Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
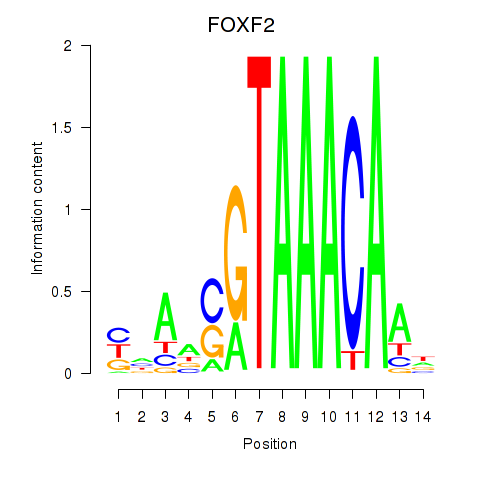
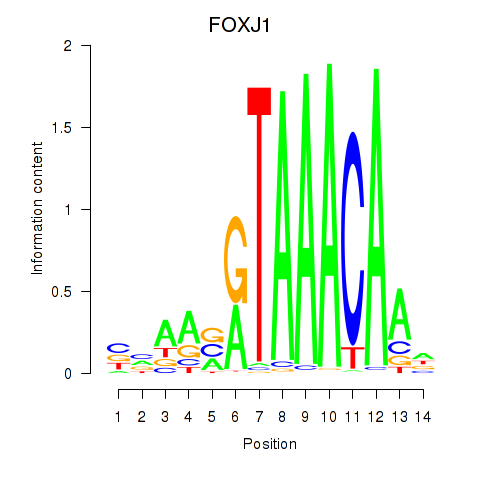
Results for FOXF2_FOXJ1
Z-value: 0.37
Transcription factors associated with FOXF2_FOXJ1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
FOXF2
|
ENSG00000137273.3 | FOXF2 |
|
FOXJ1
|
ENSG00000129654.7 | FOXJ1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| FOXF2 | hg19_v2_chr6_+_1389989_1390069 | 0.47 | 2.4e-01 | Click! |
| FOXJ1 | hg19_v2_chr17_-_74137374_74137385 | 0.31 | 4.5e-01 | Click! |
Activity profile of FOXF2_FOXJ1 motif
Sorted Z-values of FOXF2_FOXJ1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of FOXF2_FOXJ1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_-_19290483 | 0.51 |
ENST00000395592.2 ENST00000299610.4 |
MFAP4 |
microfibrillar-associated protein 4 |
| chr6_+_101846664 | 0.50 |
ENST00000421544.1 ENST00000413795.1 ENST00000369138.1 ENST00000358361.3 |
GRIK2 |
glutamate receptor, ionotropic, kainate 2 |
| chr8_+_97597148 | 0.44 |
ENST00000521590.1 |
SDC2 |
syndecan 2 |
| chr2_+_128175997 | 0.39 |
ENST00000234071.3 ENST00000429925.1 ENST00000442644.1 ENST00000453608.2 |
PROC |
protein C (inactivator of coagulation factors Va and VIIIa) |
| chr11_-_102826434 | 0.38 |
ENST00000340273.4 ENST00000260302.3 |
MMP13 |
matrix metallopeptidase 13 (collagenase 3) |
| chr18_-_56296182 | 0.37 |
ENST00000361673.3 |
ALPK2 |
alpha-kinase 2 |
| chr18_-_52626622 | 0.34 |
ENST00000591504.1 |
CCDC68 |
coiled-coil domain containing 68 |
| chr10_+_70847852 | 0.31 |
ENST00000242465.3 |
SRGN |
serglycin |
| chr6_-_56707943 | 0.29 |
ENST00000370769.4 ENST00000421834.2 ENST00000312431.6 ENST00000361203.3 ENST00000523817.1 |
DST |
dystonin |
| chr1_+_65730385 | 0.26 |
ENST00000263441.7 ENST00000395325.3 |
DNAJC6 |
DnaJ (Hsp40) homolog, subfamily C, member 6 |
| chr5_-_19988339 | 0.26 |
ENST00000382275.1 |
CDH18 |
cadherin 18, type 2 |
| chr5_+_156712372 | 0.25 |
ENST00000541131.1 |
CYFIP2 |
cytoplasmic FMR1 interacting protein 2 |
| chr11_+_71938925 | 0.25 |
ENST00000538751.1 |
INPPL1 |
inositol polyphosphate phosphatase-like 1 |
| chr12_-_92539614 | 0.25 |
ENST00000256015.3 |
BTG1 |
B-cell translocation gene 1, anti-proliferative |
| chr6_+_72922590 | 0.25 |
ENST00000523963.1 |
RIMS1 |
regulating synaptic membrane exocytosis 1 |
| chr6_+_72922505 | 0.24 |
ENST00000401910.3 |
RIMS1 |
regulating synaptic membrane exocytosis 1 |
| chr12_-_71551652 | 0.21 |
ENST00000546561.1 |
TSPAN8 |
tetraspanin 8 |
| chr6_+_114178512 | 0.21 |
ENST00000368635.4 |
MARCKS |
myristoylated alanine-rich protein kinase C substrate |
| chr2_+_33661382 | 0.20 |
ENST00000402538.3 |
RASGRP3 |
RAS guanyl releasing protein 3 (calcium and DAG-regulated) |
| chr6_+_108882069 | 0.20 |
ENST00000406360.1 |
FOXO3 |
forkhead box O3 |
| chr12_-_15038779 | 0.19 |
ENST00000228938.5 ENST00000539261.1 |
MGP |
matrix Gla protein |
| chr18_-_25616519 | 0.19 |
ENST00000399380.3 |
CDH2 |
cadherin 2, type 1, N-cadherin (neuronal) |
| chr2_+_233527443 | 0.19 |
ENST00000410095.1 |
EFHD1 |
EF-hand domain family, member D1 |
| chr10_-_4285923 | 0.18 |
ENST00000418372.1 ENST00000608792.1 |
LINC00702 |
long intergenic non-protein coding RNA 702 |
| chr2_+_12857043 | 0.18 |
ENST00000381465.2 |
TRIB2 |
tribbles pseudokinase 2 |
| chr6_-_131321863 | 0.17 |
ENST00000528282.1 |
EPB41L2 |
erythrocyte membrane protein band 4.1-like 2 |
| chr12_+_59989918 | 0.16 |
ENST00000547379.1 ENST00000549465.1 |
SLC16A7 |
solute carrier family 16 (monocarboxylate transporter), member 7 |
| chr17_+_67410832 | 0.16 |
ENST00000590474.1 |
MAP2K6 |
mitogen-activated protein kinase kinase 6 |
| chr6_+_12717892 | 0.16 |
ENST00000379350.1 |
PHACTR1 |
phosphatase and actin regulator 1 |
| chr9_+_120466610 | 0.15 |
ENST00000394487.4 |
TLR4 |
toll-like receptor 4 |
| chr12_+_79258444 | 0.15 |
ENST00000261205.4 |
SYT1 |
synaptotagmin I |
| chr13_-_39564993 | 0.14 |
ENST00000423210.1 |
STOML3 |
stomatin (EPB72)-like 3 |
| chr9_+_120466650 | 0.14 |
ENST00000355622.6 |
TLR4 |
toll-like receptor 4 |
| chrX_+_85969626 | 0.14 |
ENST00000484479.1 |
DACH2 |
dachshund homolog 2 (Drosophila) |
| chr20_+_44035200 | 0.13 |
ENST00000372717.1 ENST00000360981.4 |
DBNDD2 |
dysbindin (dystrobrevin binding protein 1) domain containing 2 |
| chr12_-_15374343 | 0.12 |
ENST00000256953.2 ENST00000546331.1 |
RERG |
RAS-like, estrogen-regulated, growth inhibitor |
| chr4_-_70626430 | 0.11 |
ENST00000310613.3 |
SULT1B1 |
sulfotransferase family, cytosolic, 1B, member 1 |
| chrX_+_57618269 | 0.11 |
ENST00000374888.1 |
ZXDB |
zinc finger, X-linked, duplicated B |
| chr4_+_71108300 | 0.11 |
ENST00000304954.3 |
CSN3 |
casein kappa |
| chr15_-_31283618 | 0.11 |
ENST00000563714.1 |
MTMR10 |
myotubularin related protein 10 |
| chr2_+_175260451 | 0.11 |
ENST00000458563.1 ENST00000409673.3 ENST00000272732.6 ENST00000435964.1 |
SCRN3 |
secernin 3 |
| chr6_-_25830785 | 0.11 |
ENST00000468082.1 |
SLC17A1 |
solute carrier family 17 (organic anion transporter), member 1 |
| chr8_+_79428539 | 0.11 |
ENST00000352966.5 |
PKIA |
protein kinase (cAMP-dependent, catalytic) inhibitor alpha |
| chr17_-_8263538 | 0.11 |
ENST00000535173.1 |
AC135178.1 |
HCG1985372; Uncharacterized protein; cDNA FLJ37541 fis, clone BRCAN2026340 |
| chr19_-_36523709 | 0.11 |
ENST00000592017.1 ENST00000360535.4 |
CLIP3 |
CAP-GLY domain containing linker protein 3 |
| chr8_-_133772794 | 0.11 |
ENST00000519187.1 ENST00000523829.1 ENST00000356838.3 ENST00000377901.4 ENST00000519304.1 |
TMEM71 |
transmembrane protein 71 |
| chr10_-_103578162 | 0.11 |
ENST00000361464.3 ENST00000357797.5 ENST00000370094.3 |
MGEA5 |
meningioma expressed antigen 5 (hyaluronidase) |
| chr22_-_37505449 | 0.10 |
ENST00000406725.1 |
TMPRSS6 |
transmembrane protease, serine 6 |
| chr20_+_31823792 | 0.10 |
ENST00000375413.4 ENST00000354297.4 ENST00000375422.2 |
BPIFA1 |
BPI fold containing family A, member 1 |
| chr17_+_58755184 | 0.10 |
ENST00000589222.1 ENST00000407086.3 ENST00000390652.5 |
BCAS3 |
breast carcinoma amplified sequence 3 |
| chr22_-_37505588 | 0.10 |
ENST00000406856.1 |
TMPRSS6 |
transmembrane protease, serine 6 |
| chr2_+_175260514 | 0.09 |
ENST00000424069.1 ENST00000427038.1 |
SCRN3 |
secernin 3 |
| chr11_-_27723158 | 0.09 |
ENST00000395980.2 |
BDNF |
brain-derived neurotrophic factor |
| chr12_-_96390108 | 0.09 |
ENST00000538703.1 ENST00000261208.3 |
HAL |
histidine ammonia-lyase |
| chr6_+_84563295 | 0.09 |
ENST00000369687.1 |
RIPPLY2 |
ripply transcriptional repressor 2 |
| chr6_+_112375275 | 0.08 |
ENST00000368666.2 ENST00000604763.1 ENST00000230529.5 |
WISP3 |
WNT1 inducible signaling pathway protein 3 |
| chr12_-_63328817 | 0.08 |
ENST00000228705.6 |
PPM1H |
protein phosphatase, Mg2+/Mn2+ dependent, 1H |
| chr1_+_150122034 | 0.08 |
ENST00000025469.6 ENST00000369124.4 |
PLEKHO1 |
pleckstrin homology domain containing, family O member 1 |
| chr17_-_8059638 | 0.08 |
ENST00000584202.1 ENST00000354903.5 ENST00000577253.1 |
PER1 |
period circadian clock 1 |
| chr21_+_17442799 | 0.08 |
ENST00000602580.1 ENST00000458468.1 ENST00000602935.1 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr21_+_17443434 | 0.08 |
ENST00000400178.2 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr4_-_70626314 | 0.08 |
ENST00000510821.1 |
SULT1B1 |
sulfotransferase family, cytosolic, 1B, member 1 |
| chr3_-_71179988 | 0.08 |
ENST00000491238.1 |
FOXP1 |
forkhead box P1 |
| chr1_+_246729724 | 0.08 |
ENST00000366513.4 ENST00000366512.3 |
CNST |
consortin, connexin sorting protein |
| chr2_+_109223595 | 0.08 |
ENST00000410093.1 |
LIMS1 |
LIM and senescent cell antigen-like domains 1 |
| chr14_-_74551096 | 0.08 |
ENST00000350259.4 |
ALDH6A1 |
aldehyde dehydrogenase 6 family, member A1 |
| chr1_+_22351977 | 0.08 |
ENST00000420503.1 ENST00000416769.1 ENST00000404210.2 |
LINC00339 |
long intergenic non-protein coding RNA 339 |
| chr5_-_42811986 | 0.08 |
ENST00000511224.1 ENST00000507920.1 ENST00000510965.1 |
SEPP1 |
selenoprotein P, plasma, 1 |
| chr12_-_71551868 | 0.07 |
ENST00000247829.3 |
TSPAN8 |
tetraspanin 8 |
| chr14_+_88851874 | 0.07 |
ENST00000393545.4 ENST00000356583.5 ENST00000555401.1 ENST00000553885.1 |
SPATA7 |
spermatogenesis associated 7 |
| chr17_-_26662440 | 0.07 |
ENST00000578122.1 |
IFT20 |
intraflagellar transport 20 homolog (Chlamydomonas) |
| chr8_-_124553437 | 0.07 |
ENST00000517956.1 ENST00000443022.2 |
FBXO32 |
F-box protein 32 |
| chr10_+_94594351 | 0.07 |
ENST00000371552.4 |
EXOC6 |
exocyst complex component 6 |
| chr4_-_152149033 | 0.07 |
ENST00000514152.1 |
SH3D19 |
SH3 domain containing 19 |
| chr9_-_215744 | 0.07 |
ENST00000382387.2 |
C9orf66 |
chromosome 9 open reading frame 66 |
| chr21_+_17443521 | 0.07 |
ENST00000456342.1 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr8_+_55528627 | 0.07 |
ENST00000220676.1 |
RP1 |
retinitis pigmentosa 1 (autosomal dominant) |
| chr1_+_81771806 | 0.07 |
ENST00000370721.1 ENST00000370727.1 ENST00000370725.1 ENST00000370723.1 ENST00000370728.1 ENST00000370730.1 |
LPHN2 |
latrophilin 2 |
| chrX_-_65259914 | 0.07 |
ENST00000374737.4 ENST00000455586.2 |
VSIG4 |
V-set and immunoglobulin domain containing 4 |
| chr10_-_49482907 | 0.07 |
ENST00000374201.3 ENST00000407470.4 |
FRMPD2 |
FERM and PDZ domain containing 2 |
| chr17_+_58677539 | 0.07 |
ENST00000305921.3 |
PPM1D |
protein phosphatase, Mg2+/Mn2+ dependent, 1D |
| chrX_-_65259900 | 0.07 |
ENST00000412866.2 |
VSIG4 |
V-set and immunoglobulin domain containing 4 |
| chr6_-_110964453 | 0.07 |
ENST00000413605.2 |
CDK19 |
cyclin-dependent kinase 19 |
| chr6_-_112575687 | 0.07 |
ENST00000521398.1 ENST00000424408.2 ENST00000243219.3 |
LAMA4 |
laminin, alpha 4 |
| chr5_+_140734570 | 0.07 |
ENST00000571252.1 |
PCDHGA4 |
protocadherin gamma subfamily A, 4 |
| chr3_-_48956818 | 0.06 |
ENST00000408959.2 |
ARIH2OS |
ariadne homolog 2 opposite strand |
| chrX_-_57937067 | 0.06 |
ENST00000358697.4 |
ZXDA |
zinc finger, X-linked, duplicated A |
| chr9_-_135819987 | 0.06 |
ENST00000298552.3 ENST00000403810.1 |
TSC1 |
tuberous sclerosis 1 |
| chr1_-_47131521 | 0.06 |
ENST00000542495.1 ENST00000532925.1 |
ATPAF1 |
ATP synthase mitochondrial F1 complex assembly factor 1 |
| chr15_-_40600026 | 0.06 |
ENST00000456256.2 ENST00000557821.1 |
PLCB2 |
phospholipase C, beta 2 |
| chr3_-_185826855 | 0.06 |
ENST00000306376.5 |
ETV5 |
ets variant 5 |
| chr12_-_96390063 | 0.06 |
ENST00000541929.1 |
HAL |
histidine ammonia-lyase |
| chr3_+_69928256 | 0.06 |
ENST00000394355.2 |
MITF |
microphthalmia-associated transcription factor |
| chr2_+_61404624 | 0.06 |
ENST00000394457.3 |
AHSA2 |
AHA1, activator of heat shock 90kDa protein ATPase homolog 2 (yeast) |
| chr16_-_80926457 | 0.06 |
ENST00000563626.1 ENST00000562231.1 |
RP11-314O13.1 |
RP11-314O13.1 |
| chr7_+_5229819 | 0.06 |
ENST00000288828.4 ENST00000401525.3 ENST00000404704.3 |
WIPI2 |
WD repeat domain, phosphoinositide interacting 2 |
| chr10_+_52751010 | 0.06 |
ENST00000373985.1 |
PRKG1 |
protein kinase, cGMP-dependent, type I |
| chr17_-_26662464 | 0.06 |
ENST00000579419.1 ENST00000585313.1 ENST00000395418.3 ENST00000578985.1 ENST00000577498.1 ENST00000585089.1 ENST00000357896.3 |
IFT20 |
intraflagellar transport 20 homolog (Chlamydomonas) |
| chr11_-_67374177 | 0.06 |
ENST00000333139.3 |
C11orf72 |
chromosome 11 open reading frame 72 |
| chr11_+_120973375 | 0.06 |
ENST00000264037.2 |
TECTA |
tectorin alpha |
| chr6_-_711395 | 0.06 |
ENST00000606285.1 |
RP11-532F6.3 |
RP11-532F6.3 |
| chr9_-_5185629 | 0.05 |
ENST00000381641.3 |
INSL6 |
insulin-like 6 |
| chr14_+_50234827 | 0.05 |
ENST00000554589.1 ENST00000557247.1 |
KLHDC2 |
kelch domain containing 2 |
| chr10_-_103578182 | 0.05 |
ENST00000439817.1 |
MGEA5 |
meningioma expressed antigen 5 (hyaluronidase) |
| chr5_-_36301984 | 0.05 |
ENST00000502994.1 ENST00000515759.1 ENST00000296604.3 |
RANBP3L |
RAN binding protein 3-like |
| chr14_-_74551172 | 0.05 |
ENST00000553458.1 |
ALDH6A1 |
aldehyde dehydrogenase 6 family, member A1 |
| chr3_-_164914640 | 0.05 |
ENST00000241274.3 |
SLITRK3 |
SLIT and NTRK-like family, member 3 |
| chr7_+_151038850 | 0.05 |
ENST00000355851.4 ENST00000566856.1 ENST00000470229.1 |
NUB1 |
negative regulator of ubiquitin-like proteins 1 |
| chr6_+_112375462 | 0.05 |
ENST00000361714.1 |
WISP3 |
WNT1 inducible signaling pathway protein 3 |
| chr2_+_152266604 | 0.05 |
ENST00000430328.2 |
RIF1 |
RAP1 interacting factor homolog (yeast) |
| chr3_+_136676707 | 0.05 |
ENST00000329582.4 |
IL20RB |
interleukin 20 receptor beta |
| chr3_+_136676851 | 0.05 |
ENST00000309741.5 |
IL20RB |
interleukin 20 receptor beta |
| chr12_-_58240470 | 0.05 |
ENST00000548823.1 ENST00000398073.2 |
CTDSP2 |
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase 2 |
| chr19_+_50380682 | 0.05 |
ENST00000221543.5 |
TBC1D17 |
TBC1 domain family, member 17 |
| chr19_-_10446449 | 0.05 |
ENST00000592439.1 |
ICAM3 |
intercellular adhesion molecule 3 |
| chr4_-_82393052 | 0.05 |
ENST00000335927.7 ENST00000504863.1 ENST00000264400.2 |
RASGEF1B |
RasGEF domain family, member 1B |
| chr1_+_236305826 | 0.05 |
ENST00000366592.3 ENST00000366591.4 |
GPR137B |
G protein-coupled receptor 137B |
| chr20_+_44035847 | 0.05 |
ENST00000372712.2 |
DBNDD2 |
dysbindin (dystrobrevin binding protein 1) domain containing 2 |
| chr5_-_59064458 | 0.05 |
ENST00000502575.1 ENST00000507116.1 |
PDE4D |
phosphodiesterase 4D, cAMP-specific |
| chr9_-_28026318 | 0.05 |
ENST00000308675.3 |
LINGO2 |
leucine rich repeat and Ig domain containing 2 |
| chr19_+_50380917 | 0.04 |
ENST00000535102.2 |
TBC1D17 |
TBC1 domain family, member 17 |
| chr2_+_61404545 | 0.04 |
ENST00000357022.2 |
AHSA2 |
AHA1, activator of heat shock 90kDa protein ATPase homolog 2 (yeast) |
| chr6_+_101847105 | 0.04 |
ENST00000369137.3 ENST00000318991.6 |
GRIK2 |
glutamate receptor, ionotropic, kainate 2 |
| chr17_+_56232494 | 0.04 |
ENST00000268912.5 |
OR4D1 |
olfactory receptor, family 4, subfamily D, member 1 |
| chr12_-_56352368 | 0.04 |
ENST00000549404.1 |
PMEL |
premelanosome protein |
| chr10_+_111985713 | 0.04 |
ENST00000239007.7 |
MXI1 |
MAX interactor 1, dimerization protein |
| chr5_+_140797296 | 0.04 |
ENST00000398594.2 |
PCDHGB7 |
protocadherin gamma subfamily B, 7 |
| chr10_-_118032697 | 0.04 |
ENST00000439649.3 |
GFRA1 |
GDNF family receptor alpha 1 |
| chr15_-_50411412 | 0.04 |
ENST00000284509.6 |
ATP8B4 |
ATPase, class I, type 8B, member 4 |
| chr10_-_25010795 | 0.04 |
ENST00000416305.1 ENST00000376410.2 |
ARHGAP21 |
Rho GTPase activating protein 21 |
| chr18_+_3449330 | 0.04 |
ENST00000549253.1 |
TGIF1 |
TGFB-induced factor homeobox 1 |
| chr14_-_101036119 | 0.04 |
ENST00000355173.2 |
BEGAIN |
brain-enriched guanylate kinase-associated |
| chr5_-_142782862 | 0.04 |
ENST00000415690.2 |
NR3C1 |
nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) |
| chr12_+_94071129 | 0.04 |
ENST00000552983.1 ENST00000332896.3 ENST00000552033.1 ENST00000548483.1 |
CRADD |
CASP2 and RIPK1 domain containing adaptor with death domain |
| chr6_-_112575838 | 0.04 |
ENST00000455073.1 |
LAMA4 |
laminin, alpha 4 |
| chr15_-_85197501 | 0.04 |
ENST00000434634.2 |
WDR73 |
WD repeat domain 73 |
| chr16_+_28763108 | 0.04 |
ENST00000357796.3 ENST00000550983.1 |
NPIPB9 |
nuclear pore complex interacting protein family, member B9 |
| chr4_+_100495864 | 0.04 |
ENST00000265517.5 ENST00000422897.2 |
MTTP |
microsomal triglyceride transfer protein |
| chr4_+_88754113 | 0.04 |
ENST00000560249.1 ENST00000540395.1 ENST00000511670.1 ENST00000361056.3 |
MEPE |
matrix extracellular phosphoglycoprotein |
| chr12_-_53045948 | 0.04 |
ENST00000309680.3 |
KRT2 |
keratin 2 |
| chr12_+_40787194 | 0.04 |
ENST00000425730.2 ENST00000454784.4 |
MUC19 |
mucin 19, oligomeric |
| chr7_+_5229904 | 0.03 |
ENST00000382384.2 |
WIPI2 |
WD repeat domain, phosphoinositide interacting 2 |
| chr19_-_12941469 | 0.03 |
ENST00000586969.1 ENST00000589681.1 ENST00000585384.1 ENST00000589808.1 |
RTBDN |
retbindin |
| chr4_+_88754069 | 0.03 |
ENST00000395102.4 ENST00000497649.2 |
MEPE |
matrix extracellular phosphoglycoprotein |
| chr3_+_50712672 | 0.03 |
ENST00000266037.9 |
DOCK3 |
dedicator of cytokinesis 3 |
| chr1_+_43855560 | 0.03 |
ENST00000562955.1 |
SZT2 |
seizure threshold 2 homolog (mouse) |
| chr1_-_152332480 | 0.03 |
ENST00000388718.5 |
FLG2 |
filaggrin family member 2 |
| chr4_-_70826725 | 0.03 |
ENST00000353151.3 |
CSN2 |
casein beta |
| chr11_+_71791359 | 0.03 |
ENST00000419228.1 ENST00000435085.1 ENST00000307198.7 ENST00000538413.1 |
LRTOMT |
leucine rich transmembrane and O-methyltransferase domain containing |
| chr13_-_99910673 | 0.03 |
ENST00000397473.2 ENST00000397470.2 |
GPR18 |
G protein-coupled receptor 18 |
| chr6_-_42016385 | 0.03 |
ENST00000502771.1 ENST00000508143.1 ENST00000514588.1 ENST00000510503.1 ENST00000415497.2 ENST00000372988.4 |
CCND3 |
cyclin D3 |
| chr11_-_85397167 | 0.03 |
ENST00000316398.3 |
CCDC89 |
coiled-coil domain containing 89 |
| chr1_+_92545862 | 0.03 |
ENST00000370382.3 ENST00000342818.3 |
BTBD8 |
BTB (POZ) domain containing 8 |
| chr15_+_72978521 | 0.03 |
ENST00000542334.1 ENST00000268057.4 |
BBS4 |
Bardet-Biedl syndrome 4 |
| chr10_+_124739911 | 0.03 |
ENST00000405485.1 |
PSTK |
phosphoseryl-tRNA kinase |
| chr6_-_112575758 | 0.03 |
ENST00000431543.2 ENST00000453937.2 ENST00000368638.4 ENST00000389463.4 |
LAMA4 |
laminin, alpha 4 |
| chr7_+_129906660 | 0.03 |
ENST00000222481.4 |
CPA2 |
carboxypeptidase A2 (pancreatic) |
| chr19_-_36523529 | 0.03 |
ENST00000593074.1 |
CLIP3 |
CAP-GLY domain containing linker protein 3 |
| chr17_+_16284604 | 0.03 |
ENST00000395839.1 ENST00000395837.1 |
UBB |
ubiquitin B |
| chr7_+_12250886 | 0.03 |
ENST00000444443.1 ENST00000396667.3 |
TMEM106B |
transmembrane protein 106B |
| chr11_-_5526834 | 0.03 |
ENST00000380237.1 ENST00000396895.1 ENST00000380252.1 |
HBE1 HBG2 |
hemoglobin, epsilon 1 hemoglobin, gamma G |
| chr7_-_112579869 | 0.03 |
ENST00000297145.4 |
C7orf60 |
chromosome 7 open reading frame 60 |
| chr3_-_157221128 | 0.03 |
ENST00000392833.2 ENST00000362010.2 |
VEPH1 |
ventricular zone expressed PH domain-containing 1 |
| chr15_+_72978539 | 0.03 |
ENST00000539603.1 ENST00000569338.1 |
BBS4 |
Bardet-Biedl syndrome 4 |
| chr12_+_72080253 | 0.03 |
ENST00000549735.1 |
TMEM19 |
transmembrane protein 19 |
| chr8_-_145754428 | 0.03 |
ENST00000527462.1 ENST00000313465.5 ENST00000524821.1 |
C8orf82 |
chromosome 8 open reading frame 82 |
| chr5_-_142783175 | 0.03 |
ENST00000231509.3 ENST00000394464.2 |
NR3C1 |
nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) |
| chr1_+_185703513 | 0.03 |
ENST00000271588.4 ENST00000367492.2 |
HMCN1 |
hemicentin 1 |
| chr19_-_50380536 | 0.03 |
ENST00000391832.3 ENST00000391834.2 ENST00000344175.5 |
AKT1S1 |
AKT1 substrate 1 (proline-rich) |
| chr1_-_57431679 | 0.03 |
ENST00000371237.4 ENST00000535057.1 ENST00000543257.1 |
C8B |
complement component 8, beta polypeptide |
| chr4_-_147043058 | 0.03 |
ENST00000512063.1 ENST00000507726.1 |
RP11-6L6.3 |
long intergenic non-protein coding RNA 1095 |
| chr16_-_28374829 | 0.02 |
ENST00000532254.1 |
NPIPB6 |
nuclear pore complex interacting protein family, member B6 |
| chr2_-_192711968 | 0.02 |
ENST00000304141.4 |
SDPR |
serum deprivation response |
| chr19_-_18902106 | 0.02 |
ENST00000542601.2 ENST00000425807.1 ENST00000222271.2 |
COMP |
cartilage oligomeric matrix protein |
| chr6_-_32122106 | 0.02 |
ENST00000428778.1 |
PRRT1 |
proline-rich transmembrane protein 1 |
| chr6_-_28973037 | 0.02 |
ENST00000377179.3 |
ZNF311 |
zinc finger protein 311 |
| chr16_+_6069072 | 0.02 |
ENST00000547605.1 ENST00000550418.1 ENST00000553186.1 |
RBFOX1 |
RNA binding protein, fox-1 homolog (C. elegans) 1 |
| chr19_+_41725140 | 0.02 |
ENST00000359092.3 |
AXL |
AXL receptor tyrosine kinase |
| chr21_-_19775973 | 0.02 |
ENST00000284885.3 |
TMPRSS15 |
transmembrane protease, serine 15 |
| chr16_+_84178874 | 0.02 |
ENST00000378553.5 |
DNAAF1 |
dynein, axonemal, assembly factor 1 |
| chr5_-_159846399 | 0.02 |
ENST00000297151.4 |
SLU7 |
SLU7 splicing factor homolog (S. cerevisiae) |
| chr2_-_175260368 | 0.02 |
ENST00000342016.3 ENST00000362053.5 |
CIR1 |
corepressor interacting with RBPJ, 1 |
| chrX_-_47863348 | 0.02 |
ENST00000376943.3 ENST00000396965.1 ENST00000305127.6 |
ZNF182 |
zinc finger protein 182 |
| chr2_+_220042933 | 0.02 |
ENST00000430297.2 |
FAM134A |
family with sequence similarity 134, member A |
| chr6_-_76203454 | 0.02 |
ENST00000237172.7 |
FILIP1 |
filamin A interacting protein 1 |
| chr5_-_110062349 | 0.02 |
ENST00000511883.2 ENST00000455884.2 |
TMEM232 |
transmembrane protein 232 |
| chr6_-_112575912 | 0.02 |
ENST00000522006.1 ENST00000230538.7 ENST00000519932.1 |
LAMA4 |
laminin, alpha 4 |
| chr16_-_84178728 | 0.02 |
ENST00000562224.1 ENST00000434463.3 ENST00000564998.1 ENST00000219439.4 |
HSDL1 |
hydroxysteroid dehydrogenase like 1 |
| chr19_-_47157914 | 0.02 |
ENST00000300875.4 |
DACT3 |
dishevelled-binding antagonist of beta-catenin 3 |
| chr15_+_42697018 | 0.02 |
ENST00000397204.4 |
CAPN3 |
calpain 3, (p94) |
| chr1_-_43855479 | 0.02 |
ENST00000290663.6 ENST00000372457.4 |
MED8 |
mediator complex subunit 8 |
| chr20_+_44420617 | 0.02 |
ENST00000449078.1 ENST00000456939.1 |
DNTTIP1 |
deoxynucleotidyltransferase, terminal, interacting protein 1 |
| chr4_-_90757364 | 0.02 |
ENST00000508895.1 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr1_-_149982624 | 0.02 |
ENST00000417191.1 ENST00000369135.4 |
OTUD7B |
OTU domain containing 7B |
| chr8_-_72459885 | 0.02 |
ENST00000523987.1 |
RP11-1102P16.1 |
Uncharacterized protein |
| chr6_-_161085291 | 0.02 |
ENST00000316300.5 |
LPA |
lipoprotein, Lp(a) |
| chr8_+_67344710 | 0.02 |
ENST00000379385.4 ENST00000396623.3 ENST00000415254.1 |
ADHFE1 |
alcohol dehydrogenase, iron containing, 1 |
| chr3_-_71632894 | 0.02 |
ENST00000493089.1 |
FOXP1 |
forkhead box P1 |
| chr13_+_103459704 | 0.02 |
ENST00000602836.1 |
BIVM-ERCC5 |
BIVM-ERCC5 readthrough |
| chr15_-_34610962 | 0.02 |
ENST00000290209.5 |
SLC12A6 |
solute carrier family 12 (potassium/chloride transporter), member 6 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0044537 | regulation of circulating fibrinogen levels(GO:0044537) |
| 0.1 | 0.3 | GO:0033364 | mast cell secretory granule organization(GO:0033364) |
| 0.1 | 0.3 | GO:0070428 | negative regulation of interleukin-23 production(GO:0032707) regulation of nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070428) |
| 0.1 | 0.4 | GO:0008218 | bioluminescence(GO:0008218) |
| 0.1 | 0.2 | GO:0001544 | initiation of primordial ovarian follicle growth(GO:0001544) |
| 0.1 | 0.2 | GO:0072709 | cellular response to sorbitol(GO:0072709) |
| 0.0 | 0.1 | GO:0098746 | fast, calcium ion-dependent exocytosis of neurotransmitter(GO:0098746) |
| 0.0 | 0.5 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.0 | 0.5 | GO:0051967 | negative regulation of synaptic transmission, glutamatergic(GO:0051967) |
| 0.0 | 0.2 | GO:0045074 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 0.0 | 0.1 | GO:0006208 | pyrimidine nucleobase catabolic process(GO:0006208) thymine catabolic process(GO:0006210) thymine metabolic process(GO:0019859) |
| 0.0 | 0.2 | GO:2000809 | positive regulation of synaptic vesicle clustering(GO:2000809) |
| 0.0 | 0.1 | GO:0031548 | regulation of brain-derived neurotrophic factor receptor signaling pathway(GO:0031548) |
| 0.0 | 0.1 | GO:0043606 | histidine catabolic process to glutamate and formamide(GO:0019556) histidine catabolic process to glutamate and formate(GO:0019557) formamide metabolic process(GO:0043606) |
| 0.0 | 0.3 | GO:0016191 | synaptic vesicle uncoating(GO:0016191) |
| 0.0 | 0.5 | GO:0097151 | positive regulation of inhibitory postsynaptic potential(GO:0097151) modulation of inhibitory postsynaptic potential(GO:0098828) |
| 0.0 | 0.3 | GO:0008090 | retrograde axonal transport(GO:0008090) |
| 0.0 | 0.1 | GO:0044010 | biofilm formation(GO:0042710) single-species biofilm formation(GO:0044010) single-species biofilm formation in or on host organism(GO:0044407) regulation of single-species biofilm formation(GO:1900190) negative regulation of single-species biofilm formation(GO:1900191) regulation of single-species biofilm formation in or on host organism(GO:1900228) negative regulation of single-species biofilm formation in or on host organism(GO:1900229) |
| 0.0 | 0.3 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.0 | 0.1 | GO:0001808 | negative regulation of type IV hypersensitivity(GO:0001808) |
| 0.0 | 0.3 | GO:0035845 | photoreceptor cell outer segment organization(GO:0035845) |
| 0.0 | 0.2 | GO:1901475 | pyruvate transport(GO:0006848) pyruvate transmembrane transport(GO:1901475) |
| 0.0 | 0.1 | GO:0032625 | interleukin-21 production(GO:0032625) interleukin-21 secretion(GO:0072619) |
| 0.0 | 0.4 | GO:0003417 | growth plate cartilage development(GO:0003417) |
| 0.0 | 0.1 | GO:0018231 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.0 | 0.1 | GO:0061739 | protein lipidation involved in autophagosome assembly(GO:0061739) |
| 0.0 | 0.2 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.0 | 0.1 | GO:0051029 | rRNA transport(GO:0051029) |
| 0.0 | 0.1 | GO:0033615 | mitochondrial proton-transporting ATP synthase complex assembly(GO:0033615) |
| 0.0 | 0.1 | GO:0098728 | germ-line stem cell division(GO:0042078) male germ-line stem cell asymmetric division(GO:0048133) germline stem cell asymmetric division(GO:0098728) |
| 0.0 | 0.2 | GO:0045603 | positive regulation of endothelial cell differentiation(GO:0045603) |
| 0.0 | 0.1 | GO:2000323 | negative regulation of glucocorticoid receptor signaling pathway(GO:2000323) |
| 0.0 | 0.2 | GO:1904776 | regulation of protein localization to cell cortex(GO:1904776) positive regulation of protein localization to cell cortex(GO:1904778) |
| 0.0 | 0.0 | GO:1903487 | regulation of lactation(GO:1903487) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0031673 | H zone(GO:0031673) |
| 0.1 | 0.2 | GO:0042585 | germinal vesicle(GO:0042585) |
| 0.1 | 0.3 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 0.1 | 0.5 | GO:0071953 | elastic fiber(GO:0071953) |
| 0.0 | 0.1 | GO:1902636 | kinociliary basal body(GO:1902636) |
| 0.0 | 0.1 | GO:0060201 | clathrin-sculpted acetylcholine transport vesicle(GO:0060200) clathrin-sculpted acetylcholine transport vesicle membrane(GO:0060201) |
| 0.0 | 0.1 | GO:0036195 | muscle cell projection(GO:0036194) muscle cell projection membrane(GO:0036195) |
| 0.0 | 0.1 | GO:0033596 | TSC1-TSC2 complex(GO:0033596) |
| 0.0 | 0.5 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.0 | 0.3 | GO:0042588 | zymogen granule(GO:0042588) |
| 0.0 | 0.2 | GO:0008091 | spectrin(GO:0008091) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0015277 | kainate selective glutamate receptor activity(GO:0015277) |
| 0.0 | 0.2 | GO:0005477 | pyruvate secondary active transmembrane transporter activity(GO:0005477) |
| 0.0 | 0.1 | GO:0010698 | acetyltransferase activator activity(GO:0010698) |
| 0.0 | 0.1 | GO:0005169 | neurotrophin TRKB receptor binding(GO:0005169) |
| 0.0 | 0.1 | GO:0002046 | opsin binding(GO:0002046) |
| 0.0 | 0.1 | GO:0030348 | syntaxin-3 binding(GO:0030348) |
| 0.0 | 0.2 | GO:0034485 | phosphatidylinositol-3,4,5-trisphosphate 5-phosphatase activity(GO:0034485) |
| 0.0 | 0.1 | GO:0097001 | ceramide binding(GO:0097001) |
| 0.0 | 0.1 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 0.0 | 0.2 | GO:0001875 | lipopolysaccharide receptor activity(GO:0001875) |
| 0.0 | 0.2 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.0 | 0.2 | GO:0042731 | PH domain binding(GO:0042731) |
| 0.0 | 0.1 | GO:0004883 | glucocorticoid receptor activity(GO:0004883) glucocorticoid-activated RNA polymerase II transcription factor binding transcription factor activity(GO:0038051) |
| 0.0 | 0.1 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.0 | 0.1 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.0 | 0.1 | GO:0004920 | interleukin-10 receptor activity(GO:0004920) |
| 0.0 | 0.3 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 0.4 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.0 | 0.3 | REACTOME IKK COMPLEX RECRUITMENT MEDIATED BY RIP1 | Genes involved in IKK complex recruitment mediated by RIP1 |
| 0.0 | 0.5 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.0 | 0.2 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.0 | 0.4 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.0 | 0.4 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |