Project
GNF SymAtlas + NCI-60 cancer cell lines, human (Su, 2004; Ross, 2000)
Navigation
Downloads
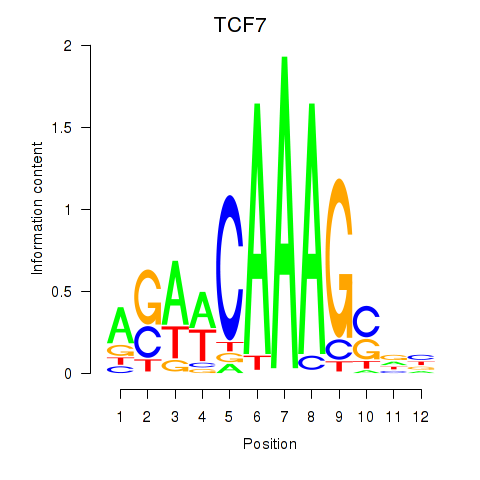
Results for TCF7
Z-value: 0.84
Transcription factors associated with TCF7
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
TCF7
|
ENSG00000081059.15 | transcription factor 7 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| TCF7 | hg19_v2_chr5_+_133477868_133477885 | -0.17 | 1.5e-02 | Click! |
Activity profile of TCF7 motif
Sorted Z-values of TCF7 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_151344172 | 28.07 |
ENST00000375734.2
ENST00000263895.4 ENST00000454202.1 |
RND3
|
Rho family GTPase 3 |
| chr5_+_34757309 | 20.05 |
ENST00000397449.1
|
RAI14
|
retinoic acid induced 14 |
| chr10_+_54074033 | 15.66 |
ENST00000373970.3
|
DKK1
|
dickkopf WNT signaling pathway inhibitor 1 |
| chr1_+_68150744 | 14.49 |
ENST00000370986.4
ENST00000370985.3 |
GADD45A
|
growth arrest and DNA-damage-inducible, alpha |
| chr22_-_42336209 | 12.48 |
ENST00000472374.2
|
CENPM
|
centromere protein M |
| chr7_-_107643674 | 11.87 |
ENST00000222399.6
|
LAMB1
|
laminin, beta 1 |
| chr12_+_69080734 | 11.40 |
ENST00000378905.2
|
NUP107
|
nucleoporin 107kDa |
| chr12_+_6309963 | 10.74 |
ENST00000382515.2
|
CD9
|
CD9 molecule |
| chr11_-_64013288 | 10.42 |
ENST00000542235.1
|
PPP1R14B
|
protein phosphatase 1, regulatory (inhibitor) subunit 14B |
| chr11_-_64013663 | 9.52 |
ENST00000392210.2
|
PPP1R14B
|
protein phosphatase 1, regulatory (inhibitor) subunit 14B |
| chr18_+_3449821 | 9.36 |
ENST00000407501.2
ENST00000405385.3 ENST00000546979.1 |
TGIF1
|
TGFB-induced factor homeobox 1 |
| chr5_+_135364584 | 9.16 |
ENST00000442011.2
ENST00000305126.8 |
TGFBI
|
transforming growth factor, beta-induced, 68kDa |
| chr15_+_74218787 | 8.85 |
ENST00000261921.7
|
LOXL1
|
lysyl oxidase-like 1 |
| chr5_+_32531893 | 8.64 |
ENST00000512913.1
|
SUB1
|
SUB1 homolog (S. cerevisiae) |
| chr12_-_54653313 | 8.40 |
ENST00000550411.1
ENST00000439541.2 |
CBX5
|
chromobox homolog 5 |
| chr2_+_234621551 | 8.02 |
ENST00000608381.1
ENST00000373414.3 |
UGT1A1
UGT1A5
|
UDP glucuronosyltransferase 1 family, polypeptide A8 UDP glucuronosyltransferase 1 family, polypeptide A5 |
| chrX_-_153599578 | 8.01 |
ENST00000360319.4
ENST00000344736.4 |
FLNA
|
filamin A, alpha |
| chr12_+_21654714 | 7.74 |
ENST00000542038.1
ENST00000540141.1 ENST00000229314.5 |
GOLT1B
|
golgi transport 1B |
| chr1_-_75198681 | 7.72 |
ENST00000370872.3
ENST00000370871.3 ENST00000340866.5 ENST00000370870.1 |
CRYZ
|
crystallin, zeta (quinone reductase) |
| chr1_-_75198940 | 7.56 |
ENST00000417775.1
|
CRYZ
|
crystallin, zeta (quinone reductase) |
| chr7_+_94023873 | 7.28 |
ENST00000297268.6
|
COL1A2
|
collagen, type I, alpha 2 |
| chr6_+_83073334 | 7.17 |
ENST00000369750.3
|
TPBG
|
trophoblast glycoprotein |
| chr12_+_69979446 | 7.06 |
ENST00000543146.2
|
CCT2
|
chaperonin containing TCP1, subunit 2 (beta) |
| chr7_+_134576317 | 6.88 |
ENST00000424922.1
ENST00000495522.1 |
CALD1
|
caldesmon 1 |
| chr1_-_209824643 | 6.88 |
ENST00000391911.1
ENST00000415782.1 |
LAMB3
|
laminin, beta 3 |
| chr4_-_139163491 | 6.83 |
ENST00000280612.5
|
SLC7A11
|
solute carrier family 7 (anionic amino acid transporter light chain, xc- system), member 11 |
| chr7_+_130131907 | 6.77 |
ENST00000223215.4
ENST00000437945.1 |
MEST
|
mesoderm specific transcript |
| chr7_+_73868439 | 6.51 |
ENST00000424337.2
|
GTF2IRD1
|
GTF2I repeat domain containing 1 |
| chr9_-_35691017 | 6.50 |
ENST00000378292.3
|
TPM2
|
tropomyosin 2 (beta) |
| chr7_+_134576151 | 6.44 |
ENST00000393118.2
|
CALD1
|
caldesmon 1 |
| chr1_-_33502528 | 6.39 |
ENST00000354858.6
|
AK2
|
adenylate kinase 2 |
| chr14_-_71107921 | 6.08 |
ENST00000553982.1
ENST00000500016.1 |
CTD-2540L5.5
CTD-2540L5.6
|
CTD-2540L5.5 CTD-2540L5.6 |
| chr7_-_80548667 | 5.49 |
ENST00000265361.3
|
SEMA3C
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3C |
| chr12_+_121131970 | 5.48 |
ENST00000535656.1
|
MLEC
|
malectin |
| chr17_-_18266660 | 5.43 |
ENST00000582653.1
ENST00000352886.6 |
SHMT1
|
serine hydroxymethyltransferase 1 (soluble) |
| chr8_+_30244580 | 5.42 |
ENST00000523115.1
ENST00000519647.1 |
RBPMS
|
RNA binding protein with multiple splicing |
| chr3_+_52719936 | 5.34 |
ENST00000418458.1
ENST00000394799.2 |
GNL3
|
guanine nucleotide binding protein-like 3 (nucleolar) |
| chr1_+_10459111 | 5.23 |
ENST00000541529.1
ENST00000270776.8 ENST00000483936.1 ENST00000538557.1 |
PGD
|
phosphogluconate dehydrogenase |
| chr10_+_17271266 | 5.11 |
ENST00000224237.5
|
VIM
|
vimentin |
| chr6_+_64281906 | 5.10 |
ENST00000370651.3
|
PTP4A1
|
protein tyrosine phosphatase type IVA, member 1 |
| chr6_+_83073952 | 5.09 |
ENST00000543496.1
|
TPBG
|
trophoblast glycoprotein |
| chr4_-_71705027 | 5.07 |
ENST00000545193.1
|
GRSF1
|
G-rich RNA sequence binding factor 1 |
| chr5_-_16936340 | 5.07 |
ENST00000507288.1
ENST00000513610.1 |
MYO10
|
myosin X |
| chr4_-_7069760 | 4.96 |
ENST00000264954.4
|
GRPEL1
|
GrpE-like 1, mitochondrial (E. coli) |
| chr3_+_159570722 | 4.96 |
ENST00000482804.1
|
SCHIP1
|
schwannomin interacting protein 1 |
| chr3_+_133502877 | 4.96 |
ENST00000466490.2
|
SRPRB
|
signal recognition particle receptor, B subunit |
| chr17_-_79818354 | 4.94 |
ENST00000576541.1
ENST00000576380.1 ENST00000571617.1 ENST00000576052.1 ENST00000576390.1 ENST00000573778.2 ENST00000439918.2 ENST00000574914.1 ENST00000331483.4 |
P4HB
|
prolyl 4-hydroxylase, beta polypeptide |
| chr7_-_27213893 | 4.93 |
ENST00000283921.4
|
HOXA10
|
homeobox A10 |
| chr7_-_16840820 | 4.90 |
ENST00000450569.1
|
AGR2
|
anterior gradient 2 |
| chr7_+_56119323 | 4.88 |
ENST00000275603.4
ENST00000335503.3 ENST00000540286.1 |
CCT6A
|
chaperonin containing TCP1, subunit 6A (zeta 1) |
| chr19_+_1026566 | 4.87 |
ENST00000348419.3
ENST00000565096.2 ENST00000562958.2 ENST00000562075.2 ENST00000607102.1 |
CNN2
|
calponin 2 |
| chr17_+_34848049 | 4.82 |
ENST00000588902.1
ENST00000591067.1 |
ZNHIT3
|
zinc finger, HIT-type containing 3 |
| chr6_+_160183492 | 4.81 |
ENST00000541436.1
|
ACAT2
|
acetyl-CoA acetyltransferase 2 |
| chr12_+_104359576 | 4.80 |
ENST00000392872.3
ENST00000436021.2 |
TDG
|
thymine-DNA glycosylase |
| chr16_-_73082274 | 4.76 |
ENST00000268489.5
|
ZFHX3
|
zinc finger homeobox 3 |
| chr13_+_29233218 | 4.69 |
ENST00000380842.4
|
POMP
|
proteasome maturation protein |
| chr5_-_157002775 | 4.66 |
ENST00000257527.4
|
ADAM19
|
ADAM metallopeptidase domain 19 |
| chr19_+_58898627 | 4.56 |
ENST00000598098.1
ENST00000598495.1 ENST00000196551.3 ENST00000596046.1 |
RPS5
|
ribosomal protein S5 |
| chr14_+_61201445 | 4.52 |
ENST00000261245.4
ENST00000539616.2 |
MNAT1
|
MNAT CDK-activating kinase assembly factor 1 |
| chr8_+_73921085 | 4.51 |
ENST00000276603.5
ENST00000276602.6 ENST00000518874.1 |
TERF1
|
telomeric repeat binding factor (NIMA-interacting) 1 |
| chr4_-_71705060 | 4.49 |
ENST00000514161.1
|
GRSF1
|
G-rich RNA sequence binding factor 1 |
| chr8_-_101962777 | 4.43 |
ENST00000395951.3
|
YWHAZ
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta |
| chr1_-_149900122 | 4.43 |
ENST00000271628.8
|
SF3B4
|
splicing factor 3b, subunit 4, 49kDa |
| chr14_+_24458093 | 4.33 |
ENST00000558753.1
ENST00000537912.1 |
DHRS4L2
|
dehydrogenase/reductase (SDR family) member 4 like 2 |
| chr17_-_18266765 | 4.30 |
ENST00000354098.3
|
SHMT1
|
serine hydroxymethyltransferase 1 (soluble) |
| chr11_-_10829851 | 4.28 |
ENST00000532082.1
|
EIF4G2
|
eukaryotic translation initiation factor 4 gamma, 2 |
| chr17_-_18266797 | 4.26 |
ENST00000316694.3
ENST00000539052.1 |
SHMT1
|
serine hydroxymethyltransferase 1 (soluble) |
| chr19_+_1026298 | 4.24 |
ENST00000263097.4
|
CNN2
|
calponin 2 |
| chr17_-_56065484 | 4.18 |
ENST00000581208.1
|
VEZF1
|
vascular endothelial zinc finger 1 |
| chr1_-_33502441 | 4.17 |
ENST00000548033.1
ENST00000487289.1 ENST00000373449.2 ENST00000480134.1 ENST00000467905.1 |
AK2
|
adenylate kinase 2 |
| chr4_-_52904425 | 4.10 |
ENST00000535450.1
|
SGCB
|
sarcoglycan, beta (43kDa dystrophin-associated glycoprotein) |
| chr2_+_189157536 | 3.92 |
ENST00000409580.1
ENST00000409637.3 |
GULP1
|
GULP, engulfment adaptor PTB domain containing 1 |
| chr13_+_102104980 | 3.86 |
ENST00000545560.2
|
ITGBL1
|
integrin, beta-like 1 (with EGF-like repeat domains) |
| chr19_+_41257084 | 3.86 |
ENST00000601393.1
|
SNRPA
|
small nuclear ribonucleoprotein polypeptide A |
| chr6_+_64282447 | 3.84 |
ENST00000370650.2
ENST00000578299.1 |
PTP4A1
|
protein tyrosine phosphatase type IVA, member 1 |
| chr7_-_148725733 | 3.64 |
ENST00000286091.4
|
PDIA4
|
protein disulfide isomerase family A, member 4 |
| chr22_+_39052632 | 3.54 |
ENST00000411557.1
ENST00000396811.2 ENST00000216029.3 ENST00000416285.1 |
CBY1
|
chibby homolog 1 (Drosophila) |
| chrX_+_102883620 | 3.50 |
ENST00000372626.3
|
TCEAL1
|
transcription elongation factor A (SII)-like 1 |
| chr2_+_172378757 | 3.41 |
ENST00000409484.1
ENST00000321348.4 ENST00000375252.3 |
CYBRD1
|
cytochrome b reductase 1 |
| chr6_+_32146131 | 3.39 |
ENST00000375094.3
|
RNF5
|
ring finger protein 5, E3 ubiquitin protein ligase |
| chr1_+_955448 | 3.37 |
ENST00000379370.2
|
AGRN
|
agrin |
| chr17_-_17485731 | 3.33 |
ENST00000395783.1
|
PEMT
|
phosphatidylethanolamine N-methyltransferase |
| chr2_-_217560248 | 3.29 |
ENST00000233813.4
|
IGFBP5
|
insulin-like growth factor binding protein 5 |
| chr2_+_46926326 | 3.24 |
ENST00000394861.2
|
SOCS5
|
suppressor of cytokine signaling 5 |
| chr1_-_110933663 | 3.24 |
ENST00000369781.4
ENST00000541986.1 ENST00000369779.4 |
SLC16A4
|
solute carrier family 16, member 4 |
| chrX_+_100645812 | 3.20 |
ENST00000427805.2
ENST00000553110.3 ENST00000392994.3 ENST00000409338.1 ENST00000409170.3 |
RPL36A
RPL36A-HNRNPH2
|
ribosomal protein L36a RPL36A-HNRNPH2 readthrough |
| chr8_-_119964434 | 3.19 |
ENST00000297350.4
|
TNFRSF11B
|
tumor necrosis factor receptor superfamily, member 11b |
| chrX_+_54835493 | 3.17 |
ENST00000396224.1
|
MAGED2
|
melanoma antigen family D, 2 |
| chr12_+_104359614 | 3.13 |
ENST00000266775.9
ENST00000544861.1 |
TDG
|
thymine-DNA glycosylase |
| chr11_-_33774944 | 3.13 |
ENST00000532057.1
ENST00000531080.1 |
FBXO3
|
F-box protein 3 |
| chrX_+_100646190 | 3.09 |
ENST00000471855.1
|
RPL36A
|
ribosomal protein L36a |
| chrX_+_12993336 | 3.07 |
ENST00000380635.1
|
TMSB4X
|
thymosin beta 4, X-linked |
| chr1_-_110933611 | 3.04 |
ENST00000472422.2
ENST00000437429.2 |
SLC16A4
|
solute carrier family 16, member 4 |
| chr1_-_147142557 | 2.98 |
ENST00000369238.6
|
ACP6
|
acid phosphatase 6, lysophosphatidic |
| chr17_+_79953310 | 2.97 |
ENST00000582355.2
|
ASPSCR1
|
alveolar soft part sarcoma chromosome region, candidate 1 |
| chr4_-_129209944 | 2.93 |
ENST00000520121.1
|
PGRMC2
|
progesterone receptor membrane component 2 |
| chr22_+_39077947 | 2.85 |
ENST00000216034.4
|
TOMM22
|
translocase of outer mitochondrial membrane 22 homolog (yeast) |
| chr17_-_20946338 | 2.84 |
ENST00000261497.4
|
USP22
|
ubiquitin specific peptidase 22 |
| chr1_-_211752073 | 2.84 |
ENST00000367001.4
|
SLC30A1
|
solute carrier family 30 (zinc transporter), member 1 |
| chr4_-_106629796 | 2.81 |
ENST00000416543.1
ENST00000515819.1 ENST00000420368.2 ENST00000503746.1 ENST00000340139.5 ENST00000433009.1 |
INTS12
|
integrator complex subunit 12 |
| chr11_-_10830463 | 2.77 |
ENST00000527419.1
ENST00000530211.1 ENST00000530702.1 ENST00000524932.1 ENST00000532570.1 |
EIF4G2
|
eukaryotic translation initiation factor 4 gamma, 2 |
| chr13_+_30002846 | 2.73 |
ENST00000542829.1
|
MTUS2
|
microtubule associated tumor suppressor candidate 2 |
| chr13_+_102142296 | 2.72 |
ENST00000376162.3
|
ITGBL1
|
integrin, beta-like 1 (with EGF-like repeat domains) |
| chr17_-_7297519 | 2.71 |
ENST00000576362.1
ENST00000571078.1 |
TMEM256-PLSCR3
|
TMEM256-PLSCR3 readthrough (NMD candidate) |
| chr19_-_2328572 | 2.67 |
ENST00000252622.10
|
LSM7
|
LSM7 homolog, U6 small nuclear RNA associated (S. cerevisiae) |
| chr2_+_177134201 | 2.63 |
ENST00000452865.1
|
MTX2
|
metaxin 2 |
| chr2_+_69240302 | 2.59 |
ENST00000303714.4
|
ANTXR1
|
anthrax toxin receptor 1 |
| chr1_-_53793584 | 2.56 |
ENST00000354412.3
ENST00000347547.2 ENST00000306052.6 |
LRP8
|
low density lipoprotein receptor-related protein 8, apolipoprotein e receptor |
| chr11_-_122930121 | 2.51 |
ENST00000524552.1
|
HSPA8
|
heat shock 70kDa protein 8 |
| chr17_+_68100989 | 2.47 |
ENST00000585558.1
ENST00000392670.1 |
KCNJ16
|
potassium inwardly-rectifying channel, subfamily J, member 16 |
| chr2_-_27592867 | 2.45 |
ENST00000451130.2
|
EIF2B4
|
eukaryotic translation initiation factor 2B, subunit 4 delta, 67kDa |
| chr1_+_169079823 | 2.44 |
ENST00000367813.3
|
ATP1B1
|
ATPase, Na+/K+ transporting, beta 1 polypeptide |
| chr6_+_125540951 | 2.42 |
ENST00000524679.1
|
TPD52L1
|
tumor protein D52-like 1 |
| chr2_-_238499303 | 2.41 |
ENST00000409576.1
|
RAB17
|
RAB17, member RAS oncogene family |
| chr5_-_148930960 | 2.38 |
ENST00000261798.5
ENST00000377843.2 |
CSNK1A1
|
casein kinase 1, alpha 1 |
| chr17_-_37009882 | 2.36 |
ENST00000378096.3
ENST00000394332.1 ENST00000394333.1 ENST00000577407.1 ENST00000479035.2 |
RPL23
|
ribosomal protein L23 |
| chr19_-_20844368 | 2.33 |
ENST00000595094.1
ENST00000601440.1 ENST00000291750.6 |
CTC-513N18.7
ZNF626
|
CTC-513N18.7 zinc finger protein 626 |
| chr9_-_21187598 | 2.28 |
ENST00000421715.1
|
IFNA4
|
interferon, alpha 4 |
| chr8_-_145047688 | 2.17 |
ENST00000356346.3
|
PLEC
|
plectin |
| chr2_-_88427568 | 2.16 |
ENST00000393750.3
ENST00000295834.3 |
FABP1
|
fatty acid binding protein 1, liver |
| chr12_+_104359641 | 2.15 |
ENST00000537100.1
|
TDG
|
thymine-DNA glycosylase |
| chr5_-_82969405 | 2.15 |
ENST00000510978.1
|
HAPLN1
|
hyaluronan and proteoglycan link protein 1 |
| chr17_-_46623441 | 2.14 |
ENST00000330070.4
|
HOXB2
|
homeobox B2 |
| chr2_+_46926048 | 2.14 |
ENST00000306503.5
|
SOCS5
|
suppressor of cytokine signaling 5 |
| chr2_-_238499337 | 2.12 |
ENST00000411462.1
ENST00000409822.1 |
RAB17
|
RAB17, member RAS oncogene family |
| chr5_+_140579162 | 2.11 |
ENST00000536699.1
ENST00000354757.3 |
PCDHB11
|
protocadherin beta 11 |
| chr12_-_49582593 | 2.11 |
ENST00000295766.5
|
TUBA1A
|
tubulin, alpha 1a |
| chr14_-_35344093 | 2.10 |
ENST00000382422.2
|
BAZ1A
|
bromodomain adjacent to zinc finger domain, 1A |
| chr1_+_182808474 | 2.07 |
ENST00000367549.3
|
DHX9
|
DEAH (Asp-Glu-Ala-His) box helicase 9 |
| chr4_-_146019693 | 2.03 |
ENST00000514390.1
|
ANAPC10
|
anaphase promoting complex subunit 10 |
| chr17_+_41150290 | 2.02 |
ENST00000589037.1
ENST00000253788.5 |
RPL27
|
ribosomal protein L27 |
| chr2_-_29093132 | 1.96 |
ENST00000306108.5
|
TRMT61B
|
tRNA methyltransferase 61 homolog B (S. cerevisiae) |
| chr1_-_19426149 | 1.95 |
ENST00000429347.2
|
UBR4
|
ubiquitin protein ligase E3 component n-recognin 4 |
| chr1_+_15479054 | 1.95 |
ENST00000376014.3
ENST00000451326.2 |
TMEM51
|
transmembrane protein 51 |
| chr1_+_15479021 | 1.94 |
ENST00000428417.1
|
TMEM51
|
transmembrane protein 51 |
| chr1_-_28527152 | 1.91 |
ENST00000321830.5
|
AL353354.1
|
Uncharacterized protein |
| chr17_-_26989136 | 1.90 |
ENST00000247020.4
|
SDF2
|
stromal cell-derived factor 2 |
| chr13_+_45694583 | 1.86 |
ENST00000340473.6
|
GTF2F2
|
general transcription factor IIF, polypeptide 2, 30kDa |
| chr1_+_11072696 | 1.85 |
ENST00000240185.3
ENST00000476201.1 |
TARDBP
|
TAR DNA binding protein |
| chr5_-_130500922 | 1.84 |
ENST00000513012.1
ENST00000508488.1 ENST00000506908.1 ENST00000304043.5 |
HINT1
|
histidine triad nucleotide binding protein 1 |
| chr13_+_102104952 | 1.81 |
ENST00000376180.3
|
ITGBL1
|
integrin, beta-like 1 (with EGF-like repeat domains) |
| chr6_-_112575912 | 1.81 |
ENST00000522006.1
ENST00000230538.7 ENST00000519932.1 |
LAMA4
|
laminin, alpha 4 |
| chr12_-_49351148 | 1.78 |
ENST00000398092.4
ENST00000539611.1 |
RP11-302B13.5
ARF3
|
ADP-ribosylation factor 3 ADP-ribosylation factor 3 |
| chr7_+_23286182 | 1.77 |
ENST00000258733.4
ENST00000381990.2 ENST00000409458.3 ENST00000539136.1 ENST00000453162.2 |
GPNMB
|
glycoprotein (transmembrane) nmb |
| chr1_-_151345159 | 1.75 |
ENST00000458566.1
ENST00000447402.3 ENST00000426705.2 ENST00000435071.1 ENST00000368868.5 |
SELENBP1
|
selenium binding protein 1 |
| chr12_+_49717019 | 1.74 |
ENST00000549275.1
ENST00000551245.1 ENST00000380327.5 ENST00000548311.1 ENST00000550346.1 ENST00000550709.1 ENST00000549534.1 ENST00000257909.3 |
TROAP
|
trophinin associated protein |
| chr17_+_74372662 | 1.74 |
ENST00000591651.1
ENST00000545180.1 |
SPHK1
|
sphingosine kinase 1 |
| chr11_-_87908600 | 1.70 |
ENST00000531138.1
ENST00000526372.1 ENST00000243662.6 |
RAB38
|
RAB38, member RAS oncogene family |
| chr17_+_75283973 | 1.66 |
ENST00000431235.2
ENST00000449803.2 |
SEPT9
|
septin 9 |
| chr16_+_68678739 | 1.66 |
ENST00000264012.4
|
CDH3
|
cadherin 3, type 1, P-cadherin (placental) |
| chr4_-_7873981 | 1.66 |
ENST00000360265.4
|
AFAP1
|
actin filament associated protein 1 |
| chr3_+_118892362 | 1.65 |
ENST00000497685.1
ENST00000264234.3 |
UPK1B
|
uroplakin 1B |
| chrX_-_102757802 | 1.63 |
ENST00000372633.1
|
RAB40A
|
RAB40A, member RAS oncogene family |
| chr2_+_1417228 | 1.62 |
ENST00000382269.3
ENST00000337415.3 ENST00000345913.4 ENST00000346956.3 ENST00000349624.3 ENST00000539820.1 ENST00000329066.4 ENST00000382201.3 |
TPO
|
thyroid peroxidase |
| chr16_-_46865047 | 1.62 |
ENST00000394806.2
|
C16orf87
|
chromosome 16 open reading frame 87 |
| chr15_-_44116873 | 1.61 |
ENST00000267812.3
|
MFAP1
|
microfibrillar-associated protein 1 |
| chr19_+_13134772 | 1.59 |
ENST00000587760.1
ENST00000585575.1 |
NFIX
|
nuclear factor I/X (CCAAT-binding transcription factor) |
| chrX_+_146993449 | 1.59 |
ENST00000218200.8
ENST00000370471.3 ENST00000370477.1 |
FMR1
|
fragile X mental retardation 1 |
| chr17_+_75447326 | 1.57 |
ENST00000591088.1
|
SEPT9
|
septin 9 |
| chr19_+_41256764 | 1.54 |
ENST00000243563.3
ENST00000601253.1 ENST00000597353.1 ENST00000599362.1 |
SNRPA
|
small nuclear ribonucleoprotein polypeptide A |
| chr6_+_52285046 | 1.53 |
ENST00000371068.5
|
EFHC1
|
EF-hand domain (C-terminal) containing 1 |
| chr19_+_45281118 | 1.51 |
ENST00000270279.3
ENST00000341505.4 |
CBLC
|
Cbl proto-oncogene C, E3 ubiquitin protein ligase |
| chr15_+_74466012 | 1.50 |
ENST00000249842.3
|
ISLR
|
immunoglobulin superfamily containing leucine-rich repeat |
| chr9_-_86593238 | 1.50 |
ENST00000351839.3
|
HNRNPK
|
heterogeneous nuclear ribonucleoprotein K |
| chr3_+_197677379 | 1.47 |
ENST00000442341.1
|
RPL35A
|
ribosomal protein L35a |
| chr2_-_231989808 | 1.46 |
ENST00000258400.3
|
HTR2B
|
5-hydroxytryptamine (serotonin) receptor 2B, G protein-coupled |
| chr17_+_26989109 | 1.46 |
ENST00000314616.6
ENST00000347486.4 |
SUPT6H
|
suppressor of Ty 6 homolog (S. cerevisiae) |
| chr17_+_48503519 | 1.45 |
ENST00000300441.4
ENST00000541920.1 ENST00000506582.1 ENST00000504392.1 ENST00000427954.2 |
ACSF2
|
acyl-CoA synthetase family member 2 |
| chr9_+_91003271 | 1.44 |
ENST00000375859.3
ENST00000541629.1 |
SPIN1
|
spindlin 1 |
| chr2_-_36825281 | 1.42 |
ENST00000405912.3
ENST00000379245.4 |
FEZ2
|
fasciculation and elongation protein zeta 2 (zygin II) |
| chr3_+_36421826 | 1.40 |
ENST00000273183.3
|
STAC
|
SH3 and cysteine rich domain |
| chr2_-_172750733 | 1.36 |
ENST00000392592.4
ENST00000422440.2 |
SLC25A12
|
solute carrier family 25 (aspartate/glutamate carrier), member 12 |
| chr1_-_79472365 | 1.34 |
ENST00000370742.3
|
ELTD1
|
EGF, latrophilin and seven transmembrane domain containing 1 |
| chr2_-_27593180 | 1.33 |
ENST00000493344.2
ENST00000445933.2 |
EIF2B4
|
eukaryotic translation initiation factor 2B, subunit 4 delta, 67kDa |
| chr3_+_197677047 | 1.28 |
ENST00000448864.1
|
RPL35A
|
ribosomal protein L35a |
| chr4_+_87515454 | 1.27 |
ENST00000427191.2
ENST00000436978.1 ENST00000502971.1 |
PTPN13
|
protein tyrosine phosphatase, non-receptor type 13 (APO-1/CD95 (Fas)-associated phosphatase) |
| chrX_+_100663243 | 1.26 |
ENST00000316594.5
|
HNRNPH2
|
heterogeneous nuclear ribonucleoprotein H2 (H') |
| chr8_+_77593448 | 1.20 |
ENST00000521891.2
|
ZFHX4
|
zinc finger homeobox 4 |
| chr8_-_90996459 | 1.18 |
ENST00000517337.1
ENST00000409330.1 |
NBN
|
nibrin |
| chr17_+_79679299 | 1.17 |
ENST00000331531.5
|
SLC25A10
|
solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 |
| chr17_+_79679369 | 1.15 |
ENST00000350690.5
|
SLC25A10
|
solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 |
| chr2_+_234580525 | 1.12 |
ENST00000609637.1
|
UGT1A1
|
UDP glucuronosyltransferase 1 family, polypeptide A8 |
| chr17_+_41150479 | 1.12 |
ENST00000589913.1
|
RPL27
|
ribosomal protein L27 |
| chr12_+_100594557 | 1.12 |
ENST00000546902.1
ENST00000552376.1 ENST00000551617.1 |
ACTR6
|
ARP6 actin-related protein 6 homolog (yeast) |
| chrX_+_135618258 | 1.11 |
ENST00000440515.1
ENST00000456412.1 |
VGLL1
|
vestigial like 1 (Drosophila) |
| chr7_+_111846643 | 1.07 |
ENST00000361822.3
|
ZNF277
|
zinc finger protein 277 |
| chr1_-_53793725 | 1.03 |
ENST00000371454.2
|
LRP8
|
low density lipoprotein receptor-related protein 8, apolipoprotein e receptor |
| chr11_-_108422926 | 1.00 |
ENST00000428840.1
ENST00000526312.1 |
EXPH5
|
exophilin 5 |
| chr1_-_16345245 | 0.97 |
ENST00000311890.9
|
HSPB7
|
heat shock 27kDa protein family, member 7 (cardiovascular) |
| chr5_-_157286104 | 0.95 |
ENST00000530742.1
ENST00000523908.1 ENST00000523094.1 ENST00000296951.5 ENST00000411809.2 |
CLINT1
|
clathrin interactor 1 |
| chr17_-_39580775 | 0.90 |
ENST00000225550.3
|
KRT37
|
keratin 37 |
| chr12_-_71003568 | 0.89 |
ENST00000547715.1
ENST00000451516.2 ENST00000538708.1 ENST00000550857.1 ENST00000261266.5 |
PTPRB
|
protein tyrosine phosphatase, receptor type, B |
| chrX_+_12993202 | 0.89 |
ENST00000451311.2
ENST00000380636.1 |
TMSB4X
|
thymosin beta 4, X-linked |
| chr14_+_24458021 | 0.88 |
ENST00000397071.1
ENST00000559411.1 ENST00000335125.6 |
DHRS4L2
|
dehydrogenase/reductase (SDR family) member 4 like 2 |
| chr2_+_234580499 | 0.82 |
ENST00000354728.4
|
UGT1A9
|
UDP glucuronosyltransferase 1 family, polypeptide A9 |
| chr5_+_150040403 | 0.81 |
ENST00000517768.1
ENST00000297130.4 |
MYOZ3
|
myozenin 3 |
| chr11_+_46740730 | 0.79 |
ENST00000311907.5
ENST00000530231.1 ENST00000442468.1 |
F2
|
coagulation factor II (thrombin) |
| chr12_-_133405288 | 0.76 |
ENST00000204726.3
|
GOLGA3
|
golgin A3 |
| chr10_-_61122220 | 0.75 |
ENST00000422313.2
ENST00000435852.2 ENST00000442566.3 ENST00000373868.2 ENST00000277705.6 ENST00000373867.3 ENST00000419214.2 |
FAM13C
|
family with sequence similarity 13, member C |
| chr5_+_43603229 | 0.74 |
ENST00000344920.4
ENST00000512996.2 |
NNT
|
nicotinamide nucleotide transhydrogenase |
Network of associatons between targets according to the STRING database.
First level regulatory network of TCF7
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.2 | 15.7 | GO:0009996 | negative regulation of Wnt signaling pathway involved in heart development(GO:0003308) negative regulation of cell fate specification(GO:0009996) Wnt signaling pathway involved in somitogenesis(GO:0090244) |
| 4.7 | 14.0 | GO:1904481 | response to tetrahydrofolate(GO:1904481) cellular response to tetrahydrofolate(GO:1904482) |
| 3.5 | 10.6 | GO:0006172 | ADP biosynthetic process(GO:0006172) |
| 3.0 | 11.9 | GO:0021812 | neuronal-glial interaction involved in cerebral cortex radial glia guided migration(GO:0021812) |
| 2.5 | 10.1 | GO:1902544 | oxidative DNA demethylation(GO:0035511) regulation of DNA N-glycosylase activity(GO:1902544) |
| 2.4 | 7.1 | GO:0051086 | chaperone mediated protein folding independent of cofactor(GO:0051086) |
| 2.3 | 11.4 | GO:0000973 | posttranscriptional tethering of RNA polymerase II gene DNA at nuclear periphery(GO:0000973) |
| 2.0 | 8.0 | GO:0016479 | negative regulation of transcription from RNA polymerase I promoter(GO:0016479) |
| 1.8 | 9.1 | GO:0070980 | biphenyl catabolic process(GO:0070980) |
| 1.7 | 5.2 | GO:0009051 | pentose-phosphate shunt, oxidative branch(GO:0009051) |
| 1.5 | 4.5 | GO:0031627 | telomeric loop formation(GO:0031627) negative regulation of protein localization to chromosome, telomeric region(GO:1904815) |
| 1.3 | 6.5 | GO:0014886 | transition between slow and fast fiber(GO:0014886) |
| 1.2 | 4.8 | GO:1904059 | regulation of locomotor rhythm(GO:1904059) |
| 1.1 | 3.3 | GO:1904204 | regulation of skeletal muscle hypertrophy(GO:1904204) |
| 1.0 | 15.3 | GO:0042178 | xenobiotic catabolic process(GO:0042178) |
| 0.9 | 5.5 | GO:0003350 | pulmonary myocardium development(GO:0003350) |
| 0.9 | 4.5 | GO:0002415 | immunoglobulin transcytosis in epithelial cells mediated by polymeric immunoglobulin receptor(GO:0002415) |
| 0.9 | 5.4 | GO:0045629 | negative regulation of T-helper 2 cell differentiation(GO:0045629) |
| 0.9 | 2.6 | GO:1901202 | negative regulation of extracellular matrix assembly(GO:1901202) |
| 0.8 | 3.4 | GO:1904398 | positive regulation of neuromuscular junction development(GO:1904398) |
| 0.8 | 4.9 | GO:1903899 | positive regulation of PERK-mediated unfolded protein response(GO:1903899) |
| 0.8 | 3.2 | GO:0042483 | negative regulation of odontogenesis(GO:0042483) |
| 0.8 | 2.3 | GO:1902356 | thiosulfate transport(GO:0015709) oxaloacetate transport(GO:0015729) malate transport(GO:0015743) malate transmembrane transport(GO:0071423) oxaloacetate(2-) transmembrane transport(GO:1902356) |
| 0.8 | 10.7 | GO:0030913 | paranodal junction assembly(GO:0030913) |
| 0.8 | 14.5 | GO:0071850 | mitotic cell cycle arrest(GO:0071850) |
| 0.7 | 3.6 | GO:0038026 | reelin-mediated signaling pathway(GO:0038026) |
| 0.7 | 2.1 | GO:0021569 | rhombomere 3 development(GO:0021569) |
| 0.7 | 10.2 | GO:1904816 | positive regulation of protein localization to chromosome, telomeric region(GO:1904816) |
| 0.7 | 4.0 | GO:2000483 | negative regulation of interleukin-8 secretion(GO:2000483) |
| 0.7 | 2.0 | GO:0070901 | mitochondrial tRNA methylation(GO:0070901) |
| 0.6 | 4.9 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) |
| 0.6 | 2.4 | GO:1903288 | protein transport into plasma membrane raft(GO:0044861) positive regulation of potassium ion import(GO:1903288) |
| 0.6 | 1.7 | GO:0046521 | sphingoid catabolic process(GO:0046521) |
| 0.6 | 2.8 | GO:0061088 | regulation of sequestering of zinc ion(GO:0061088) |
| 0.6 | 3.3 | GO:0050747 | positive regulation of lipoprotein metabolic process(GO:0050747) |
| 0.6 | 1.7 | GO:0051796 | regulation of monophenol monooxygenase activity(GO:0032771) positive regulation of monophenol monooxygenase activity(GO:0032773) negative regulation of catagen(GO:0051796) regulation of hair cycle by canonical Wnt signaling pathway(GO:0060901) positive regulation of melanosome transport(GO:1902910) |
| 0.5 | 7.8 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.5 | 2.5 | GO:1904764 | negative regulation of fibril organization(GO:1902904) chaperone-mediated autophagy translocation complex disassembly(GO:1904764) |
| 0.5 | 7.3 | GO:0043589 | skin morphogenesis(GO:0043589) |
| 0.5 | 3.8 | GO:0032057 | negative regulation of translational initiation in response to stress(GO:0032057) |
| 0.5 | 2.4 | GO:0072717 | cellular response to actinomycin D(GO:0072717) |
| 0.5 | 5.1 | GO:0060020 | Bergmann glial cell differentiation(GO:0060020) |
| 0.5 | 1.4 | GO:0043490 | malate-aspartate shuttle(GO:0043490) |
| 0.5 | 3.2 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.4 | 2.1 | GO:0035549 | positive regulation of interferon-beta secretion(GO:0035549) |
| 0.4 | 12.5 | GO:0061641 | CENP-A containing nucleosome assembly(GO:0034080) CENP-A containing chromatin organization(GO:0061641) |
| 0.4 | 6.8 | GO:0089711 | L-glutamate transmembrane transport(GO:0089711) |
| 0.4 | 1.2 | GO:0031860 | telomeric 3' overhang formation(GO:0031860) |
| 0.4 | 1.5 | GO:0072369 | regulation of lipid transport by positive regulation of transcription from RNA polymerase II promoter(GO:0072369) |
| 0.4 | 1.5 | GO:0061086 | negative regulation of histone H3-K27 methylation(GO:0061086) |
| 0.3 | 2.4 | GO:1904885 | beta-catenin destruction complex assembly(GO:1904885) |
| 0.3 | 4.5 | GO:0007512 | adult heart development(GO:0007512) |
| 0.3 | 1.6 | GO:2000301 | regulation of intracellular transport of viral material(GO:1901252) negative regulation of synaptic vesicle exocytosis(GO:2000301) |
| 0.3 | 4.1 | GO:0097084 | vascular smooth muscle cell development(GO:0097084) |
| 0.3 | 9.0 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.3 | 4.4 | GO:0090168 | Golgi reassembly(GO:0090168) |
| 0.3 | 4.4 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.3 | 4.7 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.3 | 1.5 | GO:0010513 | positive regulation of phosphatidylinositol biosynthetic process(GO:0010513) |
| 0.3 | 10.5 | GO:0060261 | positive regulation of transcription initiation from RNA polymerase II promoter(GO:0060261) |
| 0.3 | 5.4 | GO:0060391 | positive regulation of SMAD protein import into nucleus(GO:0060391) |
| 0.2 | 5.4 | GO:1900363 | regulation of mRNA polyadenylation(GO:1900363) |
| 0.2 | 3.9 | GO:0071712 | ER-associated misfolded protein catabolic process(GO:0071712) |
| 0.2 | 1.7 | GO:1903232 | melanosome assembly(GO:1903232) |
| 0.2 | 2.2 | GO:0032000 | positive regulation of fatty acid beta-oxidation(GO:0032000) |
| 0.2 | 1.4 | GO:1902902 | negative regulation of autophagosome assembly(GO:1902902) |
| 0.2 | 0.9 | GO:0006740 | NADPH regeneration(GO:0006740) |
| 0.2 | 4.6 | GO:0006450 | regulation of translational fidelity(GO:0006450) |
| 0.2 | 1.9 | GO:0071763 | nuclear membrane organization(GO:0071763) |
| 0.2 | 4.9 | GO:0060065 | uterus development(GO:0060065) |
| 0.2 | 7.4 | GO:0006378 | mRNA polyadenylation(GO:0006378) |
| 0.2 | 1.5 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.2 | 0.5 | GO:0051684 | maintenance of Golgi location(GO:0051684) |
| 0.2 | 0.8 | GO:0051552 | flavone metabolic process(GO:0051552) |
| 0.2 | 3.2 | GO:0033141 | positive regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033141) |
| 0.2 | 0.8 | GO:0051801 | cytolysis in other organism involved in symbiotic interaction(GO:0051801) positive regulation of phospholipase C-activating G-protein coupled receptor signaling pathway(GO:1900738) |
| 0.1 | 2.8 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.1 | 11.2 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.1 | 0.6 | GO:0060392 | negative regulation of SMAD protein import into nucleus(GO:0060392) |
| 0.1 | 0.6 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.1 | 6.8 | GO:0035904 | aorta development(GO:0035904) |
| 0.1 | 0.4 | GO:0009804 | coumarin metabolic process(GO:0009804) |
| 0.1 | 2.5 | GO:1902857 | positive regulation of nonmotile primary cilium assembly(GO:1902857) |
| 0.1 | 4.8 | GO:0006509 | membrane protein ectodomain proteolysis(GO:0006509) |
| 0.1 | 2.8 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 0.1 | 5.1 | GO:0051489 | regulation of filopodium assembly(GO:0051489) |
| 0.1 | 8.3 | GO:0071349 | interleukin-12-mediated signaling pathway(GO:0035722) cellular response to interleukin-12(GO:0071349) |
| 0.1 | 0.6 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.1 | 1.6 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.1 | 0.4 | GO:0008582 | regulation of synaptic growth at neuromuscular junction(GO:0008582) synaptic growth at neuromuscular junction(GO:0051124) |
| 0.1 | 7.1 | GO:0006446 | regulation of translational initiation(GO:0006446) |
| 0.1 | 2.7 | GO:0043928 | exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 0.1 | 9.0 | GO:0002181 | cytoplasmic translation(GO:0002181) |
| 0.1 | 9.0 | GO:0035335 | peptidyl-tyrosine dephosphorylation(GO:0035335) |
| 0.1 | 0.3 | GO:0038180 | nerve growth factor signaling pathway(GO:0038180) |
| 0.1 | 0.6 | GO:0007288 | sperm axoneme assembly(GO:0007288) |
| 0.1 | 1.6 | GO:0006590 | thyroid hormone generation(GO:0006590) |
| 0.1 | 0.7 | GO:0070072 | proton-transporting V-type ATPase complex assembly(GO:0070070) vacuolar proton-transporting V-type ATPase complex assembly(GO:0070072) |
| 0.1 | 0.7 | GO:0042905 | 9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.1 | 8.0 | GO:0002062 | chondrocyte differentiation(GO:0002062) |
| 0.1 | 6.5 | GO:0043462 | regulation of ATPase activity(GO:0043462) |
| 0.1 | 0.4 | GO:0070307 | lens fiber cell development(GO:0070307) |
| 0.1 | 0.5 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.1 | 1.0 | GO:0071985 | multivesicular body sorting pathway(GO:0071985) |
| 0.1 | 0.4 | GO:1903286 | regulation of potassium ion import(GO:1903286) |
| 0.1 | 1.5 | GO:0031954 | positive regulation of protein autophosphorylation(GO:0031954) |
| 0.1 | 0.4 | GO:0033629 | negative regulation of cell adhesion mediated by integrin(GO:0033629) |
| 0.1 | 4.0 | GO:0001885 | endothelial cell development(GO:0001885) |
| 0.1 | 2.0 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.1 | 4.8 | GO:0006695 | cholesterol biosynthetic process(GO:0006695) |
| 0.1 | 2.2 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.0 | 5.0 | GO:0036498 | IRE1-mediated unfolded protein response(GO:0036498) |
| 0.0 | 3.6 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.0 | 1.4 | GO:0009303 | rRNA transcription(GO:0009303) |
| 0.0 | 0.3 | GO:0070574 | cadmium ion transport(GO:0015691) cadmium ion transmembrane transport(GO:0070574) |
| 0.0 | 3.4 | GO:0006879 | cellular iron ion homeostasis(GO:0006879) |
| 0.0 | 2.1 | GO:0007416 | synapse assembly(GO:0007416) |
| 0.0 | 1.8 | GO:0009154 | purine ribonucleotide catabolic process(GO:0009154) |
| 0.0 | 1.5 | GO:0021795 | cerebral cortex cell migration(GO:0021795) |
| 0.0 | 3.0 | GO:0046324 | regulation of glucose import(GO:0046324) |
| 0.0 | 0.7 | GO:0006098 | pentose-phosphate shunt(GO:0006098) |
| 0.0 | 4.4 | GO:0007498 | mesoderm development(GO:0007498) |
| 0.0 | 1.0 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.0 | 1.4 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
| 0.0 | 2.9 | GO:0045995 | regulation of embryonic development(GO:0045995) |
| 0.0 | 4.0 | GO:0015718 | monocarboxylic acid transport(GO:0015718) |
| 0.0 | 2.7 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.0 | 0.7 | GO:0006309 | apoptotic DNA fragmentation(GO:0006309) |
| 0.0 | 5.7 | GO:0006936 | muscle contraction(GO:0006936) |
| 0.0 | 2.1 | GO:0055007 | cardiac muscle cell differentiation(GO:0055007) |
| 0.0 | 0.2 | GO:0015760 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.0 | 0.7 | GO:0042220 | response to cocaine(GO:0042220) |
| 0.0 | 0.2 | GO:0086073 | cardiac muscle cell-cardiac muscle cell adhesion(GO:0086042) bundle of His cell-Purkinje myocyte adhesion involved in cell communication(GO:0086073) |
| 0.0 | 3.2 | GO:0006338 | chromatin remodeling(GO:0006338) |
| 0.0 | 0.5 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.0 | 4.8 | GO:0006457 | protein folding(GO:0006457) |
| 0.0 | 0.3 | GO:0055072 | iron ion homeostasis(GO:0055072) |
| 0.0 | 0.3 | GO:0007064 | mitotic sister chromatid cohesion(GO:0007064) |
| 0.0 | 0.8 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.0 | 0.2 | GO:0046007 | negative regulation of activated T cell proliferation(GO:0046007) |
| 0.0 | 1.8 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.0 | 0.1 | GO:0030220 | platelet formation(GO:0030220) |
| 0.0 | 3.8 | GO:0043123 | positive regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043123) |
| 0.0 | 1.0 | GO:0006986 | response to unfolded protein(GO:0006986) |
| 0.0 | 1.5 | GO:0006626 | protein targeting to mitochondrion(GO:0006626) |
| 0.0 | 4.1 | GO:0000086 | G2/M transition of mitotic cell cycle(GO:0000086) |
| 0.0 | 0.4 | GO:0030049 | muscle filament sliding(GO:0030049) actin-myosin filament sliding(GO:0033275) |
| 0.0 | 0.6 | GO:0007274 | neuromuscular synaptic transmission(GO:0007274) |
| 0.0 | 0.6 | GO:0001961 | positive regulation of cytokine-mediated signaling pathway(GO:0001961) |
| 0.0 | 1.7 | GO:0051493 | regulation of cytoskeleton organization(GO:0051493) |
| 0.0 | 0.1 | GO:0015705 | iodide transport(GO:0015705) |
| 0.0 | 0.0 | GO:0034165 | regulation of toll-like receptor 9 signaling pathway(GO:0034163) positive regulation of toll-like receptor 9 signaling pathway(GO:0034165) |
| 0.0 | 0.1 | GO:0000717 | nucleotide-excision repair, DNA duplex unwinding(GO:0000717) UV-damage excision repair(GO:0070914) |
| 0.0 | 1.6 | GO:0042787 | protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0042787) |
| 0.0 | 0.3 | GO:0052646 | alditol phosphate metabolic process(GO:0052646) |
| 0.0 | 6.1 | GO:0030036 | actin cytoskeleton organization(GO:0030036) |
| 0.0 | 6.9 | GO:0045087 | innate immune response(GO:0045087) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.0 | 11.9 | GO:0005607 | laminin-2 complex(GO:0005607) |
| 2.7 | 8.0 | GO:0031523 | Myb complex(GO:0031523) |
| 1.6 | 4.9 | GO:0016222 | procollagen-proline 4-dioxygenase complex(GO:0016222) |
| 1.2 | 5.0 | GO:0001405 | presequence translocase-associated import motor(GO:0001405) |
| 0.9 | 6.9 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.8 | 3.9 | GO:0005726 | perichromatin fibrils(GO:0005726) |
| 0.7 | 11.4 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.7 | 5.0 | GO:0005785 | signal recognition particle receptor complex(GO:0005785) |
| 0.7 | 10.4 | GO:0030478 | actin cap(GO:0030478) |
| 0.7 | 2.0 | GO:0031515 | tRNA (m1A) methyltransferase complex(GO:0031515) |
| 0.6 | 11.9 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.6 | 1.9 | GO:0005674 | transcription factor TFIIF complex(GO:0005674) |
| 0.6 | 7.3 | GO:0098643 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.6 | 8.4 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.5 | 9.1 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.5 | 1.6 | GO:0019034 | viral replication complex(GO:0019034) |
| 0.5 | 4.1 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 0.5 | 3.4 | GO:0036513 | Derlin-1 retrotranslocation complex(GO:0036513) |
| 0.5 | 3.8 | GO:0005851 | eukaryotic translation initiation factor 2B complex(GO:0005851) |
| 0.5 | 4.5 | GO:0070187 | telosome(GO:0070187) |
| 0.4 | 2.8 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.4 | 2.7 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 0.4 | 3.3 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 0.4 | 2.5 | GO:0098575 | lumenal side of lysosomal membrane(GO:0098575) |
| 0.3 | 6.5 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.3 | 1.7 | GO:0031905 | early endosome lumen(GO:0031905) |
| 0.3 | 10.8 | GO:0005685 | U1 snRNP(GO:0005685) |
| 0.3 | 7.1 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.3 | 2.8 | GO:0032039 | integrator complex(GO:0032039) |
| 0.3 | 4.5 | GO:0005675 | holo TFIIH complex(GO:0005675) |
| 0.3 | 5.1 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.3 | 4.4 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.3 | 2.2 | GO:0045179 | apical cortex(GO:0045179) |
| 0.3 | 0.8 | GO:0090498 | extrinsic component of Golgi membrane(GO:0090498) |
| 0.3 | 7.8 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.2 | 1.2 | GO:0030870 | Mre11 complex(GO:0030870) |
| 0.2 | 25.7 | GO:0005604 | basement membrane(GO:0005604) |
| 0.2 | 2.1 | GO:0031010 | ISWI-type complex(GO:0031010) |
| 0.2 | 2.8 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.2 | 9.3 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.2 | 2.4 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.1 | 3.2 | GO:0031105 | septin complex(GO:0031105) |
| 0.1 | 2.2 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.1 | 11.1 | GO:0036126 | sperm flagellum(GO:0036126) |
| 0.1 | 12.5 | GO:0000777 | condensed chromosome kinetochore(GO:0000777) |
| 0.1 | 14.1 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.1 | 2.4 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.1 | 6.9 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.1 | 9.0 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.1 | 2.6 | GO:0031527 | filopodium membrane(GO:0031527) |
| 0.1 | 15.1 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.1 | 14.4 | GO:0042470 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.1 | 13.8 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.1 | 6.6 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.1 | 1.6 | GO:0043205 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.1 | 4.3 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.1 | 2.0 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.1 | 8.1 | GO:0035579 | specific granule membrane(GO:0035579) |
| 0.1 | 4.6 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.1 | 5.1 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.1 | 27.9 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.1 | 1.5 | GO:0035327 | transcriptionally active chromatin(GO:0035327) |
| 0.1 | 2.9 | GO:0030315 | T-tubule(GO:0030315) |
| 0.0 | 6.0 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 19.7 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.0 | 0.5 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.0 | 3.0 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.0 | 0.2 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.0 | 10.2 | GO:0016604 | nuclear body(GO:0016604) |
| 0.0 | 0.5 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.0 | 2.2 | GO:0005901 | caveola(GO:0005901) |
| 0.0 | 1.8 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 1.5 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.0 | 5.8 | GO:0009898 | cytoplasmic side of plasma membrane(GO:0009898) |
| 0.0 | 1.7 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.0 | 0.1 | GO:0000110 | nucleotide-excision repair factor 1 complex(GO:0000110) |
| 0.0 | 0.6 | GO:0005865 | striated muscle thin filament(GO:0005865) |
| 0.0 | 0.7 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 0.4 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 0.5 | GO:0030667 | secretory granule membrane(GO:0030667) |
| 0.0 | 3.0 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 7.8 | GO:0009986 | cell surface(GO:0009986) |
| 0.0 | 0.9 | GO:0005746 | mitochondrial respiratory chain(GO:0005746) |
| 0.0 | 3.7 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 1.0 | GO:0030136 | clathrin-coated vesicle(GO:0030136) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.4 | 10.1 | GO:0043739 | G/U mismatch-specific uracil-DNA glycosylase activity(GO:0043739) |
| 3.1 | 15.3 | GO:0070404 | NADH binding(GO:0070404) |
| 2.8 | 14.0 | GO:0070905 | serine binding(GO:0070905) |
| 2.0 | 8.0 | GO:0034988 | Fc-gamma receptor I complex binding(GO:0034988) |
| 1.7 | 5.2 | GO:0004616 | phosphogluconate dehydrogenase (decarboxylating) activity(GO:0004616) |
| 1.7 | 5.0 | GO:0005047 | signal recognition particle binding(GO:0005047) |
| 1.5 | 10.8 | GO:0035614 | snRNA stem-loop binding(GO:0035614) |
| 1.5 | 16.0 | GO:0039706 | co-receptor binding(GO:0039706) |
| 1.4 | 6.8 | GO:0005294 | neutral L-amino acid secondary active transmembrane transporter activity(GO:0005294) |
| 1.2 | 4.9 | GO:0071987 | WD40-repeat domain binding(GO:0071987) |
| 1.2 | 4.8 | GO:0003985 | acetyl-CoA C-acetyltransferase activity(GO:0003985) |
| 1.1 | 3.3 | GO:0000773 | phosphatidyl-N-methylethanolamine N-methyltransferase activity(GO:0000773) phosphatidylethanolamine N-methyltransferase activity(GO:0004608) phosphatidyl-N-dimethylethanolamine N-methyltransferase activity(GO:0080101) |
| 1.0 | 4.9 | GO:0004656 | procollagen-proline 4-dioxygenase activity(GO:0004656) procollagen-proline dioxygenase activity(GO:0019798) |
| 0.9 | 3.6 | GO:0038025 | reelin receptor activity(GO:0038025) |
| 0.9 | 5.1 | GO:1990254 | keratin filament binding(GO:1990254) |
| 0.8 | 9.6 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.8 | 2.3 | GO:0015117 | thiosulfate transmembrane transporter activity(GO:0015117) oxaloacetate transmembrane transporter activity(GO:0015131) |
| 0.7 | 2.0 | GO:0016429 | tRNA (adenine) methyltransferase activity(GO:0016426) tRNA (adenine-N1-)-methyltransferase activity(GO:0016429) |
| 0.6 | 4.5 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 0.6 | 4.5 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.6 | 2.4 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.6 | 3.4 | GO:0000293 | ferric-chelate reductase activity(GO:0000293) |
| 0.6 | 5.1 | GO:0060002 | plus-end directed microfilament motor activity(GO:0060002) |
| 0.6 | 3.4 | GO:0035374 | chondroitin sulfate binding(GO:0035374) |
| 0.6 | 8.9 | GO:0016641 | oxidoreductase activity, acting on the CH-NH2 group of donors, oxygen as acceptor(GO:0016641) |
| 0.5 | 7.3 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.5 | 13.9 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.4 | 1.7 | GO:0035651 | AP-1 adaptor complex binding(GO:0035650) AP-3 adaptor complex binding(GO:0035651) |
| 0.4 | 19.9 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.4 | 11.4 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.4 | 5.0 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.4 | 2.1 | GO:0061676 | importin-alpha family protein binding(GO:0061676) |
| 0.4 | 5.5 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.4 | 3.3 | GO:0000990 | transcription factor activity, core RNA polymerase binding(GO:0000990) |
| 0.3 | 9.4 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.3 | 1.6 | GO:0034046 | poly(G) binding(GO:0034046) |
| 0.3 | 1.9 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.3 | 2.2 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.3 | 3.3 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.3 | 1.7 | GO:0008481 | sphinganine kinase activity(GO:0008481) D-erythro-sphingosine kinase activity(GO:0017050) |
| 0.3 | 3.4 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.3 | 1.4 | GO:0015183 | L-aspartate transmembrane transporter activity(GO:0015183) |
| 0.3 | 1.6 | GO:0004447 | iodide peroxidase activity(GO:0004447) |
| 0.3 | 10.0 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.3 | 2.8 | GO:0015266 | protein channel activity(GO:0015266) |
| 0.2 | 0.7 | GO:0004531 | deoxyribonuclease II activity(GO:0004531) |
| 0.2 | 3.2 | GO:0005132 | type I interferon receptor binding(GO:0005132) |
| 0.2 | 2.5 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.2 | 10.5 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.2 | 0.7 | GO:0017057 | glucose-6-phosphate dehydrogenase activity(GO:0004345) 6-phosphogluconolactonase activity(GO:0017057) |
| 0.2 | 3.0 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.2 | 4.9 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.2 | 7.1 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.2 | 1.8 | GO:0045545 | syndecan binding(GO:0045545) |
| 0.2 | 8.9 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.2 | 3.6 | GO:0016864 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.2 | 2.8 | GO:0019855 | calcium channel inhibitor activity(GO:0019855) |
| 0.2 | 0.7 | GO:0022850 | serotonin-gated cation channel activity(GO:0022850) |
| 0.2 | 5.3 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.2 | 0.8 | GO:0051373 | FATZ binding(GO:0051373) |
| 0.2 | 1.7 | GO:0008430 | selenium binding(GO:0008430) |
| 0.2 | 9.6 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.2 | 2.4 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.1 | 2.8 | GO:0010485 | H4 histone acetyltransferase activity(GO:0010485) |
| 0.1 | 14.5 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 10.9 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.1 | 1.3 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.1 | 2.7 | GO:0017070 | U6 snRNA binding(GO:0017070) |
| 0.1 | 9.2 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.1 | 0.6 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.1 | 1.5 | GO:0043176 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.1 | 1.4 | GO:0003996 | acyl-CoA ligase activity(GO:0003996) |
| 0.1 | 6.9 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.1 | 11.7 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.1 | 4.8 | GO:0046966 | thyroid hormone receptor binding(GO:0046966) |
| 0.1 | 15.7 | GO:0047485 | protein N-terminus binding(GO:0047485) |
| 0.1 | 2.0 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.1 | 6.3 | GO:0008028 | monocarboxylic acid transmembrane transporter activity(GO:0008028) |
| 0.1 | 3.5 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.1 | 31.6 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.1 | 4.0 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 4.6 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.1 | 2.2 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.1 | 11.8 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 0.7 | GO:0001758 | retinal dehydrogenase activity(GO:0001758) |
| 0.1 | 0.3 | GO:0010465 | nerve growth factor receptor activity(GO:0010465) |
| 0.1 | 2.1 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.1 | 1.8 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 4.9 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 9.5 | GO:0003729 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.0 | 0.6 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.0 | 5.1 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.2 | GO:0086083 | cell adhesive protein binding involved in bundle of His cell-Purkinje myocyte communication(GO:0086083) |
| 0.0 | 0.4 | GO:0097016 | L27 domain binding(GO:0097016) |
| 0.0 | 2.4 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 1.0 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.0 | 0.2 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
| 0.0 | 5.9 | GO:0005516 | calmodulin binding(GO:0005516) |
| 0.0 | 2.9 | GO:0005496 | steroid binding(GO:0005496) |
| 0.0 | 2.2 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 0.8 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.0 | 0.4 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.0 | 0.3 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 1.3 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.0 | 0.5 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 0.9 | GO:0050661 | NADP binding(GO:0050661) |
| 0.0 | 1.5 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.0 | 1.3 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.0 | 3.8 | GO:0005198 | structural molecule activity(GO:0005198) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 18.7 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.5 | 15.7 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.3 | 13.9 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.3 | 21.5 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.3 | 10.4 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 0.3 | 8.9 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.2 | 19.9 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.2 | 13.3 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.2 | 4.0 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.2 | 9.7 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.1 | 2.6 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.1 | 5.1 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.1 | 4.5 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.1 | 14.3 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.1 | 3.4 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 11.6 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.1 | 2.6 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.1 | 3.9 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.1 | 0.8 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.1 | 4.5 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.1 | 5.3 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.0 | 2.4 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.0 | 0.9 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.0 | 9.6 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 2.4 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 2.1 | PID P73PATHWAY | p73 transcription factor network |
| 0.0 | 4.9 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 1.7 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 1.7 | ST INTEGRIN SIGNALING PATHWAY | Integrin Signaling Pathway |
| 0.0 | 3.4 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 1.3 | ST FAS SIGNALING PATHWAY | Fas Signaling Pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 10.1 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.5 | 14.0 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.4 | 4.5 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.4 | 10.0 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.3 | 19.8 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.3 | 11.4 | REACTOME NEP NS2 INTERACTS WITH THE CELLULAR EXPORT MACHINERY | Genes involved in NEP/NS2 Interacts with the Cellular Export Machinery |
| 0.3 | 9.6 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.3 | 5.5 | REACTOME N GLYCAN TRIMMING IN THE ER AND CALNEXIN CALRETICULIN CYCLE | Genes involved in N-glycan trimming in the ER and Calnexin/Calreticulin cycle |
| 0.2 | 2.4 | REACTOME CTNNB1 PHOSPHORYLATION CASCADE | Genes involved in Beta-catenin phosphorylation cascade |
| 0.2 | 3.5 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.2 | 4.9 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.2 | 9.0 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.2 | 14.0 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.2 | 3.2 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.2 | 9.4 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.2 | 11.8 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.2 | 4.5 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.2 | 4.4 | REACTOME DESTABILIZATION OF MRNA BY KSRP | Genes involved in Destabilization of mRNA by KSRP |
| 0.2 | 4.1 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.2 | 3.6 | REACTOME PLATELET SENSITIZATION BY LDL | Genes involved in Platelet sensitization by LDL |
| 0.2 | 30.1 | REACTOME INFLUENZA VIRAL RNA TRANSCRIPTION AND REPLICATION | Genes involved in Influenza Viral RNA Transcription and Replication |
| 0.2 | 12.5 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.1 | 3.4 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.1 | 5.0 | REACTOME SRP DEPENDENT COTRANSLATIONAL PROTEIN TARGETING TO MEMBRANE | Genes involved in SRP-dependent cotranslational protein targeting to membrane |
| 0.1 | 3.2 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.1 | 4.4 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.1 | 4.4 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.1 | 3.3 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.1 | 2.0 | REACTOME INHIBITION OF THE PROTEOLYTIC ACTIVITY OF APC C REQUIRED FOR THE ONSET OF ANAPHASE BY MITOTIC SPINDLE CHECKPOINT COMPONENTS | Genes involved in Inhibition of the proteolytic activity of APC/C required for the onset of anaphase by mitotic spindle checkpoint components |
| 0.1 | 1.5 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.1 | 7.1 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.1 | 9.2 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.1 | 10.2 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.1 | 5.1 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.1 | 11.4 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.1 | 1.7 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.1 | 1.8 | REACTOME SYNTHESIS OF PIPS AT THE GOLGI MEMBRANE | Genes involved in Synthesis of PIPs at the Golgi membrane |
| 0.1 | 3.3 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 0.1 | 2.7 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.1 | 3.8 | REACTOME TRANSLATION | Genes involved in Translation |
| 0.1 | 1.6 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.1 | 3.2 | REACTOME IRON UPTAKE AND TRANSPORT | Genes involved in Iron uptake and transport |
| 0.1 | 6.8 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.1 | 8.4 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.1 | 7.4 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.1 | 2.3 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.1 | 2.0 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.0 | 1.2 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.0 | 0.9 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 0.4 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.0 | 1.8 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.0 | 3.4 | REACTOME METABOLISM OF CARBOHYDRATES | Genes involved in Metabolism of carbohydrates |