Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
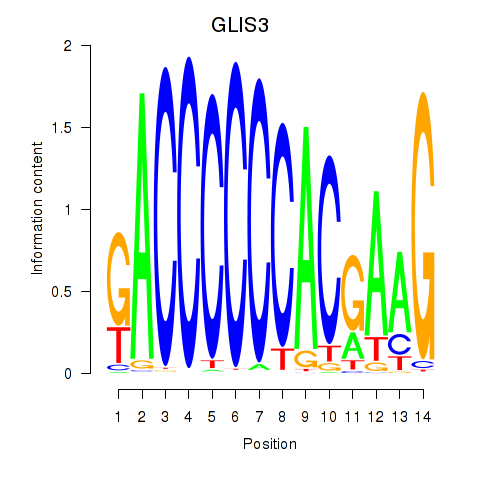
Results for GLIS3
Z-value: 0.33
Transcription factors associated with GLIS3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
GLIS3
|
ENSG00000107249.17 | GLIS family zinc finger 3 |
Activity profile of GLIS3 motif
Sorted Z-values of GLIS3 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_+_172386517 | 0.99 |
ENST00000519522.1
|
RPL26L1
|
ribosomal protein L26-like 1 |
| chr7_+_65958693 | 0.48 |
ENST00000445681.1
ENST00000452565.1 |
GS1-124K5.4
|
GS1-124K5.4 |
| chr20_-_23969416 | 0.33 |
ENST00000335694.4
|
GGTLC1
|
gamma-glutamyltransferase light chain 1 |
| chr6_+_89791507 | 0.29 |
ENST00000354922.3
|
PNRC1
|
proline-rich nuclear receptor coactivator 1 |
| chr8_-_29120580 | 0.25 |
ENST00000524189.1
|
KIF13B
|
kinesin family member 13B |
| chr19_-_38397228 | 0.25 |
ENST00000447313.2
|
WDR87
|
WD repeat domain 87 |
| chr11_-_68671264 | 0.24 |
ENST00000362034.2
|
MRPL21
|
mitochondrial ribosomal protein L21 |
| chr19_-_49828438 | 0.23 |
ENST00000454748.3
ENST00000598828.1 ENST00000335875.4 |
SLC6A16
|
solute carrier family 6, member 16 |
| chr17_-_33446820 | 0.22 |
ENST00000592577.1
ENST00000590016.1 ENST00000345365.6 ENST00000360276.3 ENST00000357906.3 |
RAD51D
|
RAD51 paralog D |
| chr21_+_38071430 | 0.19 |
ENST00000290399.6
|
SIM2
|
single-minded family bHLH transcription factor 2 |
| chr6_+_44191507 | 0.18 |
ENST00000371724.1
ENST00000371713.1 |
SLC29A1
|
solute carrier family 29 (equilibrative nucleoside transporter), member 1 |
| chr6_+_44191290 | 0.18 |
ENST00000371755.3
ENST00000371740.5 ENST00000371731.1 ENST00000393841.1 |
SLC29A1
|
solute carrier family 29 (equilibrative nucleoside transporter), member 1 |
| chr1_-_157108266 | 0.18 |
ENST00000326786.4
|
ETV3
|
ets variant 3 |
| chr4_-_74904398 | 0.18 |
ENST00000296026.4
|
CXCL3
|
chemokine (C-X-C motif) ligand 3 |
| chr17_+_48912744 | 0.17 |
ENST00000311378.4
|
WFIKKN2
|
WAP, follistatin/kazal, immunoglobulin, kunitz and netrin domain containing 2 |
| chr15_+_84116106 | 0.16 |
ENST00000535412.1
ENST00000324537.5 |
SH3GL3
|
SH3-domain GRB2-like 3 |
| chr8_+_118147498 | 0.16 |
ENST00000519688.1
ENST00000456015.2 |
SLC30A8
|
solute carrier family 30 (zinc transporter), member 8 |
| chrX_-_132549506 | 0.16 |
ENST00000370828.3
|
GPC4
|
glypican 4 |
| chr6_+_24495067 | 0.15 |
ENST00000357578.3
ENST00000546278.1 ENST00000491546.1 |
ALDH5A1
|
aldehyde dehydrogenase 5 family, member A1 |
| chr17_-_77925806 | 0.15 |
ENST00000574241.2
|
TBC1D16
|
TBC1 domain family, member 16 |
| chr17_-_5015129 | 0.14 |
ENST00000575898.1
ENST00000416429.2 |
ZNF232
|
zinc finger protein 232 |
| chr7_+_150758304 | 0.14 |
ENST00000482950.1
ENST00000463414.1 ENST00000310317.5 |
SLC4A2
|
solute carrier family 4 (anion exchanger), member 2 |
| chr1_-_92371839 | 0.14 |
ENST00000370399.2
|
TGFBR3
|
transforming growth factor, beta receptor III |
| chr1_+_107599267 | 0.14 |
ENST00000361318.5
ENST00000370078.1 |
PRMT6
|
protein arginine methyltransferase 6 |
| chr12_+_1929421 | 0.14 |
ENST00000543818.1
|
LRTM2
|
leucine-rich repeats and transmembrane domains 2 |
| chr6_+_24495185 | 0.13 |
ENST00000348925.2
|
ALDH5A1
|
aldehyde dehydrogenase 5 family, member A1 |
| chr8_-_74791051 | 0.13 |
ENST00000453587.2
ENST00000602969.1 ENST00000602593.1 ENST00000419880.3 ENST00000517608.1 |
UBE2W
|
ubiquitin-conjugating enzyme E2W (putative) |
| chr5_+_172386419 | 0.13 |
ENST00000265100.2
ENST00000519239.1 |
RPL26L1
|
ribosomal protein L26-like 1 |
| chr19_+_44488330 | 0.13 |
ENST00000591532.1
ENST00000407951.2 ENST00000270014.2 ENST00000590615.1 ENST00000586454.1 |
ZNF155
|
zinc finger protein 155 |
| chr18_+_3451584 | 0.13 |
ENST00000551541.1
|
TGIF1
|
TGFB-induced factor homeobox 1 |
| chr11_+_68671310 | 0.12 |
ENST00000255078.3
ENST00000539224.1 |
IGHMBP2
|
immunoglobulin mu binding protein 2 |
| chr19_-_56092187 | 0.12 |
ENST00000325421.4
ENST00000592239.1 |
ZNF579
|
zinc finger protein 579 |
| chr19_+_13875316 | 0.12 |
ENST00000319545.8
ENST00000593245.1 ENST00000040663.6 |
MRI1
|
methylthioribose-1-phosphate isomerase 1 |
| chr7_+_23636992 | 0.12 |
ENST00000307471.3
ENST00000409765.1 |
CCDC126
|
coiled-coil domain containing 126 |
| chr20_+_61867235 | 0.10 |
ENST00000342412.6
ENST00000217169.3 |
BIRC7
|
baculoviral IAP repeat containing 7 |
| chr3_+_120626919 | 0.10 |
ENST00000273666.6
ENST00000471454.1 ENST00000472879.1 ENST00000497029.1 ENST00000492541.1 |
STXBP5L
|
syntaxin binding protein 5-like |
| chr20_-_44718538 | 0.10 |
ENST00000290231.6
ENST00000372291.3 |
NCOA5
|
nuclear receptor coactivator 5 |
| chr19_-_48673552 | 0.09 |
ENST00000536218.1
ENST00000596549.1 |
LIG1
|
ligase I, DNA, ATP-dependent |
| chr16_-_3285144 | 0.09 |
ENST00000431561.3
ENST00000396870.4 |
ZNF200
|
zinc finger protein 200 |
| chr2_-_211168332 | 0.09 |
ENST00000341685.4
|
MYL1
|
myosin, light chain 1, alkali; skeletal, fast |
| chr22_-_50964849 | 0.09 |
ENST00000543927.1
ENST00000423348.1 |
SCO2
|
SCO2 cytochrome c oxidase assembly protein |
| chr12_+_132568814 | 0.09 |
ENST00000454179.2
ENST00000389560.2 ENST00000443539.2 ENST00000392352.1 |
EP400NL
|
EP400 N-terminal like |
| chr20_+_18447771 | 0.09 |
ENST00000377603.4
|
POLR3F
|
polymerase (RNA) III (DNA directed) polypeptide F, 39 kDa |
| chr8_+_11627148 | 0.08 |
ENST00000436750.3
|
NEIL2
|
nei endonuclease VIII-like 2 (E. coli) |
| chr1_+_1950763 | 0.08 |
ENST00000378585.4
|
GABRD
|
gamma-aminobutyric acid (GABA) A receptor, delta |
| chr22_+_18834324 | 0.08 |
ENST00000342005.4
|
AC008132.13
|
Uncharacterized protein |
| chr5_-_137090028 | 0.08 |
ENST00000314940.4
|
HNRNPA0
|
heterogeneous nuclear ribonucleoprotein A0 |
| chr20_-_18447667 | 0.08 |
ENST00000262547.5
ENST00000329494.5 ENST00000357236.4 |
DZANK1
|
double zinc ribbon and ankyrin repeat domains 1 |
| chr9_-_116061476 | 0.08 |
ENST00000441031.3
|
RNF183
|
ring finger protein 183 |
| chr2_+_220379052 | 0.08 |
ENST00000347842.3
ENST00000358078.4 |
ASIC4
|
acid-sensing (proton-gated) ion channel family member 4 |
| chr7_-_50518022 | 0.07 |
ENST00000356889.4
ENST00000420829.1 ENST00000448788.1 ENST00000395556.2 ENST00000422854.1 ENST00000435566.1 ENST00000433017.1 |
FIGNL1
|
fidgetin-like 1 |
| chr8_+_11627205 | 0.07 |
ENST00000455213.2
ENST00000403422.3 ENST00000528323.1 ENST00000284503.6 |
NEIL2
|
nei endonuclease VIII-like 2 (E. coli) |
| chr10_+_105314881 | 0.07 |
ENST00000437579.1
|
NEURL
|
neuralized E3 ubiquitin protein ligase 1 |
| chr18_+_3451646 | 0.07 |
ENST00000345133.5
ENST00000330513.5 ENST00000549546.1 |
TGIF1
|
TGFB-induced factor homeobox 1 |
| chr19_+_46850251 | 0.07 |
ENST00000012443.4
|
PPP5C
|
protein phosphatase 5, catalytic subunit |
| chr16_+_30006615 | 0.07 |
ENST00000563197.1
|
INO80E
|
INO80 complex subunit E |
| chr19_+_46850320 | 0.06 |
ENST00000391919.1
|
PPP5C
|
protein phosphatase 5, catalytic subunit |
| chr22_-_31742218 | 0.06 |
ENST00000266269.5
ENST00000405309.3 ENST00000351933.4 |
PATZ1
|
POZ (BTB) and AT hook containing zinc finger 1 |
| chr1_-_53018654 | 0.05 |
ENST00000257177.4
ENST00000355809.4 ENST00000528642.1 ENST00000470626.1 ENST00000371544.3 |
ZCCHC11
|
zinc finger, CCHC domain containing 11 |
| chr17_-_3819751 | 0.05 |
ENST00000225538.3
|
P2RX1
|
purinergic receptor P2X, ligand-gated ion channel, 1 |
| chr3_-_160167540 | 0.05 |
ENST00000496222.1
ENST00000471396.1 ENST00000471155.1 ENST00000309784.4 |
TRIM59
|
tripartite motif containing 59 |
| chr1_-_160254913 | 0.05 |
ENST00000440949.3
ENST00000368072.5 ENST00000608310.1 ENST00000556710.1 |
PEX19
DCAF8
DCAF8
|
peroxisomal biogenesis factor 19 DDB1 and CUL4 associated factor 8 DDB1- and CUL4-associated factor 8 |
| chr13_-_20767037 | 0.04 |
ENST00000382848.4
|
GJB2
|
gap junction protein, beta 2, 26kDa |
| chr22_+_31489344 | 0.04 |
ENST00000404574.1
|
SMTN
|
smoothelin |
| chr22_-_21482352 | 0.03 |
ENST00000329949.3
|
POM121L7
|
POM121 transmembrane nucleoporin-like 7 |
| chr1_-_157108130 | 0.03 |
ENST00000368192.4
|
ETV3
|
ets variant 3 |
| chr20_-_6103666 | 0.02 |
ENST00000536936.1
|
FERMT1
|
fermitin family member 1 |
| chr1_+_110527308 | 0.02 |
ENST00000369799.5
|
AHCYL1
|
adenosylhomocysteinase-like 1 |
| chr17_-_73178599 | 0.02 |
ENST00000578238.1
|
SUMO2
|
small ubiquitin-like modifier 2 |
| chr17_-_73179046 | 0.01 |
ENST00000314523.7
ENST00000420826.2 |
SUMO2
|
small ubiquitin-like modifier 2 |
| chr17_+_7792101 | 0.01 |
ENST00000358181.4
ENST00000330494.7 |
CHD3
|
chromodomain helicase DNA binding protein 3 |
| chrX_+_118370288 | 0.01 |
ENST00000535419.1
|
PGRMC1
|
progesterone receptor membrane component 1 |
| chr4_+_164265035 | 0.01 |
ENST00000338566.3
|
NPY5R
|
neuropeptide Y receptor Y5 |
| chr3_+_173116225 | 0.01 |
ENST00000457714.1
|
NLGN1
|
neuroligin 1 |
| chr15_+_91478493 | 0.01 |
ENST00000418476.2
|
UNC45A
|
unc-45 homolog A (C. elegans) |
| chr17_+_46018872 | 0.01 |
ENST00000583599.1
ENST00000434554.2 ENST00000225573.4 ENST00000544840.1 ENST00000534893.1 |
PNPO
|
pyridoxamine 5'-phosphate oxidase |
| chrX_+_118370211 | 0.01 |
ENST00000217971.7
|
PGRMC1
|
progesterone receptor membrane component 1 |
| chr20_-_62103862 | 0.01 |
ENST00000344462.4
ENST00000357249.2 ENST00000359125.2 ENST00000360480.3 ENST00000370224.1 ENST00000344425.5 ENST00000354587.3 ENST00000359689.1 |
KCNQ2
|
potassium voltage-gated channel, KQT-like subfamily, member 2 |
| chr1_-_45988542 | 0.01 |
ENST00000424390.1
|
PRDX1
|
peroxiredoxin 1 |
| chr2_+_219824357 | 0.00 |
ENST00000302625.4
|
CDK5R2
|
cyclin-dependent kinase 5, regulatory subunit 2 (p39) |
| chr22_+_50628999 | 0.00 |
ENST00000395827.1
|
TRABD
|
TraB domain containing |
| chr1_-_45987526 | 0.00 |
ENST00000372079.1
ENST00000262746.1 ENST00000447184.1 ENST00000319248.8 |
PRDX1
|
peroxiredoxin 1 |
| chr12_-_57400227 | 0.00 |
ENST00000300101.2
|
ZBTB39
|
zinc finger and BTB domain containing 39 |
| chr17_+_5389605 | 0.00 |
ENST00000576988.1
ENST00000576570.1 ENST00000573759.1 |
MIS12
|
MIS12 kinetochore complex component |
| chr19_+_11909329 | 0.00 |
ENST00000323169.5
ENST00000450087.1 |
ZNF491
|
zinc finger protein 491 |
| chr2_-_241737128 | 0.00 |
ENST00000404283.3
|
KIF1A
|
kinesin family member 1A |
Network of associatons between targets according to the STRING database.
First level regulatory network of GLIS3
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0015862 | uridine transport(GO:0015862) |
| 0.0 | 0.3 | GO:0006540 | acetate metabolic process(GO:0006083) glutamate decarboxylation to succinate(GO:0006540) gamma-aminobutyric acid catabolic process(GO:0009450) |
| 0.0 | 0.1 | GO:0034970 | histone H3-R2 methylation(GO:0034970) |
| 0.0 | 0.1 | GO:0007181 | transforming growth factor beta receptor complex assembly(GO:0007181) |
| 0.0 | 0.3 | GO:0031087 | deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.0 | 0.2 | GO:0000707 | meiotic DNA recombinase assembly(GO:0000707) |
| 0.0 | 0.3 | GO:0072383 | plus-end-directed vesicle transport along microtubule(GO:0072383) |
| 0.0 | 0.1 | GO:2000324 | positive regulation of glucocorticoid receptor signaling pathway(GO:2000324) |
| 0.0 | 0.1 | GO:0033567 | DNA replication, Okazaki fragment processing(GO:0033567) |
| 0.0 | 0.3 | GO:1901748 | leukotriene D4 metabolic process(GO:1901748) leukotriene D4 biosynthetic process(GO:1901750) |
| 0.0 | 0.2 | GO:0097011 | cellular response to granulocyte macrophage colony-stimulating factor stimulus(GO:0097011) response to granulocyte macrophage colony-stimulating factor(GO:0097012) |
| 0.0 | 0.2 | GO:0061088 | regulation of sequestering of zinc ion(GO:0061088) |
| 0.0 | 0.1 | GO:0019509 | L-methionine biosynthetic process from methylthioadenosine(GO:0019509) |
| 0.0 | 1.1 | GO:0042273 | ribosomal large subunit biogenesis(GO:0042273) |
| 0.0 | 0.1 | GO:0002554 | serotonin production involved in inflammatory response(GO:0002351) serotonin secretion involved in inflammatory response(GO:0002442) serotonin secretion by platelet(GO:0002554) |
| 0.0 | 0.0 | GO:0044752 | response to human chorionic gonadotropin(GO:0044752) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0090571 | RNA polymerase II transcription repressor complex(GO:0090571) |
| 0.0 | 0.1 | GO:0034673 | inhibin-betaglycan-ActRII complex(GO:0034673) |
| 0.0 | 0.2 | GO:0033063 | Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.0 | 0.1 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.0 | 0.3 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.0 | 0.1 | GO:0032797 | SMN complex(GO:0032797) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0035241 | protein-arginine omega-N monomethyltransferase activity(GO:0035241) |
| 0.0 | 0.2 | GO:1904929 | coreceptor activity involved in Wnt signaling pathway, planar cell polarity pathway(GO:1904929) |
| 0.0 | 0.1 | GO:0030144 | alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase activity(GO:0030144) |
| 0.0 | 0.3 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 0.0 | 0.2 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.0 | 0.1 | GO:0043141 | ATP-dependent 5'-3' DNA helicase activity(GO:0043141) |
| 0.0 | 0.1 | GO:0050265 | RNA uridylyltransferase activity(GO:0050265) |
| 0.0 | 0.2 | GO:0017151 | DEAD/H-box RNA helicase binding(GO:0017151) |
| 0.0 | 0.2 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.0 | 0.1 | GO:0005114 | type II transforming growth factor beta receptor binding(GO:0005114) |
| 0.0 | 0.4 | GO:0005337 | nucleoside transmembrane transporter activity(GO:0005337) |