Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
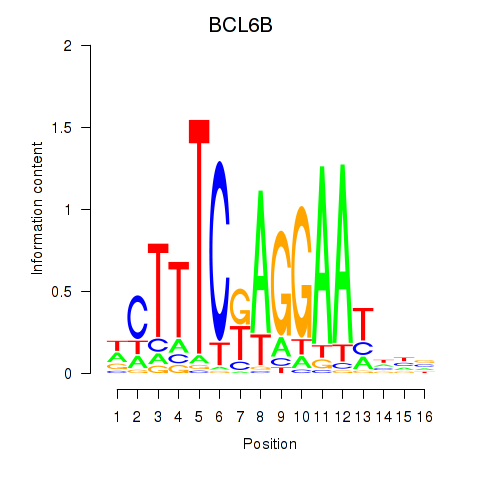
Results for BCL6B
Z-value: 0.20
Transcription factors associated with BCL6B
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
BCL6B
|
ENSG00000161940.6 | BCL6B |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| BCL6B | hg19_v2_chr17_+_6926339_6926377 | -0.18 | 6.7e-01 | Click! |
Activity profile of BCL6B motif
Sorted Z-values of BCL6B motif
Network of associatons between targets according to the STRING database.
First level regulatory network of BCL6B
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_+_7541845 | 0.33 |
ENST00000418664.2 |
DSP |
desmoplakin |
| chr2_-_31361543 | 0.29 |
ENST00000349752.5 |
GALNT14 |
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 14 (GalNAc-T14) |
| chr12_+_8975061 | 0.28 |
ENST00000299698.7 |
A2ML1 |
alpha-2-macroglobulin-like 1 |
| chr6_+_7541808 | 0.26 |
ENST00000379802.3 |
DSP |
desmoplakin |
| chr12_-_123187890 | 0.26 |
ENST00000328880.5 |
HCAR2 |
hydroxycarboxylic acid receptor 2 |
| chr12_-_123201337 | 0.25 |
ENST00000528880.2 |
HCAR3 |
hydroxycarboxylic acid receptor 3 |
| chr6_-_136788001 | 0.23 |
ENST00000544465.1 |
MAP7 |
microtubule-associated protein 7 |
| chr6_-_29343068 | 0.22 |
ENST00000396806.3 |
OR12D3 |
olfactory receptor, family 12, subfamily D, member 3 |
| chr4_+_69313145 | 0.22 |
ENST00000305363.4 |
TMPRSS11E |
transmembrane protease, serine 11E |
| chr18_-_66382289 | 0.19 |
ENST00000443099.2 ENST00000562706.1 ENST00000544714.2 |
TMX3 |
thioredoxin-related transmembrane protein 3 |
| chr12_-_42877764 | 0.19 |
ENST00000455697.1 |
PRICKLE1 |
prickle homolog 1 (Drosophila) |
| chr9_-_117880477 | 0.18 |
ENST00000534839.1 ENST00000340094.3 ENST00000535648.1 ENST00000346706.3 ENST00000345230.3 ENST00000350763.4 |
TNC |
tenascin C |
| chr12_-_42877726 | 0.18 |
ENST00000548696.1 |
PRICKLE1 |
prickle homolog 1 (Drosophila) |
| chr6_+_44194762 | 0.18 |
ENST00000371708.1 |
SLC29A1 |
solute carrier family 29 (equilibrative nucleoside transporter), member 1 |
| chr14_+_95078714 | 0.17 |
ENST00000393078.3 ENST00000393080.4 ENST00000467132.1 |
SERPINA3 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 3 |
| chr8_-_91095099 | 0.17 |
ENST00000265431.3 |
CALB1 |
calbindin 1, 28kDa |
| chr20_+_58251716 | 0.16 |
ENST00000355648.4 |
PHACTR3 |
phosphatase and actin regulator 3 |
| chr18_-_52989217 | 0.15 |
ENST00000570287.2 |
TCF4 |
transcription factor 4 |
| chr18_-_52989525 | 0.15 |
ENST00000457482.3 |
TCF4 |
transcription factor 4 |
| chr19_+_11200038 | 0.13 |
ENST00000558518.1 ENST00000557933.1 ENST00000455727.2 ENST00000535915.1 ENST00000545707.1 ENST00000558013.1 |
LDLR |
low density lipoprotein receptor |
| chr7_-_29186008 | 0.13 |
ENST00000396276.3 ENST00000265394.5 |
CPVL |
carboxypeptidase, vitellogenic-like |
| chr3_-_123512688 | 0.12 |
ENST00000475616.1 |
MYLK |
myosin light chain kinase |
| chr12_+_19358228 | 0.12 |
ENST00000424268.1 ENST00000543806.1 |
PLEKHA5 |
pleckstrin homology domain containing, family A member 5 |
| chr12_-_122985067 | 0.12 |
ENST00000540586.1 ENST00000543897.1 |
ZCCHC8 |
zinc finger, CCHC domain containing 8 |
| chr6_-_155776966 | 0.12 |
ENST00000159060.2 |
NOX3 |
NADPH oxidase 3 |
| chr7_+_21582638 | 0.12 |
ENST00000409508.3 ENST00000328843.6 |
DNAH11 |
dynein, axonemal, heavy chain 11 |
| chr12_-_55367361 | 0.11 |
ENST00000532804.1 ENST00000531122.1 ENST00000533446.1 |
TESPA1 |
thymocyte expressed, positive selection associated 1 |
| chr16_-_75498308 | 0.11 |
ENST00000569540.1 |
TMEM170A |
transmembrane protein 170A |
| chr6_+_42584847 | 0.10 |
ENST00000372883.3 |
UBR2 |
ubiquitin protein ligase E3 component n-recognin 2 |
| chr11_+_113779704 | 0.10 |
ENST00000537778.1 |
HTR3B |
5-hydroxytryptamine (serotonin) receptor 3B, ionotropic |
| chr5_+_80529104 | 0.10 |
ENST00000254035.4 ENST00000511719.1 ENST00000437669.1 ENST00000424301.2 ENST00000505060.1 |
CKMT2 |
creatine kinase, mitochondrial 2 (sarcomeric) |
| chr19_-_13227463 | 0.10 |
ENST00000437766.1 ENST00000221504.8 |
TRMT1 |
tRNA methyltransferase 1 homolog (S. cerevisiae) |
| chr19_-_13227534 | 0.10 |
ENST00000588229.1 ENST00000357720.4 |
TRMT1 |
tRNA methyltransferase 1 homolog (S. cerevisiae) |
| chr3_-_123603137 | 0.10 |
ENST00000360304.3 ENST00000359169.1 ENST00000346322.5 ENST00000360772.3 |
MYLK |
myosin light chain kinase |
| chr16_-_67517716 | 0.09 |
ENST00000290953.2 |
AGRP |
agouti related protein homolog (mouse) |
| chr3_+_35683651 | 0.09 |
ENST00000187397.4 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chr12_-_122985494 | 0.08 |
ENST00000336229.4 |
ZCCHC8 |
zinc finger, CCHC domain containing 8 |
| chr13_-_33112823 | 0.08 |
ENST00000504114.1 |
N4BP2L2 |
NEDD4 binding protein 2-like 2 |
| chr10_-_62493223 | 0.08 |
ENST00000373827.2 |
ANK3 |
ankyrin 3, node of Ranvier (ankyrin G) |
| chr12_+_104324112 | 0.08 |
ENST00000299767.5 |
HSP90B1 |
heat shock protein 90kDa beta (Grp94), member 1 |
| chr12_-_10151773 | 0.08 |
ENST00000298527.6 ENST00000348658.4 |
CLEC1B |
C-type lectin domain family 1, member B |
| chr19_+_45147098 | 0.08 |
ENST00000425690.3 ENST00000344956.4 ENST00000403059.4 |
PVR |
poliovirus receptor |
| chr15_+_40886199 | 0.07 |
ENST00000346991.5 ENST00000528975.1 ENST00000527044.1 |
CASC5 |
cancer susceptibility candidate 5 |
| chr2_+_120189422 | 0.07 |
ENST00000306406.4 |
TMEM37 |
transmembrane protein 37 |
| chr1_-_41131326 | 0.07 |
ENST00000372684.3 |
RIMS3 |
regulating synaptic membrane exocytosis 3 |
| chr17_-_42907564 | 0.07 |
ENST00000592524.1 |
GJC1 |
gap junction protein, gamma 1, 45kDa |
| chr1_-_153521714 | 0.07 |
ENST00000368713.3 |
S100A3 |
S100 calcium binding protein A3 |
| chr13_-_33112899 | 0.07 |
ENST00000267068.3 ENST00000357505.6 ENST00000399396.3 |
N4BP2L2 |
NEDD4 binding protein 2-like 2 |
| chr8_-_130799134 | 0.07 |
ENST00000276708.4 |
GSDMC |
gasdermin C |
| chr9_+_88556036 | 0.07 |
ENST00000361671.5 ENST00000416045.1 |
NAA35 |
N(alpha)-acetyltransferase 35, NatC auxiliary subunit |
| chr17_+_4981535 | 0.07 |
ENST00000318833.3 |
ZFP3 |
ZFP3 zinc finger protein |
| chr1_-_153521597 | 0.07 |
ENST00000368712.1 |
S100A3 |
S100 calcium binding protein A3 |
| chr4_-_76957214 | 0.06 |
ENST00000306621.3 |
CXCL11 |
chemokine (C-X-C motif) ligand 11 |
| chr12_-_122238913 | 0.06 |
ENST00000537157.1 |
AC084018.1 |
AC084018.1 |
| chr5_+_140579162 | 0.06 |
ENST00000536699.1 ENST00000354757.3 |
PCDHB11 |
protocadherin beta 11 |
| chr17_+_79761997 | 0.06 |
ENST00000400723.3 ENST00000570996.1 |
GCGR |
glucagon receptor |
| chr1_+_158815588 | 0.06 |
ENST00000438394.1 |
MNDA |
myeloid cell nuclear differentiation antigen |
| chrX_+_139791917 | 0.06 |
ENST00000607004.1 ENST00000370535.3 |
LINC00632 |
long intergenic non-protein coding RNA 632 |
| chr15_+_65843130 | 0.06 |
ENST00000569894.1 |
PTPLAD1 |
protein tyrosine phosphatase-like A domain containing 1 |
| chr6_+_31588478 | 0.06 |
ENST00000376007.4 ENST00000376033.2 |
PRRC2A |
proline-rich coiled-coil 2A |
| chr1_+_179699318 | 0.06 |
ENST00000423879.1 |
RP11-12M5.1 |
RP11-12M5.1 |
| chr19_+_45147313 | 0.06 |
ENST00000406449.4 |
PVR |
poliovirus receptor |
| chr19_-_47616992 | 0.06 |
ENST00000253048.5 |
ZC3H4 |
zinc finger CCCH-type containing 4 |
| chr1_-_150669604 | 0.06 |
ENST00000427665.1 ENST00000540514.1 |
GOLPH3L |
golgi phosphoprotein 3-like |
| chr11_-_104480019 | 0.06 |
ENST00000536529.1 ENST00000545630.1 ENST00000538641.1 |
RP11-886D15.1 |
RP11-886D15.1 |
| chr2_+_239756671 | 0.05 |
ENST00000448943.2 |
TWIST2 |
twist family bHLH transcription factor 2 |
| chr8_+_67039278 | 0.05 |
ENST00000276573.7 ENST00000350034.4 |
TRIM55 |
tripartite motif containing 55 |
| chr6_-_32083106 | 0.05 |
ENST00000442721.1 |
TNXB |
tenascin XB |
| chrX_-_134936550 | 0.05 |
ENST00000494421.1 |
CT45A4 |
cancer/testis antigen family 45, member A4 |
| chr12_-_10007448 | 0.05 |
ENST00000538152.1 |
CLEC2B |
C-type lectin domain family 2, member B |
| chr3_-_187455680 | 0.05 |
ENST00000438077.1 |
BCL6 |
B-cell CLL/lymphoma 6 |
| chr6_+_125540951 | 0.05 |
ENST00000524679.1 |
TPD52L1 |
tumor protein D52-like 1 |
| chr9_-_86322831 | 0.05 |
ENST00000257468.7 |
UBQLN1 |
ubiquilin 1 |
| chrX_+_95939638 | 0.05 |
ENST00000373061.3 ENST00000373054.4 ENST00000355827.4 |
DIAPH2 |
diaphanous-related formin 2 |
| chr4_+_70796784 | 0.05 |
ENST00000246891.4 ENST00000444405.3 |
CSN1S1 |
casein alpha s1 |
| chrX_+_134847370 | 0.05 |
ENST00000482795.1 |
CT45A1 |
cancer/testis antigen family 45, member A1 |
| chr10_+_18240814 | 0.05 |
ENST00000377374.4 |
SLC39A12 |
solute carrier family 39 (zinc transporter), member 12 |
| chr2_-_55237484 | 0.05 |
ENST00000394609.2 |
RTN4 |
reticulon 4 |
| chr11_+_64059464 | 0.05 |
ENST00000394525.2 |
KCNK4 |
potassium channel, subfamily K, member 4 |
| chr1_-_212004090 | 0.04 |
ENST00000366997.4 |
LPGAT1 |
lysophosphatidylglycerol acyltransferase 1 |
| chrX_-_135338503 | 0.04 |
ENST00000370663.5 |
MAP7D3 |
MAP7 domain containing 3 |
| chr2_+_169659121 | 0.04 |
ENST00000397206.2 ENST00000397209.2 ENST00000421711.2 |
NOSTRIN |
nitric oxide synthase trafficking |
| chr15_+_45003675 | 0.04 |
ENST00000558401.1 ENST00000559916.1 ENST00000544417.1 |
B2M |
beta-2-microglobulin |
| chr8_+_104427581 | 0.04 |
ENST00000521716.1 ENST00000521971.1 ENST00000519682.1 |
DCAF13 |
DDB1 and CUL4 associated factor 13 |
| chr13_-_73356234 | 0.04 |
ENST00000545453.1 |
DIS3 |
DIS3 mitotic control homolog (S. cerevisiae) |
| chr13_+_52436111 | 0.04 |
ENST00000242819.4 |
CCDC70 |
coiled-coil domain containing 70 |
| chr17_-_42908155 | 0.04 |
ENST00000426548.1 ENST00000590758.1 ENST00000591424.1 |
GJC1 |
gap junction protein, gamma 1, 45kDa |
| chr5_+_140501581 | 0.04 |
ENST00000194152.1 |
PCDHB4 |
protocadherin beta 4 |
| chr3_+_102153859 | 0.04 |
ENST00000306176.1 ENST00000466937.1 |
ZPLD1 |
zona pellucida-like domain containing 1 |
| chr7_+_23719749 | 0.04 |
ENST00000409192.3 ENST00000344962.4 ENST00000409653.1 ENST00000409994.3 |
FAM221A |
family with sequence similarity 221, member A |
| chr12_-_43945724 | 0.04 |
ENST00000389420.3 ENST00000553158.1 |
ADAMTS20 |
ADAM metallopeptidase with thrombospondin type 1 motif, 20 |
| chr8_-_107782463 | 0.04 |
ENST00000311955.3 |
ABRA |
actin-binding Rho activating protein |
| chr5_-_58295712 | 0.04 |
ENST00000317118.8 |
PDE4D |
phosphodiesterase 4D, cAMP-specific |
| chr2_+_169658928 | 0.04 |
ENST00000317647.7 ENST00000445023.2 |
NOSTRIN |
nitric oxide synthase trafficking |
| chr12_-_62997214 | 0.04 |
ENST00000408887.2 |
C12orf61 |
chromosome 12 open reading frame 61 |
| chr11_+_58874701 | 0.03 |
ENST00000529618.1 ENST00000534403.1 ENST00000343597.3 |
FAM111B |
family with sequence similarity 111, member B |
| chr15_+_69854027 | 0.03 |
ENST00000498938.2 |
RP11-279F6.1 |
RP11-279F6.1 |
| chr13_-_28194541 | 0.03 |
ENST00000316334.3 |
LNX2 |
ligand of numb-protein X 2 |
| chr15_-_38519066 | 0.03 |
ENST00000561320.1 ENST00000561161.1 |
RP11-346D14.1 |
RP11-346D14.1 |
| chr4_-_153303658 | 0.03 |
ENST00000296555.5 |
FBXW7 |
F-box and WD repeat domain containing 7, E3 ubiquitin protein ligase |
| chr14_+_101123580 | 0.03 |
ENST00000556697.1 ENST00000360899.2 ENST00000553623.1 |
LINC00523 |
long intergenic non-protein coding RNA 523 |
| chr4_+_76995855 | 0.03 |
ENST00000355810.4 ENST00000349321.3 |
ART3 |
ADP-ribosyltransferase 3 |
| chr12_+_16500037 | 0.03 |
ENST00000536371.1 ENST00000010404.2 |
MGST1 |
microsomal glutathione S-transferase 1 |
| chr20_-_18447667 | 0.03 |
ENST00000262547.5 ENST00000329494.5 ENST00000357236.4 |
DZANK1 |
double zinc ribbon and ankyrin repeat domains 1 |
| chr12_-_11175219 | 0.03 |
ENST00000390673.2 |
TAS2R19 |
taste receptor, type 2, member 19 |
| chr12_+_20522179 | 0.03 |
ENST00000359062.3 |
PDE3A |
phosphodiesterase 3A, cGMP-inhibited |
| chrX_+_79675965 | 0.03 |
ENST00000308293.5 |
FAM46D |
family with sequence similarity 46, member D |
| chr2_+_143635222 | 0.03 |
ENST00000375773.2 ENST00000409512.1 ENST00000410015.2 |
KYNU |
kynureninase |
| chr2_+_143635067 | 0.03 |
ENST00000264170.4 |
KYNU |
kynureninase |
| chr14_-_94970559 | 0.03 |
ENST00000556881.1 |
SERPINA12 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 12 |
| chr18_+_72167096 | 0.03 |
ENST00000324301.8 |
CNDP2 |
CNDP dipeptidase 2 (metallopeptidase M20 family) |
| chrX_-_134971059 | 0.03 |
ENST00000472834.1 |
CT45A6 |
cancer/testis antigen family 45, member A6 |
| chr10_+_11207438 | 0.03 |
ENST00000609692.1 ENST00000354897.3 |
CELF2 |
CUGBP, Elav-like family member 2 |
| chrX_-_139866723 | 0.03 |
ENST00000370532.2 |
CDR1 |
cerebellar degeneration-related protein 1, 34kDa |
| chr11_-_18258342 | 0.03 |
ENST00000278222.4 |
SAA4 |
serum amyloid A4, constitutive |
| chr9_+_70971815 | 0.03 |
ENST00000396392.1 ENST00000396396.1 |
PGM5 |
phosphoglucomutase 5 |
| chr7_+_872107 | 0.03 |
ENST00000405266.1 ENST00000401592.1 ENST00000403868.1 ENST00000425407.2 |
SUN1 |
Sad1 and UNC84 domain containing 1 |
| chr3_-_69101461 | 0.03 |
ENST00000543976.1 |
TMF1 |
TATA element modulatory factor 1 |
| chr3_-_69101413 | 0.03 |
ENST00000398559.2 |
TMF1 |
TATA element modulatory factor 1 |
| chr13_-_33112956 | 0.03 |
ENST00000505213.1 |
N4BP2L2 |
NEDD4 binding protein 2-like 2 |
| chr13_-_73356009 | 0.03 |
ENST00000377780.4 ENST00000377767.4 |
DIS3 |
DIS3 mitotic control homolog (S. cerevisiae) |
| chr2_-_89266286 | 0.03 |
ENST00000464162.1 |
IGKV1-6 |
immunoglobulin kappa variable 1-6 |
| chr2_-_216257849 | 0.03 |
ENST00000456923.1 |
FN1 |
fibronectin 1 |
| chr8_+_50824233 | 0.02 |
ENST00000522124.1 |
SNTG1 |
syntrophin, gamma 1 |
| chr11_+_61159832 | 0.02 |
ENST00000334888.5 ENST00000398979.3 |
TMEM216 |
transmembrane protein 216 |
| chr17_+_76183432 | 0.02 |
ENST00000591256.1 ENST00000589256.1 ENST00000588800.1 ENST00000591952.1 ENST00000327898.5 ENST00000586542.1 ENST00000586731.1 |
AFMID |
arylformamidase |
| chr6_-_28973037 | 0.02 |
ENST00000377179.3 |
ZNF311 |
zinc finger protein 311 |
| chr19_-_460996 | 0.02 |
ENST00000264554.6 |
SHC2 |
SHC (Src homology 2 domain containing) transforming protein 2 |
| chr11_+_120081475 | 0.02 |
ENST00000328965.4 |
OAF |
OAF homolog (Drosophila) |
| chr17_-_48474828 | 0.02 |
ENST00000576448.1 ENST00000225972.7 |
LRRC59 |
leucine rich repeat containing 59 |
| chr6_+_150690133 | 0.02 |
ENST00000392255.3 ENST00000500320.3 |
IYD |
iodotyrosine deiodinase |
| chr1_-_153321301 | 0.02 |
ENST00000368739.3 |
PGLYRP4 |
peptidoglycan recognition protein 4 |
| chr10_+_18240834 | 0.02 |
ENST00000377371.3 ENST00000539911.1 |
SLC39A12 |
solute carrier family 39 (zinc transporter), member 12 |
| chr5_+_140535577 | 0.02 |
ENST00000539533.1 |
PCDHB17 |
Protocadherin-psi1; Uncharacterized protein |
| chr6_-_29324054 | 0.02 |
ENST00000543825.1 |
OR5V1 |
olfactory receptor, family 5, subfamily V, member 1 |
| chr17_-_39296739 | 0.02 |
ENST00000345847.4 |
KRTAP4-6 |
keratin associated protein 4-6 |
| chrX_-_134936735 | 0.02 |
ENST00000487941.2 ENST00000434966.2 |
CT45A4 |
cancer/testis antigen family 45, member A4 |
| chr19_-_4831701 | 0.02 |
ENST00000248244.5 |
TICAM1 |
toll-like receptor adaptor molecule 1 |
| chrX_+_134847185 | 0.02 |
ENST00000370741.3 ENST00000497301.2 |
CT45A1 |
cancer/testis antigen family 45, member A1 |
| chr14_+_31028348 | 0.02 |
ENST00000550944.1 ENST00000438909.2 ENST00000553504.1 |
G2E3 |
G2/M-phase specific E3 ubiquitin protein ligase |
| chr17_+_66539369 | 0.02 |
ENST00000600820.1 |
AC079210.1 |
Uncharacterized protein; cDNA FLJ45097 fis, clone BRAWH3031054 |
| chrX_+_153775821 | 0.02 |
ENST00000263518.6 ENST00000470142.1 ENST00000393549.2 ENST00000455588.2 ENST00000369602.3 |
IKBKG |
inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase gamma |
| chr4_-_99578789 | 0.02 |
ENST00000511651.1 ENST00000505184.1 |
TSPAN5 |
tetraspanin 5 |
| chr3_-_50649192 | 0.02 |
ENST00000443053.2 ENST00000348721.3 |
CISH |
cytokine inducible SH2-containing protein |
| chr8_+_67039131 | 0.02 |
ENST00000315962.4 ENST00000353317.5 |
TRIM55 |
tripartite motif containing 55 |
| chr4_+_89378261 | 0.02 |
ENST00000264350.3 |
HERC5 |
HECT and RLD domain containing E3 ubiquitin protein ligase 5 |
| chr1_-_230850043 | 0.02 |
ENST00000366667.4 |
AGT |
angiotensinogen (serpin peptidase inhibitor, clade A, member 8) |
| chr6_-_108145499 | 0.02 |
ENST00000369020.3 ENST00000369022.2 |
SCML4 |
sex comb on midleg-like 4 (Drosophila) |
| chr15_-_49913126 | 0.02 |
ENST00000561064.1 ENST00000299338.6 |
FAM227B |
family with sequence similarity 227, member B |
| chr16_+_16481598 | 0.02 |
ENST00000327792.5 |
NPIPA7 |
nuclear pore complex interacting protein family, member A7 |
| chr20_-_4229721 | 0.01 |
ENST00000379453.4 |
ADRA1D |
adrenoceptor alpha 1D |
| chr15_-_64673665 | 0.01 |
ENST00000300035.4 |
KIAA0101 |
KIAA0101 |
| chr20_+_56136136 | 0.01 |
ENST00000319441.4 ENST00000543666.1 |
PCK1 |
phosphoenolpyruvate carboxykinase 1 (soluble) |
| chr8_-_81787006 | 0.01 |
ENST00000327835.3 |
ZNF704 |
zinc finger protein 704 |
| chr1_+_209878182 | 0.01 |
ENST00000367027.3 |
HSD11B1 |
hydroxysteroid (11-beta) dehydrogenase 1 |
| chrX_-_134971244 | 0.01 |
ENST00000491002.2 ENST00000448053.2 |
CT45A6 |
cancer/testis antigen family 45, member A6 |
| chr4_+_95972822 | 0.01 |
ENST00000509540.1 ENST00000440890.2 |
BMPR1B |
bone morphogenetic protein receptor, type IB |
| chr19_-_7812446 | 0.01 |
ENST00000394173.4 ENST00000301357.8 |
CD209 |
CD209 molecule |
| chr9_+_139702363 | 0.01 |
ENST00000371663.4 ENST00000371671.4 ENST00000311502.7 |
RABL6 |
RAB, member RAS oncogene family-like 6 |
| chr12_+_70132632 | 0.01 |
ENST00000378815.6 ENST00000483530.2 ENST00000325555.9 |
RAB3IP |
RAB3A interacting protein |
| chr3_+_155860751 | 0.01 |
ENST00000471742.1 |
KCNAB1 |
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chr20_+_43990576 | 0.01 |
ENST00000372727.1 ENST00000414310.1 |
SYS1 |
SYS1 Golgi-localized integral membrane protein homolog (S. cerevisiae) |
| chr5_+_140480083 | 0.01 |
ENST00000231130.2 |
PCDHB3 |
protocadherin beta 3 |
| chr1_-_108735440 | 0.01 |
ENST00000370041.4 |
SLC25A24 |
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 24 |
| chr2_+_173686303 | 0.01 |
ENST00000397087.3 |
RAPGEF4 |
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr13_+_73356197 | 0.01 |
ENST00000326291.6 |
PIBF1 |
progesterone immunomodulatory binding factor 1 |
| chr3_+_23958632 | 0.01 |
ENST00000412097.1 ENST00000415719.1 ENST00000435882.1 |
RPL15 |
ribosomal protein L15 |
| chr13_-_41345277 | 0.01 |
ENST00000323563.6 |
MRPS31 |
mitochondrial ribosomal protein S31 |
| chr7_-_122840015 | 0.00 |
ENST00000194130.2 |
SLC13A1 |
solute carrier family 13 (sodium/sulfate symporter), member 1 |
| chr3_+_44690211 | 0.00 |
ENST00000396056.2 ENST00000432115.2 ENST00000415571.2 ENST00000399560.2 ENST00000296092.3 ENST00000542250.1 ENST00000453164.1 |
ZNF35 |
zinc finger protein 35 |
| chr5_+_72251793 | 0.00 |
ENST00000430046.2 ENST00000341845.6 |
FCHO2 |
FCH domain only 2 |
| chr2_-_69098566 | 0.00 |
ENST00000295379.1 |
BMP10 |
bone morphogenetic protein 10 |
| chr7_+_2636522 | 0.00 |
ENST00000423196.1 |
IQCE |
IQ motif containing E |
| chr4_-_13546632 | 0.00 |
ENST00000382438.5 |
NKX3-2 |
NK3 homeobox 2 |
| chr19_-_38183201 | 0.00 |
ENST00000590008.1 ENST00000358582.4 |
ZNF781 |
zinc finger protein 781 |
| chr17_-_60883993 | 0.00 |
ENST00000583803.1 ENST00000456609.2 |
MARCH10 |
membrane-associated ring finger (C3HC4) 10, E3 ubiquitin protein ligase |
| chr11_+_73358594 | 0.00 |
ENST00000227214.6 ENST00000398494.4 ENST00000543085.1 |
PLEKHB1 |
pleckstrin homology domain containing, family B (evectins) member 1 |
| chr16_-_18441131 | 0.00 |
ENST00000339303.5 |
NPIPA8 |
nuclear pore complex interacting protein family, member A8 |
| chr19_-_7812397 | 0.00 |
ENST00000593660.1 ENST00000354397.6 ENST00000593821.1 ENST00000602261.1 ENST00000315591.8 ENST00000394161.5 ENST00000204801.8 ENST00000601256.1 ENST00000601951.1 ENST00000315599.7 |
CD209 |
CD209 molecule |
| chr3_+_46618727 | 0.00 |
ENST00000296145.5 |
TDGF1 |
teratocarcinoma-derived growth factor 1 |
| chr22_+_45725524 | 0.00 |
ENST00000405548.3 |
FAM118A |
family with sequence similarity 118, member A |
| chr7_+_74072011 | 0.00 |
ENST00000324896.4 ENST00000353920.4 ENST00000346152.4 ENST00000416070.1 |
GTF2I |
general transcription factor IIi |
| chrX_+_95939711 | 0.00 |
ENST00000373049.4 ENST00000324765.8 |
DIAPH2 |
diaphanous-related formin 2 |
| chr3_+_35682913 | 0.00 |
ENST00000449196.1 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chr13_+_20207782 | 0.00 |
ENST00000414242.2 ENST00000361479.5 |
MPHOSPH8 |
M-phase phosphoprotein 8 |
| chr8_-_102216925 | 0.00 |
ENST00000517844.1 |
ZNF706 |
zinc finger protein 706 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:2000690 | negative regulation of cardioblast differentiation(GO:0051892) regulation of cardiac muscle cell myoblast differentiation(GO:2000690) negative regulation of cardiac muscle cell myoblast differentiation(GO:2000691) |
| 0.1 | 0.6 | GO:0071896 | protein localization to adherens junction(GO:0071896) bundle of His cell-Purkinje myocyte adhesion involved in cell communication(GO:0086073) |
| 0.0 | 0.2 | GO:0018171 | peptidyl-cysteine oxidation(GO:0018171) |
| 0.0 | 0.1 | GO:0060370 | susceptibility to T cell mediated cytotoxicity(GO:0060370) |
| 0.0 | 0.3 | GO:0033031 | positive regulation of neutrophil apoptotic process(GO:0033031) |
| 0.0 | 0.2 | GO:0060447 | bud outgrowth involved in lung branching(GO:0060447) |
| 0.0 | 0.2 | GO:0015862 | uridine transport(GO:0015862) |
| 0.0 | 0.1 | GO:0090118 | receptor-mediated endocytosis of low-density lipoprotein particle involved in cholesterol transport(GO:0090118) positive regulation of lysosomal protein catabolic process(GO:1905167) |
| 0.0 | 0.1 | GO:0086021 | SA node cell to atrial cardiac muscle cell communication by electrical coupling(GO:0086021) |
| 0.0 | 0.1 | GO:0010387 | COP9 signalosome assembly(GO:0010387) |
| 0.0 | 0.1 | GO:0031247 | actin rod assembly(GO:0031247) |
| 0.0 | 0.2 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 0.0 | 0.2 | GO:1902035 | positive regulation of hematopoietic stem cell proliferation(GO:1902035) |
| 0.0 | 0.1 | GO:0071596 | ubiquitin-dependent protein catabolic process via the N-end rule pathway(GO:0071596) |
| 0.0 | 0.1 | GO:0072658 | maintenance of protein location in membrane(GO:0072658) maintenance of protein location in plasma membrane(GO:0072660) positive regulation of membrane depolarization during cardiac muscle cell action potential(GO:1900827) |
| 0.0 | 0.1 | GO:0019442 | tryptophan catabolic process to acetyl-CoA(GO:0019442) |
| 0.0 | 0.2 | GO:0035502 | metanephric part of ureteric bud development(GO:0035502) |
| 0.0 | 0.1 | GO:0032764 | negative regulation of mast cell cytokine production(GO:0032764) |
| 0.0 | 0.0 | GO:1904432 | positive regulation of iron ion transport(GO:0034758) positive regulation of iron ion transmembrane transport(GO:0034761) regulation of iron ion import(GO:1900390) regulation of ferrous iron import into cell(GO:1903989) positive regulation of ferrous iron import into cell(GO:1903991) regulation of ferrous iron binding(GO:1904432) positive regulation of ferrous iron binding(GO:1904434) regulation of transferrin receptor binding(GO:1904435) positive regulation of transferrin receptor binding(GO:1904437) regulation of ferrous iron import across plasma membrane(GO:1904438) positive regulation of ferrous iron import across plasma membrane(GO:1904440) |
| 0.0 | 0.1 | GO:0048840 | otolith development(GO:0048840) |
| 0.0 | 0.1 | GO:0071034 | CUT catabolic process(GO:0071034) CUT metabolic process(GO:0071043) |
| 0.0 | 0.1 | GO:0034140 | negative regulation of toll-like receptor 3 signaling pathway(GO:0034140) |
| 0.0 | 0.0 | GO:0071469 | detection of mechanical stimulus involved in sensory perception of touch(GO:0050976) cellular response to alkaline pH(GO:0071469) |
| 0.0 | 0.0 | GO:0071449 | cellular response to lipid hydroperoxide(GO:0071449) |
| 0.0 | 0.1 | GO:2000845 | testosterone secretion(GO:0035936) regulation of testosterone secretion(GO:2000843) positive regulation of testosterone secretion(GO:2000845) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:1990666 | PCSK9-LDLR complex(GO:1990666) |
| 0.0 | 0.6 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.0 | 0.1 | GO:0031417 | NatC complex(GO:0031417) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0086083 | cell adhesive protein binding involved in bundle of His cell-Purkinje myocyte communication(GO:0086083) |
| 0.0 | 0.2 | GO:0004809 | tRNA (guanine-N2-)-methyltransferase activity(GO:0004809) |
| 0.0 | 0.2 | GO:0016972 | thiol oxidase activity(GO:0016972) |
| 0.0 | 0.1 | GO:0086020 | gap junction channel activity involved in SA node cell-atrial cardiac muscle cell electrical coupling(GO:0086020) |
| 0.0 | 0.2 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.0 | 0.3 | GO:0001087 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.0 | 0.2 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.0 | 0.1 | GO:0004967 | glucagon receptor activity(GO:0004967) |
| 0.0 | 0.1 | GO:0030229 | very-low-density lipoprotein particle receptor activity(GO:0030229) |
| 0.0 | 0.2 | GO:0045545 | syndecan binding(GO:0045545) |
| 0.0 | 0.1 | GO:0022850 | serotonin-gated cation channel activity(GO:0022850) |
| 0.0 | 0.0 | GO:0098782 | mechanically-gated potassium channel activity(GO:0098782) |
| 0.0 | 0.1 | GO:0070728 | leucine binding(GO:0070728) |
| 0.0 | 0.1 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.0 | 0.1 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 0.0 | 0.3 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.0 | 0.0 | GO:0004119 | cGMP-inhibited cyclic-nucleotide phosphodiesterase activity(GO:0004119) |
| 0.0 | 0.1 | GO:0016823 | hydrolase activity, acting on acid carbon-carbon bonds(GO:0016822) hydrolase activity, acting on acid carbon-carbon bonds, in ketonic substances(GO:0016823) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |