Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
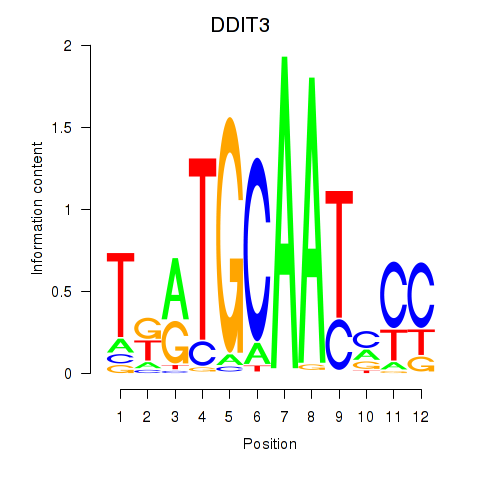
Results for DDIT3
Z-value: 0.32
Transcription factors associated with DDIT3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
DDIT3
|
ENSG00000175197.6 | DDIT3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| DDIT3 | hg19_v2_chr12_-_57914275_57914304 | 0.08 | 8.6e-01 | Click! |
Activity profile of DDIT3 motif
Sorted Z-values of DDIT3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of DDIT3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr14_-_92413353 | 0.34 |
ENST00000556154.1 |
FBLN5 |
fibulin 5 |
| chr3_+_157154578 | 0.31 |
ENST00000295927.3 |
PTX3 |
pentraxin 3, long |
| chr14_-_92413727 | 0.28 |
ENST00000267620.10 |
FBLN5 |
fibulin 5 |
| chr7_-_56101826 | 0.23 |
ENST00000421626.1 |
PSPH |
phosphoserine phosphatase |
| chr3_+_12392971 | 0.18 |
ENST00000287820.6 |
PPARG |
peroxisome proliferator-activated receptor gamma |
| chr9_+_80912059 | 0.18 |
ENST00000347159.2 ENST00000376588.3 |
PSAT1 |
phosphoserine aminotransferase 1 |
| chr8_-_93029865 | 0.16 |
ENST00000422361.2 |
RUNX1T1 |
runt-related transcription factor 1; translocated to, 1 (cyclin D-related) |
| chr20_+_54987305 | 0.16 |
ENST00000371336.3 ENST00000434344.1 |
CASS4 |
Cas scaffolding protein family member 4 |
| chr8_-_93029520 | 0.14 |
ENST00000521553.1 |
RUNX1T1 |
runt-related transcription factor 1; translocated to, 1 (cyclin D-related) |
| chr20_+_54987168 | 0.14 |
ENST00000360314.3 |
CASS4 |
Cas scaffolding protein family member 4 |
| chr15_+_41245160 | 0.13 |
ENST00000444189.2 ENST00000446533.3 |
CHAC1 |
ChaC, cation transport regulator homolog 1 (E. coli) |
| chr4_-_119274121 | 0.13 |
ENST00000296498.3 |
PRSS12 |
protease, serine, 12 (neurotrypsin, motopsin) |
| chr12_-_25101920 | 0.12 |
ENST00000539780.1 ENST00000546285.1 ENST00000342945.5 |
BCAT1 |
branched chain amino-acid transaminase 1, cytosolic |
| chr11_+_20044096 | 0.11 |
ENST00000533917.1 |
NAV2 |
neuron navigator 2 |
| chr12_-_25102252 | 0.10 |
ENST00000261192.7 |
BCAT1 |
branched chain amino-acid transaminase 1, cytosolic |
| chr8_+_27629459 | 0.10 |
ENST00000523566.1 |
ESCO2 |
establishment of sister chromatid cohesion N-acetyltransferase 2 |
| chr7_-_130598059 | 0.08 |
ENST00000432045.2 |
MIR29B1 |
microRNA 29a |
| chr4_-_99578789 | 0.08 |
ENST00000511651.1 ENST00000505184.1 |
TSPAN5 |
tetraspanin 5 |
| chr6_+_31895254 | 0.08 |
ENST00000299367.5 ENST00000442278.2 |
C2 |
complement component 2 |
| chr19_-_6604094 | 0.08 |
ENST00000597430.2 |
CD70 |
CD70 molecule |
| chr1_-_20250110 | 0.07 |
ENST00000375116.3 |
PLA2G2E |
phospholipase A2, group IIE |
| chr2_-_63815628 | 0.07 |
ENST00000409562.3 |
WDPCP |
WD repeat containing planar cell polarity effector |
| chr3_+_44840679 | 0.07 |
ENST00000425755.1 |
KIF15 |
kinesin family member 15 |
| chr2_+_227700652 | 0.07 |
ENST00000341329.3 ENST00000392062.2 ENST00000437454.1 ENST00000443477.1 ENST00000423616.1 ENST00000448992.1 |
RHBDD1 |
rhomboid domain containing 1 |
| chr19_+_12917364 | 0.06 |
ENST00000221486.4 |
RNASEH2A |
ribonuclease H2, subunit A |
| chr8_-_18744528 | 0.06 |
ENST00000523619.1 |
PSD3 |
pleckstrin and Sec7 domain containing 3 |
| chr5_-_178017355 | 0.06 |
ENST00000390654.3 |
COL23A1 |
collagen, type XXIII, alpha 1 |
| chr12_-_12491608 | 0.06 |
ENST00000545735.1 |
MANSC1 |
MANSC domain containing 1 |
| chr5_+_72143988 | 0.06 |
ENST00000506351.2 |
TNPO1 |
transportin 1 |
| chr11_-_33913708 | 0.06 |
ENST00000257818.2 |
LMO2 |
LIM domain only 2 (rhombotin-like 1) |
| chr8_+_24151620 | 0.06 |
ENST00000437154.2 |
ADAM28 |
ADAM metallopeptidase domain 28 |
| chr1_-_112032284 | 0.05 |
ENST00000414219.1 |
ADORA3 |
adenosine A3 receptor |
| chr15_-_83837983 | 0.05 |
ENST00000562702.1 |
HDGFRP3 |
Hepatoma-derived growth factor-related protein 3 |
| chr20_-_31592113 | 0.05 |
ENST00000420875.1 ENST00000375519.2 ENST00000375523.3 ENST00000356173.3 |
SUN5 |
Sad1 and UNC84 domain containing 5 |
| chr4_+_157997273 | 0.05 |
ENST00000541722.1 ENST00000512619.1 |
GLRB |
glycine receptor, beta |
| chr4_+_157997209 | 0.05 |
ENST00000264428.4 |
GLRB |
glycine receptor, beta |
| chr2_-_43453734 | 0.05 |
ENST00000282388.3 |
ZFP36L2 |
ZFP36 ring finger protein-like 2 |
| chr17_-_77813186 | 0.05 |
ENST00000448310.1 ENST00000269397.4 |
CBX4 |
chromobox homolog 4 |
| chr9_+_112542572 | 0.05 |
ENST00000374530.3 |
PALM2-AKAP2 |
PALM2-AKAP2 readthrough |
| chr1_-_114447683 | 0.05 |
ENST00000256658.4 ENST00000369564.1 |
AP4B1 |
adaptor-related protein complex 4, beta 1 subunit |
| chr2_+_169757750 | 0.05 |
ENST00000375363.3 ENST00000429379.2 ENST00000421979.1 |
G6PC2 |
glucose-6-phosphatase, catalytic, 2 |
| chr2_+_113825547 | 0.04 |
ENST00000341010.2 ENST00000337569.3 |
IL1F10 |
interleukin 1 family, member 10 (theta) |
| chr1_+_114447763 | 0.04 |
ENST00000369563.3 |
DCLRE1B |
DNA cross-link repair 1B |
| chr13_-_24007815 | 0.04 |
ENST00000382298.3 |
SACS |
spastic ataxia of Charlevoix-Saguenay (sacsin) |
| chr11_+_47279504 | 0.04 |
ENST00000441012.2 ENST00000437276.1 ENST00000436029.1 ENST00000467728.1 ENST00000405853.3 |
NR1H3 |
nuclear receptor subfamily 1, group H, member 3 |
| chr4_+_6784401 | 0.04 |
ENST00000425103.1 ENST00000307659.5 |
KIAA0232 |
KIAA0232 |
| chr1_+_154229547 | 0.04 |
ENST00000428595.1 |
UBAP2L |
ubiquitin associated protein 2-like |
| chr2_-_40680578 | 0.04 |
ENST00000455476.1 |
SLC8A1 |
solute carrier family 8 (sodium/calcium exchanger), member 1 |
| chr1_+_174844645 | 0.04 |
ENST00000486220.1 |
RABGAP1L |
RAB GTPase activating protein 1-like |
| chr1_-_114447412 | 0.04 |
ENST00000369567.1 ENST00000369566.3 |
AP4B1 |
adaptor-related protein complex 4, beta 1 subunit |
| chr1_+_40840320 | 0.04 |
ENST00000372708.1 |
SMAP2 |
small ArfGAP2 |
| chr8_+_24151553 | 0.04 |
ENST00000265769.4 ENST00000540823.1 ENST00000397649.3 |
ADAM28 |
ADAM metallopeptidase domain 28 |
| chr22_-_27456361 | 0.04 |
ENST00000453934.1 |
CTA-992D9.6 |
CTA-992D9.6 |
| chr15_+_59730348 | 0.04 |
ENST00000288228.5 ENST00000559628.1 ENST00000557914.1 ENST00000560474.1 |
FAM81A |
family with sequence similarity 81, member A |
| chr1_+_33546714 | 0.04 |
ENST00000294517.6 ENST00000358680.3 ENST00000373443.3 ENST00000398167.1 |
ADC |
arginine decarboxylase |
| chr11_-_10315741 | 0.03 |
ENST00000256190.8 |
SBF2 |
SET binding factor 2 |
| chr5_+_112074029 | 0.03 |
ENST00000512211.2 |
APC |
adenomatous polyposis coli |
| chr10_-_17659234 | 0.03 |
ENST00000466335.1 |
PTPLA |
protein tyrosine phosphatase-like (proline instead of catalytic arginine), member A |
| chr9_+_110045537 | 0.03 |
ENST00000358015.3 |
RAD23B |
RAD23 homolog B (S. cerevisiae) |
| chr12_+_57857475 | 0.03 |
ENST00000528467.1 |
GLI1 |
GLI family zinc finger 1 |
| chr8_-_11058847 | 0.03 |
ENST00000297303.4 ENST00000416569.2 |
XKR6 |
XK, Kell blood group complex subunit-related family, member 6 |
| chr12_-_63328817 | 0.03 |
ENST00000228705.6 |
PPM1H |
protein phosphatase, Mg2+/Mn2+ dependent, 1H |
| chr15_+_85923856 | 0.02 |
ENST00000560302.1 ENST00000394518.2 ENST00000361243.2 ENST00000560256.1 |
AKAP13 |
A kinase (PRKA) anchor protein 13 |
| chrX_+_9431324 | 0.02 |
ENST00000407597.2 ENST00000424279.1 ENST00000536365.1 ENST00000441088.1 ENST00000380961.1 ENST00000415293.1 |
TBL1X |
transducin (beta)-like 1X-linked |
| chr16_-_70323422 | 0.02 |
ENST00000261772.8 |
AARS |
alanyl-tRNA synthetase |
| chr10_-_17659357 | 0.02 |
ENST00000326961.6 ENST00000361271.3 |
PTPLA |
protein tyrosine phosphatase-like (proline instead of catalytic arginine), member A |
| chr2_-_63815860 | 0.02 |
ENST00000272321.7 ENST00000431065.1 |
WDPCP |
WD repeat containing planar cell polarity effector |
| chr2_-_161350305 | 0.02 |
ENST00000348849.3 |
RBMS1 |
RNA binding motif, single stranded interacting protein 1 |
| chr14_+_22265444 | 0.02 |
ENST00000390430.2 |
TRAV8-1 |
T cell receptor alpha variable 8-1 |
| chr17_+_30594823 | 0.02 |
ENST00000536287.1 |
RHBDL3 |
rhomboid, veinlet-like 3 (Drosophila) |
| chr9_-_95055956 | 0.02 |
ENST00000375629.3 ENST00000447699.2 ENST00000375643.3 ENST00000395554.3 |
IARS |
isoleucyl-tRNA synthetase |
| chr10_-_92681033 | 0.02 |
ENST00000371697.3 |
ANKRD1 |
ankyrin repeat domain 1 (cardiac muscle) |
| chr15_-_64126084 | 0.02 |
ENST00000560316.1 ENST00000443617.2 ENST00000560462.1 ENST00000558532.1 ENST00000561400.1 |
HERC1 |
HECT and RLD domain containing E3 ubiquitin protein ligase family member 1 |
| chr22_+_17956618 | 0.02 |
ENST00000262608.8 |
CECR2 |
cat eye syndrome chromosome region, candidate 2 |
| chr14_+_32798462 | 0.02 |
ENST00000280979.4 |
AKAP6 |
A kinase (PRKA) anchor protein 6 |
| chr19_+_10736183 | 0.02 |
ENST00000590857.1 ENST00000588688.1 ENST00000586078.1 |
SLC44A2 |
solute carrier family 44 (choline transporter), member 2 |
| chr11_+_34073269 | 0.02 |
ENST00000389645.3 |
CAPRIN1 |
cell cycle associated protein 1 |
| chr10_+_12391481 | 0.02 |
ENST00000378847.3 |
CAMK1D |
calcium/calmodulin-dependent protein kinase ID |
| chr9_-_95056010 | 0.02 |
ENST00000443024.2 |
IARS |
isoleucyl-tRNA synthetase |
| chr11_+_62495997 | 0.02 |
ENST00000316461.4 |
TTC9C |
tetratricopeptide repeat domain 9C |
| chrX_-_138724994 | 0.02 |
ENST00000536274.1 |
MCF2 |
MCF.2 cell line derived transforming sequence |
| chr5_+_118812237 | 0.02 |
ENST00000513628.1 |
HSD17B4 |
hydroxysteroid (17-beta) dehydrogenase 4 |
| chr14_+_32798547 | 0.02 |
ENST00000557354.1 ENST00000557102.1 ENST00000557272.1 |
AKAP6 |
A kinase (PRKA) anchor protein 6 |
| chr9_+_112542591 | 0.01 |
ENST00000483909.1 ENST00000314527.4 ENST00000413420.1 ENST00000302798.7 ENST00000555236.1 ENST00000510514.5 |
PALM2 PALM2-AKAP2 AKAP2 |
paralemmin 2 PALM2-AKAP2 readthrough A kinase (PRKA) anchor protein 2 |
| chr19_+_33865218 | 0.01 |
ENST00000585933.2 |
CEBPG |
CCAAT/enhancer binding protein (C/EBP), gamma |
| chr19_+_41497178 | 0.01 |
ENST00000324071.4 |
CYP2B6 |
cytochrome P450, family 2, subfamily B, polypeptide 6 |
| chr19_+_49258775 | 0.01 |
ENST00000593756.1 |
FGF21 |
fibroblast growth factor 21 |
| chr5_+_118812294 | 0.01 |
ENST00000509514.1 |
HSD17B4 |
hydroxysteroid (17-beta) dehydrogenase 4 |
| chr7_+_99006232 | 0.01 |
ENST00000403633.2 |
BUD31 |
BUD31 homolog (S. cerevisiae) |
| chr14_+_21387491 | 0.01 |
ENST00000258817.2 |
RP11-84C10.2 |
RP11-84C10.2 |
| chr10_+_102756800 | 0.01 |
ENST00000370223.3 |
LZTS2 |
leucine zipper, putative tumor suppressor 2 |
| chr1_-_161277210 | 0.01 |
ENST00000491222.2 |
MPZ |
myelin protein zero |
| chr3_-_148939835 | 0.01 |
ENST00000264613.6 |
CP |
ceruloplasmin (ferroxidase) |
| chr10_+_60028818 | 0.01 |
ENST00000333926.5 |
CISD1 |
CDGSH iron sulfur domain 1 |
| chr6_+_151187074 | 0.01 |
ENST00000367308.4 |
MTHFD1L |
methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1-like |
| chr20_+_24929866 | 0.01 |
ENST00000480798.1 ENST00000376835.2 |
CST7 |
cystatin F (leukocystatin) |
| chrX_-_100129128 | 0.01 |
ENST00000372960.4 ENST00000372964.1 ENST00000217885.5 |
NOX1 |
NADPH oxidase 1 |
| chr2_-_216300784 | 0.01 |
ENST00000421182.1 ENST00000432072.2 ENST00000323926.6 ENST00000336916.4 ENST00000357867.4 ENST00000359671.1 ENST00000346544.3 ENST00000345488.5 ENST00000357009.2 ENST00000446046.1 ENST00000356005.4 ENST00000443816.1 ENST00000426059.1 ENST00000354785.4 |
FN1 |
fibronectin 1 |
| chr16_-_18908196 | 0.01 |
ENST00000565324.1 ENST00000561947.1 |
SMG1 |
SMG1 phosphatidylinositol 3-kinase-related kinase |
| chr22_-_30901637 | 0.01 |
ENST00000381982.3 ENST00000255858.7 ENST00000540456.1 ENST00000392772.2 |
SEC14L4 |
SEC14-like 4 (S. cerevisiae) |
| chr8_+_118147498 | 0.01 |
ENST00000519688.1 ENST00000456015.2 |
SLC30A8 |
solute carrier family 30 (zinc transporter), member 8 |
| chr7_-_138363824 | 0.01 |
ENST00000419765.3 |
SVOPL |
SVOP-like |
| chr2_-_179672142 | 0.01 |
ENST00000342992.6 ENST00000360870.5 ENST00000460472.2 ENST00000589042.1 ENST00000591111.1 ENST00000342175.6 ENST00000359218.5 |
TTN |
titin |
| chr4_+_111397216 | 0.00 |
ENST00000265162.5 |
ENPEP |
glutamyl aminopeptidase (aminopeptidase A) |
| chr17_-_34257771 | 0.00 |
ENST00000394529.3 ENST00000293273.6 |
RDM1 |
RAD52 motif 1 |
| chrX_-_138724677 | 0.00 |
ENST00000370573.4 ENST00000338585.6 ENST00000370576.4 |
MCF2 |
MCF.2 cell line derived transforming sequence |
| chrX_-_40506766 | 0.00 |
ENST00000378421.1 ENST00000440784.2 ENST00000327877.5 ENST00000378426.1 ENST00000378418.2 |
CXorf38 |
chromosome X open reading frame 38 |
| chr11_-_8680383 | 0.00 |
ENST00000299550.6 |
TRIM66 |
tripartite motif containing 66 |
| chr5_-_137878887 | 0.00 |
ENST00000507939.1 ENST00000572514.1 ENST00000499810.2 ENST00000360541.5 |
ETF1 |
eukaryotic translation termination factor 1 |
| chr11_+_34073195 | 0.00 |
ENST00000341394.4 |
CAPRIN1 |
cell cycle associated protein 1 |
| chr1_-_44820880 | 0.00 |
ENST00000372257.2 ENST00000457571.1 ENST00000452396.1 |
ERI3 |
ERI1 exoribonuclease family member 3 |
| chr5_+_145583107 | 0.00 |
ENST00000506502.1 |
RBM27 |
RNA binding motif protein 27 |
| chr2_+_63816087 | 0.00 |
ENST00000409908.1 ENST00000442225.1 ENST00000409476.1 ENST00000436321.1 |
MDH1 |
malate dehydrogenase 1, NAD (soluble) |
| chr17_-_34257731 | 0.00 |
ENST00000431884.2 ENST00000425909.3 ENST00000394528.3 ENST00000430160.2 |
RDM1 |
RAD52 motif 1 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0052199 | negative regulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052199) |
| 0.0 | 0.6 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.0 | 0.2 | GO:2000230 | negative regulation of pancreatic stellate cell proliferation(GO:2000230) |
| 0.0 | 0.2 | GO:0009082 | branched-chain amino acid biosynthetic process(GO:0009082) leucine biosynthetic process(GO:0009098) valine biosynthetic process(GO:0009099) |
| 0.0 | 0.4 | GO:0006564 | L-serine biosynthetic process(GO:0006564) |
| 0.0 | 0.1 | GO:0043137 | DNA replication, removal of RNA primer(GO:0043137) |
| 0.0 | 0.1 | GO:0097112 | gamma-aminobutyric acid receptor clustering(GO:0097112) |
| 0.0 | 0.1 | GO:0034421 | post-translational protein acetylation(GO:0034421) |
| 0.0 | 0.1 | GO:0090521 | regulation of embryonic cell shape(GO:0016476) septin cytoskeleton organization(GO:0032185) glomerular visceral epithelial cell migration(GO:0090521) |
| 0.0 | 0.0 | GO:0090341 | negative regulation of secretion of lysosomal enzymes(GO:0090341) |
| 0.0 | 0.1 | GO:0021564 | vagus nerve development(GO:0021564) |
| 0.0 | 0.0 | GO:0006428 | isoleucyl-tRNA aminoacylation(GO:0006428) |
| 0.0 | 0.0 | GO:0031860 | telomeric 3' overhang formation(GO:0031860) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0071953 | elastic fiber(GO:0071953) |
| 0.0 | 0.1 | GO:0016935 | glycine-gated chloride channel complex(GO:0016935) |
| 0.0 | 0.1 | GO:0005873 | plus-end kinesin complex(GO:0005873) |
| 0.0 | 0.1 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0003839 | gamma-glutamylcyclotransferase activity(GO:0003839) |
| 0.0 | 0.2 | GO:0004647 | phosphoserine phosphatase activity(GO:0004647) |
| 0.0 | 0.2 | GO:0004084 | branched-chain-amino-acid transaminase activity(GO:0004084) L-leucine transaminase activity(GO:0052654) L-valine transaminase activity(GO:0052655) L-isoleucine transaminase activity(GO:0052656) |
| 0.0 | 0.3 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.0 | 0.1 | GO:0016933 | extracellular-glycine-gated ion channel activity(GO:0016933) extracellular-glycine-gated chloride channel activity(GO:0016934) |
| 0.0 | 0.2 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.0 | 0.1 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.0 | 0.0 | GO:0004822 | isoleucine-tRNA ligase activity(GO:0004822) |
| 0.0 | 0.1 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.0 | 0.0 | GO:0004346 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |