Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
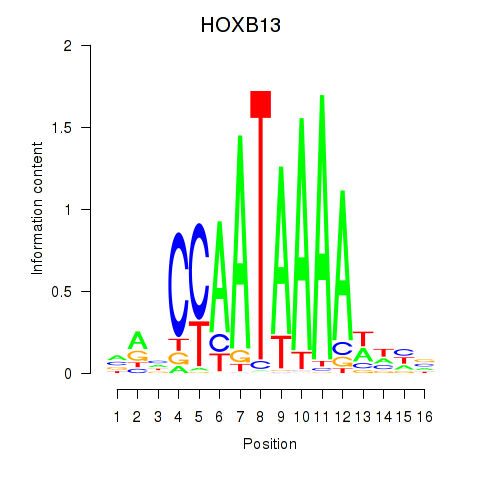
Results for HOXB13
Z-value: 0.63
Transcription factors associated with HOXB13
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HOXB13
|
ENSG00000159184.7 | HOXB13 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HOXB13 | hg19_v2_chr17_-_46806540_46806558 | 0.17 | 6.9e-01 | Click! |
Activity profile of HOXB13 motif
Sorted Z-values of HOXB13 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HOXB13
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_+_97597148 | 0.81 |
ENST00000521590.1 |
SDC2 |
syndecan 2 |
| chr1_+_78769549 | 0.74 |
ENST00000370758.1 |
PTGFR |
prostaglandin F receptor (FP) |
| chr2_-_190044480 | 0.52 |
ENST00000374866.3 |
COL5A2 |
collagen, type V, alpha 2 |
| chr8_+_77593448 | 0.31 |
ENST00000521891.2 |
ZFHX4 |
zinc finger homeobox 4 |
| chr11_-_102826434 | 0.29 |
ENST00000340273.4 ENST00000260302.3 |
MMP13 |
matrix metallopeptidase 13 (collagenase 3) |
| chr8_+_77593474 | 0.29 |
ENST00000455469.2 ENST00000050961.6 |
ZFHX4 |
zinc finger homeobox 4 |
| chr3_-_195310802 | 0.27 |
ENST00000421243.1 ENST00000453131.1 |
APOD |
apolipoprotein D |
| chr3_-_114790179 | 0.23 |
ENST00000462705.1 |
ZBTB20 |
zinc finger and BTB domain containing 20 |
| chr7_-_50860565 | 0.20 |
ENST00000403097.1 |
GRB10 |
growth factor receptor-bound protein 10 |
| chr14_-_51027838 | 0.19 |
ENST00000555216.1 |
MAP4K5 |
mitogen-activated protein kinase kinase kinase kinase 5 |
| chr5_+_135496675 | 0.18 |
ENST00000507637.1 |
SMAD5 |
SMAD family member 5 |
| chr7_-_130353553 | 0.15 |
ENST00000330992.7 ENST00000445977.2 |
COPG2 |
coatomer protein complex, subunit gamma 2 |
| chr17_+_18086392 | 0.13 |
ENST00000541285.1 |
ALKBH5 |
alkB, alkylation repair homolog 5 (E. coli) |
| chr9_+_125133315 | 0.12 |
ENST00000223423.4 ENST00000362012.2 |
PTGS1 |
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) |
| chr1_+_84630645 | 0.11 |
ENST00000394839.2 |
PRKACB |
protein kinase, cAMP-dependent, catalytic, beta |
| chr9_-_130829588 | 0.11 |
ENST00000373078.4 |
NAIF1 |
nuclear apoptosis inducing factor 1 |
| chr1_+_210501589 | 0.11 |
ENST00000413764.2 ENST00000541565.1 |
HHAT |
hedgehog acyltransferase |
| chr2_+_74685413 | 0.10 |
ENST00000233615.2 |
WBP1 |
WW domain binding protein 1 |
| chr12_+_8309630 | 0.09 |
ENST00000396570.3 |
ZNF705A |
zinc finger protein 705A |
| chr2_+_74685527 | 0.08 |
ENST00000393972.3 ENST00000409737.1 ENST00000428943.1 |
WBP1 |
WW domain binding protein 1 |
| chr16_-_24942273 | 0.08 |
ENST00000571406.1 |
ARHGAP17 |
Rho GTPase activating protein 17 |
| chr16_-_24942411 | 0.08 |
ENST00000571843.1 |
ARHGAP17 |
Rho GTPase activating protein 17 |
| chr1_+_52682052 | 0.08 |
ENST00000371591.1 |
ZFYVE9 |
zinc finger, FYVE domain containing 9 |
| chr1_-_155880672 | 0.07 |
ENST00000609492.1 ENST00000368322.3 |
RIT1 |
Ras-like without CAAX 1 |
| chr6_-_43021612 | 0.07 |
ENST00000535468.1 |
CUL7 |
cullin 7 |
| chr1_+_23037323 | 0.07 |
ENST00000544305.1 ENST00000374630.3 ENST00000400191.3 ENST00000374632.3 |
EPHB2 |
EPH receptor B2 |
| chr5_-_145214893 | 0.07 |
ENST00000394450.2 |
PRELID2 |
PRELI domain containing 2 |
| chr3_-_123710199 | 0.06 |
ENST00000184183.4 |
ROPN1 |
rhophilin associated tail protein 1 |
| chrM_-_14670 | 0.06 |
ENST00000361681.2 |
MT-ND6 |
mitochondrially encoded NADH dehydrogenase 6 |
| chr2_-_175629135 | 0.06 |
ENST00000409542.1 ENST00000409219.1 |
CHRNA1 |
cholinergic receptor, nicotinic, alpha 1 (muscle) |
| chr2_-_136288113 | 0.06 |
ENST00000401392.1 |
ZRANB3 |
zinc finger, RAN-binding domain containing 3 |
| chr17_-_46806540 | 0.06 |
ENST00000290295.7 |
HOXB13 |
homeobox B13 |
| chr7_+_132937820 | 0.06 |
ENST00000393161.2 ENST00000253861.4 |
EXOC4 |
exocyst complex component 4 |
| chr21_-_35899113 | 0.05 |
ENST00000492600.1 ENST00000481448.1 ENST00000381132.2 |
RCAN1 |
regulator of calcineurin 1 |
| chr5_-_140053152 | 0.05 |
ENST00000542735.1 |
DND1 |
DND microRNA-mediated repression inhibitor 1 |
| chr11_-_102714534 | 0.05 |
ENST00000299855.5 |
MMP3 |
matrix metallopeptidase 3 (stromelysin 1, progelatinase) |
| chr3_+_89156799 | 0.05 |
ENST00000452448.2 ENST00000494014.1 |
EPHA3 |
EPH receptor A3 |
| chr1_+_84630053 | 0.05 |
ENST00000394838.2 ENST00000370682.3 ENST00000432111.1 |
PRKACB |
protein kinase, cAMP-dependent, catalytic, beta |
| chr1_-_36863481 | 0.04 |
ENST00000315732.2 |
LSM10 |
LSM10, U7 small nuclear RNA associated |
| chr6_-_27782548 | 0.04 |
ENST00000333151.3 |
HIST1H2AJ |
histone cluster 1, H2aj |
| chr19_-_5903714 | 0.04 |
ENST00000586349.1 ENST00000585661.1 ENST00000308961.4 ENST00000592634.1 ENST00000418389.2 ENST00000252675.5 |
AC024592.12 NDUFA11 FUT5 |
Uncharacterized protein NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 11, 14.7kDa fucosyltransferase 5 (alpha (1,3) fucosyltransferase) |
| chr19_+_48497901 | 0.04 |
ENST00000339841.2 |
ELSPBP1 |
epididymal sperm binding protein 1 |
| chr19_+_48497962 | 0.04 |
ENST00000596043.1 ENST00000597519.1 |
ELSPBP1 |
epididymal sperm binding protein 1 |
| chr2_-_176046391 | 0.04 |
ENST00000392541.3 ENST00000409194.1 |
ATP5G3 |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit C3 (subunit 9) |
| chr22_+_35462129 | 0.04 |
ENST00000404699.2 ENST00000308700.6 |
ISX |
intestine-specific homeobox |
| chr16_-_80926457 | 0.04 |
ENST00000563626.1 ENST00000562231.1 |
RP11-314O13.1 |
RP11-314O13.1 |
| chr2_+_67624430 | 0.04 |
ENST00000272342.5 |
ETAA1 |
Ewing tumor-associated antigen 1 |
| chr15_-_55541227 | 0.03 |
ENST00000566877.1 |
RAB27A |
RAB27A, member RAS oncogene family |
| chr18_+_3252265 | 0.03 |
ENST00000580887.1 ENST00000536605.1 |
MYL12A |
myosin, light chain 12A, regulatory, non-sarcomeric |
| chr3_+_89156674 | 0.03 |
ENST00000336596.2 |
EPHA3 |
EPH receptor A3 |
| chr12_-_81992111 | 0.03 |
ENST00000443686.3 ENST00000407050.4 |
PPFIA2 |
protein tyrosine phosphatase, receptor type, f polypeptide (PTPRF), interacting protein (liprin), alpha 2 |
| chr9_+_125132803 | 0.03 |
ENST00000540753.1 |
PTGS1 |
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) |
| chr5_-_145214848 | 0.03 |
ENST00000505416.1 ENST00000334744.4 ENST00000358004.2 ENST00000511435.1 |
PRELID2 |
PRELI domain containing 2 |
| chr9_+_103947311 | 0.03 |
ENST00000395056.2 |
LPPR1 |
Lipid phosphate phosphatase-related protein type 1 |
| chr6_+_122793058 | 0.03 |
ENST00000392491.2 |
PKIB |
protein kinase (cAMP-dependent, catalytic) inhibitor beta |
| chr1_-_45956822 | 0.02 |
ENST00000372086.3 ENST00000341771.6 |
TESK2 |
testis-specific kinase 2 |
| chr4_+_68424434 | 0.02 |
ENST00000265404.2 ENST00000396225.1 |
STAP1 |
signal transducing adaptor family member 1 |
| chr12_-_52715179 | 0.02 |
ENST00000293670.3 |
KRT83 |
keratin 83 |
| chr16_-_71610985 | 0.02 |
ENST00000355962.4 |
TAT |
tyrosine aminotransferase |
| chr2_-_175629164 | 0.02 |
ENST00000409323.1 ENST00000261007.5 ENST00000348749.5 |
CHRNA1 |
cholinergic receptor, nicotinic, alpha 1 (muscle) |
| chr4_+_110749143 | 0.02 |
ENST00000317735.4 |
RRH |
retinal pigment epithelium-derived rhodopsin homolog |
| chr6_+_27782788 | 0.02 |
ENST00000359465.4 |
HIST1H2BM |
histone cluster 1, H2bm |
| chr6_-_116833500 | 0.01 |
ENST00000356128.4 |
TRAPPC3L |
trafficking protein particle complex 3-like |
| chr1_-_45956800 | 0.01 |
ENST00000538496.1 |
TESK2 |
testis-specific kinase 2 |
| chr1_-_151148492 | 0.01 |
ENST00000295314.4 |
TMOD4 |
tropomodulin 4 (muscle) |
| chr18_+_3252206 | 0.01 |
ENST00000578562.2 |
MYL12A |
myosin, light chain 12A, regulatory, non-sarcomeric |
| chr3_-_150690786 | 0.01 |
ENST00000327047.1 |
CLRN1 |
clarin 1 |
| chr14_-_68000442 | 0.00 |
ENST00000554278.1 |
TMEM229B |
transmembrane protein 229B |
| chr22_-_38240412 | 0.00 |
ENST00000215941.4 |
ANKRD54 |
ankyrin repeat domain 54 |
| chr8_+_119294456 | 0.00 |
ENST00000366457.2 |
AC023590.1 |
Uncharacterized protein |
| chr4_-_69111401 | 0.00 |
ENST00000332644.5 |
TMPRSS11B |
transmembrane protease, serine 11B |
| chr12_+_70219052 | 0.00 |
ENST00000552032.2 ENST00000547771.2 |
MYRFL |
myelin regulatory factor-like |
| chr15_+_66994561 | 0.00 |
ENST00000288840.5 |
SMAD6 |
SMAD family member 6 |
| chr19_+_35634146 | 0.00 |
ENST00000586063.1 ENST00000270310.2 ENST00000588265.1 |
FXYD7 |
FXYD domain containing ion transport regulator 7 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.1 | 0.8 | GO:0008218 | bioluminescence(GO:0008218) |
| 0.1 | 0.3 | GO:2000097 | regulation of smooth muscle cell-matrix adhesion(GO:2000097) |
| 0.1 | 0.7 | GO:0071798 | response to prostaglandin D(GO:0071798) cellular response to prostaglandin D stimulus(GO:0071799) |
| 0.0 | 0.1 | GO:0035553 | oxidative RNA demethylation(GO:0035513) oxidative single-stranded RNA demethylation(GO:0035553) |
| 0.0 | 0.2 | GO:0001880 | Mullerian duct regression(GO:0001880) |
| 0.0 | 0.1 | GO:0097156 | fasciculation of motor neuron axon(GO:0097156) |
| 0.0 | 0.1 | GO:0036292 | DNA rewinding(GO:0036292) |
| 0.0 | 0.1 | GO:0060743 | epithelial cell maturation involved in prostate gland development(GO:0060743) |
| 0.0 | 0.0 | GO:1904479 | negative regulation of intestinal absorption(GO:1904479) |
| 0.0 | 0.2 | GO:0046325 | negative regulation of glucose import(GO:0046325) |
| 0.0 | 0.3 | GO:0003417 | growth plate cartilage development(GO:0003417) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.0 | 0.2 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 0.0 | 0.1 | GO:1990393 | 3M complex(GO:1990393) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0004958 | prostaglandin F receptor activity(GO:0004958) |
| 0.0 | 0.2 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.0 | 0.1 | GO:0035515 | oxidative RNA demethylase activity(GO:0035515) |
| 0.0 | 0.1 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.0 | 0.2 | GO:0030618 | transforming growth factor beta receptor, pathway-specific cytoplasmic mediator activity(GO:0030618) |
| 0.0 | 0.0 | GO:0071209 | U7 snRNA binding(GO:0071209) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.9 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | REACTOME PROSTANOID LIGAND RECEPTORS | Genes involved in Prostanoid ligand receptors |
| 0.0 | 0.8 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |