Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
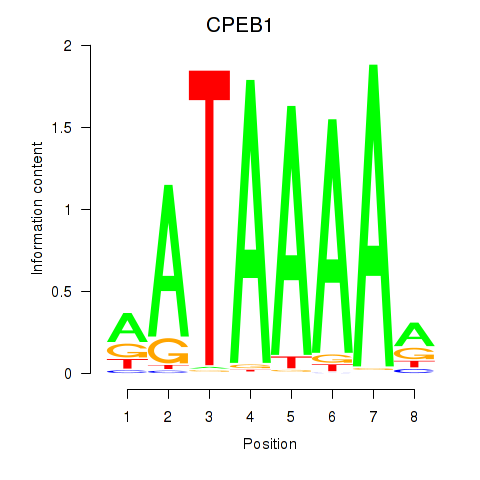
Results for CPEB1
Z-value: 0.05
Transcription factors associated with CPEB1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
CPEB1
|
ENSG00000214575.5 | CPEB1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| CPEB1 | hg19_v2_chr15_-_83316087_83316137, hg19_v2_chr15_-_83316711_83316728 | -0.64 | 9.0e-02 | Click! |
Activity profile of CPEB1 motif
Sorted Z-values of CPEB1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of CPEB1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_-_100027943 | 0.63 |
ENST00000260702.3 |
LOXL4 |
lysyl oxidase-like 4 |
| chr10_-_106240032 | 0.53 |
ENST00000447860.1 |
RP11-127O4.3 |
RP11-127O4.3 |
| chr1_+_82266053 | 0.39 |
ENST00000370715.1 ENST00000370713.1 ENST00000319517.6 ENST00000370717.2 ENST00000394879.1 ENST00000271029.4 ENST00000335786.5 |
LPHN2 |
latrophilin 2 |
| chr22_+_29876197 | 0.37 |
ENST00000310624.6 |
NEFH |
neurofilament, heavy polypeptide |
| chr3_-_155394152 | 0.33 |
ENST00000494598.1 |
PLCH1 |
phospholipase C, eta 1 |
| chr17_-_53800217 | 0.31 |
ENST00000424486.2 |
TMEM100 |
transmembrane protein 100 |
| chr8_+_104831472 | 0.30 |
ENST00000262231.10 ENST00000507740.1 |
RIMS2 |
regulating synaptic membrane exocytosis 2 |
| chr8_-_93029865 | 0.29 |
ENST00000422361.2 |
RUNX1T1 |
runt-related transcription factor 1; translocated to, 1 (cyclin D-related) |
| chr7_-_83824169 | 0.28 |
ENST00000265362.4 |
SEMA3A |
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3A |
| chr4_+_74735102 | 0.27 |
ENST00000395761.3 |
CXCL1 |
chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) |
| chr11_-_102651343 | 0.26 |
ENST00000279441.4 ENST00000539681.1 |
MMP10 |
matrix metallopeptidase 10 (stromelysin 2) |
| chr18_+_19749386 | 0.25 |
ENST00000269216.3 |
GATA6 |
GATA binding protein 6 |
| chr6_-_46293378 | 0.24 |
ENST00000330430.6 |
RCAN2 |
regulator of calcineurin 2 |
| chr18_+_61554932 | 0.24 |
ENST00000299502.4 ENST00000457692.1 ENST00000413956.1 |
SERPINB2 |
serpin peptidase inhibitor, clade B (ovalbumin), member 2 |
| chr6_+_12290586 | 0.23 |
ENST00000379375.5 |
EDN1 |
endothelin 1 |
| chr5_+_36608422 | 0.23 |
ENST00000381918.3 |
SLC1A3 |
solute carrier family 1 (glial high affinity glutamate transporter), member 3 |
| chr18_-_52989217 | 0.22 |
ENST00000570287.2 |
TCF4 |
transcription factor 4 |
| chr5_+_32788945 | 0.21 |
ENST00000326958.1 |
AC026703.1 |
AC026703.1 |
| chr12_-_123201337 | 0.21 |
ENST00000528880.2 |
HCAR3 |
hydroxycarboxylic acid receptor 3 |
| chr12_+_4385230 | 0.20 |
ENST00000536537.1 |
CCND2 |
cyclin D2 |
| chr18_-_3219847 | 0.19 |
ENST00000261606.7 |
MYOM1 |
myomesin 1 |
| chr18_+_22040620 | 0.19 |
ENST00000426880.2 |
HRH4 |
histamine receptor H4 |
| chr17_-_39538550 | 0.18 |
ENST00000394001.1 |
KRT34 |
keratin 34 |
| chr17_-_39646116 | 0.18 |
ENST00000328119.6 |
KRT36 |
keratin 36 |
| chr4_-_74904398 | 0.17 |
ENST00000296026.4 |
CXCL3 |
chemokine (C-X-C motif) ligand 3 |
| chr13_-_103719196 | 0.17 |
ENST00000245312.3 |
SLC10A2 |
solute carrier family 10 (sodium/bile acid cotransporter), member 2 |
| chrX_+_117957741 | 0.17 |
ENST00000310164.2 |
ZCCHC12 |
zinc finger, CCHC domain containing 12 |
| chr8_+_77593474 | 0.17 |
ENST00000455469.2 ENST00000050961.6 |
ZFHX4 |
zinc finger homeobox 4 |
| chr1_+_197886461 | 0.17 |
ENST00000367388.3 ENST00000337020.2 ENST00000367387.4 |
LHX9 |
LIM homeobox 9 |
| chr6_+_15401075 | 0.16 |
ENST00000541660.1 |
JARID2 |
jumonji, AT rich interactive domain 2 |
| chr8_-_108510224 | 0.16 |
ENST00000517746.1 ENST00000297450.3 |
ANGPT1 |
angiopoietin 1 |
| chrX_-_114252193 | 0.16 |
ENST00000243213.1 |
IL13RA2 |
interleukin 13 receptor, alpha 2 |
| chr14_+_56078695 | 0.16 |
ENST00000416613.1 |
KTN1 |
kinectin 1 (kinesin receptor) |
| chr3_-_114035026 | 0.15 |
ENST00000570269.1 |
RP11-553L6.5 |
RP11-553L6.5 |
| chr2_+_114256661 | 0.15 |
ENST00000306507.5 |
FOXD4L1 |
forkhead box D4-like 1 |
| chr15_-_85197501 | 0.15 |
ENST00000434634.2 |
WDR73 |
WD repeat domain 73 |
| chr17_+_42264322 | 0.15 |
ENST00000446571.3 ENST00000357984.3 ENST00000538716.2 |
TMUB2 |
transmembrane and ubiquitin-like domain containing 2 |
| chr19_+_50191921 | 0.15 |
ENST00000420022.3 |
ADM5 |
adrenomedullin 5 (putative) |
| chr11_+_69924639 | 0.14 |
ENST00000538023.1 ENST00000398543.2 |
ANO1 |
anoctamin 1, calcium activated chloride channel |
| chr6_-_9933500 | 0.14 |
ENST00000492169.1 |
OFCC1 |
orofacial cleft 1 candidate 1 |
| chr7_+_116654935 | 0.14 |
ENST00000432298.1 ENST00000422922.1 |
ST7 |
suppression of tumorigenicity 7 |
| chr17_-_37308824 | 0.14 |
ENST00000415163.1 ENST00000441877.1 ENST00000444911.2 |
PLXDC1 |
plexin domain containing 1 |
| chr18_+_61575200 | 0.14 |
ENST00000238508.3 |
SERPINB10 |
serpin peptidase inhibitor, clade B (ovalbumin), member 10 |
| chr8_-_57123815 | 0.14 |
ENST00000316981.3 ENST00000423799.2 ENST00000429357.2 |
PLAG1 |
pleiomorphic adenoma gene 1 |
| chrX_-_3264682 | 0.14 |
ENST00000217939.6 |
MXRA5 |
matrix-remodelling associated 5 |
| chr4_-_74864386 | 0.14 |
ENST00000296027.4 |
CXCL5 |
chemokine (C-X-C motif) ligand 5 |
| chr11_+_69924397 | 0.14 |
ENST00000355303.5 |
ANO1 |
anoctamin 1, calcium activated chloride channel |
| chr2_-_151344172 | 0.14 |
ENST00000375734.2 ENST00000263895.4 ENST00000454202.1 |
RND3 |
Rho family GTPase 3 |
| chr22_-_41258074 | 0.14 |
ENST00000307221.4 |
DNAJB7 |
DnaJ (Hsp40) homolog, subfamily B, member 7 |
| chr1_+_23037323 | 0.13 |
ENST00000544305.1 ENST00000374630.3 ENST00000400191.3 ENST00000374632.3 |
EPHB2 |
EPH receptor B2 |
| chrM_+_5824 | 0.13 |
ENST00000361624.2 |
MT-CO1 |
mitochondrially encoded cytochrome c oxidase I |
| chr12_+_79258444 | 0.13 |
ENST00000261205.4 |
SYT1 |
synaptotagmin I |
| chr17_+_42264556 | 0.13 |
ENST00000319511.6 ENST00000589785.1 ENST00000592825.1 ENST00000589184.1 |
TMUB2 |
transmembrane and ubiquitin-like domain containing 2 |
| chr18_-_53177984 | 0.13 |
ENST00000543082.1 |
TCF4 |
transcription factor 4 |
| chr12_+_79258547 | 0.13 |
ENST00000457153.2 |
SYT1 |
synaptotagmin I |
| chr12_+_41221794 | 0.13 |
ENST00000547849.1 |
CNTN1 |
contactin 1 |
| chr6_-_82462425 | 0.13 |
ENST00000369754.3 ENST00000320172.6 ENST00000369756.3 |
FAM46A |
family with sequence similarity 46, member A |
| chrX_+_105937068 | 0.13 |
ENST00000324342.3 |
RNF128 |
ring finger protein 128, E3 ubiquitin protein ligase |
| chr3_-_21792838 | 0.12 |
ENST00000281523.2 |
ZNF385D |
zinc finger protein 385D |
| chr1_+_107683436 | 0.12 |
ENST00000370068.1 |
NTNG1 |
netrin G1 |
| chr12_+_113229737 | 0.12 |
ENST00000551052.1 ENST00000415485.3 |
RPH3A |
rabphilin 3A homolog (mouse) |
| chr2_+_32502952 | 0.12 |
ENST00000238831.4 |
YIPF4 |
Yip1 domain family, member 4 |
| chr8_+_104892639 | 0.12 |
ENST00000436393.2 |
RIMS2 |
regulating synaptic membrane exocytosis 2 |
| chr17_+_42264395 | 0.12 |
ENST00000587989.1 ENST00000590235.1 |
TMUB2 |
transmembrane and ubiquitin-like domain containing 2 |
| chr1_+_161676983 | 0.12 |
ENST00000367957.2 |
FCRLA |
Fc receptor-like A |
| chr2_-_51259292 | 0.12 |
ENST00000401669.2 |
NRXN1 |
neurexin 1 |
| chr2_-_51259229 | 0.12 |
ENST00000405472.3 |
NRXN1 |
neurexin 1 |
| chr4_+_174089904 | 0.12 |
ENST00000265000.4 |
GALNT7 |
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 7 (GalNAc-T7) |
| chr2_-_30144432 | 0.12 |
ENST00000389048.3 |
ALK |
anaplastic lymphoma receptor tyrosine kinase |
| chr1_+_241695424 | 0.12 |
ENST00000366558.3 ENST00000366559.4 |
KMO |
kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) |
| chr17_-_3337135 | 0.12 |
ENST00000248384.1 |
OR1E2 |
olfactory receptor, family 1, subfamily E, member 2 |
| chr14_+_23842018 | 0.12 |
ENST00000397242.2 ENST00000329715.2 |
IL25 |
interleukin 25 |
| chr7_+_31569365 | 0.11 |
ENST00000454513.1 ENST00000451887.2 |
CCDC129 |
coiled-coil domain containing 129 |
| chr18_-_35145728 | 0.11 |
ENST00000361795.5 ENST00000603232.1 |
CELF4 |
CUGBP, Elav-like family member 4 |
| chr1_+_161677034 | 0.11 |
ENST00000349527.4 ENST00000309691.6 ENST00000294796.4 ENST00000367953.3 ENST00000367950.1 |
FCRLA |
Fc receptor-like A |
| chrX_-_100662881 | 0.11 |
ENST00000218516.3 |
GLA |
galactosidase, alpha |
| chr1_+_107683644 | 0.11 |
ENST00000370067.1 |
NTNG1 |
netrin G1 |
| chrX_+_9217932 | 0.11 |
ENST00000432442.1 |
GS1-519E5.1 |
GS1-519E5.1 |
| chr4_+_74702214 | 0.11 |
ENST00000226317.5 ENST00000515050.1 |
CXCL6 |
chemokine (C-X-C motif) ligand 6 |
| chr13_-_20805109 | 0.11 |
ENST00000241124.6 |
GJB6 |
gap junction protein, beta 6, 30kDa |
| chr17_-_39471947 | 0.11 |
ENST00000334202.3 |
KRTAP17-1 |
keratin associated protein 17-1 |
| chr8_+_38831683 | 0.11 |
ENST00000302495.4 |
HTRA4 |
HtrA serine peptidase 4 |
| chr18_-_52989525 | 0.11 |
ENST00000457482.3 |
TCF4 |
transcription factor 4 |
| chr14_-_61748550 | 0.11 |
ENST00000555868.1 |
TMEM30B |
transmembrane protein 30B |
| chr1_+_241695670 | 0.11 |
ENST00000366557.4 |
KMO |
kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) |
| chr9_-_122131696 | 0.11 |
ENST00000373964.2 ENST00000265922.3 |
BRINP1 |
bone morphogenetic protein/retinoic acid inducible neural-specific 1 |
| chr17_+_7533439 | 0.10 |
ENST00000441599.2 ENST00000380450.4 ENST00000416273.3 ENST00000575903.1 ENST00000576830.1 ENST00000571153.1 ENST00000575618.1 ENST00000576152.1 |
SHBG |
sex hormone-binding globulin |
| chr3_+_108541545 | 0.10 |
ENST00000295756.6 |
TRAT1 |
T cell receptor associated transmembrane adaptor 1 |
| chr8_+_110099653 | 0.10 |
ENST00000311762.2 |
TRHR |
thyrotropin-releasing hormone receptor |
| chr4_-_107957454 | 0.10 |
ENST00000285311.3 |
DKK2 |
dickkopf WNT signaling pathway inhibitor 2 |
| chr1_+_209848749 | 0.10 |
ENST00000367029.4 |
G0S2 |
G0/G1switch 2 |
| chr18_-_3220106 | 0.10 |
ENST00000356443.4 ENST00000400569.3 |
MYOM1 |
myomesin 1 |
| chr7_+_129906660 | 0.10 |
ENST00000222481.4 |
CPA2 |
carboxypeptidase A2 (pancreatic) |
| chr11_-_102826434 | 0.10 |
ENST00000340273.4 ENST00000260302.3 |
MMP13 |
matrix metallopeptidase 13 (collagenase 3) |
| chr18_+_29027696 | 0.10 |
ENST00000257189.4 |
DSG3 |
desmoglein 3 |
| chr3_+_108541608 | 0.10 |
ENST00000426646.1 |
TRAT1 |
T cell receptor associated transmembrane adaptor 1 |
| chr3_+_187086120 | 0.10 |
ENST00000259030.2 |
RTP4 |
receptor (chemosensory) transporter protein 4 |
| chr10_+_114709999 | 0.10 |
ENST00000355995.4 ENST00000545257.1 ENST00000543371.1 ENST00000536810.1 ENST00000355717.4 ENST00000538897.1 ENST00000534894.1 |
TCF7L2 |
transcription factor 7-like 2 (T-cell specific, HMG-box) |
| chr6_+_10556215 | 0.10 |
ENST00000316170.3 |
GCNT2 |
glucosaminyl (N-acetyl) transferase 2, I-branching enzyme (I blood group) |
| chr8_+_71485681 | 0.10 |
ENST00000391684.1 |
AC120194.1 |
AC120194.1 |
| chrY_+_26997726 | 0.10 |
ENST00000382296.2 |
DAZ4 |
deleted in azoospermia 4 |
| chr9_+_2717502 | 0.10 |
ENST00000382082.3 |
KCNV2 |
potassium channel, subfamily V, member 2 |
| chr4_-_41750922 | 0.10 |
ENST00000226382.2 |
PHOX2B |
paired-like homeobox 2b |
| chr8_+_110098850 | 0.10 |
ENST00000518632.1 |
TRHR |
thyrotropin-releasing hormone receptor |
| chr18_-_25616519 | 0.10 |
ENST00000399380.3 |
CDH2 |
cadherin 2, type 1, N-cadherin (neuronal) |
| chr2_+_210444748 | 0.09 |
ENST00000392194.1 |
MAP2 |
microtubule-associated protein 2 |
| chr11_+_112832202 | 0.09 |
ENST00000534015.1 |
NCAM1 |
neural cell adhesion molecule 1 |
| chr3_+_101546827 | 0.09 |
ENST00000461724.1 ENST00000483180.1 ENST00000394054.2 |
NFKBIZ |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, zeta |
| chr11_+_61129456 | 0.09 |
ENST00000278826.6 |
TMEM138 |
transmembrane protein 138 |
| chr12_-_91451758 | 0.09 |
ENST00000266719.3 |
KERA |
keratocan |
| chr1_-_116311402 | 0.09 |
ENST00000261448.5 |
CASQ2 |
calsequestrin 2 (cardiac muscle) |
| chr14_+_56127989 | 0.09 |
ENST00000555573.1 |
KTN1 |
kinectin 1 (kinesin receptor) |
| chr14_-_35344093 | 0.09 |
ENST00000382422.2 |
BAZ1A |
bromodomain adjacent to zinc finger domain, 1A |
| chr12_-_96184533 | 0.09 |
ENST00000343702.4 ENST00000344911.4 |
NTN4 |
netrin 4 |
| chr4_-_65275162 | 0.09 |
ENST00000381210.3 ENST00000507440.1 |
TECRL |
trans-2,3-enoyl-CoA reductase-like |
| chrX_-_19689106 | 0.09 |
ENST00000379716.1 |
SH3KBP1 |
SH3-domain kinase binding protein 1 |
| chr15_-_55790515 | 0.09 |
ENST00000448430.2 ENST00000457155.2 |
DYX1C1 |
dyslexia susceptibility 1 candidate 1 |
| chr18_-_77276057 | 0.09 |
ENST00000597412.1 |
AC018445.1 |
Uncharacterized protein |
| chr2_-_190927447 | 0.09 |
ENST00000260950.4 |
MSTN |
myostatin |
| chr18_-_53070913 | 0.09 |
ENST00000568186.1 ENST00000564228.1 |
TCF4 |
transcription factor 4 |
| chr19_+_11350278 | 0.09 |
ENST00000252453.8 |
C19orf80 |
chromosome 19 open reading frame 80 |
| chr16_-_56459354 | 0.09 |
ENST00000290649.5 |
AMFR |
autocrine motility factor receptor, E3 ubiquitin protein ligase |
| chr5_-_146258205 | 0.09 |
ENST00000394413.3 |
PPP2R2B |
protein phosphatase 2, regulatory subunit B, beta |
| chr3_-_155394099 | 0.09 |
ENST00000414191.1 |
PLCH1 |
phospholipase C, eta 1 |
| chr19_+_1440838 | 0.09 |
ENST00000594262.1 |
AC027307.3 |
Uncharacterized protein |
| chr1_-_200379129 | 0.09 |
ENST00000367353.1 |
ZNF281 |
zinc finger protein 281 |
| chr1_-_168513229 | 0.09 |
ENST00000367819.2 |
XCL2 |
chemokine (C motif) ligand 2 |
| chr8_-_95220775 | 0.09 |
ENST00000441892.2 ENST00000521491.1 ENST00000027335.3 |
CDH17 |
cadherin 17, LI cadherin (liver-intestine) |
| chr12_-_52828147 | 0.09 |
ENST00000252245.5 |
KRT75 |
keratin 75 |
| chr2_+_11679963 | 0.09 |
ENST00000263834.5 |
GREB1 |
growth regulation by estrogen in breast cancer 1 |
| chr1_-_119530428 | 0.09 |
ENST00000369429.3 |
TBX15 |
T-box 15 |
| chr17_-_39197699 | 0.08 |
ENST00000306271.4 |
KRTAP1-1 |
keratin associated protein 1-1 |
| chr11_+_17281900 | 0.08 |
ENST00000530527.1 |
NUCB2 |
nucleobindin 2 |
| chr11_-_125773085 | 0.08 |
ENST00000227474.3 ENST00000534158.1 ENST00000529801.1 |
PUS3 |
pseudouridylate synthase 3 |
| chr8_-_101734308 | 0.08 |
ENST00000519004.1 ENST00000519363.1 ENST00000520142.1 |
PABPC1 |
poly(A) binding protein, cytoplasmic 1 |
| chr18_+_47088401 | 0.08 |
ENST00000261292.4 ENST00000427224.2 ENST00000580036.1 |
LIPG |
lipase, endothelial |
| chr16_-_51185172 | 0.08 |
ENST00000251020.4 |
SALL1 |
spalt-like transcription factor 1 |
| chr8_+_28748765 | 0.08 |
ENST00000355231.5 |
HMBOX1 |
homeobox containing 1 |
| chr8_-_7681411 | 0.08 |
ENST00000334773.6 |
DEFB105A |
defensin, beta 105A |
| chr6_+_6588902 | 0.08 |
ENST00000230568.4 |
LY86 |
lymphocyte antigen 86 |
| chr2_-_228244013 | 0.08 |
ENST00000304568.3 |
TM4SF20 |
transmembrane 4 L six family member 20 |
| chr14_+_73563735 | 0.08 |
ENST00000532192.1 |
RBM25 |
RNA binding motif protein 25 |
| chr20_-_46415341 | 0.08 |
ENST00000484875.1 ENST00000361612.4 |
SULF2 |
sulfatase 2 |
| chr8_-_120651020 | 0.08 |
ENST00000522826.1 ENST00000520066.1 ENST00000259486.6 ENST00000075322.6 |
ENPP2 |
ectonucleotide pyrophosphatase/phosphodiesterase 2 |
| chr7_+_143174966 | 0.08 |
ENST00000408916.1 |
TAS2R41 |
taste receptor, type 2, member 41 |
| chr8_+_7345191 | 0.08 |
ENST00000335510.6 |
DEFB105B |
defensin, beta 105B |
| chr10_+_47894023 | 0.08 |
ENST00000358474.5 |
FAM21B |
family with sequence similarity 21, member B |
| chr8_+_77593448 | 0.08 |
ENST00000521891.2 |
ZFHX4 |
zinc finger homeobox 4 |
| chr8_+_85095769 | 0.08 |
ENST00000518566.1 |
RALYL |
RALY RNA binding protein-like |
| chr17_-_1531635 | 0.08 |
ENST00000571650.1 |
SLC43A2 |
solute carrier family 43 (amino acid system L transporter), member 2 |
| chr6_+_26021869 | 0.08 |
ENST00000359907.3 |
HIST1H4A |
histone cluster 1, H4a |
| chr11_+_22696314 | 0.08 |
ENST00000532398.1 ENST00000433790.1 |
GAS2 |
growth arrest-specific 2 |
| chr6_+_108487245 | 0.08 |
ENST00000368986.4 |
NR2E1 |
nuclear receptor subfamily 2, group E, member 1 |
| chr6_+_73331918 | 0.08 |
ENST00000402622.2 ENST00000355635.3 ENST00000403813.2 ENST00000414165.2 |
KCNQ5 |
potassium voltage-gated channel, KQT-like subfamily, member 5 |
| chr20_-_46415297 | 0.08 |
ENST00000467815.1 ENST00000359930.4 |
SULF2 |
sulfatase 2 |
| chr4_-_111119804 | 0.08 |
ENST00000394607.3 ENST00000302274.3 |
ELOVL6 |
ELOVL fatty acid elongase 6 |
| chr1_+_197881592 | 0.08 |
ENST00000367391.1 ENST00000367390.3 |
LHX9 |
LIM homeobox 9 |
| chr8_-_25281747 | 0.08 |
ENST00000421054.2 |
GNRH1 |
gonadotropin-releasing hormone 1 (luteinizing-releasing hormone) |
| chr16_-_67217844 | 0.08 |
ENST00000563902.1 ENST00000561621.1 ENST00000290881.7 |
KIAA0895L |
KIAA0895-like |
| chr10_-_105845674 | 0.08 |
ENST00000353479.5 ENST00000369733.3 |
COL17A1 |
collagen, type XVII, alpha 1 |
| chr1_-_152297679 | 0.08 |
ENST00000368799.1 |
FLG |
filaggrin |
| chr1_-_152386732 | 0.08 |
ENST00000271835.3 |
CRNN |
cornulin |
| chr9_-_139137648 | 0.07 |
ENST00000358701.5 |
QSOX2 |
quiescin Q6 sulfhydryl oxidase 2 |
| chr9_-_118417 | 0.07 |
ENST00000382500.2 |
FOXD4 |
forkhead box D4 |
| chrX_-_138914394 | 0.07 |
ENST00000327569.3 ENST00000361648.2 ENST00000370543.1 ENST00000359686.2 |
ATP11C |
ATPase, class VI, type 11C |
| chr8_-_101734170 | 0.07 |
ENST00000522387.1 ENST00000518196.1 |
PABPC1 |
poly(A) binding protein, cytoplasmic 1 |
| chr8_+_85095497 | 0.07 |
ENST00000522455.1 ENST00000521695.1 |
RALYL |
RALY RNA binding protein-like |
| chr7_+_39017504 | 0.07 |
ENST00000403058.1 |
POU6F2 |
POU class 6 homeobox 2 |
| chr14_+_23727694 | 0.07 |
ENST00000399905.1 ENST00000470456.1 |
C14orf164 |
chromosome 14 open reading frame 164 |
| chr21_+_17792672 | 0.07 |
ENST00000602620.1 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr5_-_73937244 | 0.07 |
ENST00000302351.4 ENST00000510316.1 ENST00000508331.1 |
ENC1 |
ectodermal-neural cortex 1 (with BTB domain) |
| chr3_-_108248169 | 0.07 |
ENST00000273353.3 |
MYH15 |
myosin, heavy chain 15 |
| chr12_+_130646999 | 0.07 |
ENST00000539839.1 ENST00000229030.4 |
FZD10 |
frizzled family receptor 10 |
| chr4_+_71494461 | 0.07 |
ENST00000396073.3 |
ENAM |
enamelin |
| chr9_+_5450503 | 0.07 |
ENST00000381573.4 ENST00000381577.3 |
CD274 |
CD274 molecule |
| chr4_-_41884582 | 0.07 |
ENST00000499082.2 |
LINC00682 |
long intergenic non-protein coding RNA 682 |
| chr4_+_74347400 | 0.07 |
ENST00000226355.3 |
AFM |
afamin |
| chr7_-_27205136 | 0.07 |
ENST00000396345.1 ENST00000343483.6 |
HOXA9 |
homeobox A9 |
| chr1_+_161736072 | 0.07 |
ENST00000367942.3 |
ATF6 |
activating transcription factor 6 |
| chr2_+_207308220 | 0.07 |
ENST00000264377.3 |
ADAM23 |
ADAM metallopeptidase domain 23 |
| chr7_-_81399411 | 0.07 |
ENST00000423064.2 |
HGF |
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr2_+_28718921 | 0.07 |
ENST00000327757.5 ENST00000422425.2 ENST00000404858.1 |
PLB1 |
phospholipase B1 |
| chr14_-_24036943 | 0.07 |
ENST00000556843.1 ENST00000397120.3 ENST00000557189.1 |
AP1G2 |
adaptor-related protein complex 1, gamma 2 subunit |
| chr4_-_52904425 | 0.07 |
ENST00000535450.1 |
SGCB |
sarcoglycan, beta (43kDa dystrophin-associated glycoprotein) |
| chr17_-_56494882 | 0.07 |
ENST00000584437.1 |
RNF43 |
ring finger protein 43 |
| chr11_-_13517565 | 0.07 |
ENST00000282091.1 ENST00000529816.1 |
PTH |
parathyroid hormone |
| chr3_+_159706537 | 0.07 |
ENST00000305579.2 ENST00000480787.1 ENST00000466512.1 |
IL12A |
interleukin 12A (natural killer cell stimulatory factor 1, cytotoxic lymphocyte maturation factor 1, p35) |
| chr9_-_215744 | 0.07 |
ENST00000382387.2 |
C9orf66 |
chromosome 9 open reading frame 66 |
| chr11_-_57194948 | 0.07 |
ENST00000533235.1 ENST00000526621.1 ENST00000352187.1 |
SLC43A3 |
solute carrier family 43, member 3 |
| chrX_+_105445717 | 0.07 |
ENST00000372552.1 |
MUM1L1 |
melanoma associated antigen (mutated) 1-like 1 |
| chr1_-_200379104 | 0.07 |
ENST00000367352.3 |
ZNF281 |
zinc finger protein 281 |
| chr12_+_40787194 | 0.07 |
ENST00000425730.2 ENST00000454784.4 |
MUC19 |
mucin 19, oligomeric |
| chr17_-_56494713 | 0.07 |
ENST00000407977.2 |
RNF43 |
ring finger protein 43 |
| chr7_-_20826504 | 0.07 |
ENST00000418710.2 ENST00000361443.4 |
SP8 |
Sp8 transcription factor |
| chr17_-_56494908 | 0.07 |
ENST00000577716.1 |
RNF43 |
ring finger protein 43 |
| chr2_-_55277654 | 0.07 |
ENST00000337526.6 ENST00000317610.7 ENST00000357732.4 |
RTN4 |
reticulon 4 |
| chr5_+_112073544 | 0.07 |
ENST00000257430.4 ENST00000508376.2 |
APC |
adenomatous polyposis coli |
| chr20_-_10654639 | 0.07 |
ENST00000254958.5 |
JAG1 |
jagged 1 |
| chr18_-_53253323 | 0.07 |
ENST00000540999.1 ENST00000563888.2 |
TCF4 |
transcription factor 4 |
| chr12_+_14518598 | 0.07 |
ENST00000261168.4 ENST00000538511.1 ENST00000545723.1 ENST00000543189.1 ENST00000536444.1 |
ATF7IP |
activating transcription factor 7 interacting protein |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0048936 | neurofilament bundle assembly(GO:0033693) peripheral nervous system neuron axonogenesis(GO:0048936) |
| 0.1 | 0.4 | GO:0048880 | sensory system development(GO:0048880) |
| 0.1 | 0.3 | GO:0098746 | fast, calcium ion-dependent exocytosis of neurotransmitter(GO:0098746) |
| 0.1 | 0.3 | GO:0032911 | negative regulation of transforming growth factor beta1 production(GO:0032911) |
| 0.1 | 0.2 | GO:0060584 | regulation of prostaglandin-endoperoxide synthase activity(GO:0060584) positive regulation of prostaglandin-endoperoxide synthase activity(GO:0060585) |
| 0.1 | 0.2 | GO:0035262 | gonad morphogenesis(GO:0035262) |
| 0.1 | 0.4 | GO:0098814 | spontaneous neurotransmitter secretion(GO:0061669) spontaneous synaptic transmission(GO:0098814) |
| 0.0 | 0.4 | GO:0002074 | extraocular skeletal muscle development(GO:0002074) |
| 0.0 | 0.2 | GO:0038018 | Wnt receptor catabolic process(GO:0038018) |
| 0.0 | 0.3 | GO:0060842 | arterial endothelial cell differentiation(GO:0060842) |
| 0.0 | 0.0 | GO:0086073 | bundle of His cell-Purkinje myocyte adhesion involved in cell communication(GO:0086073) |
| 0.0 | 0.2 | GO:0006537 | glutamate biosynthetic process(GO:0006537) gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 0.0 | 0.2 | GO:0019805 | quinolinate biosynthetic process(GO:0019805) |
| 0.0 | 0.3 | GO:0015705 | iodide transport(GO:0015705) |
| 0.0 | 0.1 | GO:0061031 | endodermal digestive tract morphogenesis(GO:0061031) |
| 0.0 | 0.1 | GO:1902725 | negative regulation of satellite cell differentiation(GO:1902725) |
| 0.0 | 0.1 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.0 | 0.1 | GO:0060164 | regulation of timing of neuron differentiation(GO:0060164) |
| 0.0 | 0.1 | GO:0072092 | ureteric bud invasion(GO:0072092) |
| 0.0 | 0.7 | GO:0090023 | positive regulation of neutrophil chemotaxis(GO:0090023) |
| 0.0 | 0.1 | GO:0090290 | positive regulation of osteoclast proliferation(GO:0090290) negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 0.0 | 0.2 | GO:0001920 | negative regulation of receptor recycling(GO:0001920) |
| 0.0 | 0.2 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.0 | 0.1 | GO:0061073 | ciliary body morphogenesis(GO:0061073) |
| 0.0 | 0.2 | GO:0097118 | neuroligin clustering involved in postsynaptic membrane assembly(GO:0097118) |
| 0.0 | 0.1 | GO:0002314 | germinal center B cell differentiation(GO:0002314) |
| 0.0 | 0.1 | GO:0010982 | regulation of high-density lipoprotein particle clearance(GO:0010982) |
| 0.0 | 0.0 | GO:0021529 | spinal cord oligodendrocyte cell differentiation(GO:0021529) spinal cord oligodendrocyte cell fate specification(GO:0021530) |
| 0.0 | 0.1 | GO:0070781 | arginine biosynthetic process via ornithine(GO:0042450) response to biotin(GO:0070781) |
| 0.0 | 0.1 | GO:1990637 | response to prolactin(GO:1990637) |
| 0.0 | 0.0 | GO:0010621 | negative regulation of transcription by transcription factor localization(GO:0010621) |
| 0.0 | 0.1 | GO:0048867 | ganglion mother cell fate determination(GO:0007402) stem cell fate determination(GO:0048867) |
| 0.0 | 0.1 | GO:1903413 | cellular response to bile acid(GO:1903413) negative regulation of intestinal absorption(GO:1904479) response to iron ion starvation(GO:1990641) |
| 0.0 | 0.1 | GO:0035426 | extracellular matrix-cell signaling(GO:0035426) |
| 0.0 | 0.1 | GO:0033861 | negative regulation of NAD(P)H oxidase activity(GO:0033861) |
| 0.0 | 0.2 | GO:2000622 | regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
| 0.0 | 0.2 | GO:0033007 | negative regulation of mast cell activation involved in immune response(GO:0033007) negative regulation of mast cell degranulation(GO:0043305) |
| 0.0 | 0.1 | GO:2000502 | negative regulation of natural killer cell chemotaxis(GO:2000502) |
| 0.0 | 0.1 | GO:0007113 | endomitotic cell cycle(GO:0007113) |
| 0.0 | 0.1 | GO:1904783 | positive regulation of NMDA glutamate receptor activity(GO:1904783) |
| 0.0 | 0.1 | GO:1904885 | beta-catenin destruction complex assembly(GO:1904885) |
| 0.0 | 0.1 | GO:0009624 | response to nematode(GO:0009624) eosinophil differentiation(GO:0030222) |
| 0.0 | 0.1 | GO:0045653 | negative regulation of megakaryocyte differentiation(GO:0045653) |
| 0.0 | 0.0 | GO:0030185 | nitric oxide transport(GO:0030185) |
| 0.0 | 0.0 | GO:1901207 | regulation of heart looping(GO:1901207) |
| 0.0 | 0.1 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.0 | 0.1 | GO:1902866 | regulation of retina development in camera-type eye(GO:1902866) |
| 0.0 | 0.1 | GO:1990539 | fructose transport(GO:0015755) fructose import(GO:0032445) carbohydrate import into cell(GO:0097319) carbohydrate import across plasma membrane(GO:0098704) fructose import across plasma membrane(GO:1990539) |
| 0.0 | 0.1 | GO:0006990 | positive regulation of transcription from RNA polymerase II promoter involved in unfolded protein response(GO:0006990) |
| 0.0 | 0.1 | GO:1990481 | mRNA pseudouridine synthesis(GO:1990481) |
| 0.0 | 0.2 | GO:0042761 | very long-chain fatty acid biosynthetic process(GO:0042761) |
| 0.0 | 0.0 | GO:0036091 | positive regulation of transcription from RNA polymerase II promoter in response to oxidative stress(GO:0036091) |
| 0.0 | 0.0 | GO:0035526 | retrograde transport, plasma membrane to Golgi(GO:0035526) |
| 0.0 | 0.1 | GO:0002436 | immune complex clearance(GO:0002434) immune complex clearance by monocytes and macrophages(GO:0002436) response to high density lipoprotein particle(GO:0055099) regulation of immune complex clearance by monocytes and macrophages(GO:0090264) |
| 0.0 | 0.1 | GO:0003335 | corneocyte development(GO:0003335) |
| 0.0 | 0.1 | GO:0007039 | protein catabolic process in the vacuole(GO:0007039) |
| 0.0 | 0.1 | GO:1904352 | positive regulation of protein catabolic process in the vacuole(GO:1904352) |
| 0.0 | 0.1 | GO:0036438 | maintenance of lens transparency(GO:0036438) |
| 0.0 | 0.1 | GO:0043950 | positive regulation of cAMP-mediated signaling(GO:0043950) |
| 0.0 | 0.0 | GO:0080154 | regulation of fertilization(GO:0080154) |
| 0.0 | 0.1 | GO:0042488 | positive regulation of odontogenesis of dentin-containing tooth(GO:0042488) |
| 0.0 | 0.1 | GO:2001181 | positive regulation of interleukin-10 secretion(GO:2001181) |
| 0.0 | 0.1 | GO:0046477 | glycosylceramide catabolic process(GO:0046477) |
| 0.0 | 0.0 | GO:1905205 | positive regulation of connective tissue replacement(GO:1905205) |
| 0.0 | 0.1 | GO:0002329 | pre-B cell differentiation(GO:0002329) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0005863 | striated muscle myosin thick filament(GO:0005863) |
| 0.1 | 0.2 | GO:0048237 | rough endoplasmic reticulum lumen(GO:0048237) |
| 0.1 | 0.3 | GO:0060200 | clathrin-sculpted acetylcholine transport vesicle(GO:0060200) clathrin-sculpted acetylcholine transport vesicle membrane(GO:0060201) |
| 0.0 | 0.2 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 0.0 | 0.4 | GO:0005883 | neurofilament(GO:0005883) |
| 0.0 | 0.1 | GO:0097442 | CA3 pyramidal cell dendrite(GO:0097442) |
| 0.0 | 0.1 | GO:0090571 | RNA polymerase II transcription repressor complex(GO:0090571) |
| 0.0 | 0.1 | GO:0033150 | cytoskeletal calyx(GO:0033150) |
| 0.0 | 0.2 | GO:0016342 | catenin complex(GO:0016342) |
| 0.0 | 0.1 | GO:0009330 | DNA topoisomerase complex (ATP-hydrolyzing)(GO:0009330) |
| 0.0 | 0.1 | GO:0005587 | collagen type IV trimer(GO:0005587) |
| 0.0 | 0.1 | GO:0030121 | AP-1 adaptor complex(GO:0030121) |
| 0.0 | 0.1 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.0 | 0.1 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 0.0 | 0.1 | GO:0045275 | mitochondrial respiratory chain complex III(GO:0005750) respiratory chain complex III(GO:0045275) |
| 0.0 | 0.0 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0050429 | calcium-dependent phospholipase C activity(GO:0050429) |
| 0.1 | 0.3 | GO:0015111 | iodide transmembrane transporter activity(GO:0015111) |
| 0.1 | 0.7 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.0 | 0.2 | GO:0031708 | endothelin B receptor binding(GO:0031708) |
| 0.0 | 0.7 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.0 | 0.3 | GO:0030348 | syntaxin-3 binding(GO:0030348) |
| 0.0 | 0.2 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.0 | 0.2 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.0 | 0.2 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.0 | 0.6 | GO:0016641 | oxidoreductase activity, acting on the CH-NH2 group of donors, oxygen as acceptor(GO:0016641) |
| 0.0 | 0.1 | GO:1904288 | BAT3 complex binding(GO:1904288) |
| 0.0 | 0.1 | GO:0016936 | galactoside binding(GO:0016936) |
| 0.0 | 0.1 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.0 | 0.1 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.0 | 0.1 | GO:0042835 | BRE binding(GO:0042835) |
| 0.0 | 0.1 | GO:0035673 | oligopeptide transmembrane transporter activity(GO:0035673) |
| 0.0 | 0.3 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.0 | 0.1 | GO:0047391 | alkylglycerophosphoethanolamine phosphodiesterase activity(GO:0047391) |
| 0.0 | 0.3 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 0.1 | GO:0008431 | vitamin E binding(GO:0008431) |
| 0.0 | 0.1 | GO:0061505 | DNA topoisomerase type II (ATP-hydrolyzing) activity(GO:0003918) DNA topoisomerase II activity(GO:0061505) |
| 0.0 | 0.2 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.0 | 0.1 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.0 | 0.1 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.0 | 0.2 | GO:0016174 | NAD(P)H oxidase activity(GO:0016174) |
| 0.0 | 0.1 | GO:0005353 | fructose transmembrane transporter activity(GO:0005353) |
| 0.0 | 0.1 | GO:0050253 | retinyl-palmitate esterase activity(GO:0050253) |
| 0.0 | 0.1 | GO:0008109 | N-acetyllactosaminide beta-1,6-N-acetylglucosaminyltransferase activity(GO:0008109) |
| 0.0 | 0.0 | GO:0004157 | dihydropyrimidinase activity(GO:0004157) |
| 0.0 | 0.1 | GO:0005497 | androgen binding(GO:0005497) |
| 0.0 | 0.5 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.0 | 0.3 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.0 | 0.0 | GO:0016503 | pheromone receptor activity(GO:0016503) |
| 0.0 | 0.2 | GO:0042301 | phosphate ion binding(GO:0042301) |
| 0.0 | 0.1 | GO:0004704 | NF-kappaB-inducing kinase activity(GO:0004704) |
| 0.0 | 0.0 | GO:0080101 | phosphatidyl-N-methylethanolamine N-methyltransferase activity(GO:0000773) phosphatidylethanolamine N-methyltransferase activity(GO:0004608) phosphatidyl-N-dimethylethanolamine N-methyltransferase activity(GO:0080101) |
| 0.0 | 0.3 | GO:0097493 | structural molecule activity conferring elasticity(GO:0097493) |
| 0.0 | 0.1 | GO:0004465 | lipoprotein lipase activity(GO:0004465) |
| 0.0 | 0.0 | GO:0070698 | type I activin receptor binding(GO:0070698) |
| 0.0 | 0.1 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.0 | 0.1 | GO:0043237 | laminin-1 binding(GO:0043237) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
| 0.0 | 0.0 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | REACTOME DOPAMINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Dopamine Neurotransmitter Release Cycle |
| 0.0 | 0.3 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 0.9 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.7 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 0.2 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.0 | 0.2 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |