000429829.1
|
XIST
|
X inactive specific transcript (non-protein coding) |
| chr21_+_35445827 | 4.40 |
ENST00000608209.1
ENST00000381151.3 |
SLC5A3
SLC5A3
|
sodium/myo-inositol cotransporter solute carrier family 5 (sodium/myo-inositol cotransporter), member 3 |
| chr16_+_1583567 | 4.37 |
ENST00000566264.1
|
TMEM204
|
transmembrane protein 204 |
| chr16_+_28943260 | 4.33 |
ENST00000538922.1
ENST00000324662.3 ENST00000567541.1 |
CD19
|
CD19 molecule |
| chr5_+_40909354 | 4.28 |
ENST00000313164.9
|
C7
|
complement component 7 |
| chr10_+_114133773 | 4.27 |
ENST00000354655.4
|
ACSL5
|
acyl-CoA synthetase long-chain family member 5 |
| chrX_-_47518498 | 4.20 |
ENST00000335890.2
|
UXT
|
ubiquitously-expressed, prefoldin-like chaperone |
| chr20_-_44455976 | 4.18 |
ENST00000372555.3
|
TNNC2
|
troponin C type 2 (fast) |
| chr2_+_219247021 | 4.16 |
ENST00000539932.1
|
SLC11A1
|
solute carrier family 11 (proton-coupled divalent metal ion transporter), member 1 |
| chr6_+_31795506 | 4.16 |
ENST00000375650.3
|
HSPA1B
|
heat shock 70kDa protein 1B |
| chrX_+_64887512 | 4.16 |
ENST00000360270.5
|
MSN
|
moesin |
| chr11_+_65190245 | 4.13 |
ENST00000499732.1
ENST00000501122.2 ENST00000601801.1 |
NEAT1
|
nuclear paraspeckle assembly transcript 1 (non-protein coding) |
| chr17_-_56350797 | 4.12 |
ENST00000577220.1
|
MPO
|
myeloperoxidase |
| chr19_+_55014085 | 4.08 |
ENST00000351841.2
|
LAIR2
|
leukocyte-associated immunoglobulin-like receptor 2 |
| chr7_+_102715315 | 4.06 |
ENST00000428183.2
ENST00000323716.3 ENST00000441711.2 ENST00000454559.1 ENST00000425331.1 ENST00000541300.1 |
ARMC10
|
armadillo repeat containing 10 |
| chrX_+_13752832 | 4.04 |
ENST00000380550.3
ENST00000398395.3 ENST00000340096.6 ENST00000380567.1 |
OFD1
|
oral-facial-digital syndrome 1 |
| chr21_-_43735446 | 3.98 |
ENST00000398431.2
|
TFF3
|
trefoil factor 3 (intestinal) |
| chr2_+_102624977 | 3.96 |
ENST00000441002.1
|
IL1R2
|
interleukin 1 receptor, type II |
| chr11_-_1785139 | 3.94 |
ENST00000236671.2
|
CTSD
|
cathepsin D |
| chr13_-_79979919 | 3.94 |
ENST00000267229.7
|
RBM26
|
RNA binding motif protein 26 |
| chr19_-_14629224 | 3.92 |
ENST00000254322.2
|
DNAJB1
|
DnaJ (Hsp40) homolog, subfamily B, member 1 |
| chr16_-_15180257 | 3.92 |
ENST00000540462.1
|
RRN3
|
RRN3 RNA polymerase I transcription factor homolog (S. cerevisiae) |
| chr21_-_34144157 | 3.90 |
ENST00000331923.4
|
PAXBP1
|
PAX3 and PAX7 binding protein 1 |
| chr16_+_28996572 | 3.85 |
ENST00000360872.5
ENST00000566177.1 ENST00000354453.4 |
LAT
|
linker for activation of T cells |
| chr1_-_225616515 | 3.73 |
ENST00000338179.2
ENST00000425080.1 |
LBR
|
lamin B receptor |
| chr1_+_22307592 | 3.72 |
ENST00000400277.2
|
CELA3B
|
chymotrypsin-like elastase family, member 3B |
| chr14_-_106518922 | 3.69 |
ENST00000390598.2
|
IGHV3-7
|
immunoglobulin heavy variable 3-7 |
| chr12_-_52911718 | 3.68 |
ENST00000548409.1
|
KRT5
|
keratin 5 |
| chr17_-_3595181 | 3.63 |
ENST00000552050.1
|
P2RX5
|
purinergic receptor P2X, ligand-gated ion channel, 5 |
| chr16_-_20339123 | 3.62 |
ENST00000381360.5
|
GP2
|
glycoprotein 2 (zymogen granule membrane) |
| chr13_-_79979952 | 3.62 |
ENST00000438724.1
|
RBM26
|
RNA binding motif protein 26 |
| chr16_-_20338748 | 3.61 |
ENST00000575582.1
ENST00000341642.5 ENST00000381362.4 ENST00000572347.1 ENST00000572478.1 ENST00000302555.5 |
GP2
|
glycoprotein 2 (zymogen granule membrane) |
| chr20_+_46130671 | 3.58 |
ENST00000371998.3
ENST00000371997.3 |
NCOA3
|
nuclear receptor coactivator 3 |
| chr3_-_48936272 | 3.57 |
ENST00000544097.1
ENST00000430379.1 ENST00000319017.4 |
SLC25A20
|
solute carrier family 25 (carnitine/acylcarnitine translocase), member 20 |
| chrX_+_15767971 | 3.57 |
ENST00000479740.1
ENST00000454127.2 |
CA5B
|
carbonic anhydrase VB, mitochondrial |
| chr7_+_80275752 | 3.56 |
ENST00000419819.2
|
CD36
|
CD36 molecule (thrombospondin receptor) |
| chr15_-_55563072 | 3.55 |
ENST00000567380.1
ENST00000565972.1 ENST00000569493.1 |
RAB27A
|
RAB27A, member RAS oncogene family |
| chrX_-_154563889 | 3.54 |
ENST00000369449.2
ENST00000321926.4 |
CLIC2
|
chloride intracellular channel 2 |
| chr9_-_97402531 | 3.54 |
ENST00000415431.1
|
FBP1
|
fructose-1,6-bisphosphatase 1 |
| chr17_+_8339189 | 3.53 |
ENST00000585098.1
ENST00000380025.4 ENST00000402554.3 ENST00000584866.1 ENST00000582490.1 |
NDEL1
|
nudE neurodevelopment protein 1-like 1 |
| chr7_+_114562172 | 3.53 |
ENST00000393486.1
ENST00000257724.3 |
MDFIC
|
MyoD family inhibitor domain containing |
| chr8_-_61880248 | 3.49 |
ENST00000525556.1
|
AC022182.3
|
AC022182.3 |
| chr19_+_55014013 | 3.47 |
ENST00000301202.2
|
LAIR2
|
leukocyte-associated immunoglobulin-like receptor 2 |
| chrX_-_230884 | 3.46 |
ENST00000400701.3
ENST00000326153.4 |
GTPBP6
|
GTP binding protein 6 (putative) |
| chr8_-_101321584 | 3.46 |
ENST00000523167.1
|
RNF19A
|
ring finger protein 19A, RBR E3 ubiquitin protein ligase |
| chr13_-_79980315 | 3.44 |
ENST00000438737.2
|
RBM26
|
RNA binding motif protein 26 |
| chr4_-_48136217 | 3.44 |
ENST00000264316.4
|
TXK
|
TXK tyrosine kinase |
| chr6_-_131949305 | 3.35 |
ENST00000368053.4
ENST00000354577.4 ENST00000403834.3 ENST00000540546.1 ENST00000368068.3 ENST00000368060.3 |
MED23
|
mediator complex subunit 23 |
| chr2_-_225811747 | 3.34 |
ENST00000409592.3
|
DOCK10
|
dedicator of cytokinesis 10 |
| chr3_-_150920979 | 3.34 |
ENST00000309180.5
ENST00000480322.1 |
GPR171
|
G protein-coupled receptor 171 |
| chr7_-_99573640 | 3.34 |
ENST00000411734.1
|
AZGP1
|
alpha-2-glycoprotein 1, zinc-binding |
| chr1_+_67773044 | 3.33 |
ENST00000262345.1
ENST00000371000.1 |
IL12RB2
|
interleukin 12 receptor, beta 2 |
| chr18_+_9136758 | 3.31 |
ENST00000383440.2
ENST00000262126.4 ENST00000577992.1 |
ANKRD12
|
ankyrin repeat domain 12 |
| chr16_-_79634595 | 3.27 |
ENST00000326043.4
ENST00000393350.1 |
MAF
|
v-maf avian musculoaponeurotic fibrosarcoma oncogene homolog |
| chr7_-_99573677 | 3.22 |
ENST00000292401.4
|
AZGP1
|
alpha-2-glycoprotein 1, zinc-binding |
| chr1_-_160832642 | 3.21 |
ENST00000368034.4
|
CD244
|
CD244 molecule, natural killer cell receptor 2B4 |
| chr3_+_101443476 | 3.20 |
ENST00000327230.4
ENST00000494050.1 |
CEP97
|
centrosomal protein 97kDa |
| chr20_-_23731569 | 3.20 |
ENST00000304749.2
|
CST1
|
cystatin SN |
| chr8_+_142402089 | 3.19 |
ENST00000521578.1
ENST00000520105.1 ENST00000523147.1 |
PTP4A3
|
protein tyrosine phosphatase type IVA, member 3 |
| chr3_-_187454281 | 3.17 |
ENST00000232014.4
|
BCL6
|
B-cell CLL/lymphoma 6 |
| chr11_+_46354455 | 3.16 |
ENST00000343674.6
|
DGKZ
|
diacylglycerol kinase, zeta |
| chr12_-_108154925 | 3.15 |
ENST00000228437.5
|
PRDM4
|
PR domain containing 4 |
| chrY_-_180884 | 3.15 |
ENSTR0000400701.3
ENSTR0000326153.4 |
GTPBP6
|
GTP binding protein 6 (putative) |
| chrX_+_108779870 | 3.12 |
ENST00000372107.1
|
NXT2
|
nuclear transport factor 2-like export factor 2 |
| chr9_+_123970052 | 3.11 |
ENST00000373823.3
|
GSN
|
gelsolin |
| chr9_+_140513438 | 3.11 |
ENST00000462484.1
ENST00000334856.6 ENST00000460843.1 |
EHMT1
|
euchromatic histone-lysine N-methyltransferase 1 |
| chr13_+_31191920 | 3.10 |
ENST00000255304.4
|
USPL1
|
ubiquitin specific peptidase like 1 |
| chr14_-_106586656 | 3.07 |
ENST00000390602.2
|
IGHV3-13
|
immunoglobulin heavy variable 3-13 |
| chr18_+_13611763 | 3.07 |
ENST00000585931.1
|
LDLRAD4
|
low density lipoprotein receptor class A domain containing 4 |
| chrX_-_47518527 | 3.03 |
ENST00000333119.3
|
UXT
|
ubiquitously-expressed, prefoldin-like chaperone |
| chr10_-_125851961 | 3.01 |
ENST00000346248.5
|
CHST15
|
carbohydrate (N-acetylgalactosamine 4-sulfate 6-O) sulfotransferase 15 |
| chr2_-_191878874 | 2.99 |
ENST00000392322.3
ENST00000392323.2 ENST00000424722.1 ENST00000361099.3 |
STAT1
|
signal transducer and activator of transcription 1, 91kDa |
| chr16_+_28996416 | 2.98 |
ENST00000395456.2
ENST00000454369.2 |
LAT
|
linker for activation of T cells |
| chr22_+_41253080 | 2.98 |
ENST00000541156.1
ENST00000414396.1 ENST00000357137.4 |
XPNPEP3
|
X-prolyl aminopeptidase (aminopeptidase P) 3, putative |
| chr2_+_58655461 | 2.98 |
ENST00000429095.1
ENST00000429664.1 ENST00000452840.1 |
AC007092.1
|
long intergenic non-protein coding RNA 1122 |
| chr12_-_123011536 | 2.94 |
ENST00000331738.7
ENST00000354654.2 |
RSRC2
|
arginine/serine-rich coiled-coil 2 |
| chr8_+_38677850 | 2.92 |
ENST00000518809.1
ENST00000520611.1 |
TACC1
|
transforming, acidic coiled-coil containing protein 1 |
| chr7_+_80275621 | 2.91 |
ENST00000426978.1
ENST00000432207.1 |
CD36
|
CD36 molecule (thrombospondin receptor) |
| chr6_-_32784687 | 2.90 |
ENST00000447394.1
ENST00000438763.2 |
HLA-DOB
|
major histocompatibility complex, class II, DO beta |
| chr12_+_51632600 | 2.88 |
ENST00000549555.1
ENST00000439799.2 ENST00000425012.2 |
DAZAP2
|
DAZ associated protein 2 |
| chr6_+_32936353 | 2.87 |
ENST00000374825.4
|
BRD2
|
bromodomain containing 2 |
| chr7_-_115608304 | 2.82 |
ENST00000457268.1
|
TFEC
|
transcription factor EC |
| chr1_-_228613026 | 2.81 |
ENST00000366696.1
|
HIST3H3
|
histone cluster 3, H3 |
| chr11_+_64323098 | 2.78 |
ENST00000301891.4
|
SLC22A11
|
solute carrier family 22 (organic anion/urate transporter), member 11 |
| chr19_-_5680499 | 2.76 |
ENST00000587589.1
|
C19orf70
|
chromosome 19 open reading frame 70 |
| chr12_-_123011476 | 2.71 |
ENST00000528279.1
ENST00000344591.4 ENST00000526560.2 |
RSRC2
|
arginine/serine-rich coiled-coil 2 |
| chr7_+_130020180 | 2.69 |
ENST00000481342.1
ENST00000011292.3 ENST00000604896.1 |
CPA1
|
carboxypeptidase A1 (pancreatic) |
| chr10_+_93683519 | 2.69 |
ENST00000265990.6
|
BTAF1
|
BTAF1 RNA polymerase II, B-TFIID transcription factor-associated, 170kDa |
| chr12_+_51632638 | 2.67 |
ENST00000549732.2
|
DAZAP2
|
DAZ associated protein 2 |
| chr20_+_43803517 | 2.67 |
ENST00000243924.3
|
PI3
|
peptidase inhibitor 3, skin-derived |
| chr6_-_46620522 | 2.65 |
ENST00000275016.2
|
CYP39A1
|
cytochrome P450, family 39, subfamily A, polypeptide 1 |
| chr14_+_39734482 | 2.64 |
ENST00000554392.1
ENST00000555716.1 ENST00000341749.3 ENST00000557038.1 |
CTAGE5
|
CTAGE family, member 5 |
| chr8_-_56987057 | 2.61 |
ENST00000518875.1
|
RPS20
|
ribosomal protein S20 |
| chr17_-_43138357 | 2.58 |
ENST00000342350.5
|
DCAKD
|
dephospho-CoA kinase domain containing |
| chr9_-_98079965 | 2.57 |
ENST00000289081.3
|
FANCC
|
Fanconi anemia, complementation group C |
| chr2_-_190627481 | 2.55 |
ENST00000264151.5
ENST00000520350.1 ENST00000521630.1 ENST00000517895.1 |
OSGEPL1
|
O-sialoglycoprotein endopeptidase-like 1 |
| chr1_+_152635854 | 2.51 |
ENST00000368784.1
|
LCE2D
|
late cornified envelope 2D |
| chr7_+_43622664 | 2.50 |
ENST00000319357.5
|
STK17A
|
serine/threonine kinase 17a |
| chr21_+_39644214 | 2.49 |
ENST00000438657.1
|
KCNJ15
|
potassium inwardly-rectifying channel, subfamily J, member 15 |
| chr6_-_131949200 | 2.48 |
ENST00000539158.1
ENST00000368058.1 |
MED23
|
mediator complex subunit 23 |
| chr10_+_102222798 | 2.48 |
ENST00000343737.5
|
WNT8B
|
wingless-type MMTV integration site family, member 8B |
| chr8_-_29940464 | 2.48 |
ENST00000521265.1
ENST00000536273.1 |
TMEM66
|
transmembrane protein 66 |
| chr17_+_56315936 | 2.45 |
ENST00000543544.1
|
LPO
|
lactoperoxidase |
| chr19_+_5681011 | 2.43 |
ENST00000581893.1
ENST00000411793.2 ENST00000301382.4 ENST00000581773.1 ENST00000423665.2 ENST00000583928.1 ENST00000342970.2 ENST00000422535.2 ENST00000581521.1 ENST00000339423.2 |
HSD11B1L
|
hydroxysteroid (11-beta) dehydrogenase 1-like |
| chr19_+_782755 | 2.43 |
ENST00000606242.1
ENST00000586061.1 |
AC006273.5
|
AC006273.5 |
| chr1_-_6260896 | 2.39 |
ENST00000497965.1
|
RPL22
|
ribosomal protein L22 |
| chr6_+_88182643 | 2.35 |
ENST00000369556.3
ENST00000544441.1 ENST00000369552.4 ENST00000369557.5 |
SLC35A1
|
solute carrier family 35 (CMP-sialic acid transporter), member A1 |
| chr17_-_34313685 | 2.35 |
ENST00000435911.2
ENST00000586216.1 ENST00000394509.4 |
CCL14
|
chemokine (C-C motif) ligand 14 |
| chr16_-_33647696 | 2.32 |
ENST00000558425.1
ENST00000569103.2 |
RP11-812E19.9
|
Uncharacterized protein |
Network of associatons between targets according to the STRING database.
First level regulatory network of HSF1
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.8 | 23.3 | GO:1900159 | iron assimilation(GO:0033212) iron assimilation by chelation and transport(GO:0033214) positive regulation of bone mineralization involved in bone maturation(GO:1900159) negative regulation of tumor necrosis factor (ligand) superfamily member 11 production(GO:2000308) |
| 5.0 | 14.9 | GO:0006478 | peptidyl-tyrosine sulfation(GO:0006478) |
| 4.3 | 21.4 | GO:0010286 | heat acclimation(GO:0010286) cellular heat acclimation(GO:0070370) |
| 3.9 | 11.7 | GO:0061394 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) |
| 3.7 | 18.3 | GO:0002399 | MHC class II protein complex assembly(GO:0002399) |
| 3.3 | 9.9 | GO:0002818 | intracellular defense response(GO:0002818) |
| 2.7 | 19.1 | GO:0032119 | sequestering of zinc ion(GO:0032119) |
| 1.9 | 5.8 | GO:0038178 | complement component C5a signaling pathway(GO:0038178) |
| 1.9 | 5.8 | GO:0035470 | positive regulation of vascular wound healing(GO:0035470) regulation of lactation(GO:1903487) |
| 1.9 | 11.4 | GO:1903435 | positive regulation of constitutive secretory pathway(GO:1903435) |
| 1.7 | 6.7 | GO:0005986 | sucrose biosynthetic process(GO:0005986) |
| 1.6 | 6.5 | GO:0002296 | T-helper 1 cell lineage commitment(GO:0002296) |
| 1.5 | 6.2 | GO:2000697 | negative regulation by virus of viral protein levels in host cell(GO:0046725) negative regulation of nephron tubule epithelial cell differentiation(GO:0072183) negative regulation of metanephric nephron tubule epithelial cell differentiation(GO:0072308) negative regulation of epithelial cell differentiation involved in kidney development(GO:2000697) |
| 1.5 | 4.6 | GO:0033364 | mast cell secretory granule organization(GO:0033364) |
| 1.5 | 4.6 | GO:0038043 | interleukin-5-mediated signaling pathway(GO:0038043) |
| 1.5 | 13.3 | GO:1903265 | positive regulation of tumor necrosis factor-mediated signaling pathway(GO:1903265) |
| 1.3 | 29.2 | GO:0044146 | negative regulation of growth of symbiont involved in interaction with host(GO:0044146) |
| 1.2 | 4.8 | GO:0060335 | positive regulation of response to interferon-gamma(GO:0060332) positive regulation of interferon-gamma-mediated signaling pathway(GO:0060335) |
| 1.2 | 21.4 | GO:0040015 | negative regulation of multicellular organism growth(GO:0040015) |
| 1.1 | 9.8 | GO:0002480 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-independent(GO:0002480) |
| 1.1 | 3.2 | GO:2000566 | positive regulation of CD8-positive, alpha-beta T cell proliferation(GO:2000566) |
| 1.1 | 3.2 | GO:0048294 | negative regulation of isotype switching to IgE isotypes(GO:0048294) |
| 1.0 | 22.7 | GO:1901741 | positive regulation of myoblast fusion(GO:1901741) |
| 1.0 | 4.0 | GO:0032690 | negative regulation of interleukin-1 alpha production(GO:0032690) negative regulation of interleukin-1 alpha secretion(GO:0050712) |
| 1.0 | 3.9 | GO:0003335 | corneocyte development(GO:0003335) |
| 1.0 | 2.9 | GO:0002581 | negative regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002581) |
| 0.9 | 1.9 | GO:1904100 | regulation of protein O-linked glycosylation(GO:1904098) positive regulation of protein O-linked glycosylation(GO:1904100) |
| 0.9 | 1.9 | GO:0070662 | mast cell proliferation(GO:0070662) |
| 0.8 | 2.5 | GO:0002949 | tRNA threonylcarbamoyladenosine modification(GO:0002949) |
| 0.8 | 4.2 | GO:0071350 | interleukin-15-mediated signaling pathway(GO:0035723) cellular response to interleukin-15(GO:0071350) |
| 0.8 | 4.2 | GO:1902023 | L-arginine import(GO:0043091) arginine import(GO:0090467) L-arginine transport(GO:1902023) |
| 0.8 | 6.1 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 0.7 | 4.5 | GO:0006196 | AMP catabolic process(GO:0006196) |
| 0.7 | 8.0 | GO:0043587 | tongue morphogenesis(GO:0043587) |
| 0.7 | 87.6 | GO:0031295 | T cell costimulation(GO:0031295) |
| 0.7 | 5.0 | GO:0033227 | dsRNA transport(GO:0033227) |
| 0.7 | 4.2 | GO:0022614 | membrane to membrane docking(GO:0022614) |
| 0.6 | 2.5 | GO:1903237 | negative regulation of leukocyte tethering or rolling(GO:1903237) |
| 0.6 | 40.9 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.6 | 3.1 | GO:0044856 | plasma membrane raft distribution(GO:0044855) plasma membrane raft localization(GO:0044856) plasma membrane raft polarization(GO:0044858) regulation of plasma membrane raft polarization(GO:1903906) |
| 0.6 | 3.6 | GO:0035624 | receptor transactivation(GO:0035624) |
| 0.6 | 15.9 | GO:0001580 | detection of chemical stimulus involved in sensory perception of bitter taste(GO:0001580) |
| 0.6 | 25.6 | GO:0045737 | positive regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045737) |
| 0.6 | 1.7 | GO:0071529 | cementum mineralization(GO:0071529) |
| 0.6 | 2.3 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.6 | 1.7 | GO:0003220 | left ventricular cardiac muscle tissue morphogenesis(GO:0003220) cell proliferation involved in heart valve morphogenesis(GO:0003249) regulation of cell proliferation involved in heart valve morphogenesis(GO:0003250) |
| 0.5 | 2.1 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.5 | 5.3 | GO:0010917 | negative regulation of mitochondrial membrane potential(GO:0010917) |
| 0.5 | 5.7 | GO:2000288 | positive regulation of myoblast proliferation(GO:2000288) |
| 0.5 | 8.5 | GO:1902894 | negative regulation of pri-miRNA transcription from RNA polymerase II promoter(GO:1902894) |
| 0.5 | 5.4 | GO:0003373 | dynamin polymerization involved in membrane fission(GO:0003373) dynamin polymerization involved in mitochondrial fission(GO:0003374) |
| 0.5 | 2.0 | GO:0061762 | CAMKK-AMPK signaling cascade(GO:0061762) |
| 0.5 | 8.8 | GO:0071243 | cellular response to arsenic-containing substance(GO:0071243) |
| 0.5 | 0.9 | GO:0002071 | glandular epithelial cell maturation(GO:0002071) type B pancreatic cell maturation(GO:0072560) |
| 0.5 | 1.4 | GO:0052250 | modulation of signal transduction in other organism(GO:0044501) modulation by symbiont of host signal transduction pathway(GO:0052027) modulation of signal transduction in other organism involved in symbiotic interaction(GO:0052250) modulation by symbiont of host I-kappaB kinase/NF-kappaB cascade(GO:0085032) |
| 0.5 | 4.6 | GO:1901223 | negative regulation of NIK/NF-kappaB signaling(GO:1901223) |
| 0.5 | 3.2 | GO:0043985 | histone H4-R3 methylation(GO:0043985) |
| 0.4 | 3.1 | GO:0060992 | response to fungicide(GO:0060992) |
| 0.4 | 1.3 | GO:0002894 | positive regulation of type IIa hypersensitivity(GO:0001798) positive regulation of type II hypersensitivity(GO:0002894) |
| 0.4 | 4.4 | GO:0015791 | polyol transport(GO:0015791) |
| 0.4 | 5.9 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
| 0.4 | 1.3 | GO:0014734 | skeletal muscle hypertrophy(GO:0014734) |
| 0.4 | 3.3 | GO:0032649 | regulation of interferon-gamma production(GO:0032649) |
| 0.4 | 2.0 | GO:0046449 | creatinine metabolic process(GO:0046449) |
| 0.4 | 1.2 | GO:0044793 | negative regulation by host of viral process(GO:0044793) |
| 0.4 | 4.0 | GO:0060287 | epithelial cilium movement involved in determination of left/right asymmetry(GO:0060287) |
| 0.4 | 3.9 | GO:0007000 | nucleolus organization(GO:0007000) |
| 0.4 | 2.7 | GO:0045163 | clustering of voltage-gated potassium channels(GO:0045163) |
| 0.4 | 3.1 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.4 | 2.3 | GO:0019532 | oxalate transport(GO:0019532) |
| 0.4 | 1.1 | GO:0098937 | dendritic transport(GO:0098935) anterograde dendritic transport(GO:0098937) |
| 0.4 | 2.6 | GO:0034316 | negative regulation of Arp2/3 complex-mediated actin nucleation(GO:0034316) |
| 0.4 | 34.6 | GO:0002479 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-dependent(GO:0002479) |
| 0.4 | 3.5 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.3 | 34.2 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.3 | 1.4 | GO:0035995 | detection of muscle stretch(GO:0035995) |
| 0.3 | 6.0 | GO:0034374 | low-density lipoprotein particle remodeling(GO:0034374) |
| 0.3 | 3.5 | GO:0060315 | negative regulation of ryanodine-sensitive calcium-release channel activity(GO:0060315) |
| 0.3 | 1.6 | GO:0051182 | coenzyme transport(GO:0051182) |
| 0.3 | 3.1 | GO:0015781 | pyrimidine nucleotide-sugar transport(GO:0015781) |
| 0.3 | 0.9 | GO:2000687 | negative regulation of rubidium ion transport(GO:2000681) negative regulation of rubidium ion transmembrane transporter activity(GO:2000687) |
| 0.3 | 2.8 | GO:0015747 | urate transport(GO:0015747) |
| 0.3 | 2.7 | GO:0035562 | negative regulation of chromatin binding(GO:0035562) |
| 0.3 | 3.2 | GO:1902018 | negative regulation of cilium assembly(GO:1902018) |
| 0.3 | 3.2 | GO:0043117 | positive regulation of vascular permeability(GO:0043117) |
| 0.3 | 0.9 | GO:0000472 | endonucleolytic cleavage to generate mature 5'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000472) rRNA 5'-end processing(GO:0000967) ncRNA 5'-end processing(GO:0034471) |
| 0.3 | 5.6 | GO:0034643 | establishment of mitochondrion localization, microtubule-mediated(GO:0034643) mitochondrion transport along microtubule(GO:0047497) |
| 0.3 | 3.3 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 0.3 | 0.8 | GO:2000724 | nuclear fragmentation involved in apoptotic nuclear change(GO:0030264) positive regulation of cardiac vascular smooth muscle cell differentiation(GO:2000724) |
| 0.3 | 0.5 | GO:1901624 | negative regulation of lymphocyte chemotaxis(GO:1901624) |
| 0.3 | 3.1 | GO:0010991 | negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.3 | 1.8 | GO:0042126 | nitrate metabolic process(GO:0042126) |
| 0.3 | 5.3 | GO:0050930 | induction of positive chemotaxis(GO:0050930) |
| 0.2 | 1.5 | GO:0051970 | negative regulation of transmission of nerve impulse(GO:0051970) |
| 0.2 | 2.5 | GO:0048263 | determination of dorsal identity(GO:0048263) |
| 0.2 | 1.0 | GO:0097411 | hypoxia-inducible factor-1alpha signaling pathway(GO:0097411) |
| 0.2 | 1.7 | GO:0032057 | negative regulation of translational initiation in response to stress(GO:0032057) |
| 0.2 | 1.6 | GO:0042492 | gamma-delta T cell differentiation(GO:0042492) |
| 0.2 | 1.2 | GO:0032383 | regulation of intracellular lipid transport(GO:0032377) regulation of intracellular sterol transport(GO:0032380) regulation of intracellular cholesterol transport(GO:0032383) |
| 0.2 | 7.3 | GO:2000401 | regulation of lymphocyte migration(GO:2000401) |
| 0.2 | 2.6 | GO:1905097 | regulation of guanyl-nucleotide exchange factor activity(GO:1905097) |
| 0.2 | 1.7 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.2 | 1.8 | GO:0030213 | hyaluronan biosynthetic process(GO:0030213) |
| 0.2 | 2.1 | GO:0098789 | pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.2 | 1.5 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.2 | 2.5 | GO:2000271 | positive regulation of fibroblast apoptotic process(GO:2000271) |
| 0.2 | 1.1 | GO:0014012 | peripheral nervous system axon regeneration(GO:0014012) |
| 0.2 | 1.1 | GO:0035871 | protein K11-linked deubiquitination(GO:0035871) |
| 0.2 | 0.9 | GO:1902714 | negative regulation of interferon-gamma secretion(GO:1902714) |
| 0.2 | 1.7 | GO:0006531 | aspartate metabolic process(GO:0006531) |
| 0.2 | 1.4 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 0.2 | 1.2 | GO:0001955 | blood vessel maturation(GO:0001955) |
| 0.2 | 14.2 | GO:0045576 | mast cell activation(GO:0045576) |
| 0.2 | 2.3 | GO:0090220 | meiotic telomere clustering(GO:0045141) chromosome localization to nuclear envelope involved in homologous chromosome segregation(GO:0090220) |
| 0.2 | 1.4 | GO:0006848 | pyruvate transport(GO:0006848) pyruvate transmembrane transport(GO:1901475) |
| 0.2 | 0.5 | GO:0061357 | positive regulation of Wnt protein secretion(GO:0061357) |
| 0.2 | 5.0 | GO:0035025 | positive regulation of Rho protein signal transduction(GO:0035025) |
| 0.2 | 5.0 | GO:0035590 | purinergic nucleotide receptor signaling pathway(GO:0035590) |
| 0.2 | 3.6 | GO:0051181 | cofactor transport(GO:0051181) |
| 0.2 | 11.0 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.2 | 1.7 | GO:0032516 | positive regulation of phosphoprotein phosphatase activity(GO:0032516) |
| 0.2 | 1.7 | GO:0032025 | response to cobalt ion(GO:0032025) |
| 0.1 | 1.2 | GO:0051660 | establishment of centrosome localization(GO:0051660) |
| 0.1 | 1.5 | GO:0003376 | sphingosine-1-phosphate signaling pathway(GO:0003376) sphingolipid mediated signaling pathway(GO:0090520) |
| 0.1 | 2.7 | GO:0007620 | copulation(GO:0007620) |
| 0.1 | 4.4 | GO:0001945 | lymph vessel development(GO:0001945) |
| 0.1 | 1.6 | GO:2001013 | epithelial cell proliferation involved in renal tubule morphogenesis(GO:2001013) |
| 0.1 | 1.0 | GO:1904153 | negative regulation of protein exit from endoplasmic reticulum(GO:0070862) negative regulation of retrograde protein transport, ER to cytosol(GO:1904153) |
| 0.1 | 18.8 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.1 | 0.4 | GO:1902630 | regulation of membrane hyperpolarization(GO:1902630) |
| 0.1 | 1.8 | GO:0006346 | methylation-dependent chromatin silencing(GO:0006346) |
| 0.1 | 3.2 | GO:0046339 | diacylglycerol metabolic process(GO:0046339) |
| 0.1 | 2.0 | GO:0016024 | CDP-diacylglycerol biosynthetic process(GO:0016024) |
| 0.1 | 0.9 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.1 | 1.4 | GO:2000346 | negative regulation of hepatocyte proliferation(GO:2000346) |
| 0.1 | 0.4 | GO:0015860 | pyrimidine nucleobase transport(GO:0015855) purine nucleoside transmembrane transport(GO:0015860) purine nucleobase transmembrane transport(GO:1904823) |
| 0.1 | 1.0 | GO:0050689 | negative regulation of defense response to virus by host(GO:0050689) |
| 0.1 | 0.9 | GO:0002088 | lens development in camera-type eye(GO:0002088) |
| 0.1 | 1.0 | GO:0006930 | substrate-dependent cell migration, cell extension(GO:0006930) |
| 0.1 | 2.6 | GO:0097150 | neuronal stem cell population maintenance(GO:0097150) |
| 0.1 | 1.7 | GO:2001275 | positive regulation of glucose import in response to insulin stimulus(GO:2001275) |
| 0.1 | 3.0 | GO:0030206 | chondroitin sulfate biosynthetic process(GO:0030206) |
| 0.1 | 1.9 | GO:1990118 | sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.1 | 4.6 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.1 | 0.9 | GO:1902219 | negative regulation of intrinsic apoptotic signaling pathway in response to osmotic stress(GO:1902219) |
| 0.1 | 0.2 | GO:0045626 | negative regulation of T-helper 1 cell differentiation(GO:0045626) |
| 0.1 | 0.8 | GO:0006307 | DNA dealkylation involved in DNA repair(GO:0006307) |
| 0.1 | 1.8 | GO:0090161 | Golgi ribbon formation(GO:0090161) |
| 0.1 | 0.2 | GO:0032911 | negative regulation of transforming growth factor beta1 production(GO:0032911) |
| 0.1 | 1.9 | GO:0071498 | cellular response to fluid shear stress(GO:0071498) |
| 0.1 | 0.5 | GO:0031642 | negative regulation of myelination(GO:0031642) |
| 0.1 | 11.6 | GO:0002181 | cytoplasmic translation(GO:0002181) |
| 0.1 | 1.2 | GO:0021554 | optic nerve development(GO:0021554) |
| 0.1 | 3.1 | GO:0007257 | activation of JUN kinase activity(GO:0007257) |
| 0.1 | 2.4 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 0.1 | 5.8 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.1 | 2.1 | GO:0030214 | hyaluronan catabolic process(GO:0030214) |
| 0.1 | 2.2 | GO:0051895 | negative regulation of focal adhesion assembly(GO:0051895) |
| 0.1 | 3.0 | GO:0043154 | negative regulation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0043154) |
| 0.1 | 0.9 | GO:0021859 | pyramidal neuron differentiation(GO:0021859) pyramidal neuron development(GO:0021860) |
| 0.1 | 0.3 | GO:0031999 | negative regulation of fatty acid beta-oxidation(GO:0031999) |
| 0.1 | 1.0 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.1 | 0.8 | GO:0046951 | ketone body biosynthetic process(GO:0046951) |
| 0.1 | 0.2 | GO:0045212 | negative regulation of synaptic transmission, cholinergic(GO:0032223) neurotransmitter receptor biosynthetic process(GO:0045212) |
| 0.1 | 3.6 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.1 | 3.1 | GO:0001510 | RNA methylation(GO:0001510) |
| 0.1 | 0.3 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.1 | 2.7 | GO:0006699 | bile acid biosynthetic process(GO:0006699) |
| 0.1 | 1.3 | GO:0008203 | cholesterol metabolic process(GO:0008203) |
| 0.1 | 4.3 | GO:0006968 | cellular defense response(GO:0006968) |
| 0.1 | 0.5 | GO:0045023 | G0 to G1 transition(GO:0045023) |
| 0.1 | 2.1 | GO:0007140 | male meiosis(GO:0007140) |
| 0.1 | 2.8 | GO:0016233 | telomere capping(GO:0016233) |
| 0.1 | 0.9 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.1 | 3.0 | GO:0002228 | natural killer cell mediated immunity(GO:0002228) |
| 0.1 | 4.0 | GO:0006501 | C-terminal protein lipidation(GO:0006501) |
| 0.1 | 1.3 | GO:0098743 | cell aggregation(GO:0098743) |
| 0.1 | 0.9 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.1 | 0.6 | GO:0006776 | vitamin A metabolic process(GO:0006776) |
| 0.1 | 5.8 | GO:0042157 | lipoprotein metabolic process(GO:0042157) |
| 0.1 | 2.8 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.1 | 0.9 | GO:0021984 | adenohypophysis development(GO:0021984) |
| 0.1 | 1.6 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 0.1 | 1.1 | GO:0019054 | modulation by virus of host process(GO:0019054) |
| 0.1 | 3.0 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.1 | 1.7 | GO:0070306 | lens fiber cell differentiation(GO:0070306) |
| 0.1 | 0.9 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.1 | 0.8 | GO:0099514 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.0 | 1.8 | GO:0006298 | mismatch repair(GO:0006298) |
| 0.0 | 3.3 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.0 | 0.7 | GO:0009437 | carnitine metabolic process(GO:0009437) |
| 0.0 | 3.3 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.0 | 1.2 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.0 | 1.3 | GO:0046039 | GTP metabolic process(GO:0046039) |
| 0.0 | 1.6 | GO:0006656 | phosphatidylcholine biosynthetic process(GO:0006656) |
| 0.0 | 1.0 | GO:0046676 | negative regulation of insulin secretion(GO:0046676) |
| 0.0 | 0.1 | GO:0002925 | positive regulation of humoral immune response mediated by circulating immunoglobulin(GO:0002925) |
| 0.0 | 2.5 | GO:0016579 | protein deubiquitination(GO:0016579) |
| 0.0 | 0.1 | GO:0051533 | positive regulation of NFAT protein import into nucleus(GO:0051533) |
| 0.0 | 0.3 | GO:0009249 | protein lipoylation(GO:0009249) |
| 0.0 | 2.9 | GO:0018107 | peptidyl-threonine phosphorylation(GO:0018107) |
| 0.0 | 0.3 | GO:0045793 | positive regulation of cell size(GO:0045793) |
| 0.0 | 0.7 | GO:0006337 | nucleosome disassembly(GO:0006337) |
| 0.0 | 1.9 | GO:0006826 | iron ion transport(GO:0006826) |
| 0.0 | 0.2 | GO:0006682 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 0.0 | 0.2 | GO:0009642 | response to light intensity(GO:0009642) |
| 0.0 | 1.2 | GO:0002456 | T cell mediated immunity(GO:0002456) |
| 0.0 | 0.6 | GO:0060334 | regulation of interferon-gamma-mediated signaling pathway(GO:0060334) |
| 0.0 | 0.8 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.0 | 1.4 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.0 | 0.7 | GO:0019433 | triglyceride catabolic process(GO:0019433) |
| 0.0 | 1.4 | GO:0006446 | regulation of translational initiation(GO:0006446) |
| 0.0 | 1.7 | GO:0090630 | activation of GTPase activity(GO:0090630) |
| 0.0 | 0.1 | GO:0055129 | proline biosynthetic process(GO:0006561) L-proline biosynthetic process(GO:0055129) |
| 0.0 | 1.6 | GO:0010811 | positive regulation of cell-substrate adhesion(GO:0010811) |
| 0.0 | 0.3 | GO:0090360 | platelet-derived growth factor production(GO:0090360) regulation of platelet-derived growth factor production(GO:0090361) |
| 0.0 | 0.2 | GO:0060285 | cilium-dependent cell motility(GO:0060285) |
| 0.0 | 0.4 | GO:0000042 | protein targeting to Golgi(GO:0000042) |
| 0.0 | 2.1 | GO:0000209 | protein polyubiquitination(GO:0000209) |
| 0.0 | 0.6 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.0 | 0.1 | GO:0033353 | S-adenosylmethionine cycle(GO:0033353) |
| 0.0 | 0.9 | GO:0030574 | collagen catabolic process(GO:0030574) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.2 | 108.4 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 4.7 | 33.1 | GO:0097013 | phagocytic vesicle lumen(GO:0097013) |
| 3.9 | 11.7 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 3.9 | 31.0 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 1.7 | 10.1 | GO:0072557 | IPAF inflammasome complex(GO:0072557) |
| 1.5 | 6.1 | GO:1990031 | pinceau fiber(GO:1990031) |
| 1.4 | 6.8 | GO:0036398 | TCR signalosome(GO:0036398) |
| 1.2 | 3.5 | GO:0060053 | neurofilament cytoskeleton(GO:0060053) |
| 1.0 | 11.4 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 0.9 | 9.8 | GO:0042611 | MHC protein complex(GO:0042611) |
| 0.8 | 41.5 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.6 | 11.9 | GO:0031229 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) nuclear membrane part(GO:0044453) |
| 0.5 | 30.7 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.5 | 2.5 | GO:0000408 | EKC/KEOPS complex(GO:0000408) |
| 0.5 | 4.3 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.4 | 1.3 | GO:0097057 | TRAF2-GSTP1 complex(GO:0097057) |
| 0.4 | 2.6 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.4 | 26.1 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.4 | 1.7 | GO:0043293 | apoptosome(GO:0043293) |
| 0.4 | 4.6 | GO:0005664 | origin recognition complex(GO:0000808) nuclear origin of replication recognition complex(GO:0005664) |
| 0.3 | 1.7 | GO:0071546 | pi-body(GO:0071546) |
| 0.3 | 6.5 | GO:0005849 | mRNA cleavage factor complex(GO:0005849) |
| 0.3 | 1.7 | GO:0065010 | extracellular membrane-bounded organelle(GO:0065010) |
| 0.3 | 1.7 | GO:0032444 | activin responsive factor complex(GO:0032444) |
| 0.3 | 7.2 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.3 | 11.2 | GO:0045095 | keratin filament(GO:0045095) |
| 0.3 | 28.3 | GO:0035578 | azurophil granule lumen(GO:0035578) |
| 0.2 | 5.5 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.2 | 1.0 | GO:0005967 | mitochondrial pyruvate dehydrogenase complex(GO:0005967) |
| 0.2 | 1.5 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.2 | 13.1 | GO:0016235 | aggresome(GO:0016235) |
| 0.2 | 3.1 | GO:0030478 | actin cap(GO:0030478) |
| 0.2 | 4.2 | GO:0031254 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 0.2 | 1.7 | GO:0032389 | MutLalpha complex(GO:0032389) |
| 0.2 | 1.7 | GO:0005851 | eukaryotic translation initiation factor 2B complex(GO:0005851) |
| 0.2 | 6.4 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.2 | 2.8 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.2 | 7.3 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.2 | 4.0 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.2 | 2.0 | GO:0097433 | dense body(GO:0097433) |
| 0.2 | 1.9 | GO:0030123 | AP-3 adaptor complex(GO:0030123) |
| 0.2 | 5.2 | GO:0042629 | mast cell granule(GO:0042629) |
| 0.1 | 17.3 | GO:0045111 | intermediate filament cytoskeleton(GO:0045111) |
| 0.1 | 0.9 | GO:0016600 | flotillin complex(GO:0016600) |
| 0.1 | 4.6 | GO:0000777 | condensed chromosome kinetochore(GO:0000777) |
| 0.1 | 2.8 | GO:0005861 | troponin complex(GO:0005861) |
| 0.1 | 0.4 | GO:1990876 | cytoplasmic side of nuclear pore(GO:1990876) |
| 0.1 | 13.1 | GO:0035579 | specific granule membrane(GO:0035579) |
| 0.1 | 10.9 | GO:0005814 | centriole(GO:0005814) |
| 0.1 | 2.2 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.1 | 0.3 | GO:0048269 | methionine adenosyltransferase complex(GO:0048269) |
| 0.1 | 2.6 | GO:0070971 | endoplasmic reticulum exit site(GO:0070971) |
| 0.1 | 10.7 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.1 | 1.2 | GO:0043190 | ATP-binding cassette (ABC) transporter complex(GO:0043190) |
| 0.1 | 1.1 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.1 | 23.6 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.1 | 1.8 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.1 | 1.3 | GO:0000177 | cytoplasmic exosome (RNase complex)(GO:0000177) |
| 0.1 | 8.9 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.1 | 1.3 | GO:0033268 | node of Ranvier(GO:0033268) |
| 0.1 | 4.5 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 4.4 | GO:0035097 | histone methyltransferase complex(GO:0035097) |
| 0.0 | 4.4 | GO:0030175 | filopodium(GO:0030175) |
| 0.0 | 3.1 | GO:0005657 | replication fork(GO:0005657) |
| 0.0 | 3.5 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.0 | 0.6 | GO:0031011 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.0 | 2.6 | GO:0015030 | Cajal body(GO:0015030) |
| 0.0 | 3.4 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.0 | 0.7 | GO:0016514 | SWI/SNF complex(GO:0016514) |
| 0.0 | 1.8 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 0.8 | GO:0031105 | septin complex(GO:0031105) |
| 0.0 | 2.2 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.0 | 7.1 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 1.9 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.0 | 1.8 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.0 | 3.5 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 1.0 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 0.5 | GO:0032590 | dendrite membrane(GO:0032590) |
| 0.0 | 2.1 | GO:0034705 | voltage-gated potassium channel complex(GO:0008076) potassium channel complex(GO:0034705) |
| 0.0 | 2.4 | GO:0098802 | plasma membrane receptor complex(GO:0098802) |
| 0.0 | 0.9 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.0 | 1.0 | GO:0030672 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.0 | 0.9 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 0.9 | GO:0035580 | specific granule lumen(GO:0035580) |
| 0.0 | 0.6 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.4 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.6 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.0 | 2.3 | GO:0005681 | spliceosomal complex(GO:0005681) |
| 0.0 | 3.5 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.0 | 0.7 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.0 | 2.6 | GO:0000790 | nuclear chromatin(GO:0000790) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.0 | 18.0 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 5.2 | 72.6 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 5.0 | 14.9 | GO:0008476 | protein-tyrosine sulfotransferase activity(GO:0008476) |
| 2.7 | 8.2 | GO:0050613 | delta14-sterol reductase activity(GO:0050613) |
| 2.3 | 11.5 | GO:0023024 | MHC class I protein complex binding(GO:0023024) |
| 1.9 | 5.8 | GO:0004878 | complement component C5a receptor activity(GO:0004878) |
| 1.7 | 6.7 | GO:0042132 | fructose 1,6-bisphosphate 1-phosphatase activity(GO:0042132) |
| 1.5 | 4.6 | GO:0004914 | interleukin-5 receptor activity(GO:0004914) |
| 1.5 | 45.6 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 1.4 | 4.3 | GO:0015226 | amino-acid betaine transmembrane transporter activity(GO:0015199) carnitine transmembrane transporter activity(GO:0015226) |
| 1.4 | 4.2 | GO:0051139 | metal ion:proton antiporter activity(GO:0051139) |
| 1.4 | 5.5 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 1.3 | 3.9 | GO:0001181 | transcription factor activity, core RNA polymerase I binding(GO:0001181) |
| 1.3 | 6.3 | GO:0052740 | 1-acyl-2-lysophosphatidylserine acylhydrolase activity(GO:0052740) |
| 1.1 | 6.5 | GO:0070892 | lipoteichoic acid receptor activity(GO:0070892) |
| 1.0 | 31.0 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.9 | 3.7 | GO:0004140 | dephospho-CoA kinase activity(GO:0004140) |
| 0.9 | 12.0 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.8 | 5.0 | GO:0051033 | nucleic acid transmembrane transporter activity(GO:0051032) RNA transmembrane transporter activity(GO:0051033) |
| 0.8 | 3.2 | GO:0004727 | prenylated protein tyrosine phosphatase activity(GO:0004727) |
| 0.8 | 4.0 | GO:0004910 | interleukin-1, Type II, blocking receptor activity(GO:0004910) |
| 0.8 | 2.3 | GO:0061505 | DNA topoisomerase type II (ATP-hydrolyzing) activity(GO:0003918) DNA topoisomerase II activity(GO:0061505) |
| 0.8 | 41.5 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.7 | 2.9 | GO:0005502 | 11-cis retinal binding(GO:0005502) |
| 0.6 | 1.9 | GO:0030350 | iron-responsive element binding(GO:0030350) |
| 0.6 | 2.5 | GO:0031735 | CCR10 chemokine receptor binding(GO:0031735) |
| 0.6 | 3.1 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.6 | 1.8 | GO:0052596 | tryptamine:oxygen oxidoreductase (deaminating) activity(GO:0052593) aminoacetone:oxygen oxidoreductase(deaminating) activity(GO:0052594) aliphatic-amine oxidase activity(GO:0052595) phenethylamine:oxygen oxidoreductase (deaminating) activity(GO:0052596) |
| 0.6 | 5.0 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.6 | 1.7 | GO:0004515 | nicotinate-nucleotide adenylyltransferase activity(GO:0004515) |
| 0.5 | 4.4 | GO:0015166 | polyol transmembrane transporter activity(GO:0015166) |
| 0.5 | 6.0 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.5 | 31.9 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.5 | 5.3 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.5 | 1.6 | GO:0051185 | coenzyme transporter activity(GO:0051185) |
| 0.5 | 3.0 | GO:0050656 | 3'-phosphoadenosine 5'-phosphosulfate binding(GO:0050656) |
| 0.5 | 4.5 | GO:0047623 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.5 | 9.3 | GO:0010857 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) calcium-dependent protein kinase activity(GO:0010857) |
| 0.5 | 16.6 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.5 | 1.4 | GO:0016434 | rRNA (cytosine) methyltransferase activity(GO:0016434) |
| 0.5 | 2.3 | GO:0015111 | iodide transmembrane transporter activity(GO:0015111) |
| 0.4 | 1.8 | GO:0008940 | nitrate reductase activity(GO:0008940) |
| 0.4 | 11.4 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.4 | 1.3 | GO:0005174 | CD40 receptor binding(GO:0005174) |
| 0.4 | 3.1 | GO:0016929 | SUMO-specific protease activity(GO:0016929) |
| 0.4 | 3.6 | GO:0004931 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.3 | 6.2 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.3 | 1.4 | GO:0005477 | pyruvate secondary active transmembrane transporter activity(GO:0005477) |
| 0.3 | 6.6 | GO:0008320 | protein transmembrane transporter activity(GO:0008320) |
| 0.3 | 4.8 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.3 | 2.7 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.3 | 2.6 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.3 | 1.9 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.3 | 2.4 | GO:0015165 | pyrimidine nucleotide-sugar transmembrane transporter activity(GO:0015165) |
| 0.3 | 1.5 | GO:0016427 | tRNA (cytosine) methyltransferase activity(GO:0016427) |
| 0.3 | 0.9 | GO:0005330 | dopamine:sodium symporter activity(GO:0005330) |
| 0.3 | 2.8 | GO:0015143 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.3 | 0.8 | GO:0017082 | mineralocorticoid receptor activity(GO:0017082) |
| 0.3 | 3.5 | GO:0030957 | Tat protein binding(GO:0030957) |
| 0.3 | 0.8 | GO:0003858 | 3-hydroxybutyrate dehydrogenase activity(GO:0003858) |
| 0.3 | 4.2 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.3 | 3.1 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.3 | 0.5 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) |
| 0.3 | 4.6 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.3 | 5.3 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 0.2 | 1.7 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.2 | 1.5 | GO:0045125 | bioactive lipid receptor activity(GO:0045125) |
| 0.2 | 1.0 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.2 | 10.0 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.2 | 7.6 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.2 | 1.8 | GO:0003696 | satellite DNA binding(GO:0003696) |
| 0.2 | 8.8 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.2 | 6.9 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.2 | 9.3 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.2 | 3.2 | GO:0001161 | intronic transcription regulatory region sequence-specific DNA binding(GO:0001161) intronic transcription regulatory region DNA binding(GO:0044213) |
| 0.2 | 3.6 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.2 | 1.5 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.2 | 8.0 | GO:0000983 | transcription factor activity, RNA polymerase II core promoter sequence-specific(GO:0000983) |
| 0.2 | 15.9 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.2 | 7.2 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.2 | 1.9 | GO:0015385 | sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.2 | 0.9 | GO:0005152 | interleukin-1 receptor antagonist activity(GO:0005152) |
| 0.2 | 6.1 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 0.2 | 2.2 | GO:0043225 | anion transmembrane-transporting ATPase activity(GO:0043225) |
| 0.2 | 3.9 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.2 | 2.1 | GO:0043176 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.2 | 1.9 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.2 | 1.9 | GO:0003951 | NAD+ kinase activity(GO:0003951) |
| 0.2 | 29.5 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.2 | 0.9 | GO:0016402 | pristanoyl-CoA oxidase activity(GO:0016402) |
| 0.2 | 0.7 | GO:0032558 | adenyl deoxyribonucleotide binding(GO:0032558) dATP binding(GO:0032564) |
| 0.2 | 21.5 | GO:0003823 | antigen binding(GO:0003823) |
| 0.2 | 1.6 | GO:0019784 | NEDD8-specific protease activity(GO:0019784) |
| 0.2 | 1.9 | GO:0004716 | receptor signaling protein tyrosine kinase activity(GO:0004716) |
| 0.2 | 1.7 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.2 | 1.2 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.1 | 1.1 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.1 | 1.1 | GO:0005497 | androgen binding(GO:0005497) |
| 0.1 | 1.4 | GO:0031433 | telethonin binding(GO:0031433) |
| 0.1 | 1.7 | GO:0043199 | sulfate binding(GO:0043199) |
| 0.1 | 2.9 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.1 | 1.5 | GO:0016494 | C-X-C chemokine receptor activity(GO:0016494) |
| 0.1 | 3.6 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.1 | 0.4 | GO:0005350 | pyrimidine nucleobase transmembrane transporter activity(GO:0005350) pyrimidine- and adenine-specific:sodium symporter activity(GO:0015389) |
| 0.1 | 1.3 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 0.1 | 1.0 | GO:0046703 | natural killer cell lectin-like receptor binding(GO:0046703) |
| 0.1 | 3.5 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.1 | 0.6 | GO:0004906 | interferon-gamma receptor activity(GO:0004906) |
| 0.1 | 0.8 | GO:0009008 | DNA-methyltransferase activity(GO:0009008) |
| 0.1 | 1.1 | GO:0004579 | dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.1 | 1.1 | GO:1990380 | Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.1 | 0.5 | GO:0004668 | protein-arginine deiminase activity(GO:0004668) |
| 0.1 | 3.1 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.1 | 10.2 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.1 | 0.1 | GO:0001025 | RNA polymerase III transcription factor binding(GO:0001025) |
| 0.1 | 3.2 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.1 | 0.6 | GO:0004993 | G-protein coupled serotonin receptor activity(GO:0004993) |
| 0.1 | 1.7 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.1 | 1.6 | GO:0016493 | C-C chemokine receptor activity(GO:0016493) |
| 0.1 | 2.4 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.1 | 1.0 | GO:0070403 | NAD+ binding(GO:0070403) |
| 0.1 | 6.3 | GO:0000979 | RNA polymerase II core promoter sequence-specific DNA binding(GO:0000979) |
| 0.1 | 0.2 | GO:0047275 | glucosaminylgalactosylglucosylceramide beta-galactosyltransferase activity(GO:0047275) |
| 0.1 | 1.3 | GO:0001608 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.1 | 2.3 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.1 | 1.8 | GO:0033558 | histone deacetylase activity(GO:0004407) protein deacetylase activity(GO:0033558) |
| 0.1 | 3.4 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.1 | 1.6 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.1 | 0.4 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
| 0.1 | 7.2 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.1 | 1.1 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.1 | 5.8 | GO:0001158 | enhancer sequence-specific DNA binding(GO:0001158) |
| 0.1 | 0.3 | GO:0003989 | acetyl-CoA carboxylase activity(GO:0003989) |
| 0.1 | 2.3 | GO:0070717 | poly-purine tract binding(GO:0070717) |
| 0.1 | 0.2 | GO:0003990 | acetylcholinesterase activity(GO:0003990) |
| 0.1 | 1.1 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.1 | 1.2 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.1 | 1.6 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.1 | 1.4 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 3.7 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 2.8 | GO:0004004 | ATP-dependent RNA helicase activity(GO:0004004) |
| 0.0 | 3.0 | GO:0046934 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) |
| 0.0 | 4.2 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.0 | 0.9 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) |
| 0.0 | 1.5 | GO:0008395 | steroid hydroxylase activity(GO:0008395) |
| 0.0 | 2.0 | GO:0016836 | hydro-lyase activity(GO:0016836) |
| 0.0 | 3.8 | GO:0031072 | heat shock protein binding(GO:0031072) |
| 0.0 | 8.0 | GO:0008201 | heparin binding(GO:0008201) |
| 0.0 | 1.5 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.0 | 1.6 | GO:0008173 | RNA methyltransferase activity(GO:0008173) |
| 0.0 | 1.4 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.0 | 1.2 | GO:0008009 | chemokine activity(GO:0008009) |
| 0.0 | 0.9 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 2.7 | GO:0004386 | helicase activity(GO:0004386) |
| 0.0 | 1.9 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 2.7 | GO:0004536 | deoxyribonuclease activity(GO:0004536) |
| 0.0 | 4.4 | GO:0003713 | transcription coactivator activity(GO:0003713) |
| 0.0 | 0.7 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 1.8 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) |
| 0.0 | 1.1 | GO:0043621 | protein self-association(GO:0043621) |
| 0.0 | 3.4 | GO:0005516 | calmodulin binding(GO:0005516) |
| 0.0 | 0.3 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.0 | 0.9 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.2 | GO:0048406 | nerve growth factor binding(GO:0048406) |
| 0.0 | 0.8 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.0 | 1.0 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 0.2 | GO:0070492 | oligosaccharide binding(GO:0070492) |
| 0.0 | 2.5 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 0.8 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.0 | 0.9 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.0 | 0.2 | GO:0005537 | mannose binding(GO:0005537) |
| 0.0 | 4.7 | GO:0000978 | RNA polymerase II core promoter proximal region sequence-specific DNA binding(GO:0000978) |
| 0.0 | 1.5 | GO:0004896 | cytokine receptor activity(GO:0004896) |
| 0.0 | 2.7 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.1 | GO:0016802 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.0 | 0.3 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.0 | 0.1 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 3.0 | GO:0005085 | guanyl-nucleotide exchange factor activity(GO:0005085) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 12.2 | ST INTERFERON GAMMA PATHWAY | Interferon gamma pathway. |
| 0.6 | 14.8 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.6 | 33.5 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.5 | 4.6 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.5 | 23.7 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.4 | 11.9 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.4 | 9.2 | ST INTERLEUKIN 4 PATHWAY | Interleukin 4 (IL-4) Pathway |
| 0.4 | 10.9 | PID GMCSF PATHWAY | GMCSF-mediated signaling events |
| 0.4 | 15.8 | PID CD8 TCR PATHWAY | TCR signaling in naïve CD8+ T cells |
| 0.3 | 5.7 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.3 | 0.9 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.3 | 3.2 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.3 | 4.4 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 0.2 | 9.3 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.2 | 2.3 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.2 | 20.1 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.2 | 1.1 | PID PI3KCI AKT PATHWAY | Class I PI3K signaling events mediated by Akt |
| 0.2 | 3.8 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.2 | 4.3 | SIG BCR SIGNALING PATHWAY | Members of the BCR signaling pathway |
| 0.1 | 1.3 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.1 | 7.3 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.1 | 10.5 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.1 | 4.6 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.1 | 3.1 | PID AVB3 OPN PATHWAY | Osteopontin-mediated events |
| 0.1 | 6.5 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.1 | 1.5 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 0.1 | 4.0 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.1 | 2.8 | PID ER NONGENOMIC PATHWAY | Plasma membrane estrogen receptor signaling |
| 0.1 | 3.9 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.1 | 5.9 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.1 | 4.1 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.1 | 3.3 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.1 | 4.8 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.1 | 1.7 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.1 | 1.3 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.1 | 1.7 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.1 | 5.7 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.1 | 1.9 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.1 | 6.3 | PID E2F PATHWAY | E2F transcription factor network |
| 0.1 | 3.9 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.1 | 14.6 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.1 | 2.5 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.1 | 2.0 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.0 | 2.7 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.0 | 1.4 | PID ATM PATHWAY | ATM pathway |
| 0.0 | 2.4 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 2.7 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 1.8 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.0 | 1.2 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 0.9 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 1.4 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 0.4 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.0 | 0.9 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.8 | 111.0 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 1.0 | 9.8 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 0.8 | 5.3 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.7 | 28.7 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 0.5 | 51.3 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.5 | 3.6 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.5 | 14.8 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.4 | 32.5 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 0.4 | 11.7 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.4 | 5.3 | REACTOME ACYL CHAIN REMODELLING OF PG | Genes involved in Acyl chain remodelling of PG |
| 0.3 | 6.2 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.3 | 2.7 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 0.3 | 12.9 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.2 | 3.6 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.2 | 5.9 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.2 | 6.8 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.2 | 17.1 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.2 | 12.7 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.2 | 4.3 | REACTOME HYALURONAN METABOLISM | Genes involved in Hyaluronan metabolism |
| 0.2 | 4.2 | REACTOME DESTABILIZATION OF MRNA BY AUF1 HNRNP D0 | Genes involved in Destabilization of mRNA by AUF1 (hnRNP D0) |
| 0.2 | 3.4 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.2 | 7.3 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.2 | 2.1 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.2 | 3.0 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.2 | 6.5 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.1 | 4.6 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.1 | 4.0 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 2.7 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.1 | 2.0 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.1 | 1.9 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.1 | 2.4 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.1 | 1.8 | REACTOME RNA POL I PROMOTER OPENING | Genes involved in RNA Polymerase I Promoter Opening |
| 0.1 | 4.3 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.1 | 5.5 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 3.3 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.1 | 18.6 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.1 | 3.1 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.1 | 3.0 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.1 | 4.2 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |
| 0.1 | 17.5 | REACTOME 3 UTR MEDIATED TRANSLATIONAL REGULATION | Genes involved in 3' -UTR-mediated translational regulation |
| 0.1 | 1.4 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.1 | 1.6 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.1 | 1.4 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.1 | 1.3 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 0.1 | 3.2 | REACTOME P75 NTR RECEPTOR MEDIATED SIGNALLING | Genes involved in p75 NTR receptor-mediated signalling |
| 0.1 | 1.6 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.1 | 3.8 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.1 | 0.8 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.1 | 2.6 | REACTOME RNA POL I TRANSCRIPTION INITIATION | Genes involved in RNA Polymerase I Transcription Initiation |
| 0.1 | 0.9 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.1 | 1.0 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.1 | 1.0 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.1 | 1.7 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.1 | 2.8 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.1 | 1.8 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.1 | 0.8 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.1 | 1.0 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.1 | 1.3 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.1 | 1.3 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.0 | 2.4 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.0 | 2.0 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 0.9 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.0 | 1.4 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.0 | 1.2 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 2.3 | REACTOME TRANSLATION | Genes involved in Translation |
| 0.0 | 4.1 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.0 | 0.9 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.0 | 0.9 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 0.9 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.0 | 1.8 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.0 | 1.7 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 2.8 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.0 | 3.3 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 0.6 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.0 | 0.1 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.0 | 0.4 | REACTOME REGULATION OF MRNA STABILITY BY PROTEINS THAT BIND AU RICH ELEMENTS | Genes involved in Regulation of mRNA Stability by Proteins that Bind AU-rich Elements |