Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
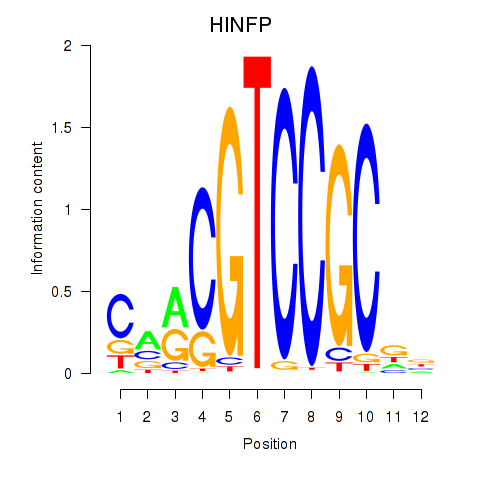
Results for HINFP
Z-value: 2.35
Transcription factors associated with HINFP
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HINFP
|
ENSG00000172273.8 | HINFP |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HINFP | hg19_v2_chr11_+_118992269_118992334 | 0.21 | 6.1e-01 | Click! |
Activity profile of HINFP motif
Sorted Z-values of HINFP motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HINFP
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_+_94023873 | 4.32 |
ENST00000297268.6 |
COL1A2 |
collagen, type I, alpha 2 |
| chr6_-_139695757 | 1.71 |
ENST00000367651.2 |
CITED2 |
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 |
| chr3_+_110790590 | 1.71 |
ENST00000485303.1 |
PVRL3 |
poliovirus receptor-related 3 |
| chr12_+_90102729 | 1.69 |
ENST00000605386.1 |
LINC00936 |
long intergenic non-protein coding RNA 936 |
| chr12_+_65672423 | 1.57 |
ENST00000355192.3 ENST00000308259.5 ENST00000540804.1 ENST00000535664.1 ENST00000541189.1 |
MSRB3 |
methionine sulfoxide reductase B3 |
| chr1_-_92351769 | 1.43 |
ENST00000212355.4 |
TGFBR3 |
transforming growth factor, beta receptor III |
| chr5_-_136834982 | 1.36 |
ENST00000510689.1 ENST00000394945.1 |
SPOCK1 |
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 1 |
| chr10_-_62761188 | 1.26 |
ENST00000357917.4 |
RHOBTB1 |
Rho-related BTB domain containing 1 |
| chr3_+_49449636 | 1.25 |
ENST00000273590.3 |
TCTA |
T-cell leukemia translocation altered |
| chr6_+_56820018 | 1.22 |
ENST00000370746.3 |
BEND6 |
BEN domain containing 6 |
| chr17_-_46115122 | 1.19 |
ENST00000006101.4 |
COPZ2 |
coatomer protein complex, subunit zeta 2 |
| chr2_+_120770581 | 1.17 |
ENST00000263713.5 |
EPB41L5 |
erythrocyte membrane protein band 4.1 like 5 |
| chr6_+_56819773 | 1.17 |
ENST00000370750.2 |
BEND6 |
BEN domain containing 6 |
| chr4_+_123747834 | 1.13 |
ENST00000264498.3 |
FGF2 |
fibroblast growth factor 2 (basic) |
| chr9_-_89562104 | 1.10 |
ENST00000298743.7 |
GAS1 |
growth arrest-specific 1 |
| chr11_+_62104897 | 1.06 |
ENST00000415229.2 ENST00000535727.1 ENST00000301776.5 |
ASRGL1 |
asparaginase like 1 |
| chr12_+_65672702 | 1.05 |
ENST00000538045.1 ENST00000535239.1 |
MSRB3 |
methionine sulfoxide reductase B3 |
| chr5_-_41510656 | 1.03 |
ENST00000377801.3 |
PLCXD3 |
phosphatidylinositol-specific phospholipase C, X domain containing 3 |
| chr5_-_41510725 | 1.01 |
ENST00000328457.3 |
PLCXD3 |
phosphatidylinositol-specific phospholipase C, X domain containing 3 |
| chr12_+_12764773 | 0.90 |
ENST00000228865.2 |
CREBL2 |
cAMP responsive element binding protein-like 2 |
| chr1_+_109102652 | 0.90 |
ENST00000370035.3 ENST00000405454.1 |
FAM102B |
family with sequence similarity 102, member B |
| chr4_+_123747979 | 0.86 |
ENST00000608478.1 |
FGF2 |
fibroblast growth factor 2 (basic) |
| chr8_-_27462822 | 0.86 |
ENST00000522098.1 |
CLU |
clusterin |
| chr11_+_64879317 | 0.86 |
ENST00000526809.1 ENST00000279263.7 ENST00000524986.1 ENST00000534371.1 ENST00000540748.1 ENST00000525385.1 ENST00000345348.5 ENST00000531321.1 ENST00000529414.1 ENST00000526085.1 ENST00000530750.1 |
TM7SF2 |
transmembrane 7 superfamily member 2 |
| chr5_-_158526693 | 0.85 |
ENST00000380654.4 |
EBF1 |
early B-cell factor 1 |
| chr5_-_158526756 | 0.80 |
ENST00000313708.6 ENST00000517373.1 |
EBF1 |
early B-cell factor 1 |
| chr16_-_54963026 | 0.79 |
ENST00000560208.1 ENST00000557792.1 |
CRNDE |
colorectal neoplasia differentially expressed (non-protein coding) |
| chr17_-_66453562 | 0.73 |
ENST00000262139.5 ENST00000546360.1 |
WIPI1 |
WD repeat domain, phosphoinositide interacting 1 |
| chr4_+_148402069 | 0.72 |
ENST00000358556.4 ENST00000339690.5 ENST00000511804.1 ENST00000324300.5 |
EDNRA |
endothelin receptor type A |
| chr15_+_43803143 | 0.71 |
ENST00000382031.1 |
MAP1A |
microtubule-associated protein 1A |
| chr2_+_10442993 | 0.69 |
ENST00000423674.1 ENST00000307845.3 |
HPCAL1 |
hippocalcin-like 1 |
| chr11_-_66336060 | 0.66 |
ENST00000310325.5 |
CTSF |
cathepsin F |
| chr9_+_37650945 | 0.65 |
ENST00000377765.3 |
FRMPD1 |
FERM and PDZ domain containing 1 |
| chr20_-_30458019 | 0.62 |
ENST00000486996.1 ENST00000398084.2 |
DUSP15 |
dual specificity phosphatase 15 |
| chr20_-_30458491 | 0.61 |
ENST00000339738.5 |
DUSP15 |
dual specificity phosphatase 15 |
| chr22_-_28197486 | 0.59 |
ENST00000302326.4 |
MN1 |
meningioma (disrupted in balanced translocation) 1 |
| chr17_-_42276574 | 0.57 |
ENST00000589805.1 |
ATXN7L3 |
ataxin 7-like 3 |
| chr12_-_90102594 | 0.55 |
ENST00000428670.3 |
ATP2B1 |
ATPase, Ca++ transporting, plasma membrane 1 |
| chr2_-_175547571 | 0.55 |
ENST00000409415.3 ENST00000359761.3 ENST00000272746.5 |
WIPF1 |
WAS/WASL interacting protein family, member 1 |
| chr6_+_24495067 | 0.55 |
ENST00000357578.3 ENST00000546278.1 ENST00000491546.1 |
ALDH5A1 |
aldehyde dehydrogenase 5 family, member A1 |
| chr1_+_61547405 | 0.54 |
ENST00000371189.4 |
NFIA |
nuclear factor I/A |
| chr3_-_16555150 | 0.52 |
ENST00000334133.4 |
RFTN1 |
raftlin, lipid raft linker 1 |
| chr2_+_120770645 | 0.51 |
ENST00000443902.2 |
EPB41L5 |
erythrocyte membrane protein band 4.1 like 5 |
| chr17_-_42277203 | 0.50 |
ENST00000587097.1 |
ATXN7L3 |
ataxin 7-like 3 |
| chr2_-_233352531 | 0.49 |
ENST00000304546.1 |
ECEL1 |
endothelin converting enzyme-like 1 |
| chr17_-_73178599 | 0.49 |
ENST00000578238.1 |
SUMO2 |
small ubiquitin-like modifier 2 |
| chr16_-_30006922 | 0.49 |
ENST00000564026.1 |
HIRIP3 |
HIRA interacting protein 3 |
| chr13_-_77460525 | 0.48 |
ENST00000377474.2 ENST00000317765.2 |
KCTD12 |
potassium channel tetramerization domain containing 12 |
| chr6_+_56819895 | 0.48 |
ENST00000370748.3 |
BEND6 |
BEN domain containing 6 |
| chrX_+_21392553 | 0.46 |
ENST00000279451.4 |
CNKSR2 |
connector enhancer of kinase suppressor of Ras 2 |
| chr1_+_22351977 | 0.44 |
ENST00000420503.1 ENST00000416769.1 ENST00000404210.2 |
LINC00339 |
long intergenic non-protein coding RNA 339 |
| chr20_+_42086525 | 0.44 |
ENST00000244020.3 |
SRSF6 |
serine/arginine-rich splicing factor 6 |
| chr14_+_52118576 | 0.44 |
ENST00000395718.2 ENST00000344768.5 |
FRMD6 |
FERM domain containing 6 |
| chr15_+_91643442 | 0.43 |
ENST00000394232.1 |
SV2B |
synaptic vesicle glycoprotein 2B |
| chr5_+_72143988 | 0.42 |
ENST00000506351.2 |
TNPO1 |
transportin 1 |
| chr17_-_46690839 | 0.42 |
ENST00000498634.2 |
HOXB8 |
homeobox B8 |
| chr16_-_54962704 | 0.42 |
ENST00000502066.2 ENST00000560912.1 ENST00000558952.1 |
CRNDE |
colorectal neoplasia differentially expressed (non-protein coding) |
| chr17_+_7341586 | 0.41 |
ENST00000575235.1 |
FGF11 |
fibroblast growth factor 11 |
| chr12_+_107349497 | 0.41 |
ENST00000548125.1 ENST00000280756.4 |
C12orf23 |
chromosome 12 open reading frame 23 |
| chr11_+_66045634 | 0.41 |
ENST00000528852.1 ENST00000311445.6 |
CNIH2 |
cornichon family AMPA receptor auxiliary protein 2 |
| chr10_+_124907638 | 0.41 |
ENST00000339992.3 |
HMX2 |
H6 family homeobox 2 |
| chr17_-_1419878 | 0.39 |
ENST00000449479.1 ENST00000477910.1 ENST00000542125.1 ENST00000575172.1 |
INPP5K |
inositol polyphosphate-5-phosphatase K |
| chr20_+_30946106 | 0.39 |
ENST00000375687.4 ENST00000542461.1 |
ASXL1 |
additional sex combs like 1 (Drosophila) |
| chr11_-_73309228 | 0.39 |
ENST00000356467.4 ENST00000064778.4 |
FAM168A |
family with sequence similarity 168, member A |
| chr2_-_232791038 | 0.39 |
ENST00000295440.2 ENST00000409852.1 |
NPPC |
natriuretic peptide C |
| chr17_+_80186908 | 0.38 |
ENST00000582743.1 ENST00000578684.1 ENST00000577650.1 ENST00000582715.1 |
SLC16A3 |
solute carrier family 16 (monocarboxylate transporter), member 3 |
| chr20_-_39946237 | 0.38 |
ENST00000441102.2 ENST00000559234.1 |
ZHX3 |
zinc fingers and homeoboxes 3 |
| chrX_-_118827113 | 0.38 |
ENST00000394617.2 |
SEPT6 |
septin 6 |
| chr2_-_43453734 | 0.38 |
ENST00000282388.3 |
ZFP36L2 |
ZFP36 ring finger protein-like 2 |
| chr10_+_94608245 | 0.38 |
ENST00000443748.2 ENST00000260762.6 |
EXOC6 |
exocyst complex component 6 |
| chr18_-_44336998 | 0.37 |
ENST00000315087.7 |
ST8SIA5 |
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 5 |
| chr13_+_21714653 | 0.37 |
ENST00000382533.4 |
SAP18 |
Sin3A-associated protein, 18kDa |
| chr17_-_1419914 | 0.37 |
ENST00000397335.3 ENST00000574561.1 |
INPP5K |
inositol polyphosphate-5-phosphatase K |
| chr16_-_54962415 | 0.36 |
ENST00000501177.3 ENST00000559598.2 |
CRNDE |
colorectal neoplasia differentially expressed (non-protein coding) |
| chr11_-_2162162 | 0.36 |
ENST00000381389.1 |
IGF2 |
insulin-like growth factor 2 (somatomedin A) |
| chr17_-_1420182 | 0.36 |
ENST00000421807.2 |
INPP5K |
inositol polyphosphate-5-phosphatase K |
| chr11_-_2162468 | 0.36 |
ENST00000434045.2 |
IGF2 |
insulin-like growth factor 2 (somatomedin A) |
| chr17_-_1420006 | 0.36 |
ENST00000320345.6 ENST00000406424.4 |
INPP5K |
inositol polyphosphate-5-phosphatase K |
| chr16_-_686235 | 0.36 |
ENST00000568773.1 ENST00000565163.1 ENST00000397665.2 ENST00000397666.2 ENST00000301686.8 ENST00000338401.4 ENST00000397664.4 ENST00000568830.1 |
C16orf13 |
chromosome 16 open reading frame 13 |
| chr19_-_14016877 | 0.35 |
ENST00000454313.1 ENST00000591586.1 ENST00000346736.2 |
C19orf57 |
chromosome 19 open reading frame 57 |
| chr16_-_54962625 | 0.35 |
ENST00000559432.1 |
CRNDE |
colorectal neoplasia differentially expressed (non-protein coding) |
| chr4_-_107237374 | 0.35 |
ENST00000361687.4 ENST00000507696.1 ENST00000394708.2 ENST00000509532.1 |
TBCK |
TBC1 domain containing kinase |
| chr18_-_268019 | 0.35 |
ENST00000261600.6 |
THOC1 |
THO complex 1 |
| chr13_+_21714711 | 0.34 |
ENST00000607003.1 ENST00000492245.1 |
SAP18 |
Sin3A-associated protein, 18kDa |
| chr5_-_142782862 | 0.34 |
ENST00000415690.2 |
NR3C1 |
nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) |
| chr1_-_1509931 | 0.33 |
ENST00000359060.4 |
SSU72 |
SSU72 RNA polymerase II CTD phosphatase homolog (S. cerevisiae) |
| chr22_-_24110063 | 0.33 |
ENST00000520222.1 ENST00000401675.3 |
CHCHD10 |
coiled-coil-helix-coiled-coil-helix domain containing 10 |
| chr3_-_49449521 | 0.33 |
ENST00000431929.1 ENST00000418115.1 |
RHOA |
ras homolog family member A |
| chr6_-_111804905 | 0.33 |
ENST00000358835.3 ENST00000435970.1 |
REV3L |
REV3-like, polymerase (DNA directed), zeta, catalytic subunit |
| chr6_-_35464817 | 0.32 |
ENST00000338863.7 |
TEAD3 |
TEA domain family member 3 |
| chr4_-_107237340 | 0.32 |
ENST00000394706.3 |
TBCK |
TBC1 domain containing kinase |
| chr11_-_6440624 | 0.32 |
ENST00000311051.3 |
APBB1 |
amyloid beta (A4) precursor protein-binding, family B, member 1 (Fe65) |
| chr11_-_6440283 | 0.32 |
ENST00000299402.6 ENST00000609360.1 ENST00000389906.2 ENST00000532020.2 |
APBB1 |
amyloid beta (A4) precursor protein-binding, family B, member 1 (Fe65) |
| chr5_-_141030943 | 0.31 |
ENST00000522783.1 ENST00000519800.1 ENST00000435817.2 |
FCHSD1 |
FCH and double SH3 domains 1 |
| chr3_-_52719810 | 0.30 |
ENST00000424867.1 ENST00000394830.3 ENST00000431678.1 ENST00000450271.1 |
PBRM1 |
polybromo 1 |
| chr6_-_35464727 | 0.30 |
ENST00000402886.3 |
TEAD3 |
TEA domain family member 3 |
| chr9_+_100174344 | 0.30 |
ENST00000422139.2 |
TDRD7 |
tudor domain containing 7 |
| chr10_+_99079008 | 0.30 |
ENST00000371021.3 |
FRAT1 |
frequently rearranged in advanced T-cell lymphomas |
| chr4_-_186733363 | 0.29 |
ENST00000393523.2 ENST00000393528.3 ENST00000449407.2 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr9_-_139922631 | 0.29 |
ENST00000341511.6 |
ABCA2 |
ATP-binding cassette, sub-family A (ABC1), member 2 |
| chr9_+_34458771 | 0.29 |
ENST00000437363.1 ENST00000242317.4 |
DNAI1 |
dynein, axonemal, intermediate chain 1 |
| chr20_-_32031680 | 0.28 |
ENST00000217381.2 |
SNTA1 |
syntrophin, alpha 1 |
| chr7_+_97910981 | 0.28 |
ENST00000297290.3 |
BRI3 |
brain protein I3 |
| chr3_+_139654018 | 0.28 |
ENST00000458420.3 |
CLSTN2 |
calsyntenin 2 |
| chr1_-_36915880 | 0.28 |
ENST00000445843.3 |
OSCP1 |
organic solute carrier partner 1 |
| chr9_-_139922726 | 0.28 |
ENST00000265662.5 ENST00000371605.3 |
ABCA2 |
ATP-binding cassette, sub-family A (ABC1), member 2 |
| chrX_-_153602991 | 0.27 |
ENST00000369850.3 ENST00000422373.1 |
FLNA |
filamin A, alpha |
| chr1_-_36916011 | 0.27 |
ENST00000356637.5 ENST00000354267.3 ENST00000235532.5 |
OSCP1 |
organic solute carrier partner 1 |
| chr16_+_30007524 | 0.26 |
ENST00000567254.1 ENST00000567705.1 |
INO80E |
INO80 complex subunit E |
| chr7_+_97910962 | 0.26 |
ENST00000539286.1 |
BRI3 |
brain protein I3 |
| chr2_+_30670077 | 0.26 |
ENST00000466477.1 ENST00000465200.1 ENST00000379509.3 ENST00000319406.4 ENST00000488144.1 ENST00000465538.1 ENST00000309052.4 ENST00000359433.1 |
LCLAT1 |
lysocardiolipin acyltransferase 1 |
| chr9_-_72374848 | 0.26 |
ENST00000377200.5 ENST00000340434.4 ENST00000472967.2 |
PTAR1 |
protein prenyltransferase alpha subunit repeat containing 1 |
| chr7_-_134143841 | 0.25 |
ENST00000285930.4 |
AKR1B1 |
aldo-keto reductase family 1, member B1 (aldose reductase) |
| chr3_-_49449350 | 0.25 |
ENST00000454011.2 ENST00000445425.1 ENST00000422781.1 |
RHOA |
ras homolog family member A |
| chr5_-_159546396 | 0.25 |
ENST00000523662.1 ENST00000456329.3 ENST00000307063.7 |
PWWP2A |
PWWP domain containing 2A |
| chr12_-_2113583 | 0.25 |
ENST00000397173.4 ENST00000280665.6 |
DCP1B |
decapping mRNA 1B |
| chr5_-_138718973 | 0.24 |
ENST00000353963.3 ENST00000348729.3 |
SLC23A1 |
solute carrier family 23 (ascorbic acid transporter), member 1 |
| chr11_-_62380199 | 0.24 |
ENST00000419857.1 ENST00000394773.2 |
EML3 |
echinoderm microtubule associated protein like 3 |
| chr12_+_49760639 | 0.24 |
ENST00000549538.1 ENST00000548654.1 ENST00000550643.1 ENST00000548710.1 ENST00000549179.1 ENST00000548377.1 |
SPATS2 |
spermatogenesis associated, serine-rich 2 |
| chr2_+_204193129 | 0.24 |
ENST00000417864.1 |
ABI2 |
abl-interactor 2 |
| chr19_+_34972543 | 0.24 |
ENST00000590071.2 |
WTIP |
Wilms tumor 1 interacting protein |
| chr12_+_53773944 | 0.23 |
ENST00000551969.1 ENST00000327443.4 |
SP1 |
Sp1 transcription factor |
| chr9_+_100174232 | 0.23 |
ENST00000355295.4 |
TDRD7 |
tudor domain containing 7 |
| chr15_-_43802769 | 0.23 |
ENST00000263801.3 |
TP53BP1 |
tumor protein p53 binding protein 1 |
| chr1_+_61547894 | 0.22 |
ENST00000403491.3 |
NFIA |
nuclear factor I/A |
| chr5_+_176873446 | 0.22 |
ENST00000507881.1 |
PRR7 |
proline rich 7 (synaptic) |
| chr8_-_11324273 | 0.22 |
ENST00000284486.4 |
FAM167A |
family with sequence similarity 167, member A |
| chr2_+_121010370 | 0.22 |
ENST00000420510.1 |
RALB |
v-ral simian leukemia viral oncogene homolog B |
| chr11_+_125462690 | 0.21 |
ENST00000392708.4 ENST00000529196.1 ENST00000531491.1 |
STT3A |
STT3A, subunit of the oligosaccharyltransferase complex (catalytic) |
| chr17_-_45056606 | 0.21 |
ENST00000322329.3 |
RPRML |
reprimo-like |
| chr6_-_24721054 | 0.21 |
ENST00000378119.4 |
C6orf62 |
chromosome 6 open reading frame 62 |
| chr9_+_137533615 | 0.21 |
ENST00000371817.3 |
COL5A1 |
collagen, type V, alpha 1 |
| chr20_-_30458432 | 0.21 |
ENST00000375966.4 ENST00000278979.3 |
DUSP15 |
dual specificity phosphatase 15 |
| chr7_+_100271446 | 0.21 |
ENST00000419828.1 ENST00000427895.1 |
GNB2 |
guanine nucleotide binding protein (G protein), beta polypeptide 2 |
| chr2_+_204193149 | 0.21 |
ENST00000422511.2 |
ABI2 |
abl-interactor 2 |
| chr8_+_42995548 | 0.21 |
ENST00000458501.2 ENST00000379644.4 |
HGSNAT |
heparan-alpha-glucosaminide N-acetyltransferase |
| chr10_-_104179682 | 0.21 |
ENST00000406432.1 |
PSD |
pleckstrin and Sec7 domain containing |
| chr9_-_94186131 | 0.21 |
ENST00000297689.3 |
NFIL3 |
nuclear factor, interleukin 3 regulated |
| chr5_-_127873496 | 0.21 |
ENST00000508989.1 |
FBN2 |
fibrillin 2 |
| chr1_+_93811438 | 0.21 |
ENST00000370272.4 ENST00000370267.1 |
DR1 |
down-regulator of transcription 1, TBP-binding (negative cofactor 2) |
| chr2_-_37384175 | 0.20 |
ENST00000411537.2 ENST00000233057.4 ENST00000395127.2 ENST00000390013.3 |
EIF2AK2 |
eukaryotic translation initiation factor 2-alpha kinase 2 |
| chr9_+_129622904 | 0.20 |
ENST00000319119.4 |
ZBTB34 |
zinc finger and BTB domain containing 34 |
| chrX_-_134232630 | 0.19 |
ENST00000535837.1 ENST00000433425.2 |
LINC00087 |
long intergenic non-protein coding RNA 87 |
| chr4_+_190861941 | 0.19 |
ENST00000226798.4 |
FRG1 |
FSHD region gene 1 |
| chr19_+_1249869 | 0.19 |
ENST00000591446.2 |
MIDN |
midnolin |
| chr21_+_35445811 | 0.19 |
ENST00000399312.2 |
MRPS6 |
mitochondrial ribosomal protein S6 |
| chr7_+_100271355 | 0.18 |
ENST00000436220.1 ENST00000424361.1 |
GNB2 |
guanine nucleotide binding protein (G protein), beta polypeptide 2 |
| chr17_-_73179046 | 0.18 |
ENST00000314523.7 ENST00000420826.2 |
SUMO2 |
small ubiquitin-like modifier 2 |
| chr19_+_6372444 | 0.18 |
ENST00000245812.3 |
ALKBH7 |
alkB, alkylation repair homolog 7 (E. coli) |
| chr5_-_142783175 | 0.18 |
ENST00000231509.3 ENST00000394464.2 |
NR3C1 |
nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) |
| chr16_-_28074822 | 0.18 |
ENST00000395724.3 ENST00000380898.2 ENST00000447459.2 |
GSG1L |
GSG1-like |
| chr1_+_6845578 | 0.17 |
ENST00000467404.2 ENST00000439411.2 |
CAMTA1 |
calmodulin binding transcription activator 1 |
| chr8_-_144241432 | 0.17 |
ENST00000430474.2 |
LY6H |
lymphocyte antigen 6 complex, locus H |
| chr19_+_14017116 | 0.17 |
ENST00000589606.1 |
CC2D1A |
coiled-coil and C2 domain containing 1A |
| chr17_+_17206635 | 0.17 |
ENST00000389022.4 |
NT5M |
5',3'-nucleotidase, mitochondrial |
| chr2_+_64068074 | 0.17 |
ENST00000394417.2 ENST00000484142.1 ENST00000482668.1 ENST00000467648.2 |
UGP2 |
UDP-glucose pyrophosphorylase 2 |
| chr5_-_132166579 | 0.17 |
ENST00000378679.3 |
SHROOM1 |
shroom family member 1 |
| chr2_+_120301997 | 0.17 |
ENST00000602047.1 |
PCDP1 |
Primary ciliary dyskinesia protein 1 |
| chr2_+_42396574 | 0.17 |
ENST00000401738.3 |
EML4 |
echinoderm microtubule associated protein like 4 |
| chr12_+_56137064 | 0.16 |
ENST00000257868.5 ENST00000546799.1 |
GDF11 |
growth differentiation factor 11 |
| chr2_+_120302041 | 0.16 |
ENST00000442513.3 ENST00000413369.3 |
PCDP1 |
Primary ciliary dyskinesia protein 1 |
| chr12_+_57482665 | 0.16 |
ENST00000300131.3 |
NAB2 |
NGFI-A binding protein 2 (EGR1 binding protein 2) |
| chr2_+_121010413 | 0.16 |
ENST00000404963.3 |
RALB |
v-ral simian leukemia viral oncogene homolog B |
| chr3_-_48936272 | 0.16 |
ENST00000544097.1 ENST00000430379.1 ENST00000319017.4 |
SLC25A20 |
solute carrier family 25 (carnitine/acylcarnitine translocase), member 20 |
| chr12_+_49761147 | 0.16 |
ENST00000549298.1 |
SPATS2 |
spermatogenesis associated, serine-rich 2 |
| chr2_+_121010324 | 0.16 |
ENST00000272519.5 |
RALB |
v-ral simian leukemia viral oncogene homolog B |
| chr2_-_97304105 | 0.15 |
ENST00000599854.1 ENST00000441706.2 |
KANSL3 |
KAT8 regulatory NSL complex subunit 3 |
| chr16_+_30006997 | 0.15 |
ENST00000304516.7 |
INO80E |
INO80 complex subunit E |
| chr19_+_14017003 | 0.15 |
ENST00000318003.7 |
CC2D1A |
coiled-coil and C2 domain containing 1A |
| chr3_-_49377499 | 0.15 |
ENST00000265560.4 |
USP4 |
ubiquitin specific peptidase 4 (proto-oncogene) |
| chr13_+_48877895 | 0.15 |
ENST00000267163.4 |
RB1 |
retinoblastoma 1 |
| chrX_+_154299690 | 0.15 |
ENST00000340647.4 ENST00000330045.7 |
BRCC3 |
BRCA1/BRCA2-containing complex, subunit 3 |
| chr2_-_97304009 | 0.15 |
ENST00000431828.1 ENST00000435669.1 ENST00000440133.1 ENST00000448075.1 |
KANSL3 |
KAT8 regulatory NSL complex subunit 3 |
| chr15_-_35047166 | 0.14 |
ENST00000290374.4 |
GJD2 |
gap junction protein, delta 2, 36kDa |
| chr10_+_94352956 | 0.14 |
ENST00000260731.3 |
KIF11 |
kinesin family member 11 |
| chr6_+_24495185 | 0.14 |
ENST00000348925.2 |
ALDH5A1 |
aldehyde dehydrogenase 5 family, member A1 |
| chr3_-_196295437 | 0.14 |
ENST00000429115.1 |
WDR53 |
WD repeat domain 53 |
| chr19_-_45004556 | 0.14 |
ENST00000587047.1 ENST00000391956.4 ENST00000221327.4 ENST00000586637.1 ENST00000591064.1 ENST00000592529.1 |
ZNF180 |
zinc finger protein 180 |
| chr19_-_39881777 | 0.14 |
ENST00000595564.1 ENST00000221265.3 |
PAF1 |
Paf1, RNA polymerase II associated factor, homolog (S. cerevisiae) |
| chr14_+_64971292 | 0.13 |
ENST00000358738.3 ENST00000394712.2 |
ZBTB1 |
zinc finger and BTB domain containing 1 |
| chr7_+_65540853 | 0.13 |
ENST00000380839.4 ENST00000395332.3 ENST00000362000.5 ENST00000395331.3 |
ASL |
argininosuccinate lyase |
| chr1_+_6845497 | 0.13 |
ENST00000473578.1 ENST00000557126.1 |
CAMTA1 |
calmodulin binding transcription activator 1 |
| chr4_-_183838372 | 0.13 |
ENST00000503820.1 ENST00000503988.1 |
DCTD |
dCMP deaminase |
| chr2_-_33824382 | 0.13 |
ENST00000238823.8 |
FAM98A |
family with sequence similarity 98, member A |
| chr13_-_22178284 | 0.12 |
ENST00000468222.2 ENST00000382374.4 |
MICU2 |
mitochondrial calcium uptake 2 |
| chr8_+_26240414 | 0.12 |
ENST00000380629.2 |
BNIP3L |
BCL2/adenovirus E1B 19kDa interacting protein 3-like |
| chr2_+_157292859 | 0.12 |
ENST00000438166.2 |
GPD2 |
glycerol-3-phosphate dehydrogenase 2 (mitochondrial) |
| chr7_+_65540780 | 0.12 |
ENST00000304874.9 |
ASL |
argininosuccinate lyase |
| chr8_-_144241664 | 0.12 |
ENST00000342752.4 |
LY6H |
lymphocyte antigen 6 complex, locus H |
| chrX_-_119695279 | 0.12 |
ENST00000336592.6 |
CUL4B |
cullin 4B |
| chr2_-_33824336 | 0.12 |
ENST00000431950.1 ENST00000403368.1 ENST00000441530.2 |
FAM98A |
family with sequence similarity 98, member A |
| chr19_-_36545649 | 0.12 |
ENST00000292894.1 |
THAP8 |
THAP domain containing 8 |
| chr10_-_131909071 | 0.11 |
ENST00000456581.1 |
LINC00959 |
long intergenic non-protein coding RNA 959 |
| chr4_-_18023350 | 0.11 |
ENST00000539056.1 ENST00000382226.5 ENST00000326877.4 |
LCORL |
ligand dependent nuclear receptor corepressor-like |
| chr11_-_6633799 | 0.11 |
ENST00000299424.4 |
TAF10 |
TAF10 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 30kDa |
| chr17_-_79650818 | 0.11 |
ENST00000397498.4 |
ARL16 |
ADP-ribosylation factor-like 16 |
| chr20_+_3451732 | 0.11 |
ENST00000446916.2 |
ATRN |
attractin |
| chr5_+_14143728 | 0.11 |
ENST00000344204.4 ENST00000537187.1 |
TRIO |
trio Rho guanine nucleotide exchange factor |
| chr18_+_67068228 | 0.11 |
ENST00000382713.5 |
DOK6 |
docking protein 6 |
| chr22_+_32871224 | 0.10 |
ENST00000452138.1 ENST00000382058.3 ENST00000397426.1 |
FBXO7 |
F-box protein 7 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.7 | GO:0061428 | negative regulation of transcription from RNA polymerase II promoter in response to hypoxia(GO:0061428) |
| 0.4 | 1.5 | GO:2001151 | regulation of renal water transport(GO:2001151) positive regulation of renal water transport(GO:2001153) |
| 0.4 | 1.4 | GO:0007181 | transforming growth factor beta receptor complex assembly(GO:0007181) response to luteinizing hormone(GO:0034699) |
| 0.3 | 2.0 | GO:2000546 | positive regulation of cell chemotaxis to fibroblast growth factor(GO:1904849) positive regulation of endothelial cell chemotaxis to fibroblast growth factor(GO:2000546) |
| 0.3 | 4.3 | GO:0043589 | skin morphogenesis(GO:0043589) |
| 0.2 | 0.7 | GO:0048203 | vesicle targeting, trans-Golgi to endosome(GO:0048203) |
| 0.2 | 2.6 | GO:0030091 | protein repair(GO:0030091) |
| 0.2 | 0.6 | GO:0050760 | negative regulation of thymidylate synthase biosynthetic process(GO:0050760) |
| 0.2 | 1.2 | GO:0072674 | multinuclear osteoclast differentiation(GO:0072674) osteoclast fusion(GO:0072675) |
| 0.2 | 0.5 | GO:0001928 | regulation of exocyst assembly(GO:0001928) regulation of exocyst localization(GO:0060178) |
| 0.1 | 0.6 | GO:1990868 | response to chemokine(GO:1990868) cellular response to chemokine(GO:1990869) |
| 0.1 | 0.9 | GO:0061518 | macrophage proliferation(GO:0061517) microglial cell proliferation(GO:0061518) regulation of neuronal signal transduction(GO:1902847) positive regulation of neurofibrillary tangle assembly(GO:1902998) |
| 0.1 | 1.0 | GO:0003383 | apical constriction(GO:0003383) |
| 0.1 | 0.4 | GO:1902683 | regulation of receptor localization to synapse(GO:1902683) |
| 0.1 | 0.4 | GO:0035359 | negative regulation of peroxisome proliferator activated receptor signaling pathway(GO:0035359) |
| 0.1 | 1.1 | GO:0006528 | asparagine metabolic process(GO:0006528) |
| 0.1 | 0.7 | GO:0006540 | acetate metabolic process(GO:0006083) glutamate decarboxylation to succinate(GO:0006540) gamma-aminobutyric acid catabolic process(GO:0009450) |
| 0.1 | 0.6 | GO:0036072 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.1 | 0.4 | GO:1900194 | negative regulation of oocyte maturation(GO:1900194) |
| 0.1 | 0.6 | GO:0032380 | regulation of intracellular lipid transport(GO:0032377) regulation of intracellular sterol transport(GO:0032380) regulation of intracellular cholesterol transport(GO:0032383) |
| 0.1 | 0.3 | GO:0042450 | arginine biosynthetic process via ornithine(GO:0042450) |
| 0.1 | 0.3 | GO:0006059 | hexitol metabolic process(GO:0006059) inner medullary collecting duct development(GO:0072061) |
| 0.1 | 0.2 | GO:0070837 | L-ascorbic acid transport(GO:0015882) dehydroascorbic acid transport(GO:0070837) transepithelial L-ascorbic acid transport(GO:0070904) |
| 0.1 | 0.7 | GO:0038028 | insulin receptor signaling pathway via phosphatidylinositol 3-kinase(GO:0038028) |
| 0.1 | 0.6 | GO:0055059 | asymmetric neuroblast division(GO:0055059) |
| 0.1 | 0.7 | GO:0014824 | artery smooth muscle contraction(GO:0014824) |
| 0.1 | 0.2 | GO:0043686 | co-translational protein modification(GO:0043686) |
| 0.1 | 0.2 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.1 | 2.9 | GO:0045746 | negative regulation of Notch signaling pathway(GO:0045746) |
| 0.1 | 0.3 | GO:0045829 | negative regulation of isotype switching(GO:0045829) |
| 0.1 | 0.6 | GO:1990034 | cellular response to corticosterone stimulus(GO:0071386) calcium ion export from cell(GO:1990034) |
| 0.1 | 0.5 | GO:0043402 | glucocorticoid mediated signaling pathway(GO:0043402) |
| 0.1 | 1.1 | GO:0060628 | regulation of ER to Golgi vesicle-mediated transport(GO:0060628) |
| 0.1 | 0.5 | GO:0002457 | T cell antigen processing and presentation(GO:0002457) |
| 0.1 | 0.4 | GO:0035879 | plasma membrane lactate transport(GO:0035879) |
| 0.1 | 0.4 | GO:0045617 | negative regulation of keratinocyte differentiation(GO:0045617) |
| 0.1 | 0.2 | GO:0016185 | synaptic vesicle budding from presynaptic endocytic zone membrane(GO:0016185) |
| 0.1 | 0.2 | GO:1902616 | acyl carnitine transport(GO:0006844) acyl carnitine transmembrane transport(GO:1902616) |
| 0.1 | 0.5 | GO:0046078 | dUMP metabolic process(GO:0046078) |
| 0.1 | 1.5 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 0.1 | 0.2 | GO:0071921 | establishment of sister chromatid cohesion(GO:0034085) cohesin loading(GO:0071921) regulation of cohesin loading(GO:0071922) negative regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0071930) |
| 0.0 | 0.4 | GO:1904628 | response to phorbol 13-acetate 12-myristate(GO:1904627) cellular response to phorbol 13-acetate 12-myristate(GO:1904628) |
| 0.0 | 0.3 | GO:0070940 | dephosphorylation of RNA polymerase II C-terminal domain(GO:0070940) |
| 0.0 | 0.3 | GO:0016479 | negative regulation of transcription from RNA polymerase I promoter(GO:0016479) |
| 0.0 | 0.4 | GO:2000601 | positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 0.0 | 0.2 | GO:1902949 | positive regulation of tau-protein kinase activity(GO:1902949) |
| 0.0 | 0.3 | GO:0035965 | cardiolipin acyl-chain remodeling(GO:0035965) |
| 0.0 | 0.2 | GO:0019255 | glucose 1-phosphate metabolic process(GO:0019255) |
| 0.0 | 0.1 | GO:0002572 | pro-T cell differentiation(GO:0002572) |
| 0.0 | 0.1 | GO:0051771 | negative regulation of nitric-oxide synthase biosynthetic process(GO:0051771) |
| 0.0 | 0.2 | GO:1903300 | negative regulation of glucokinase activity(GO:0033132) negative regulation of hexokinase activity(GO:1903300) |
| 0.0 | 0.1 | GO:0051562 | negative regulation of mitochondrial calcium ion concentration(GO:0051562) |
| 0.0 | 0.2 | GO:1902445 | regulation of mitochondrial membrane permeability involved in programmed necrotic cell death(GO:1902445) |
| 0.0 | 1.4 | GO:0048713 | regulation of oligodendrocyte differentiation(GO:0048713) |
| 0.0 | 0.8 | GO:0072189 | ureter development(GO:0072189) |
| 0.0 | 0.2 | GO:0019049 | evasion or tolerance of host defenses by virus(GO:0019049) avoidance of host defenses(GO:0044413) evasion or tolerance of host defenses(GO:0044415) avoidance of defenses of other organism involved in symbiotic interaction(GO:0051832) evasion or tolerance of defenses of other organism involved in symbiotic interaction(GO:0051834) |
| 0.0 | 0.1 | GO:0021526 | medial motor column neuron differentiation(GO:0021526) |
| 0.0 | 0.4 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 0.0 | 0.1 | GO:0006127 | glycerophosphate shuttle(GO:0006127) |
| 0.0 | 0.2 | GO:0031087 | nuclear-transcribed mRNA catabolic process, deadenylation-independent decay(GO:0031086) deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.0 | 0.2 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 0.0 | 0.1 | GO:0060406 | positive regulation of penile erection(GO:0060406) |
| 0.0 | 0.4 | GO:0021516 | dorsal spinal cord development(GO:0021516) |
| 0.0 | 0.2 | GO:0035583 | sequestering of TGFbeta in extracellular matrix(GO:0035583) |
| 0.0 | 0.5 | GO:0002089 | lens morphogenesis in camera-type eye(GO:0002089) |
| 0.0 | 0.3 | GO:0097354 | protein prenylation(GO:0018342) prenylation(GO:0097354) |
| 0.0 | 0.1 | GO:1904327 | protein localization to cytosolic proteasome complex(GO:1904327) protein localization to cytosolic proteasome complex involved in ERAD pathway(GO:1904379) |
| 0.0 | 0.2 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.0 | 0.1 | GO:1902988 | neurofibrillary tangle assembly(GO:1902988) |
| 0.0 | 0.7 | GO:0048025 | negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.0 | 0.1 | GO:1900239 | phenotypic switching(GO:0036166) regulation of phenotypic switching(GO:1900239) |
| 0.0 | 0.1 | GO:0007185 | transmembrane receptor protein tyrosine phosphatase signaling pathway(GO:0007185) |
| 0.0 | 0.2 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination(GO:2000042) |
| 0.0 | 0.1 | GO:0035694 | mitochondrial protein catabolic process(GO:0035694) |
| 0.0 | 0.4 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.0 | 0.3 | GO:0043981 | histone H4-K5 acetylation(GO:0043981) histone H4-K8 acetylation(GO:0043982) |
| 0.0 | 0.3 | GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) |
| 0.0 | 1.4 | GO:0010812 | negative regulation of cell-substrate adhesion(GO:0010812) |
| 0.0 | 0.1 | GO:0007195 | adenylate cyclase-inhibiting dopamine receptor signaling pathway(GO:0007195) |
| 0.0 | 0.8 | GO:0070911 | global genome nucleotide-excision repair(GO:0070911) |
| 0.0 | 0.3 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 0.0 | 0.4 | GO:0097503 | glycosphingolipid biosynthetic process(GO:0006688) sialylation(GO:0097503) |
| 0.0 | 0.3 | GO:0061049 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.0 | 0.3 | GO:0042276 | error-prone translesion synthesis(GO:0042276) |
| 0.0 | 0.1 | GO:2000825 | positive regulation of androgen receptor activity(GO:2000825) |
| 0.0 | 0.5 | GO:0003016 | respiratory system process(GO:0003016) |
| 0.0 | 0.1 | GO:1904424 | regulation of GTP binding(GO:1904424) |
| 0.0 | 1.2 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.0 | 0.9 | GO:0046326 | positive regulation of glucose import(GO:0046326) |
| 0.0 | 0.1 | GO:1903599 | positive regulation of mitophagy(GO:1903599) |
| 0.0 | 0.3 | GO:1904886 | beta-catenin destruction complex disassembly(GO:1904886) |
| 0.0 | 0.0 | GO:0036353 | histone H2A-K119 monoubiquitination(GO:0036353) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.4 | GO:0034673 | inhibin-betaglycan-ActRII complex(GO:0034673) |
| 0.3 | 4.5 | GO:0005583 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.1 | 0.4 | GO:0005940 | septin ring(GO:0005940) septin collar(GO:0032173) |
| 0.1 | 1.2 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 0.1 | 0.7 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.1 | 0.6 | GO:1990812 | growth cone lamellipodium(GO:1990761) growth cone filopodium(GO:1990812) |
| 0.1 | 1.1 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.1 | 0.2 | GO:0031523 | Myb complex(GO:0031523) |
| 0.1 | 0.9 | GO:0097418 | neurofibrillary tangle(GO:0097418) |
| 0.1 | 0.3 | GO:0097454 | Schwann cell microvillus(GO:0097454) |
| 0.1 | 0.3 | GO:0016035 | zeta DNA polymerase complex(GO:0016035) |
| 0.1 | 0.6 | GO:0043190 | ATP-binding cassette (ABC) transporter complex(GO:0043190) |
| 0.1 | 0.2 | GO:0035189 | Rb-E2F complex(GO:0035189) |
| 0.0 | 1.3 | GO:0033268 | node of Ranvier(GO:0033268) |
| 0.0 | 0.3 | GO:0000347 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.0 | 0.1 | GO:0005953 | CAAX-protein geranylgeranyltransferase complex(GO:0005953) |
| 0.0 | 0.7 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.0 | 0.5 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.0 | 0.3 | GO:0016013 | syntrophin complex(GO:0016013) |
| 0.0 | 0.4 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.0 | 0.3 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.0 | 0.1 | GO:1990246 | uniplex complex(GO:1990246) |
| 0.0 | 0.2 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.0 | 0.2 | GO:0072669 | tRNA-splicing ligase complex(GO:0072669) |
| 0.0 | 0.6 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.0 | 0.1 | GO:0000125 | PCAF complex(GO:0000125) |
| 0.0 | 0.1 | GO:0016939 | kinesin II complex(GO:0016939) |
| 0.0 | 0.1 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.0 | 0.2 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.0 | 0.9 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.0 | 0.6 | GO:0000145 | exocyst(GO:0000145) |
| 0.0 | 0.1 | GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 0.0 | 0.1 | GO:1990037 | Lewy body core(GO:1990037) |
| 0.0 | 0.3 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.0 | 2.7 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 0.1 | GO:0030934 | anchoring collagen complex(GO:0030934) |
| 0.0 | 0.4 | GO:0031011 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.0 | 0.1 | GO:0000322 | storage vacuole(GO:0000322) |
| 0.0 | 0.2 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.0 | 0.2 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.0 | 0.4 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 2.6 | GO:0033743 | peptide-methionine (R)-S-oxide reductase activity(GO:0033743) |
| 0.4 | 1.1 | GO:0008798 | beta-aspartyl-peptidase activity(GO:0008798) |
| 0.3 | 4.5 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.3 | 0.9 | GO:0050613 | delta14-sterol reductase activity(GO:0050613) |
| 0.2 | 1.5 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.1 | 1.4 | GO:0005114 | type II transforming growth factor beta receptor binding(GO:0005114) |
| 0.1 | 1.7 | GO:0050693 | LBD domain binding(GO:0050693) |
| 0.1 | 0.5 | GO:0038051 | glucocorticoid receptor activity(GO:0004883) glucocorticoid-activated RNA polymerase II transcription factor binding transcription factor activity(GO:0038051) |
| 0.1 | 0.6 | GO:0051022 | GDP-dissociation inhibitor binding(GO:0051021) Rho GDP-dissociation inhibitor binding(GO:0051022) |
| 0.1 | 0.3 | GO:0004132 | dCMP deaminase activity(GO:0004132) |
| 0.1 | 0.2 | GO:0033300 | L-ascorbate:sodium symporter activity(GO:0008520) L-ascorbic acid transporter activity(GO:0015229) dehydroascorbic acid transporter activity(GO:0033300) sodium-dependent L-ascorbate transmembrane transporter activity(GO:0070890) |
| 0.1 | 0.3 | GO:0034988 | Fc-gamma receptor I complex binding(GO:0034988) |
| 0.1 | 2.0 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.1 | 0.2 | GO:0051748 | UTP:glucose-1-phosphate uridylyltransferase activity(GO:0003983) UTP-monosaccharide-1-phosphate uridylyltransferase activity(GO:0051748) |
| 0.1 | 0.7 | GO:0031386 | protein tag(GO:0031386) |
| 0.1 | 0.2 | GO:0015227 | acyl carnitine transmembrane transporter activity(GO:0015227) |
| 0.1 | 0.2 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 0.0 | 1.6 | GO:0070742 | C2H2 zinc finger domain binding(GO:0070742) |
| 0.0 | 1.4 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 2.6 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.0 | 0.4 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.0 | 0.9 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.0 | 0.2 | GO:0031685 | adenosine receptor binding(GO:0031685) |
| 0.0 | 0.4 | GO:0008318 | protein prenyltransferase activity(GO:0008318) |
| 0.0 | 0.4 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.0 | 0.6 | GO:0048156 | tau protein binding(GO:0048156) |
| 0.0 | 0.2 | GO:0061649 | ubiquitinated histone binding(GO:0061649) |
| 0.0 | 0.3 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 0.0 | 0.3 | GO:0008420 | CTD phosphatase activity(GO:0008420) |
| 0.0 | 0.1 | GO:0052591 | sn-glycerol-3-phosphate:ubiquinone oxidoreductase activity(GO:0052590) sn-glycerol-3-phosphate:ubiquinone-8 oxidoreductase activity(GO:0052591) |
| 0.0 | 0.5 | GO:0005522 | profilin binding(GO:0005522) |
| 0.0 | 0.2 | GO:0061676 | importin-alpha family protein binding(GO:0061676) |
| 0.0 | 0.4 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.0 | 0.2 | GO:0030023 | extracellular matrix constituent conferring elasticity(GO:0030023) |
| 0.0 | 0.1 | GO:0004140 | dephospho-CoA kinase activity(GO:0004140) |
| 0.0 | 0.3 | GO:0004032 | alditol:NADP+ 1-oxidoreductase activity(GO:0004032) |
| 0.0 | 0.3 | GO:0046972 | histone acetyltransferase activity (H4-K5 specific)(GO:0043995) histone acetyltransferase activity (H4-K8 specific)(GO:0043996) histone acetyltransferase activity (H4-K16 specific)(GO:0046972) |
| 0.0 | 0.7 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.0 | 0.8 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 0.4 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.0 | 0.7 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) |
| 0.0 | 0.1 | GO:0099609 | microtubule lateral binding(GO:0099609) |
| 0.0 | 0.6 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.0 | 1.2 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.0 | 1.3 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.0 | 0.4 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.0 | 0.3 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.0 | 0.2 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.0 | 0.2 | GO:0004579 | dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.0 | 0.5 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.0 | 0.7 | GO:0016620 | oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor(GO:0016620) |
| 0.0 | 0.4 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.0 | 0.2 | GO:0050998 | nitric-oxide synthase binding(GO:0050998) |
| 0.0 | 0.7 | GO:0004407 | histone deacetylase activity(GO:0004407) |
| 0.0 | 0.1 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 0.1 | GO:0001665 | alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase activity(GO:0001665) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 4.3 | PID LYMPH ANGIOGENESIS PATHWAY | VEGFR3 signaling in lymphatic endothelium |
| 0.0 | 1.7 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.0 | 2.6 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 1.8 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 0.8 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 1.1 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.3 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 1.4 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 0.7 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.0 | 0.4 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.0 | 0.7 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.0 | 0.6 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.0 | 1.2 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 4.3 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.1 | 2.0 | REACTOME SIGNALING BY FGFR3 MUTANTS | Genes involved in Signaling by FGFR3 mutants |
| 0.0 | 1.7 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 1.5 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.0 | 0.7 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.0 | 0.9 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 1.0 | REACTOME G BETA GAMMA SIGNALLING THROUGH PI3KGAMMA | Genes involved in G beta:gamma signalling through PI3Kgamma |
| 0.0 | 0.6 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.0 | 0.7 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.0 | 0.7 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 0.2 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.0 | 1.6 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.4 | REACTOME DESTABILIZATION OF MRNA BY TRISTETRAPROLIN TTP | Genes involved in Destabilization of mRNA by Tristetraprolin (TTP) |
| 0.0 | 0.5 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.0 | 0.4 | REACTOME P38MAPK EVENTS | Genes involved in p38MAPK events |
| 0.0 | 0.4 | REACTOME PYRUVATE METABOLISM | Genes involved in Pyruvate metabolism |
| 0.0 | 0.5 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.0 | 0.4 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 0.1 | REACTOME ADP SIGNALLING THROUGH P2RY1 | Genes involved in ADP signalling through P2Y purinoceptor 1 |