Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
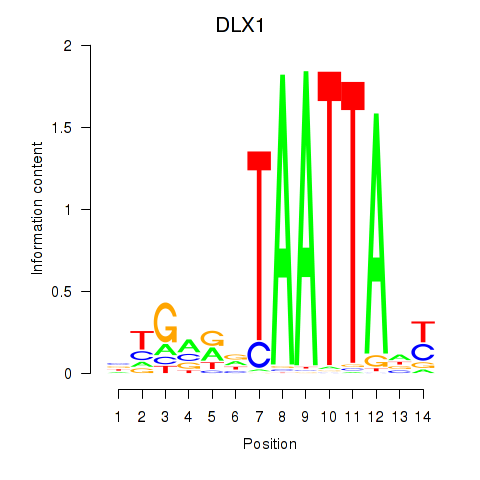
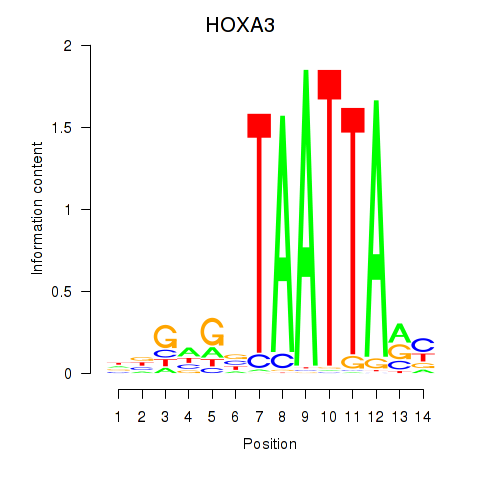
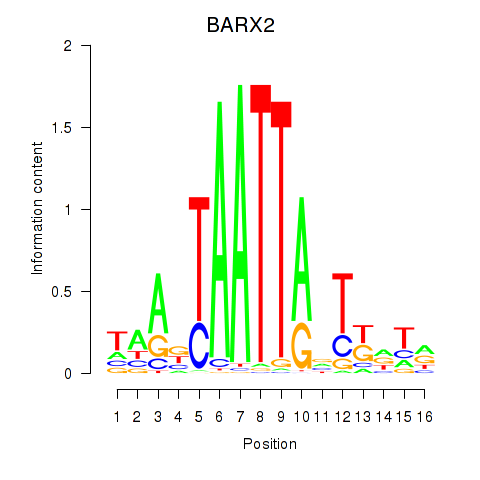
Results for DLX1_HOXA3_BARX2
Z-value: 0.33
Transcription factors associated with DLX1_HOXA3_BARX2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
DLX1
|
ENSG00000144355.10 | DLX1 |
|
HOXA3
|
ENSG00000105997.18 | HOXA3 |
|
BARX2
|
ENSG00000043039.5 | BARX2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| BARX2 | hg19_v2_chr11_+_129245796_129245835 | 0.81 | 1.4e-02 | Click! |
| DLX1 | hg19_v2_chr2_+_172949468_172949522 | 0.36 | 3.8e-01 | Click! |
Activity profile of DLX1_HOXA3_BARX2 motif
Sorted Z-values of DLX1_HOXA3_BARX2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of DLX1_HOXA3_BARX2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_-_36001113 | 0.64 |
ENST00000434389.1 |
DMKN |
dermokine |
| chr8_-_41166953 | 0.52 |
ENST00000220772.3 |
SFRP1 |
secreted frizzled-related protein 1 |
| chr10_-_135379132 | 0.48 |
ENST00000343131.5 |
SYCE1 |
synaptonemal complex central element protein 1 |
| chr10_+_6779326 | 0.47 |
ENST00000417112.1 |
RP11-554I8.2 |
RP11-554I8.2 |
| chr1_+_160370344 | 0.44 |
ENST00000368061.2 |
VANGL2 |
VANGL planar cell polarity protein 2 |
| chr3_-_151034734 | 0.43 |
ENST00000260843.4 |
GPR87 |
G protein-coupled receptor 87 |
| chr10_-_105845674 | 0.33 |
ENST00000353479.5 ENST00000369733.3 |
COL17A1 |
collagen, type XVII, alpha 1 |
| chr4_-_74486217 | 0.31 |
ENST00000335049.5 ENST00000307439.5 |
RASSF6 |
Ras association (RalGDS/AF-6) domain family member 6 |
| chr18_+_34124507 | 0.31 |
ENST00000591635.1 |
FHOD3 |
formin homology 2 domain containing 3 |
| chr2_+_228736335 | 0.29 |
ENST00000440997.1 ENST00000545118.1 |
DAW1 |
dynein assembly factor with WDR repeat domains 1 |
| chr4_-_25865159 | 0.28 |
ENST00000502949.1 ENST00000264868.5 ENST00000513691.1 ENST00000514872.1 |
SEL1L3 |
sel-1 suppressor of lin-12-like 3 (C. elegans) |
| chr13_+_32313658 | 0.27 |
ENST00000380314.1 ENST00000298386.2 |
RXFP2 |
relaxin/insulin-like family peptide receptor 2 |
| chr5_+_66300446 | 0.26 |
ENST00000261569.7 |
MAST4 |
microtubule associated serine/threonine kinase family member 4 |
| chr17_-_2996290 | 0.26 |
ENST00000331459.1 |
OR1D2 |
olfactory receptor, family 1, subfamily D, member 2 |
| chr14_+_61654271 | 0.25 |
ENST00000555185.1 ENST00000557294.1 ENST00000556778.1 |
PRKCH |
protein kinase C, eta |
| chr3_+_111718036 | 0.24 |
ENST00000455401.2 |
TAGLN3 |
transgelin 3 |
| chr1_-_242612779 | 0.24 |
ENST00000427495.1 |
PLD5 |
phospholipase D family, member 5 |
| chr3_+_111717600 | 0.24 |
ENST00000273368.4 |
TAGLN3 |
transgelin 3 |
| chr3_+_111718173 | 0.24 |
ENST00000494932.1 |
TAGLN3 |
transgelin 3 |
| chr1_+_28261492 | 0.23 |
ENST00000373894.3 |
SMPDL3B |
sphingomyelin phosphodiesterase, acid-like 3B |
| chr4_+_119810134 | 0.23 |
ENST00000434046.2 |
SYNPO2 |
synaptopodin 2 |
| chr15_+_43886057 | 0.22 |
ENST00000441322.1 ENST00000413657.2 ENST00000453733.1 |
CKMT1B |
creatine kinase, mitochondrial 1B |
| chr11_-_26593779 | 0.22 |
ENST00000529533.1 |
MUC15 |
mucin 15, cell surface associated |
| chr3_+_111717511 | 0.22 |
ENST00000478951.1 ENST00000393917.2 |
TAGLN3 |
transgelin 3 |
| chr11_-_26593677 | 0.21 |
ENST00000527569.1 |
MUC15 |
mucin 15, cell surface associated |
| chr2_+_228736321 | 0.21 |
ENST00000309931.2 |
DAW1 |
dynein assembly factor with WDR repeat domains 1 |
| chr15_+_43985725 | 0.21 |
ENST00000413453.2 |
CKMT1A |
creatine kinase, mitochondrial 1A |
| chr9_-_5304432 | 0.21 |
ENST00000416837.1 ENST00000308420.3 |
RLN2 |
relaxin 2 |
| chr1_+_81771806 | 0.21 |
ENST00000370721.1 ENST00000370727.1 ENST00000370725.1 ENST00000370723.1 ENST00000370728.1 ENST00000370730.1 |
LPHN2 |
latrophilin 2 |
| chr16_-_28634874 | 0.20 |
ENST00000395609.1 ENST00000350842.4 |
SULT1A1 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chr13_-_86373536 | 0.20 |
ENST00000400286.2 |
SLITRK6 |
SLIT and NTRK-like family, member 6 |
| chr12_-_95510743 | 0.20 |
ENST00000551521.1 |
FGD6 |
FYVE, RhoGEF and PH domain containing 6 |
| chr5_+_55149150 | 0.18 |
ENST00000297015.3 |
IL31RA |
interleukin 31 receptor A |
| chr4_-_87028478 | 0.18 |
ENST00000515400.1 ENST00000395157.3 |
MAPK10 |
mitogen-activated protein kinase 10 |
| chr6_-_9933500 | 0.18 |
ENST00000492169.1 |
OFCC1 |
orofacial cleft 1 candidate 1 |
| chr10_-_14590644 | 0.17 |
ENST00000378470.1 |
FAM107B |
family with sequence similarity 107, member B |
| chr6_-_28411241 | 0.16 |
ENST00000289788.4 |
ZSCAN23 |
zinc finger and SCAN domain containing 23 |
| chr1_+_62439037 | 0.16 |
ENST00000545929.1 |
INADL |
InaD-like (Drosophila) |
| chr11_-_13517565 | 0.16 |
ENST00000282091.1 ENST00000529816.1 |
PTH |
parathyroid hormone |
| chr6_+_130339710 | 0.16 |
ENST00000526087.1 ENST00000533560.1 ENST00000361794.2 |
L3MBTL3 |
l(3)mbt-like 3 (Drosophila) |
| chr11_+_101918153 | 0.16 |
ENST00000434758.2 ENST00000526781.1 ENST00000534360.1 |
C11orf70 |
chromosome 11 open reading frame 70 |
| chr17_-_39274606 | 0.15 |
ENST00000391413.2 |
KRTAP4-11 |
keratin associated protein 4-11 |
| chr18_+_61254570 | 0.15 |
ENST00000344731.5 |
SERPINB13 |
serpin peptidase inhibitor, clade B (ovalbumin), member 13 |
| chr5_+_36608422 | 0.15 |
ENST00000381918.3 |
SLC1A3 |
solute carrier family 1 (glial high affinity glutamate transporter), member 3 |
| chr18_+_61254534 | 0.15 |
ENST00000269489.5 |
SERPINB13 |
serpin peptidase inhibitor, clade B (ovalbumin), member 13 |
| chr5_+_110427983 | 0.14 |
ENST00000513710.2 ENST00000505303.1 |
WDR36 |
WD repeat domain 36 |
| chr1_-_185597619 | 0.14 |
ENST00000608417.1 ENST00000436955.1 |
GS1-204I12.1 |
GS1-204I12.1 |
| chr18_+_55888767 | 0.14 |
ENST00000431212.2 ENST00000586268.1 ENST00000587190.1 |
NEDD4L |
neural precursor cell expressed, developmentally down-regulated 4-like, E3 ubiquitin protein ligase |
| chr17_-_38956205 | 0.14 |
ENST00000306658.7 |
KRT28 |
keratin 28 |
| chr14_+_56584414 | 0.14 |
ENST00000559044.1 |
PELI2 |
pellino E3 ubiquitin protein ligase family member 2 |
| chr9_-_5339873 | 0.14 |
ENST00000223862.1 ENST00000223858.4 |
RLN1 |
relaxin 1 |
| chr12_-_22063787 | 0.14 |
ENST00000544039.1 |
ABCC9 |
ATP-binding cassette, sub-family C (CFTR/MRP), member 9 |
| chr3_-_112565703 | 0.13 |
ENST00000488794.1 |
CD200R1L |
CD200 receptor 1-like |
| chrX_-_50386648 | 0.13 |
ENST00000460112.3 |
SHROOM4 |
shroom family member 4 |
| chr14_-_23652849 | 0.13 |
ENST00000316902.7 ENST00000469263.1 ENST00000525062.1 ENST00000524758.1 |
SLC7A8 |
solute carrier family 7 (amino acid transporter light chain, L system), member 8 |
| chr8_+_105235572 | 0.13 |
ENST00000523362.1 |
RIMS2 |
regulating synaptic membrane exocytosis 2 |
| chr4_+_88896819 | 0.13 |
ENST00000237623.7 ENST00000395080.3 ENST00000508233.1 ENST00000360804.4 |
SPP1 |
secreted phosphoprotein 1 |
| chr8_-_66474884 | 0.13 |
ENST00000520902.1 |
CTD-3025N20.2 |
CTD-3025N20.2 |
| chr1_+_207226574 | 0.12 |
ENST00000367080.3 ENST00000367079.2 |
PFKFB2 |
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2 |
| chr19_+_9296279 | 0.12 |
ENST00000344248.2 |
OR7D2 |
olfactory receptor, family 7, subfamily D, member 2 |
| chr3_-_112127981 | 0.12 |
ENST00000486726.2 |
RP11-231E6.1 |
RP11-231E6.1 |
| chr6_+_125540951 | 0.12 |
ENST00000524679.1 |
TPD52L1 |
tumor protein D52-like 1 |
| chrX_+_130192318 | 0.12 |
ENST00000370922.1 |
ARHGAP36 |
Rho GTPase activating protein 36 |
| chr3_-_191000172 | 0.12 |
ENST00000427544.2 |
UTS2B |
urotensin 2B |
| chr6_-_32908765 | 0.11 |
ENST00000416244.2 |
HLA-DMB |
major histocompatibility complex, class II, DM beta |
| chr9_+_12693336 | 0.11 |
ENST00000381137.2 ENST00000388918.5 |
TYRP1 |
tyrosinase-related protein 1 |
| chr17_-_4938712 | 0.11 |
ENST00000254853.5 ENST00000424747.1 |
SLC52A1 |
solute carrier family 52 (riboflavin transporter), member 1 |
| chr9_-_95166884 | 0.11 |
ENST00000375561.5 |
OGN |
osteoglycin |
| chr2_-_101925055 | 0.11 |
ENST00000295317.3 |
RNF149 |
ring finger protein 149 |
| chr1_+_101003687 | 0.11 |
ENST00000315033.4 |
GPR88 |
G protein-coupled receptor 88 |
| chr5_-_139726181 | 0.10 |
ENST00000507104.1 ENST00000230990.6 |
HBEGF |
heparin-binding EGF-like growth factor |
| chr4_-_74486109 | 0.10 |
ENST00000395777.2 |
RASSF6 |
Ras association (RalGDS/AF-6) domain family member 6 |
| chr5_+_140227048 | 0.10 |
ENST00000532602.1 |
PCDHA9 |
protocadherin alpha 9 |
| chr11_-_102651343 | 0.10 |
ENST00000279441.4 ENST00000539681.1 |
MMP10 |
matrix metallopeptidase 10 (stromelysin 2) |
| chr11_-_107729887 | 0.10 |
ENST00000525815.1 |
SLC35F2 |
solute carrier family 35, member F2 |
| chr15_+_58430567 | 0.10 |
ENST00000536493.1 |
AQP9 |
aquaporin 9 |
| chr1_-_197115818 | 0.10 |
ENST00000367409.4 ENST00000294732.7 |
ASPM |
asp (abnormal spindle) homolog, microcephaly associated (Drosophila) |
| chr9_-_95166841 | 0.10 |
ENST00000262551.4 |
OGN |
osteoglycin |
| chr2_+_102615416 | 0.10 |
ENST00000393414.2 |
IL1R2 |
interleukin 1 receptor, type II |
| chr1_-_24741525 | 0.10 |
ENST00000374409.1 |
STPG1 |
sperm-tail PG-rich repeat containing 1 |
| chr7_+_142982023 | 0.09 |
ENST00000359333.3 ENST00000409244.1 ENST00000409541.1 ENST00000410004.1 |
TMEM139 |
transmembrane protein 139 |
| chr2_+_202047596 | 0.09 |
ENST00000286186.6 ENST00000360132.3 |
CASP10 |
caspase 10, apoptosis-related cysteine peptidase |
| chr11_+_118398178 | 0.09 |
ENST00000302783.4 ENST00000539546.1 |
TTC36 |
tetratricopeptide repeat domain 36 |
| chr2_-_89327228 | 0.09 |
ENST00000483158.1 |
IGKV3-11 |
immunoglobulin kappa variable 3-11 |
| chr10_-_50970382 | 0.09 |
ENST00000419399.1 ENST00000432695.1 |
OGDHL |
oxoglutarate dehydrogenase-like |
| chrX_-_130423200 | 0.09 |
ENST00000361420.3 |
IGSF1 |
immunoglobulin superfamily, member 1 |
| chr16_+_12059050 | 0.09 |
ENST00000396495.3 |
TNFRSF17 |
tumor necrosis factor receptor superfamily, member 17 |
| chr12_+_26348246 | 0.09 |
ENST00000422622.2 |
SSPN |
sarcospan |
| chr4_-_120243545 | 0.09 |
ENST00000274024.3 |
FABP2 |
fatty acid binding protein 2, intestinal |
| chr2_+_90077680 | 0.09 |
ENST00000390270.2 |
IGKV3D-20 |
immunoglobulin kappa variable 3D-20 |
| chr17_-_40337470 | 0.09 |
ENST00000293330.1 |
HCRT |
hypocretin (orexin) neuropeptide precursor |
| chr11_+_121447469 | 0.09 |
ENST00000532694.1 ENST00000534286.1 |
SORL1 |
sortilin-related receptor, L(DLR class) A repeats containing |
| chr10_-_50970322 | 0.09 |
ENST00000374103.4 |
OGDHL |
oxoglutarate dehydrogenase-like |
| chr6_+_26365443 | 0.09 |
ENST00000527422.1 ENST00000356386.2 ENST00000396934.3 ENST00000377708.2 ENST00000396948.1 ENST00000508906.2 |
BTN3A2 |
butyrophilin, subfamily 3, member A2 |
| chr9_+_108463234 | 0.09 |
ENST00000374688.1 |
TMEM38B |
transmembrane protein 38B |
| chr4_-_74486347 | 0.09 |
ENST00000342081.3 |
RASSF6 |
Ras association (RalGDS/AF-6) domain family member 6 |
| chr4_-_39979576 | 0.09 |
ENST00000303538.8 ENST00000503396.1 |
PDS5A |
PDS5, regulator of cohesion maintenance, homolog A (S. cerevisiae) |
| chr1_+_70876926 | 0.08 |
ENST00000370938.3 ENST00000346806.2 |
CTH |
cystathionase (cystathionine gamma-lyase) |
| chr1_+_70876891 | 0.08 |
ENST00000411986.2 |
CTH |
cystathionase (cystathionine gamma-lyase) |
| chr9_-_117111222 | 0.08 |
ENST00000374079.4 |
AKNA |
AT-hook transcription factor |
| chr15_+_94899183 | 0.08 |
ENST00000557742.1 |
MCTP2 |
multiple C2 domains, transmembrane 2 |
| chr17_-_73663245 | 0.08 |
ENST00000584999.1 ENST00000317905.5 ENST00000420326.2 ENST00000340830.5 |
RECQL5 |
RecQ protein-like 5 |
| chr8_+_107738343 | 0.08 |
ENST00000521592.1 |
OXR1 |
oxidation resistance 1 |
| chr15_+_35270552 | 0.08 |
ENST00000391457.2 |
AC114546.1 |
HCG37415; PRO1914; Uncharacterized protein |
| chr4_+_71200681 | 0.08 |
ENST00000273936.5 |
CABS1 |
calcium-binding protein, spermatid-specific 1 |
| chr14_+_21387508 | 0.08 |
ENST00000555624.1 |
RP11-84C10.2 |
RP11-84C10.2 |
| chr10_+_5238793 | 0.08 |
ENST00000263126.1 |
AKR1C4 |
aldo-keto reductase family 1, member C4 |
| chr17_-_48785216 | 0.08 |
ENST00000285243.6 |
ANKRD40 |
ankyrin repeat domain 40 |
| chr17_-_5321549 | 0.08 |
ENST00000572809.1 |
NUP88 |
nucleoporin 88kDa |
| chr3_-_185538849 | 0.08 |
ENST00000421047.2 |
IGF2BP2 |
insulin-like growth factor 2 mRNA binding protein 2 |
| chr4_+_77356248 | 0.08 |
ENST00000296043.6 |
SHROOM3 |
shroom family member 3 |
| chr2_-_134326009 | 0.08 |
ENST00000409261.1 ENST00000409213.1 |
NCKAP5 |
NCK-associated protein 5 |
| chr11_+_55594695 | 0.08 |
ENST00000378397.1 |
OR5L2 |
olfactory receptor, family 5, subfamily L, member 2 |
| chr12_+_8276495 | 0.08 |
ENST00000546339.1 |
CLEC4A |
C-type lectin domain family 4, member A |
| chr12_-_15114658 | 0.08 |
ENST00000542276.1 |
ARHGDIB |
Rho GDP dissociation inhibitor (GDI) beta |
| chr21_+_43619796 | 0.08 |
ENST00000398457.2 |
ABCG1 |
ATP-binding cassette, sub-family G (WHITE), member 1 |
| chr19_-_3557570 | 0.08 |
ENST00000355415.2 |
MFSD12 |
major facilitator superfamily domain containing 12 |
| chr10_+_695888 | 0.08 |
ENST00000441152.2 |
PRR26 |
proline rich 26 |
| chr16_+_58010339 | 0.08 |
ENST00000290871.5 ENST00000441824.2 |
TEPP |
testis, prostate and placenta expressed |
| chr14_-_67878917 | 0.08 |
ENST00000216446.4 |
PLEK2 |
pleckstrin 2 |
| chr20_-_50418947 | 0.08 |
ENST00000371539.3 |
SALL4 |
spalt-like transcription factor 4 |
| chr12_+_28410128 | 0.08 |
ENST00000381259.1 ENST00000381256.1 |
CCDC91 |
coiled-coil domain containing 91 |
| chr17_+_43238438 | 0.08 |
ENST00000593138.1 ENST00000586681.1 |
HEXIM2 |
hexamethylene bis-acetamide inducible 2 |
| chr14_-_78083112 | 0.08 |
ENST00000216484.2 |
SPTLC2 |
serine palmitoyltransferase, long chain base subunit 2 |
| chr20_-_50418972 | 0.08 |
ENST00000395997.3 |
SALL4 |
spalt-like transcription factor 4 |
| chr6_+_134758827 | 0.07 |
ENST00000431422.1 |
LINC01010 |
long intergenic non-protein coding RNA 1010 |
| chr11_+_18154059 | 0.07 |
ENST00000531264.1 |
MRGPRX3 |
MAS-related GPR, member X3 |
| chr3_-_33686743 | 0.07 |
ENST00000333778.6 ENST00000539981.1 |
CLASP2 |
cytoplasmic linker associated protein 2 |
| chr5_-_82969405 | 0.07 |
ENST00000510978.1 |
HAPLN1 |
hyaluronan and proteoglycan link protein 1 |
| chr20_-_50419055 | 0.07 |
ENST00000217086.4 |
SALL4 |
spalt-like transcription factor 4 |
| chr4_+_70861647 | 0.07 |
ENST00000246895.4 ENST00000381060.2 |
STATH |
statherin |
| chr5_-_41213607 | 0.07 |
ENST00000337836.5 ENST00000433294.1 |
C6 |
complement component 6 |
| chr12_-_15114603 | 0.07 |
ENST00000228945.4 |
ARHGDIB |
Rho GDP dissociation inhibitor (GDI) beta |
| chrX_-_15332665 | 0.07 |
ENST00000537676.1 ENST00000344384.4 |
ASB11 |
ankyrin repeat and SOCS box containing 11 |
| chr14_-_21566731 | 0.07 |
ENST00000360947.3 |
ZNF219 |
zinc finger protein 219 |
| chr22_-_29107919 | 0.07 |
ENST00000434810.1 ENST00000456369.1 |
CHEK2 |
checkpoint kinase 2 |
| chr16_+_82090028 | 0.07 |
ENST00000568090.1 |
HSD17B2 |
hydroxysteroid (17-beta) dehydrogenase 2 |
| chr11_-_124190184 | 0.07 |
ENST00000357438.2 |
OR8D2 |
olfactory receptor, family 8, subfamily D, member 2 |
| chr21_-_31588365 | 0.07 |
ENST00000399899.1 |
CLDN8 |
claudin 8 |
| chr3_+_147795932 | 0.07 |
ENST00000490465.1 |
RP11-639B1.1 |
RP11-639B1.1 |
| chr16_+_15489603 | 0.07 |
ENST00000568766.1 ENST00000287594.7 |
RP11-1021N1.1 MPV17L |
Uncharacterized protein MPV17 mitochondrial membrane protein-like |
| chr9_+_130911770 | 0.07 |
ENST00000372998.1 |
LCN2 |
lipocalin 2 |
| chr18_-_74839891 | 0.07 |
ENST00000581878.1 |
MBP |
myelin basic protein |
| chr5_+_89770664 | 0.07 |
ENST00000503973.1 ENST00000399107.1 |
POLR3G |
polymerase (RNA) III (DNA directed) polypeptide G (32kD) |
| chr3_+_97887544 | 0.07 |
ENST00000356526.2 |
OR5H15 |
olfactory receptor, family 5, subfamily H, member 15 |
| chr21_-_31588338 | 0.07 |
ENST00000286809.1 |
CLDN8 |
claudin 8 |
| chr4_+_71458012 | 0.07 |
ENST00000449493.2 |
AMBN |
ameloblastin (enamel matrix protein) |
| chr10_-_98031265 | 0.07 |
ENST00000224337.5 ENST00000371176.2 |
BLNK |
B-cell linker |
| chr1_+_12834984 | 0.07 |
ENST00000357726.4 |
PRAMEF12 |
PRAME family member 12 |
| chr10_-_47151341 | 0.07 |
ENST00000422732.2 |
LINC00842 |
long intergenic non-protein coding RNA 842 |
| chrX_+_144908928 | 0.07 |
ENST00000408967.2 |
TMEM257 |
transmembrane protein 257 |
| chr2_+_90273679 | 0.07 |
ENST00000423080.2 |
IGKV3D-7 |
immunoglobulin kappa variable 3D-7 |
| chr12_+_20963632 | 0.06 |
ENST00000540853.1 ENST00000261196.2 |
SLCO1B3 |
solute carrier organic anion transporter family, member 1B3 |
| chr4_+_80584903 | 0.06 |
ENST00000506460.1 |
RP11-452C8.1 |
RP11-452C8.1 |
| chr5_+_89770696 | 0.06 |
ENST00000504930.1 ENST00000514483.1 |
POLR3G |
polymerase (RNA) III (DNA directed) polypeptide G (32kD) |
| chr11_-_128894053 | 0.06 |
ENST00000392657.3 |
ARHGAP32 |
Rho GTPase activating protein 32 |
| chr10_-_135379074 | 0.06 |
ENST00000303903.6 |
SYCE1 |
synaptonemal complex central element protein 1 |
| chr21_+_17442799 | 0.06 |
ENST00000602580.1 ENST00000458468.1 ENST00000602935.1 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr3_-_151047327 | 0.06 |
ENST00000325602.5 |
P2RY13 |
purinergic receptor P2Y, G-protein coupled, 13 |
| chr17_-_10372875 | 0.06 |
ENST00000255381.2 |
MYH4 |
myosin, heavy chain 4, skeletal muscle |
| chr16_-_30122717 | 0.06 |
ENST00000566613.1 |
GDPD3 |
glycerophosphodiester phosphodiesterase domain containing 3 |
| chr3_-_33686925 | 0.06 |
ENST00000485378.2 ENST00000313350.6 ENST00000487200.1 |
CLASP2 |
cytoplasmic linker associated protein 2 |
| chr16_-_28621312 | 0.06 |
ENST00000314752.7 |
SULT1A1 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chr3_-_98235962 | 0.06 |
ENST00000513873.1 |
CLDND1 |
claudin domain containing 1 |
| chr17_-_27418537 | 0.06 |
ENST00000408971.2 |
TIAF1 |
TGFB1-induced anti-apoptotic factor 1 |
| chrX_+_150565653 | 0.06 |
ENST00000330374.6 |
VMA21 |
VMA21 vacuolar H+-ATPase homolog (S. cerevisiae) |
| chr4_-_46911223 | 0.06 |
ENST00000396533.1 |
COX7B2 |
cytochrome c oxidase subunit VIIb2 |
| chr11_-_104827425 | 0.06 |
ENST00000393150.3 |
CASP4 |
caspase 4, apoptosis-related cysteine peptidase |
| chr12_+_7014126 | 0.06 |
ENST00000415834.1 ENST00000436789.1 |
LRRC23 |
leucine rich repeat containing 23 |
| chr7_-_111032971 | 0.06 |
ENST00000450877.1 |
IMMP2L |
IMP2 inner mitochondrial membrane peptidase-like (S. cerevisiae) |
| chr4_+_86525299 | 0.06 |
ENST00000512201.1 |
ARHGAP24 |
Rho GTPase activating protein 24 |
| chr5_-_141249154 | 0.06 |
ENST00000357517.5 ENST00000536585.1 |
PCDH1 |
protocadherin 1 |
| chr4_-_79860506 | 0.06 |
ENST00000295462.3 ENST00000380645.4 ENST00000512733.1 |
PAQR3 |
progestin and adipoQ receptor family member III |
| chr2_+_201754050 | 0.06 |
ENST00000426253.1 ENST00000416651.1 ENST00000454952.1 ENST00000409020.1 ENST00000359683.4 |
NIF3L1 |
NIF3 NGG1 interacting factor 3-like 1 (S. cerevisiae) |
| chr9_+_130911723 | 0.06 |
ENST00000277480.2 ENST00000373013.2 ENST00000540948.1 |
LCN2 |
lipocalin 2 |
| chr2_+_201754135 | 0.06 |
ENST00000409357.1 ENST00000409129.2 |
NIF3L1 |
NIF3 NGG1 interacting factor 3-like 1 (S. cerevisiae) |
| chr12_-_112123524 | 0.06 |
ENST00000327551.6 |
BRAP |
BRCA1 associated protein |
| chr11_+_65554493 | 0.06 |
ENST00000335987.3 |
OVOL1 |
ovo-like zinc finger 1 |
| chr15_+_49462434 | 0.06 |
ENST00000558145.1 ENST00000543495.1 ENST00000544523.1 ENST00000560138.1 |
GALK2 |
galactokinase 2 |
| chr10_+_5454505 | 0.06 |
ENST00000355029.4 |
NET1 |
neuroepithelial cell transforming 1 |
| chr3_-_143567262 | 0.06 |
ENST00000474151.1 ENST00000316549.6 |
SLC9A9 |
solute carrier family 9, subfamily A (NHE9, cation proton antiporter 9), member 9 |
| chr1_+_75198828 | 0.06 |
ENST00000457880.2 ENST00000370867.3 ENST00000421739.2 |
TYW3 |
tRNA-yW synthesizing protein 3 homolog (S. cerevisiae) |
| chr4_+_41614909 | 0.06 |
ENST00000509454.1 ENST00000396595.3 ENST00000381753.4 |
LIMCH1 |
LIM and calponin homology domains 1 |
| chr4_+_119809984 | 0.06 |
ENST00000307142.4 ENST00000448416.2 ENST00000429713.2 |
SYNPO2 |
synaptopodin 2 |
| chr7_+_129906660 | 0.06 |
ENST00000222481.4 |
CPA2 |
carboxypeptidase A2 (pancreatic) |
| chr15_-_54051831 | 0.05 |
ENST00000557913.1 ENST00000360509.5 |
WDR72 |
WD repeat domain 72 |
| chr5_-_93077293 | 0.05 |
ENST00000510627.4 |
POU5F2 |
POU domain class 5, transcription factor 2 |
| chr7_-_37026108 | 0.05 |
ENST00000396045.3 |
ELMO1 |
engulfment and cell motility 1 |
| chr12_+_26348429 | 0.05 |
ENST00000242729.2 |
SSPN |
sarcospan |
| chr10_-_98031310 | 0.05 |
ENST00000427367.2 ENST00000413476.2 |
BLNK |
B-cell linker |
| chr1_+_86934526 | 0.05 |
ENST00000394711.1 |
CLCA1 |
chloride channel accessory 1 |
| chr2_+_228735763 | 0.05 |
ENST00000373666.2 |
DAW1 |
dynein assembly factor with WDR repeat domains 1 |
| chr19_+_15160130 | 0.05 |
ENST00000427043.3 |
CASP14 |
caspase 14, apoptosis-related cysteine peptidase |
| chr20_-_4721314 | 0.05 |
ENST00000418528.1 ENST00000326539.2 ENST00000423718.2 |
PRNT |
prion protein (testis specific) |
| chr1_+_107683644 | 0.05 |
ENST00000370067.1 |
NTNG1 |
netrin G1 |
| chrX_+_49687267 | 0.05 |
ENST00000376091.3 |
CLCN5 |
chloride channel, voltage-sensitive 5 |
| chr7_+_107224364 | 0.05 |
ENST00000491150.1 |
BCAP29 |
B-cell receptor-associated protein 29 |
| chr10_+_696000 | 0.05 |
ENST00000381489.5 |
PRR26 |
proline rich 26 |
| chr1_+_107683436 | 0.05 |
ENST00000370068.1 |
NTNG1 |
netrin G1 |
| chr1_-_222763101 | 0.05 |
ENST00000391883.2 ENST00000366890.1 |
TAF1A |
TATA box binding protein (TBP)-associated factor, RNA polymerase I, A, 48kDa |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:1904956 | regulation of midbrain dopaminergic neuron differentiation(GO:1904956) regulation of Wnt signaling pathway involved in dorsal/ventral axis specification(GO:2000053) |
| 0.1 | 0.4 | GO:0061341 | orthogonal dichotomous subdivision of terminal units involved in lung branching morphogenesis(GO:0060488) planar dichotomous subdivision of terminal units involved in lung branching morphogenesis(GO:0060489) lateral sprouting involved in lung morphogenesis(GO:0060490) non-canonical Wnt signaling pathway involved in heart development(GO:0061341) planar cell polarity pathway involved in heart morphogenesis(GO:0061346) |
| 0.1 | 0.6 | GO:1903575 | cornified envelope assembly(GO:1903575) |
| 0.1 | 0.2 | GO:2001190 | positive regulation of T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:2001190) |
| 0.1 | 0.2 | GO:1900158 | positive regulation of osteoclast proliferation(GO:0090290) negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 0.1 | 0.2 | GO:2000866 | positive regulation of estrogen secretion(GO:2000863) positive regulation of estradiol secretion(GO:2000866) |
| 0.0 | 0.2 | GO:0018352 | protein-pyridoxal-5-phosphate linkage(GO:0018352) |
| 0.0 | 0.1 | GO:1990414 | replication-born double-strand break repair via sister chromatid exchange(GO:1990414) |
| 0.0 | 0.1 | GO:0061743 | motor learning(GO:0061743) |
| 0.0 | 0.2 | GO:0060005 | vestibular reflex(GO:0060005) |
| 0.0 | 0.0 | GO:1902908 | regulation of melanosome transport(GO:1902908) regulation of melanosome organization(GO:1903056) |
| 0.0 | 0.1 | GO:0009786 | regulation of asymmetric cell division(GO:0009786) |
| 0.0 | 0.2 | GO:0009449 | gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 0.0 | 0.2 | GO:0071461 | cellular response to redox state(GO:0071461) |
| 0.0 | 0.1 | GO:1902771 | regulation of choline O-acetyltransferase activity(GO:1902769) positive regulation of choline O-acetyltransferase activity(GO:1902771) negative regulation of tau-protein kinase activity(GO:1902948) positive regulation of early endosome to recycling endosome transport(GO:1902955) negative regulation of aspartic-type endopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902960) negative regulation of neurofibrillary tangle assembly(GO:1902997) negative regulation of aspartic-type peptidase activity(GO:1905246) |
| 0.0 | 0.1 | GO:0015722 | canalicular bile acid transport(GO:0015722) |
| 0.0 | 0.1 | GO:2001288 | positive regulation of caveolin-mediated endocytosis(GO:2001288) |
| 0.0 | 0.3 | GO:0097283 | keratinocyte apoptotic process(GO:0097283) regulation of keratinocyte apoptotic process(GO:1902172) |
| 0.0 | 0.1 | GO:0015891 | iron chelate transport(GO:0015688) siderophore transport(GO:0015891) |
| 0.0 | 0.1 | GO:0006667 | sphinganine metabolic process(GO:0006667) |
| 0.0 | 0.2 | GO:0034351 | regulation of glial cell apoptotic process(GO:0034350) negative regulation of glial cell apoptotic process(GO:0034351) |
| 0.0 | 0.1 | GO:0032690 | negative regulation of interleukin-1 alpha production(GO:0032690) negative regulation of interleukin-1 alpha secretion(GO:0050712) |
| 0.0 | 0.1 | GO:0008063 | Toll signaling pathway(GO:0008063) |
| 0.0 | 0.1 | GO:1903926 | signal transduction involved in intra-S DNA damage checkpoint(GO:0072428) response to bisphenol A(GO:1903925) cellular response to bisphenol A(GO:1903926) |
| 0.0 | 0.6 | GO:0007130 | synaptonemal complex assembly(GO:0007130) |
| 0.0 | 0.2 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 0.0 | 0.1 | GO:0061713 | neural crest cell migration involved in heart formation(GO:0003147) anterior neural tube closure(GO:0061713) |
| 0.0 | 0.1 | GO:0032218 | riboflavin transport(GO:0032218) |
| 0.0 | 0.2 | GO:0006600 | creatine metabolic process(GO:0006600) |
| 0.0 | 0.1 | GO:0033133 | fructose 2,6-bisphosphate metabolic process(GO:0006003) positive regulation of glucokinase activity(GO:0033133) positive regulation of hexokinase activity(GO:1903301) |
| 0.0 | 0.1 | GO:0009753 | response to jasmonic acid(GO:0009753) cellular response to jasmonic acid stimulus(GO:0071395) |
| 0.0 | 0.1 | GO:0061300 | cerebellum vasculature development(GO:0061300) |
| 0.0 | 0.5 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.0 | 0.1 | GO:0055099 | detection of endogenous stimulus(GO:0009726) response to high density lipoprotein particle(GO:0055099) |
| 0.0 | 0.1 | GO:0044771 | meiotic cell cycle phase transition(GO:0044771) regulation of meiotic cell cycle phase transition(GO:1901993) negative regulation of meiotic cell cycle phase transition(GO:1901994) |
| 0.0 | 0.2 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.0 | 0.3 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.0 | 0.1 | GO:0001970 | positive regulation of activation of membrane attack complex(GO:0001970) |
| 0.0 | 0.1 | GO:0000294 | nuclear-transcribed mRNA catabolic process, endonucleolytic cleavage-dependent decay(GO:0000294) |
| 0.0 | 0.1 | GO:0010730 | negative regulation of hydrogen peroxide biosynthetic process(GO:0010730) |
| 0.0 | 0.0 | GO:0072200 | apoptotic process involved in endocardial cushion morphogenesis(GO:0003277) tendon cell differentiation(GO:0035990) tendon formation(GO:0035992) intermediate mesoderm morphogenesis(GO:0048390) intermediate mesoderm formation(GO:0048391) intermediate mesodermal cell differentiation(GO:0048392) regulation of cardiac muscle fiber development(GO:0055018) positive regulation of cardiac muscle fiber development(GO:0055020) bud dilation involved in lung branching(GO:0060503) BMP signaling pathway involved in ureter morphogenesis(GO:0061149) renal system segmentation(GO:0061150) BMP signaling pathway involved in renal system segmentation(GO:0061151) pulmonary artery endothelial tube morphogenesis(GO:0061155) regulation of transcription from RNA polymerase II promoter involved in mesonephros development(GO:0061216) BMP signaling pathway involved in nephric duct formation(GO:0071893) negative regulation of branch elongation involved in ureteric bud branching(GO:0072096) negative regulation of branch elongation involved in ureteric bud branching by BMP signaling pathway(GO:0072097) anterior/posterior pattern specification involved in ureteric bud development(GO:0072099) specification of ureteric bud anterior/posterior symmetry(GO:0072100) specification of ureteric bud anterior/posterior symmetry by BMP signaling pathway(GO:0072101) ureter epithelial cell differentiation(GO:0072192) negative regulation of mesenchymal cell proliferation involved in ureter development(GO:0072200) positive regulation of cell proliferation involved in outflow tract morphogenesis(GO:1901964) cardiac jelly development(GO:1905072) regulation of metanephric S-shaped body morphogenesis(GO:2000004) negative regulation of metanephric S-shaped body morphogenesis(GO:2000005) regulation of metanephric comma-shaped body morphogenesis(GO:2000006) negative regulation of metanephric comma-shaped body morphogenesis(GO:2000007) |
| 0.0 | 0.1 | GO:0061669 | spontaneous neurotransmitter secretion(GO:0061669) spontaneous synaptic transmission(GO:0098814) |
| 0.0 | 0.2 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.0 | 0.1 | GO:0051970 | negative regulation of transmission of nerve impulse(GO:0051970) |
| 0.0 | 0.1 | GO:1904685 | positive regulation of metalloendopeptidase activity(GO:1904685) |
| 0.0 | 0.0 | GO:2000502 | negative regulation of natural killer cell chemotaxis(GO:2000502) |
| 0.0 | 0.0 | GO:0002644 | negative regulation of tolerance induction(GO:0002644) positive regulation of interleukin-4 biosynthetic process(GO:0045404) |
| 0.0 | 0.2 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.0 | 0.0 | GO:0072709 | cellular response to sorbitol(GO:0072709) |
| 0.0 | 0.0 | GO:0030807 | positive regulation of cyclic nucleotide catabolic process(GO:0030807) positive regulation of cAMP catabolic process(GO:0030822) positive regulation of purine nucleotide catabolic process(GO:0033123) |
| 0.0 | 0.1 | GO:0000965 | mitochondrial RNA 3'-end processing(GO:0000965) |
| 0.0 | 0.0 | GO:0070105 | positive regulation of interleukin-6-mediated signaling pathway(GO:0070105) |
| 0.0 | 0.2 | GO:0043031 | negative regulation of macrophage activation(GO:0043031) |
| 0.0 | 0.3 | GO:0006068 | ethanol catabolic process(GO:0006068) |
| 0.0 | 0.3 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.0 | 0.0 | GO:0034310 | primary alcohol catabolic process(GO:0034310) |
| 0.0 | 0.0 | GO:1901529 | positive regulation of anion channel activity(GO:1901529) |
| 0.0 | 0.0 | GO:0006435 | threonyl-tRNA aminoacylation(GO:0006435) |
| 0.0 | 0.0 | GO:0070563 | negative regulation of vitamin D receptor signaling pathway(GO:0070563) |
| 0.0 | 0.2 | GO:0042340 | keratan sulfate catabolic process(GO:0042340) |
| 0.0 | 0.1 | GO:1900005 | positive regulation of serine-type endopeptidase activity(GO:1900005) positive regulation of serine-type peptidase activity(GO:1902573) |
| 0.0 | 0.0 | GO:0001694 | histamine biosynthetic process(GO:0001694) |
| 0.0 | 0.1 | GO:0090091 | positive regulation of extracellular matrix disassembly(GO:0090091) |
| 0.0 | 0.0 | GO:0006258 | UDP-glucose catabolic process(GO:0006258) |
| 0.0 | 0.0 | GO:0002331 | pre-B cell allelic exclusion(GO:0002331) |
| 0.0 | 0.0 | GO:1990167 | protein K27-linked deubiquitination(GO:1990167) protein K33-linked deubiquitination(GO:1990168) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0060187 | cell pole(GO:0060187) |
| 0.1 | 0.5 | GO:0000801 | central element(GO:0000801) |
| 0.0 | 0.2 | GO:0008282 | ATP-sensitive potassium channel complex(GO:0008282) |
| 0.0 | 0.1 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.0 | 0.1 | GO:0042720 | mitochondrial inner membrane peptidase complex(GO:0042720) |
| 0.0 | 0.2 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 0.0 | 0.3 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.0 | 0.1 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.0 | 0.1 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.0 | 0.1 | GO:0031074 | nucleocytoplasmic shuttling complex(GO:0031074) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0008281 | sulfonylurea receptor activity(GO:0008281) |
| 0.1 | 0.3 | GO:0047894 | flavonol 3-sulfotransferase activity(GO:0047894) |
| 0.0 | 0.2 | GO:0004699 | calcium-independent protein kinase C activity(GO:0004699) |
| 0.0 | 0.1 | GO:0047006 | 17-alpha,20-alpha-dihydroxypregn-4-en-3-one dehydrogenase activity(GO:0047006) |
| 0.0 | 0.1 | GO:0033858 | N-acetylgalactosamine kinase activity(GO:0033858) |
| 0.0 | 0.2 | GO:0004705 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.0 | 0.2 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.0 | 0.1 | GO:0047023 | androsterone dehydrogenase activity(GO:0047023) |
| 0.0 | 0.1 | GO:0034041 | sterol-transporting ATPase activity(GO:0034041) |
| 0.0 | 0.2 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 0.0 | 0.1 | GO:0005275 | amine transmembrane transporter activity(GO:0005275) |
| 0.0 | 0.1 | GO:0046848 | hydroxyapatite binding(GO:0046848) |
| 0.0 | 0.1 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
| 0.0 | 0.2 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.0 | 0.1 | GO:0032217 | riboflavin transporter activity(GO:0032217) |
| 0.0 | 0.1 | GO:0009378 | four-way junction helicase activity(GO:0009378) |
| 0.0 | 0.5 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 0.1 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) fructose-2,6-bisphosphate 2-phosphatase activity(GO:0004331) |
| 0.0 | 0.5 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.0 | 0.1 | GO:0004910 | interleukin-1, Type II, blocking receptor activity(GO:0004910) |
| 0.0 | 0.2 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.2 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 0.0 | 0.3 | GO:0005549 | odorant binding(GO:0005549) |
| 0.0 | 0.0 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.1 | GO:0061714 | folic acid receptor activity(GO:0061714) |
| 0.0 | 0.1 | GO:0019534 | toxin transporter activity(GO:0019534) |
| 0.0 | 0.0 | GO:0015173 | aromatic amino acid transmembrane transporter activity(GO:0015173) |
| 0.0 | 0.3 | GO:0016500 | protein-hormone receptor activity(GO:0016500) |
| 0.0 | 0.1 | GO:0016454 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.0 | 0.0 | GO:1904455 | ubiquitin-specific protease activity involved in negative regulation of ERAD pathway(GO:1904455) |
| 0.0 | 0.3 | GO:0031005 | filamin binding(GO:0031005) |
| 0.0 | 0.1 | GO:0034711 | inhibin binding(GO:0034711) |
| 0.0 | 0.0 | GO:0004829 | threonine-tRNA ligase activity(GO:0004829) |
| 0.0 | 0.0 | GO:0018636 | phenanthrene 9,10-monooxygenase activity(GO:0018636) ketosteroid monooxygenase activity(GO:0047086) |
| 0.0 | 0.1 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.0 | 0.0 | GO:0004657 | proline dehydrogenase activity(GO:0004657) |
| 0.0 | 0.0 | GO:0003918 | DNA topoisomerase type II (ATP-hydrolyzing) activity(GO:0003918) DNA topoisomerase II activity(GO:0061505) |
| 0.0 | 0.0 | GO:0004802 | transketolase activity(GO:0004802) |
| 0.0 | 0.2 | GO:0015125 | bile acid transmembrane transporter activity(GO:0015125) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.0 | 0.2 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.0 | 0.2 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.0 | 0.3 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |