Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
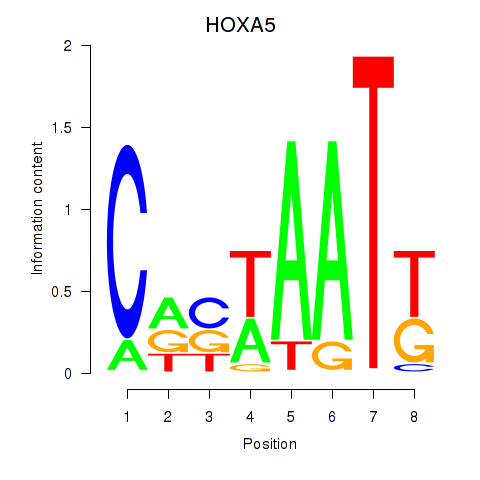
Results for HOXA5
Z-value: 1.61
Transcription factors associated with HOXA5
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HOXA5
|
ENSG00000106004.4 | HOXA5 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HOXA5 | hg19_v2_chr7_-_27183263_27183287 | 0.54 | 1.7e-01 | Click! |
Activity profile of HOXA5 motif
Sorted Z-values of HOXA5 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HOXA5
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr14_-_92413353 | 3.45 |
ENST00000556154.1 |
FBLN5 |
fibulin 5 |
| chr14_-_92413727 | 3.04 |
ENST00000267620.10 |
FBLN5 |
fibulin 5 |
| chr2_-_158300556 | 2.63 |
ENST00000264192.3 |
CYTIP |
cytohesin 1 interacting protein |
| chr12_-_91574142 | 2.37 |
ENST00000547937.1 |
DCN |
decorin |
| chr3_+_12392971 | 1.28 |
ENST00000287820.6 |
PPARG |
peroxisome proliferator-activated receptor gamma |
| chr1_-_149889382 | 1.01 |
ENST00000369145.1 ENST00000369146.3 |
SV2A |
synaptic vesicle glycoprotein 2A |
| chr2_-_160919112 | 0.97 |
ENST00000283243.7 ENST00000392771.1 |
PLA2R1 |
phospholipase A2 receptor 1, 180kDa |
| chr17_+_19437132 | 0.91 |
ENST00000436810.2 ENST00000270570.4 ENST00000457293.1 ENST00000542886.1 ENST00000575023.1 ENST00000395585.1 |
SLC47A1 |
solute carrier family 47 (multidrug and toxin extrusion), member 1 |
| chr1_-_232651312 | 0.90 |
ENST00000262861.4 |
SIPA1L2 |
signal-induced proliferation-associated 1 like 2 |
| chr13_+_76378305 | 0.88 |
ENST00000526371.1 ENST00000526528.1 |
LMO7 |
LIM domain 7 |
| chr1_-_227505289 | 0.77 |
ENST00000366765.3 |
CDC42BPA |
CDC42 binding protein kinase alpha (DMPK-like) |
| chr1_+_221051699 | 0.74 |
ENST00000366903.6 |
HLX |
H2.0-like homeobox |
| chr2_-_179343226 | 0.71 |
ENST00000434643.2 |
FKBP7 |
FK506 binding protein 7 |
| chr7_-_23510086 | 0.66 |
ENST00000258729.3 |
IGF2BP3 |
insulin-like growth factor 2 mRNA binding protein 3 |
| chr2_-_207082748 | 0.64 |
ENST00000407325.2 ENST00000411719.1 |
GPR1 |
G protein-coupled receptor 1 |
| chr20_+_33292068 | 0.63 |
ENST00000374810.3 ENST00000374809.2 ENST00000451665.1 |
TP53INP2 |
tumor protein p53 inducible nuclear protein 2 |
| chrX_+_54947229 | 0.62 |
ENST00000442098.1 ENST00000430420.1 ENST00000453081.1 ENST00000173898.7 ENST00000319167.8 ENST00000375022.4 ENST00000399736.1 ENST00000440072.1 ENST00000420798.2 ENST00000431115.1 ENST00000440759.1 ENST00000375041.2 |
TRO |
trophinin |
| chr19_+_50016610 | 0.58 |
ENST00000596975.1 |
FCGRT |
Fc fragment of IgG, receptor, transporter, alpha |
| chr6_+_124125286 | 0.57 |
ENST00000368416.1 ENST00000368417.1 ENST00000546092.1 |
NKAIN2 |
Na+/K+ transporting ATPase interacting 2 |
| chr17_-_33446735 | 0.57 |
ENST00000460118.2 ENST00000335858.7 |
RAD51D |
RAD51 paralog D |
| chr2_-_40657397 | 0.56 |
ENST00000408028.2 ENST00000332839.4 ENST00000406391.2 ENST00000542024.1 ENST00000542756.1 ENST00000405901.3 |
SLC8A1 |
solute carrier family 8 (sodium/calcium exchanger), member 1 |
| chr6_+_25963020 | 0.54 |
ENST00000357085.3 |
TRIM38 |
tripartite motif containing 38 |
| chr7_+_99717230 | 0.53 |
ENST00000262932.3 |
CNPY4 |
canopy FGF signaling regulator 4 |
| chr16_+_10971037 | 0.52 |
ENST00000324288.8 ENST00000381835.5 |
CIITA |
class II, major histocompatibility complex, transactivator |
| chr4_-_142054590 | 0.51 |
ENST00000306799.3 |
RNF150 |
ring finger protein 150 |
| chr11_+_20044096 | 0.48 |
ENST00000533917.1 |
NAV2 |
neuron navigator 2 |
| chrX_+_10124977 | 0.48 |
ENST00000380833.4 |
CLCN4 |
chloride channel, voltage-sensitive 4 |
| chr17_-_33446820 | 0.47 |
ENST00000592577.1 ENST00000590016.1 ENST00000345365.6 ENST00000360276.3 ENST00000357906.3 |
RAD51D |
RAD51 paralog D |
| chr14_+_21156915 | 0.46 |
ENST00000397990.4 ENST00000555597.1 |
ANG RNASE4 |
angiogenin, ribonuclease, RNase A family, 5 ribonuclease, RNase A family, 4 |
| chr9_+_75766763 | 0.46 |
ENST00000456643.1 ENST00000415424.1 |
ANXA1 |
annexin A1 |
| chrX_+_55478538 | 0.45 |
ENST00000342972.1 |
MAGEH1 |
melanoma antigen family H, 1 |
| chr5_+_140723601 | 0.44 |
ENST00000253812.6 |
PCDHGA3 |
protocadherin gamma subfamily A, 3 |
| chr9_-_123342415 | 0.42 |
ENST00000349780.4 ENST00000360190.4 ENST00000360822.3 ENST00000359309.3 |
CDK5RAP2 |
CDK5 regulatory subunit associated protein 2 |
| chr11_-_96076334 | 0.42 |
ENST00000524717.1 |
MAML2 |
mastermind-like 2 (Drosophila) |
| chr6_-_116575226 | 0.41 |
ENST00000420283.1 |
TSPYL4 |
TSPY-like 4 |
| chr22_+_46476192 | 0.40 |
ENST00000443490.1 |
FLJ27365 |
hsa-mir-4763 |
| chrX_-_65253506 | 0.39 |
ENST00000427538.1 |
VSIG4 |
V-set and immunoglobulin domain containing 4 |
| chr8_-_122653630 | 0.39 |
ENST00000303924.4 |
HAS2 |
hyaluronan synthase 2 |
| chr11_-_33744256 | 0.38 |
ENST00000415002.2 ENST00000437761.2 ENST00000445143.2 |
CD59 |
CD59 molecule, complement regulatory protein |
| chr17_-_295730 | 0.37 |
ENST00000329099.4 |
FAM101B |
family with sequence similarity 101, member B |
| chr6_+_155537771 | 0.37 |
ENST00000275246.7 |
TIAM2 |
T-cell lymphoma invasion and metastasis 2 |
| chr6_+_116575329 | 0.36 |
ENST00000430252.2 ENST00000540275.1 ENST00000448740.2 |
DSE RP3-486I3.7 |
dermatan sulfate epimerase RP3-486I3.7 |
| chr9_-_110251836 | 0.36 |
ENST00000374672.4 |
KLF4 |
Kruppel-like factor 4 (gut) |
| chr11_-_33744487 | 0.35 |
ENST00000426650.2 |
CD59 |
CD59 molecule, complement regulatory protein |
| chr1_-_149900122 | 0.35 |
ENST00000271628.8 |
SF3B4 |
splicing factor 3b, subunit 4, 49kDa |
| chr10_+_17270214 | 0.35 |
ENST00000544301.1 |
VIM |
vimentin |
| chrX_-_118826784 | 0.32 |
ENST00000394616.4 |
SEPT6 |
septin 6 |
| chr2_+_30670077 | 0.31 |
ENST00000466477.1 ENST00000465200.1 ENST00000379509.3 ENST00000319406.4 ENST00000488144.1 ENST00000465538.1 ENST00000309052.4 ENST00000359433.1 |
LCLAT1 |
lysocardiolipin acyltransferase 1 |
| chr6_-_34216766 | 0.31 |
ENST00000481533.1 ENST00000468145.1 ENST00000413013.2 ENST00000394990.4 ENST00000335352.3 |
C6orf1 |
chromosome 6 open reading frame 1 |
| chr20_-_34638841 | 0.29 |
ENST00000565493.1 |
LINC00657 |
long intergenic non-protein coding RNA 657 |
| chr8_-_123139423 | 0.28 |
ENST00000523792.1 |
RP11-398G24.2 |
RP11-398G24.2 |
| chr3_+_29323043 | 0.27 |
ENST00000452462.1 ENST00000456853.1 |
RBMS3 |
RNA binding motif, single stranded interacting protein 3 |
| chr1_+_84609944 | 0.27 |
ENST00000370685.3 |
PRKACB |
protein kinase, cAMP-dependent, catalytic, beta |
| chr19_-_44384291 | 0.27 |
ENST00000324394.6 |
ZNF404 |
zinc finger protein 404 |
| chr10_+_103986085 | 0.25 |
ENST00000370005.3 |
ELOVL3 |
ELOVL fatty acid elongase 3 |
| chr2_-_61697862 | 0.25 |
ENST00000398571.2 |
USP34 |
ubiquitin specific peptidase 34 |
| chr6_-_154677900 | 0.25 |
ENST00000265198.4 ENST00000520261.1 |
IPCEF1 |
interaction protein for cytohesin exchange factors 1 |
| chr9_-_127905736 | 0.24 |
ENST00000336505.6 ENST00000373549.4 |
SCAI |
suppressor of cancer cell invasion |
| chr12_-_54653313 | 0.24 |
ENST00000550411.1 ENST00000439541.2 |
CBX5 |
chromobox homolog 5 |
| chr1_+_152943122 | 0.24 |
ENST00000328051.2 |
SPRR4 |
small proline-rich protein 4 |
| chr3_-_149093499 | 0.21 |
ENST00000472441.1 |
TM4SF1 |
transmembrane 4 L six family member 1 |
| chr10_+_18240834 | 0.21 |
ENST00000377371.3 ENST00000539911.1 |
SLC39A12 |
solute carrier family 39 (zinc transporter), member 12 |
| chr11_+_33061543 | 0.20 |
ENST00000432887.1 ENST00000528898.1 ENST00000531632.2 |
TCP11L1 |
t-complex 11, testis-specific-like 1 |
| chr7_+_44084233 | 0.20 |
ENST00000448521.1 |
DBNL |
drebrin-like |
| chr10_-_127505167 | 0.19 |
ENST00000368786.1 |
UROS |
uroporphyrinogen III synthase |
| chr4_+_71063641 | 0.19 |
ENST00000514097.1 |
ODAM |
odontogenic, ameloblast asssociated |
| chr11_+_12399071 | 0.19 |
ENST00000539723.1 ENST00000550549.1 |
PARVA |
parvin, alpha |
| chr7_-_23571586 | 0.19 |
ENST00000538367.1 ENST00000392502.4 ENST00000297071.4 |
TRA2A |
transformer 2 alpha homolog (Drosophila) |
| chr14_+_24025462 | 0.18 |
ENST00000556015.1 ENST00000554970.1 ENST00000554789.1 |
THTPA |
thiamine triphosphatase |
| chr11_-_2162468 | 0.18 |
ENST00000434045.2 |
IGF2 |
insulin-like growth factor 2 (somatomedin A) |
| chr9_+_131037623 | 0.17 |
ENST00000495313.1 ENST00000372898.2 |
SWI5 |
SWI5 recombination repair homolog (yeast) |
| chr7_+_44084262 | 0.17 |
ENST00000456905.1 ENST00000440166.1 ENST00000452943.1 ENST00000468694.1 ENST00000494774.1 ENST00000490734.2 |
DBNL |
drebrin-like |
| chr1_-_182921119 | 0.16 |
ENST00000423786.1 |
SHCBP1L |
SHC SH2-domain binding protein 1-like |
| chr6_+_140175987 | 0.16 |
ENST00000414038.1 ENST00000431609.1 |
RP5-899B16.1 |
RP5-899B16.1 |
| chr19_-_35264089 | 0.15 |
ENST00000588760.1 ENST00000329285.8 ENST00000587354.2 |
ZNF599 |
zinc finger protein 599 |
| chr7_-_99717463 | 0.15 |
ENST00000437822.2 |
TAF6 |
TAF6 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 80kDa |
| chr3_+_127770455 | 0.15 |
ENST00000464451.1 |
SEC61A1 |
Sec61 alpha 1 subunit (S. cerevisiae) |
| chr7_-_99716952 | 0.15 |
ENST00000523306.1 ENST00000344095.4 ENST00000417349.1 ENST00000493322.1 ENST00000520135.1 ENST00000418432.2 ENST00000460673.2 ENST00000452041.1 ENST00000452438.2 ENST00000451699.1 ENST00000453269.2 |
TAF6 |
TAF6 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 80kDa |
| chr13_+_76378407 | 0.14 |
ENST00000447038.1 |
LMO7 |
LIM domain 7 |
| chr11_+_65686802 | 0.14 |
ENST00000376991.2 |
DRAP1 |
DR1-associated protein 1 (negative cofactor 2 alpha) |
| chr11_-_31531121 | 0.14 |
ENST00000532287.1 ENST00000526776.1 ENST00000534812.1 ENST00000529749.1 ENST00000278200.1 ENST00000530023.1 ENST00000533642.1 |
IMMP1L |
IMP1 inner mitochondrial membrane peptidase-like (S. cerevisiae) |
| chr11_-_18610246 | 0.13 |
ENST00000379387.4 ENST00000541984.1 |
UEVLD |
UEV and lactate/malate dehyrogenase domains |
| chr12_+_111051832 | 0.13 |
ENST00000550703.2 ENST00000551590.1 |
TCTN1 |
tectonic family member 1 |
| chr1_+_177140633 | 0.12 |
ENST00000361539.4 |
BRINP2 |
bone morphogenetic protein/retinoic acid inducible neural-specific 2 |
| chr8_-_79717750 | 0.12 |
ENST00000263851.4 ENST00000379113.2 |
IL7 |
interleukin 7 |
| chr1_+_154229547 | 0.12 |
ENST00000428595.1 |
UBAP2L |
ubiquitin associated protein 2-like |
| chr10_+_18549645 | 0.12 |
ENST00000396576.2 |
CACNB2 |
calcium channel, voltage-dependent, beta 2 subunit |
| chr16_-_66764119 | 0.11 |
ENST00000569320.1 |
DYNC1LI2 |
dynein, cytoplasmic 1, light intermediate chain 2 |
| chr4_+_158493642 | 0.11 |
ENST00000507108.1 ENST00000455598.1 ENST00000509450.1 |
RP11-364P22.1 |
RP11-364P22.1 |
| chr4_+_25378912 | 0.11 |
ENST00000510092.1 ENST00000505991.1 |
ANAPC4 |
anaphase promoting complex subunit 4 |
| chr12_+_111051902 | 0.11 |
ENST00000397655.3 ENST00000471804.2 ENST00000377654.3 ENST00000397659.4 |
TCTN1 |
tectonic family member 1 |
| chr3_+_186383741 | 0.11 |
ENST00000232003.4 |
HRG |
histidine-rich glycoprotein |
| chr19_-_44259136 | 0.11 |
ENST00000270066.6 |
SMG9 |
SMG9 nonsense mediated mRNA decay factor |
| chr3_+_37035289 | 0.10 |
ENST00000455445.2 ENST00000441265.1 ENST00000435176.1 ENST00000429117.1 ENST00000536378.1 |
MLH1 |
mutL homolog 1 |
| chr14_+_65878565 | 0.10 |
ENST00000556518.1 ENST00000557164.1 |
FUT8 |
fucosyltransferase 8 (alpha (1,6) fucosyltransferase) |
| chr11_+_9685604 | 0.10 |
ENST00000447399.2 ENST00000318950.6 |
SWAP70 |
SWAP switching B-cell complex 70kDa subunit |
| chr10_+_18240814 | 0.10 |
ENST00000377374.4 |
SLC39A12 |
solute carrier family 39 (zinc transporter), member 12 |
| chr1_+_171283331 | 0.09 |
ENST00000367749.3 |
FMO4 |
flavin containing monooxygenase 4 |
| chr4_+_74606223 | 0.09 |
ENST00000307407.3 ENST00000401931.1 |
IL8 |
interleukin 8 |
| chr6_+_168418553 | 0.09 |
ENST00000354419.2 ENST00000351261.3 |
KIF25 |
kinesin family member 25 |
| chr3_+_174577070 | 0.09 |
ENST00000454872.1 |
NAALADL2 |
N-acetylated alpha-linked acidic dipeptidase-like 2 |
| chr5_+_132149017 | 0.08 |
ENST00000378693.2 |
SOWAHA |
sosondowah ankyrin repeat domain family member A |
| chr17_-_38911580 | 0.08 |
ENST00000312150.4 |
KRT25 |
keratin 25 |
| chr6_-_2751146 | 0.08 |
ENST00000268446.5 ENST00000274643.7 |
MYLK4 |
myosin light chain kinase family, member 4 |
| chr14_+_93389425 | 0.08 |
ENST00000216492.5 ENST00000334654.4 |
CHGA |
chromogranin A (parathyroid secretory protein 1) |
| chr10_-_7708918 | 0.08 |
ENST00000256861.6 ENST00000397146.2 ENST00000446830.2 ENST00000397145.2 |
ITIH5 |
inter-alpha-trypsin inhibitor heavy chain family, member 5 |
| chr19_+_41768401 | 0.07 |
ENST00000352456.3 ENST00000595018.1 ENST00000597725.1 |
HNRNPUL1 |
heterogeneous nuclear ribonucleoprotein U-like 1 |
| chr4_-_103749205 | 0.07 |
ENST00000508249.1 |
UBE2D3 |
ubiquitin-conjugating enzyme E2D 3 |
| chr16_-_62070305 | 0.07 |
ENST00000584337.1 |
CDH8 |
cadherin 8, type 2 |
| chr20_+_2361545 | 0.06 |
ENST00000202625.2 ENST00000381423.1 |
TGM6 |
transglutaminase 6 |
| chr10_+_126630692 | 0.06 |
ENST00000359653.4 |
ZRANB1 |
zinc finger, RAN-binding domain containing 1 |
| chr12_-_11150474 | 0.06 |
ENST00000538986.1 |
TAS2R20 |
taste receptor, type 2, member 20 |
| chr2_+_190541153 | 0.06 |
ENST00000313581.4 ENST00000438402.2 ENST00000431575.2 ENST00000281412.6 |
ANKAR |
ankyrin and armadillo repeat containing |
| chr1_-_91487013 | 0.06 |
ENST00000347275.5 ENST00000370440.1 |
ZNF644 |
zinc finger protein 644 |
| chr9_+_33290491 | 0.05 |
ENST00000379540.3 ENST00000379521.4 ENST00000318524.6 |
NFX1 |
nuclear transcription factor, X-box binding 1 |
| chr7_-_92855762 | 0.05 |
ENST00000453812.2 ENST00000394468.2 |
HEPACAM2 |
HEPACAM family member 2 |
| chr6_+_131958436 | 0.05 |
ENST00000357639.3 ENST00000543135.1 ENST00000427148.2 ENST00000358229.5 |
ENPP3 |
ectonucleotide pyrophosphatase/phosphodiesterase 3 |
| chr3_-_157221128 | 0.05 |
ENST00000392833.2 ENST00000362010.2 |
VEPH1 |
ventricular zone expressed PH domain-containing 1 |
| chr4_+_86525299 | 0.05 |
ENST00000512201.1 |
ARHGAP24 |
Rho GTPase activating protein 24 |
| chr16_-_72128270 | 0.04 |
ENST00000426362.2 |
TXNL4B |
thioredoxin-like 4B |
| chr6_-_9977801 | 0.04 |
ENST00000316020.6 ENST00000491508.1 |
OFCC1 |
orofacial cleft 1 candidate 1 |
| chr8_+_55528627 | 0.04 |
ENST00000220676.1 |
RP1 |
retinitis pigmentosa 1 (autosomal dominant) |
| chr8_+_38065104 | 0.03 |
ENST00000521311.1 |
BAG4 |
BCL2-associated athanogene 4 |
| chr4_-_103746683 | 0.03 |
ENST00000504211.1 ENST00000508476.1 |
UBE2D3 |
ubiquitin-conjugating enzyme E2D 3 |
| chr8_+_75262629 | 0.03 |
ENST00000434412.2 |
GDAP1 |
ganglioside induced differentiation associated protein 1 |
| chr5_+_140552218 | 0.03 |
ENST00000231137.3 |
PCDHB7 |
protocadherin beta 7 |
| chr1_+_153950202 | 0.02 |
ENST00000608236.1 |
RP11-422P24.11 |
RP11-422P24.11 |
| chr22_-_41682172 | 0.02 |
ENST00000356244.3 |
RANGAP1 |
Ran GTPase activating protein 1 |
| chr9_+_140125385 | 0.02 |
ENST00000361134.2 |
SLC34A3 |
solute carrier family 34 (type II sodium/phosphate contransporter), member 3 |
| chr19_-_44259053 | 0.02 |
ENST00000601170.1 |
SMG9 |
SMG9 nonsense mediated mRNA decay factor |
| chr10_-_97200772 | 0.02 |
ENST00000371241.1 ENST00000354106.3 ENST00000371239.1 ENST00000361941.3 ENST00000277982.5 ENST00000371245.3 |
SORBS1 |
sorbin and SH3 domain containing 1 |
| chr6_+_29426230 | 0.02 |
ENST00000442615.1 |
OR2H1 |
olfactory receptor, family 2, subfamily H, member 1 |
| chr1_-_154150651 | 0.02 |
ENST00000302206.5 |
TPM3 |
tropomyosin 3 |
| chr8_+_92261516 | 0.02 |
ENST00000276609.3 ENST00000309536.2 |
SLC26A7 |
solute carrier family 26 (anion exchanger), member 7 |
| chr10_-_115904361 | 0.02 |
ENST00000428953.1 ENST00000543782.1 |
C10orf118 |
chromosome 10 open reading frame 118 |
| chr3_+_238273 | 0.01 |
ENST00000256509.2 |
CHL1 |
cell adhesion molecule L1-like |
| chr5_+_76326187 | 0.01 |
ENST00000312916.7 ENST00000506806.1 |
AGGF1 |
angiogenic factor with G patch and FHA domains 1 |
| chr13_+_30510003 | 0.01 |
ENST00000400540.1 |
LINC00544 |
long intergenic non-protein coding RNA 544 |
| chr2_+_114163945 | 0.01 |
ENST00000453673.3 |
IGKV1OR2-108 |
immunoglobulin kappa variable 1/OR2-108 (non-functional) |
| chr19_-_54604083 | 0.01 |
ENST00000391761.1 ENST00000356532.3 ENST00000359649.4 ENST00000358375.4 ENST00000391760.1 ENST00000351806.4 |
OSCAR |
osteoclast associated, immunoglobulin-like receptor |
| chr3_+_179322481 | 0.01 |
ENST00000259037.3 |
NDUFB5 |
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 5, 16kDa |
| chr2_+_74154032 | 0.00 |
ENST00000356837.6 |
DGUOK |
deoxyguanosine kinase |
| chr8_-_135522425 | 0.00 |
ENST00000521673.1 |
ZFAT |
zinc finger and AT hook domain containing |
| chr1_-_63988846 | 0.00 |
ENST00000283568.8 ENST00000371092.3 ENST00000271002.10 |
ITGB3BP |
integrin beta 3 binding protein (beta3-endonexin) |
| chr1_-_197169672 | 0.00 |
ENST00000367405.4 |
ZBTB41 |
zinc finger and BTB domain containing 41 |
| chr14_+_102276209 | 0.00 |
ENST00000445439.3 ENST00000334743.5 ENST00000557095.1 |
PPP2R5C |
protein phosphatase 2, regulatory subunit B', gamma |
| chr4_+_71296204 | 0.00 |
ENST00000413702.1 |
MUC7 |
mucin 7, secreted |
| chr21_-_35884573 | 0.00 |
ENST00000399286.2 |
KCNE1 |
potassium voltage-gated channel, Isk-related family, member 1 |
| chr6_-_24666819 | 0.00 |
ENST00000341060.3 |
TDP2 |
tyrosyl-DNA phosphodiesterase 2 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 6.5 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.4 | 1.4 | GO:1900138 | negative regulation of phospholipase A2 activity(GO:1900138) |
| 0.3 | 1.3 | GO:2000230 | negative regulation of pancreatic stellate cell proliferation(GO:2000230) |
| 0.2 | 0.7 | GO:0001971 | negative regulation of activation of membrane attack complex(GO:0001971) |
| 0.2 | 2.5 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.1 | 0.4 | GO:0046379 | extracellular polysaccharide biosynthetic process(GO:0045226) extracellular polysaccharide metabolic process(GO:0046379) |
| 0.1 | 0.7 | GO:0045629 | negative regulation of T-helper 2 cell differentiation(GO:0045629) |
| 0.1 | 0.4 | GO:0014740 | negative regulation of muscle hyperplasia(GO:0014740) |
| 0.1 | 1.0 | GO:0000707 | meiotic DNA recombinase assembly(GO:0000707) |
| 0.1 | 1.0 | GO:0014052 | regulation of gamma-aminobutyric acid secretion(GO:0014052) |
| 0.1 | 0.9 | GO:0046618 | drug export(GO:0046618) |
| 0.1 | 0.5 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 0.1 | 0.6 | GO:0002414 | immunoglobulin transcytosis in epithelial cells(GO:0002414) |
| 0.1 | 0.2 | GO:0071418 | cellular response to amine stimulus(GO:0071418) |
| 0.1 | 0.5 | GO:0021564 | vagus nerve development(GO:0021564) |
| 0.1 | 0.6 | GO:0098735 | positive regulation of the force of heart contraction(GO:0098735) |
| 0.1 | 0.5 | GO:0032966 | negative regulation of collagen biosynthetic process(GO:0032966) |
| 0.1 | 0.2 | GO:0039019 | pronephric nephron development(GO:0039019) |
| 0.0 | 0.4 | GO:0007221 | positive regulation of transcription of Notch receptor target(GO:0007221) |
| 0.0 | 0.5 | GO:0046598 | positive regulation of viral entry into host cell(GO:0046598) |
| 0.0 | 0.2 | GO:0042357 | thiamine diphosphate metabolic process(GO:0042357) |
| 0.0 | 0.3 | GO:0035965 | cardiolipin acyl-chain remodeling(GO:0035965) |
| 0.0 | 0.3 | GO:0097338 | response to clozapine(GO:0097338) |
| 0.0 | 0.2 | GO:0021523 | somatic motor neuron differentiation(GO:0021523) protein localization to ciliary transition zone(GO:1904491) |
| 0.0 | 0.1 | GO:0051311 | meiotic metaphase I plate congression(GO:0043060) meiotic spindle midzone assembly(GO:0051257) meiotic metaphase plate congression(GO:0051311) |
| 0.0 | 0.1 | GO:0036071 | N-glycan fucosylation(GO:0036071) |
| 0.0 | 0.1 | GO:1902309 | negative regulation of peptidyl-serine dephosphorylation(GO:1902309) |
| 0.0 | 0.4 | GO:0046600 | negative regulation of centriole replication(GO:0046600) |
| 0.0 | 0.2 | GO:0060054 | positive regulation of epithelial cell proliferation involved in wound healing(GO:0060054) |
| 0.0 | 0.3 | GO:0060020 | Bergmann glial cell differentiation(GO:0060020) |
| 0.0 | 0.3 | GO:0019367 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.0 | 0.2 | GO:0000730 | DNA recombinase assembly(GO:0000730) double-strand break repair via synthesis-dependent strand annealing(GO:0045003) |
| 0.0 | 0.1 | GO:1901899 | dense core granule biogenesis(GO:0061110) positive regulation of relaxation of cardiac muscle(GO:1901899) regulation of dense core granule biogenesis(GO:2000705) |
| 0.0 | 0.4 | GO:0030208 | dermatan sulfate biosynthetic process(GO:0030208) |
| 0.0 | 0.1 | GO:0061484 | hematopoietic stem cell homeostasis(GO:0061484) |
| 0.0 | 0.1 | GO:1904879 | positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 0.0 | 0.1 | GO:0071947 | protein deubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0071947) |
| 0.0 | 0.2 | GO:0038028 | insulin receptor signaling pathway via phosphatidylinositol 3-kinase(GO:0038028) |
| 0.0 | 0.0 | GO:1903215 | negative regulation of protein targeting to mitochondrion(GO:1903215) |
| 0.0 | 0.1 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.0 | 0.0 | GO:0030505 | inorganic diphosphate transport(GO:0030505) |
| 0.0 | 0.4 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.0 | 0.2 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.0 | 0.1 | GO:0046606 | negative regulation of centrosome cycle(GO:0046606) |
| 0.0 | 0.2 | GO:0071670 | smooth muscle cell chemotaxis(GO:0071670) |
| 0.0 | 0.2 | GO:0071578 | zinc II ion transmembrane import(GO:0071578) |
| 0.0 | 0.7 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 0.4 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.0 | 0.5 | GO:0006915 | apoptotic process(GO:0006915) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 6.5 | GO:0071953 | elastic fiber(GO:0071953) |
| 0.2 | 2.4 | GO:0005589 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.1 | 1.0 | GO:0033063 | Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.1 | 0.3 | GO:0032173 | septin ring(GO:0005940) septin collar(GO:0032173) |
| 0.1 | 0.5 | GO:0032311 | angiogenin-PRI complex(GO:0032311) |
| 0.1 | 0.5 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) extrinsic component of endosome membrane(GO:0031313) |
| 0.0 | 0.1 | GO:0042720 | mitochondrial inner membrane peptidase complex(GO:0042720) |
| 0.0 | 0.2 | GO:0032798 | Swi5-Sfr1 complex(GO:0032798) |
| 0.0 | 1.0 | GO:0098563 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.0 | 0.1 | GO:0005715 | late recombination nodule(GO:0005715) |
| 0.0 | 0.7 | GO:0043218 | compact myelin(GO:0043218) |
| 0.0 | 0.5 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.0 | 0.2 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.0 | 0.4 | GO:0097431 | mitotic spindle pole(GO:0097431) |
| 0.0 | 0.3 | GO:0033276 | transcription factor TFTC complex(GO:0033276) |
| 0.0 | 0.2 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.0 | 0.2 | GO:0036038 | MKS complex(GO:0036038) |
| 0.0 | 0.3 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.0 | 0.4 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.9 | GO:0015307 | drug:proton antiporter activity(GO:0015307) |
| 0.1 | 0.6 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 0.1 | 1.3 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.1 | 0.4 | GO:0050501 | hyaluronan synthase activity(GO:0050501) |
| 0.1 | 0.6 | GO:0099580 | ion antiporter activity involved in regulation of postsynaptic membrane potential(GO:0099580) |
| 0.1 | 1.0 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.1 | 0.4 | GO:0047757 | chondroitin-glucuronate 5-epimerase activity(GO:0047757) |
| 0.1 | 0.4 | GO:0001010 | transcription factor activity, sequence-specific DNA binding transcription factor recruiting(GO:0001010) |
| 0.1 | 0.5 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.1 | 0.5 | GO:0019834 | phospholipase A2 inhibitor activity(GO:0019834) |
| 0.1 | 0.2 | GO:0004852 | uroporphyrinogen-III synthase activity(GO:0004852) |
| 0.0 | 0.3 | GO:1990254 | keratin filament binding(GO:1990254) |
| 0.0 | 1.0 | GO:0043274 | phospholipase binding(GO:0043274) |
| 0.0 | 2.4 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.1 | GO:0008424 | glycoprotein 6-alpha-L-fucosyltransferase activity(GO:0008424) alpha-(1->6)-fucosyltransferase activity(GO:0046921) |
| 0.0 | 6.5 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.3 | GO:0009922 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.0 | 0.6 | GO:0042923 | neuropeptide binding(GO:0042923) |
| 0.0 | 0.7 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 0.3 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.0 | 0.7 | GO:0001848 | complement binding(GO:0001848) |
| 0.0 | 0.5 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.0 | 0.7 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.0 | 0.3 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.0 | 0.1 | GO:0086056 | voltage-gated calcium channel activity involved in AV node cell action potential(GO:0086056) |
| 0.0 | 0.1 | GO:0032137 | guanine/thymine mispair binding(GO:0032137) |
| 0.0 | 0.2 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 0.0 | 0.1 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.0 | 0.3 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 2.3 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.7 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.0 | 1.3 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 6.1 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.4 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.4 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.1 | 0.9 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 0.4 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.0 | 0.4 | REACTOME HYALURONAN METABOLISM | Genes involved in Hyaluronan metabolism |
| 0.0 | 0.7 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 0.6 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.0 | 0.4 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.0 | 1.2 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.3 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |