Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
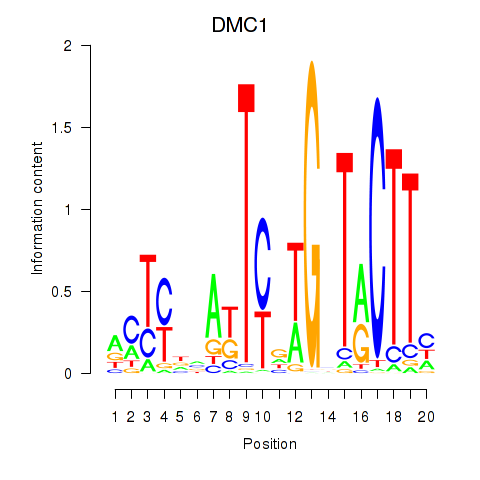
Results for DMC1
Z-value: 0.65
Transcription factors associated with DMC1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
DMC1
|
ENSG00000100206.5 | DMC1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| DMC1 | hg19_v2_chr22_-_38966172_38966291 | 0.43 | 2.9e-01 | Click! |
Activity profile of DMC1 motif
Sorted Z-values of DMC1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of DMC1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_-_8803128 | 1.15 |
ENST00000543467.1 |
MFAP5 |
microfibrillar associated protein 5 |
| chr14_+_21525981 | 0.54 |
ENST00000308227.2 |
RNASE8 |
ribonuclease, RNase A family, 8 |
| chr21_-_46348626 | 0.47 |
ENST00000517563.1 |
ITGB2 |
integrin, beta 2 (complement component 3 receptor 3 and 4 subunit) |
| chr2_-_31637560 | 0.39 |
ENST00000379416.3 |
XDH |
xanthine dehydrogenase |
| chr16_-_30122717 | 0.30 |
ENST00000566613.1 |
GDPD3 |
glycerophosphodiester phosphodiesterase domain containing 3 |
| chr2_-_75788038 | 0.29 |
ENST00000393913.3 ENST00000410113.1 |
EVA1A |
eva-1 homolog A (C. elegans) |
| chr7_-_22539771 | 0.29 |
ENST00000406890.2 ENST00000424363.1 |
STEAP1B |
STEAP family member 1B |
| chr5_-_82969405 | 0.27 |
ENST00000510978.1 |
HAPLN1 |
hyaluronan and proteoglycan link protein 1 |
| chr12_-_71031220 | 0.23 |
ENST00000334414.6 |
PTPRB |
protein tyrosine phosphatase, receptor type, B |
| chr4_-_76944621 | 0.22 |
ENST00000306602.1 |
CXCL10 |
chemokine (C-X-C motif) ligand 10 |
| chr12_-_71031185 | 0.22 |
ENST00000548122.1 ENST00000551525.1 ENST00000550358.1 |
PTPRB |
protein tyrosine phosphatase, receptor type, B |
| chr6_+_116782527 | 0.20 |
ENST00000368606.3 ENST00000368605.1 |
FAM26F |
family with sequence similarity 26, member F |
| chr7_-_150884332 | 0.18 |
ENST00000275838.1 ENST00000422024.1 ENST00000434669.1 |
ASB10 |
ankyrin repeat and SOCS box containing 10 |
| chr10_-_98031265 | 0.17 |
ENST00000224337.5 ENST00000371176.2 |
BLNK |
B-cell linker |
| chr17_+_38333263 | 0.16 |
ENST00000456989.2 ENST00000543876.1 ENST00000544503.1 ENST00000264644.6 ENST00000538884.1 |
RAPGEFL1 |
Rap guanine nucleotide exchange factor (GEF)-like 1 |
| chr10_-_98031310 | 0.15 |
ENST00000427367.2 ENST00000413476.2 |
BLNK |
B-cell linker |
| chr21_-_46348694 | 0.14 |
ENST00000355153.4 ENST00000397850.2 |
ITGB2 |
integrin, beta 2 (complement component 3 receptor 3 and 4 subunit) |
| chr11_+_71846764 | 0.13 |
ENST00000456237.1 ENST00000442948.2 ENST00000546166.1 |
FOLR3 |
folate receptor 3 (gamma) |
| chrX_-_134305322 | 0.13 |
ENST00000276241.6 ENST00000344129.2 |
CXorf48 |
cancer/testis antigen 55 |
| chr10_-_62332357 | 0.12 |
ENST00000503366.1 |
ANK3 |
ankyrin 3, node of Ranvier (ankyrin G) |
| chr5_+_55149356 | 0.12 |
ENST00000490985.1 ENST00000354961.4 |
IL31RA |
interleukin 31 receptor A |
| chr15_-_49103235 | 0.11 |
ENST00000380950.2 |
CEP152 |
centrosomal protein 152kDa |
| chr11_+_71846748 | 0.10 |
ENST00000445078.2 |
FOLR3 |
folate receptor 3 (gamma) |
| chr14_-_21491477 | 0.10 |
ENST00000298684.5 ENST00000557169.1 ENST00000553563.1 |
NDRG2 |
NDRG family member 2 |
| chr19_+_1041212 | 0.10 |
ENST00000433129.1 |
ABCA7 |
ATP-binding cassette, sub-family A (ABC1), member 7 |
| chr8_-_101734907 | 0.10 |
ENST00000318607.5 ENST00000521865.1 ENST00000520804.1 ENST00000522720.1 ENST00000521067.1 |
PABPC1 |
poly(A) binding protein, cytoplasmic 1 |
| chr6_+_80129989 | 0.09 |
ENST00000429444.1 |
RP1-232L24.3 |
RP1-232L24.3 |
| chrX_+_105855160 | 0.09 |
ENST00000372544.2 ENST00000372548.4 |
CXorf57 |
chromosome X open reading frame 57 |
| chr14_+_74318513 | 0.08 |
ENST00000555228.1 ENST00000555661.1 |
PTGR2 |
prostaglandin reductase 2 |
| chr18_+_56892724 | 0.08 |
ENST00000456142.3 ENST00000530323.1 |
GRP |
gastrin-releasing peptide |
| chrX_-_63450480 | 0.08 |
ENST00000362002.2 |
ASB12 |
ankyrin repeat and SOCS box containing 12 |
| chr17_+_57297807 | 0.08 |
ENST00000284116.4 ENST00000581140.1 ENST00000581276.1 |
GDPD1 |
glycerophosphodiester phosphodiesterase domain containing 1 |
| chr11_-_85780086 | 0.07 |
ENST00000532317.1 ENST00000528256.1 ENST00000526033.1 |
PICALM |
phosphatidylinositol binding clathrin assembly protein |
| chr2_-_163008903 | 0.07 |
ENST00000418842.2 ENST00000375497.3 |
GCG |
glucagon |
| chr14_+_74318611 | 0.07 |
ENST00000555976.1 ENST00000267568.4 |
PTGR2 |
prostaglandin reductase 2 |
| chr1_+_166958346 | 0.07 |
ENST00000367872.4 |
MAEL |
maelstrom spermatogenic transposon silencer |
| chrX_-_49965663 | 0.07 |
ENST00000376056.2 ENST00000376058.2 ENST00000358526.2 |
AKAP4 |
A kinase (PRKA) anchor protein 4 |
| chr10_+_62538089 | 0.07 |
ENST00000519078.2 ENST00000395284.3 ENST00000316629.4 |
CDK1 |
cyclin-dependent kinase 1 |
| chr14_-_21492113 | 0.06 |
ENST00000554094.1 |
NDRG2 |
NDRG family member 2 |
| chr1_+_166958497 | 0.06 |
ENST00000367870.2 |
MAEL |
maelstrom spermatogenic transposon silencer |
| chr19_-_42947121 | 0.06 |
ENST00000601181.1 |
CXCL17 |
chemokine (C-X-C motif) ligand 17 |
| chr1_-_111061797 | 0.05 |
ENST00000369771.2 |
KCNA10 |
potassium voltage-gated channel, shaker-related subfamily, member 10 |
| chrX_+_65382433 | 0.05 |
ENST00000374727.3 |
HEPH |
hephaestin |
| chr6_+_26383404 | 0.05 |
ENST00000416795.2 ENST00000494184.1 |
BTN2A2 |
butyrophilin, subfamily 2, member A2 |
| chr14_-_21492251 | 0.05 |
ENST00000554398.1 |
NDRG2 |
NDRG family member 2 |
| chr12_-_109025849 | 0.05 |
ENST00000228463.6 |
SELPLG |
selectin P ligand |
| chr2_+_61244697 | 0.05 |
ENST00000401576.1 ENST00000295030.5 ENST00000414712.2 |
PEX13 |
peroxisomal biogenesis factor 13 |
| chrX_-_49965009 | 0.05 |
ENST00000437370.1 ENST00000376064.3 ENST00000448865.1 |
AKAP4 |
A kinase (PRKA) anchor protein 4 |
| chr2_+_187454749 | 0.05 |
ENST00000261023.3 ENST00000374907.3 |
ITGAV |
integrin, alpha V |
| chr10_+_62538248 | 0.05 |
ENST00000448257.2 |
CDK1 |
cyclin-dependent kinase 1 |
| chr2_+_120187465 | 0.05 |
ENST00000409826.1 ENST00000417645.1 |
TMEM37 |
transmembrane protein 37 |
| chr3_+_101546827 | 0.05 |
ENST00000461724.1 ENST00000483180.1 ENST00000394054.2 |
NFKBIZ |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, zeta |
| chr5_-_157603408 | 0.05 |
ENST00000522769.1 |
CTC-436K13.5 |
CTC-436K13.5 |
| chr2_-_75426826 | 0.04 |
ENST00000305249.5 |
TACR1 |
tachykinin receptor 1 |
| chr6_+_26383318 | 0.04 |
ENST00000469230.1 ENST00000490025.1 ENST00000356709.4 ENST00000352867.2 ENST00000493275.1 ENST00000472507.1 ENST00000482536.1 ENST00000432533.2 ENST00000482842.1 |
BTN2A2 |
butyrophilin, subfamily 2, member A2 |
| chr7_-_150884873 | 0.04 |
ENST00000377867.3 |
ASB10 |
ankyrin repeat and SOCS box containing 10 |
| chr17_+_31318886 | 0.04 |
ENST00000269053.3 ENST00000394638.1 |
SPACA3 |
sperm acrosome associated 3 |
| chr7_+_150758304 | 0.04 |
ENST00000482950.1 ENST00000463414.1 ENST00000310317.5 |
SLC4A2 |
solute carrier family 4 (anion exchanger), member 2 |
| chr17_+_77019030 | 0.03 |
ENST00000580454.1 |
C1QTNF1 |
C1q and tumor necrosis factor related protein 1 |
| chr2_-_18741882 | 0.03 |
ENST00000381249.3 |
RDH14 |
retinol dehydrogenase 14 (all-trans/9-cis/11-cis) |
| chr1_+_18958008 | 0.03 |
ENST00000420770.2 ENST00000400661.3 |
PAX7 |
paired box 7 |
| chr4_+_15471489 | 0.03 |
ENST00000424120.1 ENST00000413206.1 ENST00000438599.2 ENST00000511544.1 ENST00000512702.1 ENST00000507954.1 ENST00000515124.1 ENST00000503292.1 ENST00000503658.1 |
CC2D2A |
coiled-coil and C2 domain containing 2A |
| chr17_+_77018896 | 0.03 |
ENST00000578229.1 |
C1QTNF1 |
C1q and tumor necrosis factor related protein 1 |
| chr7_-_6010263 | 0.02 |
ENST00000455618.2 ENST00000405415.1 ENST00000404406.1 ENST00000542644.1 |
RSPH10B |
radial spoke head 10 homolog B (Chlamydomonas) |
| chr7_-_15601595 | 0.02 |
ENST00000342526.3 |
AGMO |
alkylglycerol monooxygenase |
| chr7_+_6793740 | 0.02 |
ENST00000403107.1 ENST00000404077.1 ENST00000435395.1 ENST00000418406.1 |
RSPH10B2 |
radial spoke head 10 homolog B2 (Chlamydomonas) |
| chr14_+_22508822 | 0.01 |
ENST00000390448.3 |
TRAV20 |
T cell receptor alpha variable 20 |
| chr3_-_108672609 | 0.01 |
ENST00000393963.3 ENST00000471108.1 |
GUCA1C |
guanylate cyclase activator 1C |
| chr10_+_11047259 | 0.01 |
ENST00000379261.4 ENST00000416382.2 |
CELF2 |
CUGBP, Elav-like family member 2 |
| chr3_-_108672742 | 0.01 |
ENST00000261047.3 |
GUCA1C |
guanylate cyclase activator 1C |
| chr7_+_47694842 | 0.01 |
ENST00000408988.2 |
C7orf65 |
chromosome 7 open reading frame 65 |
| chr1_-_190446759 | 0.01 |
ENST00000367462.3 |
BRINP3 |
bone morphogenetic protein/retinoic acid inducible neural-specific 3 |
| chr11_-_73720276 | 0.01 |
ENST00000348534.4 |
UCP3 |
uncoupling protein 3 (mitochondrial, proton carrier) |
| chr14_-_47351391 | 0.00 |
ENST00000399222.3 |
MDGA2 |
MAM domain containing glycosylphosphatidylinositol anchor 2 |
| chr12_-_9268707 | 0.00 |
ENST00000318602.7 |
A2M |
alpha-2-macroglobulin |
| chr17_+_4846101 | 0.00 |
ENST00000576965.1 |
RNF167 |
ring finger protein 167 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0061760 | antifungal innate immune response(GO:0061760) |
| 0.1 | 0.4 | GO:0046110 | xanthine metabolic process(GO:0046110) |
| 0.0 | 0.3 | GO:0097461 | ferric iron import into cell(GO:0097461) ferric iron import across plasma membrane(GO:0098706) |
| 0.0 | 0.1 | GO:1902995 | regulation of phospholipid efflux(GO:1902994) positive regulation of phospholipid efflux(GO:1902995) |
| 0.0 | 0.1 | GO:0072658 | maintenance of protein location in membrane(GO:0072658) maintenance of protein location in plasma membrane(GO:0072660) positive regulation of membrane depolarization during cardiac muscle cell action potential(GO:1900827) |
| 0.0 | 0.2 | GO:0043950 | positive regulation of cAMP-mediated signaling(GO:0043950) |
| 0.0 | 0.6 | GO:0002523 | leukocyte migration involved in inflammatory response(GO:0002523) |
| 0.0 | 1.2 | GO:0060216 | definitive hemopoiesis(GO:0060216) |
| 0.0 | 0.0 | GO:0050902 | leukocyte adhesive activation(GO:0050902) |
| 0.0 | 0.1 | GO:0014038 | regulation of Schwann cell differentiation(GO:0014038) |
| 0.0 | 0.1 | GO:0035948 | positive regulation of gluconeogenesis by positive regulation of transcription from RNA polymerase II promoter(GO:0035948) |
| 0.0 | 0.0 | GO:0060152 | peroxisome localization(GO:0060151) microtubule-based peroxisome localization(GO:0060152) |
| 0.0 | 0.2 | GO:0015884 | folic acid transport(GO:0015884) |
| 0.0 | 0.1 | GO:1900738 | positive regulation of phospholipase C-activating G-protein coupled receptor signaling pathway(GO:1900738) |
| 0.0 | 0.1 | GO:0098535 | de novo centriole assembly(GO:0098535) |
| 0.0 | 0.2 | GO:0090360 | platelet-derived growth factor production(GO:0090360) regulation of platelet-derived growth factor production(GO:0090361) |
| 0.0 | 0.0 | GO:0045715 | negative regulation of low-density lipoprotein particle receptor biosynthetic process(GO:0045715) |
| 0.0 | 0.1 | GO:2000622 | regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0034688 | integrin alphaM-beta2 complex(GO:0034688) |
| 0.0 | 1.2 | GO:0043205 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.0 | 0.1 | GO:0030849 | autosome(GO:0030849) |
| 0.0 | 0.1 | GO:0097125 | cyclin B1-CDK1 complex(GO:0097125) |
| 0.0 | 0.1 | GO:0098536 | deuterosome(GO:0098536) |
| 0.0 | 0.0 | GO:0034685 | integrin alphav-beta6 complex(GO:0034685) integrin alphav-beta8 complex(GO:0034686) |
| 0.0 | 0.1 | GO:0035686 | sperm fibrous sheath(GO:0035686) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.1 | 0.6 | GO:0030369 | ICAM-3 receptor activity(GO:0030369) |
| 0.0 | 0.2 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 0.0 | 0.4 | GO:0043546 | molybdopterin cofactor binding(GO:0043546) |
| 0.0 | 0.3 | GO:0052851 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 0.0 | 0.1 | GO:0090556 | apolipoprotein A-I receptor activity(GO:0034188) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 0.0 | 0.2 | GO:0047522 | 13-prostaglandin reductase activity(GO:0036132) 15-oxoprostaglandin 13-oxidase activity(GO:0047522) |
| 0.0 | 0.4 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.0 | 0.3 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.0 | 0.2 | GO:0005542 | folic acid binding(GO:0005542) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | PID INTEGRIN CS PATHWAY | Integrin family cell surface interactions |
| 0.0 | 1.2 | PID NOTCH PATHWAY | Notch signaling pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |