Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
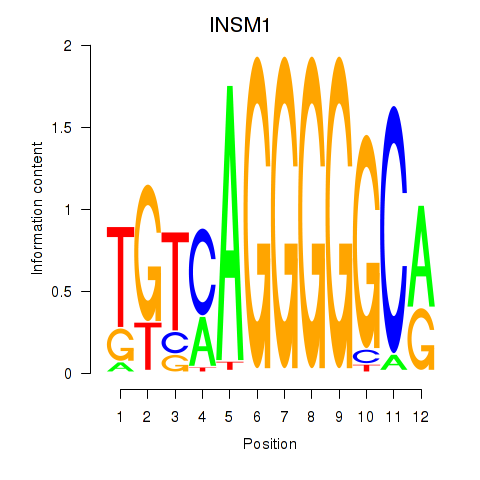
Results for INSM1
Z-value: 1.12
Transcription factors associated with INSM1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
INSM1
|
ENSG00000173404.3 | INSM1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| INSM1 | hg19_v2_chr20_+_20348740_20348765 | 0.24 | 5.7e-01 | Click! |
Activity profile of INSM1 motif
Sorted Z-values of INSM1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of INSM1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_-_52887034 | 1.46 |
ENST00000330722.6 |
KRT6A |
keratin 6A |
| chr3_+_50192537 | 0.96 |
ENST00000002829.3 |
SEMA3F |
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3F |
| chr3_+_50192499 | 0.94 |
ENST00000413852.1 |
SEMA3F |
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3F |
| chr1_+_160370344 | 0.78 |
ENST00000368061.2 |
VANGL2 |
VANGL planar cell polarity protein 2 |
| chr11_+_124932986 | 0.68 |
ENST00000407458.1 ENST00000298280.5 |
SLC37A2 |
solute carrier family 37 (glucose-6-phosphate transporter), member 2 |
| chr11_+_124932955 | 0.68 |
ENST00000403796.2 |
SLC37A2 |
solute carrier family 37 (glucose-6-phosphate transporter), member 2 |
| chr11_+_124933191 | 0.68 |
ENST00000532000.1 ENST00000308074.4 |
SLC37A2 |
solute carrier family 37 (glucose-6-phosphate transporter), member 2 |
| chr5_-_60140009 | 0.68 |
ENST00000505959.1 |
ELOVL7 |
ELOVL fatty acid elongase 7 |
| chr5_-_60140089 | 0.63 |
ENST00000507047.1 ENST00000438340.1 ENST00000425382.1 ENST00000508821.1 |
ELOVL7 |
ELOVL fatty acid elongase 7 |
| chr7_+_86273700 | 0.60 |
ENST00000546348.1 |
GRM3 |
glutamate receptor, metabotropic 3 |
| chr7_+_86273952 | 0.59 |
ENST00000536043.1 |
GRM3 |
glutamate receptor, metabotropic 3 |
| chr7_+_86274145 | 0.58 |
ENST00000439827.1 ENST00000394720.2 ENST00000421579.1 |
GRM3 |
glutamate receptor, metabotropic 3 |
| chr12_+_49372251 | 0.55 |
ENST00000293549.3 |
WNT1 |
wingless-type MMTV integration site family, member 1 |
| chr2_-_72375167 | 0.52 |
ENST00000001146.2 |
CYP26B1 |
cytochrome P450, family 26, subfamily B, polypeptide 1 |
| chr15_+_45722727 | 0.50 |
ENST00000396650.2 ENST00000558435.1 ENST00000344300.3 |
C15orf48 |
chromosome 15 open reading frame 48 |
| chr1_+_65613340 | 0.47 |
ENST00000546702.1 |
AK4 |
adenylate kinase 4 |
| chr1_+_65613217 | 0.46 |
ENST00000545314.1 |
AK4 |
adenylate kinase 4 |
| chr2_-_74667612 | 0.45 |
ENST00000305557.5 ENST00000233330.6 |
RTKN |
rhotekin |
| chr5_-_175964366 | 0.42 |
ENST00000274811.4 |
RNF44 |
ring finger protein 44 |
| chr12_+_70760056 | 0.39 |
ENST00000258111.4 |
KCNMB4 |
potassium large conductance calcium-activated channel, subfamily M, beta member 4 |
| chr12_-_54779511 | 0.36 |
ENST00000551109.1 ENST00000546970.1 |
ZNF385A |
zinc finger protein 385A |
| chr17_-_27507377 | 0.34 |
ENST00000531253.1 |
MYO18A |
myosin XVIIIA |
| chr1_+_153750622 | 0.33 |
ENST00000532853.1 |
SLC27A3 |
solute carrier family 27 (fatty acid transporter), member 3 |
| chr15_+_89181974 | 0.33 |
ENST00000306072.5 |
ISG20 |
interferon stimulated exonuclease gene 20kDa |
| chr3_+_10857885 | 0.32 |
ENST00000254488.2 ENST00000454147.1 |
SLC6A11 |
solute carrier family 6 (neurotransmitter transporter), member 11 |
| chr17_-_27507395 | 0.32 |
ENST00000354329.4 ENST00000527372.1 |
MYO18A |
myosin XVIIIA |
| chr15_+_89182178 | 0.31 |
ENST00000559876.1 |
ISG20 |
interferon stimulated exonuclease gene 20kDa |
| chrX_+_16804544 | 0.31 |
ENST00000380122.5 ENST00000398155.4 |
TXLNG |
taxilin gamma |
| chr8_-_144660485 | 0.27 |
ENST00000276844.7 ENST00000340490.3 ENST00000435154.3 ENST00000426292.3 |
NAPRT1 |
nicotinate phosphoribosyltransferase domain containing 1 |
| chr4_-_143767428 | 0.27 |
ENST00000513000.1 ENST00000509777.1 ENST00000503927.1 |
INPP4B |
inositol polyphosphate-4-phosphatase, type II, 105kDa |
| chr4_-_120548779 | 0.25 |
ENST00000264805.5 |
PDE5A |
phosphodiesterase 5A, cGMP-specific |
| chr1_-_23886285 | 0.25 |
ENST00000374561.5 |
ID3 |
inhibitor of DNA binding 3, dominant negative helix-loop-helix protein |
| chr19_-_55881741 | 0.24 |
ENST00000264563.2 ENST00000590625.1 ENST00000585513.1 |
IL11 |
interleukin 11 |
| chr14_+_73563735 | 0.24 |
ENST00000532192.1 |
RBM25 |
RNA binding motif protein 25 |
| chr1_-_1051736 | 0.23 |
ENST00000448924.1 ENST00000294576.5 ENST00000437760.1 ENST00000462097.1 ENST00000475119.1 |
C1orf159 |
chromosome 1 open reading frame 159 |
| chr2_-_73511559 | 0.23 |
ENST00000521871.1 |
FBXO41 |
F-box protein 41 |
| chr6_-_9933500 | 0.23 |
ENST00000492169.1 |
OFCC1 |
orofacial cleft 1 candidate 1 |
| chr1_-_1051455 | 0.23 |
ENST00000379339.1 ENST00000480643.1 ENST00000434641.1 ENST00000421241.2 |
C1orf159 |
chromosome 1 open reading frame 159 |
| chr14_+_76776957 | 0.23 |
ENST00000512784.1 |
ESRRB |
estrogen-related receptor beta |
| chr19_-_38916839 | 0.22 |
ENST00000433821.2 ENST00000426920.2 ENST00000587753.1 ENST00000454404.2 ENST00000293062.9 |
RASGRP4 |
RAS guanyl releasing protein 4 |
| chr2_-_27435125 | 0.22 |
ENST00000414408.1 ENST00000310574.3 |
SLC5A6 |
solute carrier family 5 (sodium/multivitamin and iodide cotransporter), member 6 |
| chr19_-_38916822 | 0.22 |
ENST00000586305.1 |
RASGRP4 |
RAS guanyl releasing protein 4 |
| chr1_-_6557156 | 0.22 |
ENST00000537245.1 |
PLEKHG5 |
pleckstrin homology domain containing, family G (with RhoGef domain) member 5 |
| chr4_-_151936416 | 0.22 |
ENST00000510413.1 ENST00000507224.1 |
LRBA |
LPS-responsive vesicle trafficking, beach and anchor containing |
| chrX_+_152907913 | 0.21 |
ENST00000370167.4 |
DUSP9 |
dual specificity phosphatase 9 |
| chr15_+_89182156 | 0.21 |
ENST00000379224.5 |
ISG20 |
interferon stimulated exonuclease gene 20kDa |
| chr21_+_46825032 | 0.20 |
ENST00000400337.2 |
COL18A1 |
collagen, type XVIII, alpha 1 |
| chr13_-_31736027 | 0.20 |
ENST00000380406.5 ENST00000320027.5 ENST00000380405.4 |
HSPH1 |
heat shock 105kDa/110kDa protein 1 |
| chr2_+_153191706 | 0.20 |
ENST00000288670.9 |
FMNL2 |
formin-like 2 |
| chr19_+_45394477 | 0.20 |
ENST00000252487.5 ENST00000405636.2 ENST00000592434.1 ENST00000426677.2 ENST00000589649.1 |
TOMM40 |
translocase of outer mitochondrial membrane 40 homolog (yeast) |
| chr3_+_14989186 | 0.20 |
ENST00000435454.1 ENST00000323373.6 |
NR2C2 |
nuclear receptor subfamily 2, group C, member 2 |
| chr8_-_145550571 | 0.18 |
ENST00000332324.4 |
DGAT1 |
diacylglycerol O-acyltransferase 1 |
| chr13_-_31736132 | 0.18 |
ENST00000429785.2 |
HSPH1 |
heat shock 105kDa/110kDa protein 1 |
| chr14_-_21566731 | 0.18 |
ENST00000360947.3 |
ZNF219 |
zinc finger protein 219 |
| chr10_-_103603523 | 0.18 |
ENST00000370046.1 |
KCNIP2 |
Kv channel interacting protein 2 |
| chr4_-_151936865 | 0.17 |
ENST00000535741.1 |
LRBA |
LPS-responsive vesicle trafficking, beach and anchor containing |
| chr19_+_14142535 | 0.17 |
ENST00000263379.2 |
IL27RA |
interleukin 27 receptor, alpha |
| chr19_-_38916802 | 0.17 |
ENST00000587738.1 |
RASGRP4 |
RAS guanyl releasing protein 4 |
| chr6_-_94129244 | 0.16 |
ENST00000369303.4 ENST00000369297.1 |
EPHA7 |
EPH receptor A7 |
| chr15_+_68924327 | 0.16 |
ENST00000543950.1 |
CORO2B |
coronin, actin binding protein, 2B |
| chr11_+_66610883 | 0.16 |
ENST00000309657.3 ENST00000524506.1 |
RCE1 |
Ras converting CAAX endopeptidase 1 |
| chr1_-_6321035 | 0.15 |
ENST00000377893.2 |
GPR153 |
G protein-coupled receptor 153 |
| chr17_-_46667594 | 0.15 |
ENST00000476342.1 ENST00000460160.1 ENST00000472863.1 |
HOXB3 |
homeobox B3 |
| chr17_-_46667628 | 0.15 |
ENST00000498678.1 |
HOXB3 |
homeobox B3 |
| chr4_-_75719896 | 0.15 |
ENST00000395743.3 |
BTC |
betacellulin |
| chr17_+_7211280 | 0.15 |
ENST00000419711.2 ENST00000571955.1 ENST00000573714.1 |
EIF5A |
eukaryotic translation initiation factor 5A |
| chr13_+_47127293 | 0.14 |
ENST00000311191.6 |
LRCH1 |
leucine-rich repeats and calponin homology (CH) domain containing 1 |
| chr9_-_34637718 | 0.14 |
ENST00000378892.1 ENST00000277010.4 |
SIGMAR1 |
sigma non-opioid intracellular receptor 1 |
| chr2_+_32582086 | 0.13 |
ENST00000421745.2 |
BIRC6 |
baculoviral IAP repeat containing 6 |
| chr6_-_146285455 | 0.13 |
ENST00000367505.2 |
SHPRH |
SNF2 histone linker PHD RING helicase, E3 ubiquitin protein ligase |
| chr6_-_146285221 | 0.13 |
ENST00000367503.3 ENST00000438092.2 ENST00000275233.7 |
SHPRH |
SNF2 histone linker PHD RING helicase, E3 ubiquitin protein ligase |
| chr7_-_105752971 | 0.13 |
ENST00000011473.2 |
SYPL1 |
synaptophysin-like 1 |
| chr15_+_90118685 | 0.13 |
ENST00000268138.7 |
TICRR |
TOPBP1-interacting checkpoint and replication regulator |
| chr9_-_140115775 | 0.13 |
ENST00000391553.1 ENST00000392827.1 |
RNF208 |
ring finger protein 208 |
| chr22_-_20104700 | 0.12 |
ENST00000439169.2 ENST00000445045.1 ENST00000404751.3 ENST00000252136.7 ENST00000403707.3 |
TRMT2A |
tRNA methyltransferase 2 homolog A (S. cerevisiae) |
| chr16_-_23568651 | 0.12 |
ENST00000563232.1 ENST00000563459.1 ENST00000449606.1 |
EARS2 |
glutamyl-tRNA synthetase 2, mitochondrial |
| chr11_-_57092381 | 0.12 |
ENST00000358252.3 |
TNKS1BP1 |
tankyrase 1 binding protein 1, 182kDa |
| chr7_-_105752651 | 0.11 |
ENST00000470347.1 ENST00000455385.2 |
SYPL1 |
synaptophysin-like 1 |
| chr14_-_104313824 | 0.11 |
ENST00000553739.1 ENST00000202556.9 |
PPP1R13B |
protein phosphatase 1, regulatory subunit 13B |
| chr1_+_210001309 | 0.11 |
ENST00000491415.2 |
DIEXF |
digestive organ expansion factor homolog (zebrafish) |
| chr3_-_69062764 | 0.11 |
ENST00000295571.5 |
EOGT |
EGF domain-specific O-linked N-acetylglucosamine (GlcNAc) transferase |
| chr4_+_56815102 | 0.11 |
ENST00000257287.4 |
CEP135 |
centrosomal protein 135kDa |
| chrX_+_69674943 | 0.11 |
ENST00000542398.1 |
DLG3 |
discs, large homolog 3 (Drosophila) |
| chr17_-_7297833 | 0.11 |
ENST00000571802.1 ENST00000576201.1 ENST00000573213.1 ENST00000324822.11 |
TMEM256-PLSCR3 |
TMEM256-PLSCR3 readthrough (NMD candidate) |
| chr7_-_91509986 | 0.11 |
ENST00000456229.1 ENST00000442961.1 ENST00000406735.2 ENST00000419292.1 ENST00000351870.3 |
MTERF |
mitochondrial transcription termination factor |
| chr17_-_42992856 | 0.10 |
ENST00000588316.1 ENST00000435360.2 ENST00000586793.1 ENST00000588735.1 ENST00000588037.1 ENST00000592320.1 ENST00000253408.5 |
GFAP |
glial fibrillary acidic protein |
| chr3_+_46924829 | 0.10 |
ENST00000313049.5 |
PTH1R |
parathyroid hormone 1 receptor |
| chr9_-_37034028 | 0.10 |
ENST00000520281.1 ENST00000446742.1 ENST00000522003.1 ENST00000523145.1 ENST00000414447.1 ENST00000377847.2 ENST00000377853.2 ENST00000377852.2 ENST00000523241.1 ENST00000520154.1 ENST00000358127.4 |
PAX5 |
paired box 5 |
| chr1_+_154474689 | 0.10 |
ENST00000368482.4 |
TDRD10 |
tudor domain containing 10 |
| chr12_+_7342178 | 0.10 |
ENST00000266563.5 ENST00000543974.1 |
PEX5 |
peroxisomal biogenesis factor 5 |
| chr9_-_34637806 | 0.10 |
ENST00000477726.1 |
SIGMAR1 |
sigma non-opioid intracellular receptor 1 |
| chr2_-_159313214 | 0.09 |
ENST00000409889.1 ENST00000283233.5 ENST00000536771.1 |
CCDC148 |
coiled-coil domain containing 148 |
| chr15_+_90118723 | 0.09 |
ENST00000560985.1 |
TICRR |
TOPBP1-interacting checkpoint and replication regulator |
| chr8_+_99076750 | 0.09 |
ENST00000545282.1 |
C8orf47 |
chromosome 8 open reading frame 47 |
| chr12_+_58148842 | 0.09 |
ENST00000266643.5 |
MARCH9 |
membrane-associated ring finger (C3HC4) 9 |
| chr11_+_46383121 | 0.09 |
ENST00000454345.1 |
DGKZ |
diacylglycerol kinase, zeta |
| chr4_-_147443043 | 0.09 |
ENST00000394059.4 ENST00000502607.1 ENST00000335472.7 ENST00000432059.2 ENST00000394062.3 |
SLC10A7 |
solute carrier family 10, member 7 |
| chr12_-_118810688 | 0.09 |
ENST00000542532.1 ENST00000392533.3 |
TAOK3 |
TAO kinase 3 |
| chr12_-_51419924 | 0.09 |
ENST00000541174.2 |
SLC11A2 |
solute carrier family 11 (proton-coupled divalent metal ion transporter), member 2 |
| chr1_-_85725316 | 0.09 |
ENST00000344356.5 ENST00000471115.1 |
C1orf52 |
chromosome 1 open reading frame 52 |
| chr12_-_51420108 | 0.09 |
ENST00000547198.1 |
SLC11A2 |
solute carrier family 11 (proton-coupled divalent metal ion transporter), member 2 |
| chr19_+_42817450 | 0.08 |
ENST00000301204.3 |
TMEM145 |
transmembrane protein 145 |
| chrX_+_146993534 | 0.08 |
ENST00000334557.6 ENST00000439526.2 ENST00000370475.4 |
FMR1 |
fragile X mental retardation 1 |
| chr18_-_51750948 | 0.08 |
ENST00000583046.1 ENST00000398398.2 |
MBD2 |
methyl-CpG binding domain protein 2 |
| chr6_-_17706618 | 0.08 |
ENST00000262077.2 ENST00000537253.1 |
NUP153 |
nucleoporin 153kDa |
| chr1_-_38230738 | 0.08 |
ENST00000427468.2 ENST00000373048.4 ENST00000319637.6 |
EPHA10 |
EPH receptor A10 |
| chr1_-_99470368 | 0.08 |
ENST00000263177.4 |
LPPR5 |
Lipid phosphate phosphatase-related protein type 5 |
| chr6_-_41863098 | 0.08 |
ENST00000373006.1 |
USP49 |
ubiquitin specific peptidase 49 |
| chr13_-_36705425 | 0.08 |
ENST00000255448.4 ENST00000360631.3 ENST00000379892.4 |
DCLK1 |
doublecortin-like kinase 1 |
| chrX_+_153059608 | 0.08 |
ENST00000370087.1 |
SSR4 |
signal sequence receptor, delta |
| chr10_+_88516396 | 0.08 |
ENST00000372037.3 |
BMPR1A |
bone morphogenetic protein receptor, type IA |
| chr11_-_118972575 | 0.07 |
ENST00000432443.2 |
DPAGT1 |
dolichyl-phosphate (UDP-N-acetylglucosamine) N-acetylglucosaminephosphotransferase 1 (GlcNAc-1-P transferase) |
| chrX_+_146993449 | 0.07 |
ENST00000218200.8 ENST00000370471.3 ENST00000370477.1 |
FMR1 |
fragile X mental retardation 1 |
| chr12_-_51420128 | 0.07 |
ENST00000262051.7 ENST00000547732.1 ENST00000262052.5 ENST00000546488.1 ENST00000550714.1 ENST00000548193.1 ENST00000547579.1 ENST00000546743.1 |
SLC11A2 |
solute carrier family 11 (proton-coupled divalent metal ion transporter), member 2 |
| chr12_-_72057571 | 0.07 |
ENST00000548100.1 |
ZFC3H1 |
zinc finger, C3H1-type containing |
| chr17_+_37793318 | 0.07 |
ENST00000336308.5 |
STARD3 |
StAR-related lipid transfer (START) domain containing 3 |
| chr14_-_85996332 | 0.07 |
ENST00000380722.1 |
RP11-497E19.1 |
RP11-497E19.1 |
| chr1_-_36851475 | 0.07 |
ENST00000373129.3 |
STK40 |
serine/threonine kinase 40 |
| chr1_-_36851489 | 0.07 |
ENST00000373130.3 ENST00000373132.3 |
STK40 |
serine/threonine kinase 40 |
| chr12_+_7342271 | 0.07 |
ENST00000434354.2 ENST00000544456.1 ENST00000545574.1 ENST00000420616.2 |
PEX5 |
peroxisomal biogenesis factor 5 |
| chr17_+_37793378 | 0.07 |
ENST00000544210.2 ENST00000581894.1 ENST00000394250.4 ENST00000579479.1 ENST00000577248.1 ENST00000580611.1 |
STARD3 |
StAR-related lipid transfer (START) domain containing 3 |
| chr12_+_7342441 | 0.06 |
ENST00000412720.2 ENST00000396637.3 |
PEX5 |
peroxisomal biogenesis factor 5 |
| chr17_-_46623441 | 0.06 |
ENST00000330070.4 |
HOXB2 |
homeobox B2 |
| chr1_+_230202936 | 0.06 |
ENST00000366672.4 |
GALNT2 |
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 2 (GalNAc-T2) |
| chr17_-_7297519 | 0.06 |
ENST00000576362.1 ENST00000571078.1 |
TMEM256-PLSCR3 |
TMEM256-PLSCR3 readthrough (NMD candidate) |
| chr17_+_19912640 | 0.06 |
ENST00000395527.4 ENST00000583482.2 ENST00000583528.1 ENST00000583463.1 |
SPECC1 |
sperm antigen with calponin homology and coiled-coil domains 1 |
| chr14_+_85996471 | 0.06 |
ENST00000330753.4 |
FLRT2 |
fibronectin leucine rich transmembrane protein 2 |
| chr12_+_53645870 | 0.06 |
ENST00000329548.4 |
MFSD5 |
major facilitator superfamily domain containing 5 |
| chr1_-_99470558 | 0.06 |
ENST00000370188.3 |
LPPR5 |
Lipid phosphate phosphatase-related protein type 5 |
| chr8_-_124286495 | 0.06 |
ENST00000297857.2 |
ZHX1 |
zinc fingers and homeoboxes 1 |
| chr10_+_91061712 | 0.06 |
ENST00000371826.3 |
IFIT2 |
interferon-induced protein with tetratricopeptide repeats 2 |
| chr16_-_19896220 | 0.06 |
ENST00000562469.1 ENST00000300571.2 |
GPRC5B |
G protein-coupled receptor, family C, group 5, member B |
| chr13_-_52027134 | 0.06 |
ENST00000311234.4 ENST00000425000.1 ENST00000463928.1 ENST00000442263.3 ENST00000398119.2 |
INTS6 |
integrator complex subunit 6 |
| chr17_-_56606664 | 0.06 |
ENST00000580844.1 |
SEPT4 |
septin 4 |
| chr9_-_130517522 | 0.05 |
ENST00000373274.3 ENST00000420366.1 |
SH2D3C |
SH2 domain containing 3C |
| chr9_-_127533519 | 0.05 |
ENST00000487099.2 ENST00000344523.4 ENST00000373584.3 |
NR6A1 |
nuclear receptor subfamily 6, group A, member 1 |
| chr17_-_56606705 | 0.05 |
ENST00000317268.3 |
SEPT4 |
septin 4 |
| chr10_-_103603568 | 0.05 |
ENST00000356640.2 |
KCNIP2 |
Kv channel interacting protein 2 |
| chr11_-_47400062 | 0.05 |
ENST00000533030.1 |
SPI1 |
spleen focus forming virus (SFFV) proviral integration oncogene |
| chr1_-_154531095 | 0.05 |
ENST00000292211.4 |
UBE2Q1 |
ubiquitin-conjugating enzyme E2Q family member 1 |
| chr10_-_104001231 | 0.05 |
ENST00000370002.3 |
PITX3 |
paired-like homeodomain 3 |
| chr21_-_46707793 | 0.05 |
ENST00000331343.7 ENST00000349485.5 |
POFUT2 |
protein O-fucosyltransferase 2 |
| chrX_+_153060090 | 0.05 |
ENST00000370086.3 ENST00000370085.3 |
SSR4 |
signal sequence receptor, delta |
| chr7_-_128695147 | 0.05 |
ENST00000482320.1 ENST00000393245.1 ENST00000471234.1 |
TNPO3 |
transportin 3 |
| chrX_-_153095945 | 0.05 |
ENST00000164640.4 |
PDZD4 |
PDZ domain containing 4 |
| chr17_-_56606639 | 0.05 |
ENST00000579371.1 |
SEPT4 |
septin 4 |
| chr1_-_160040038 | 0.05 |
ENST00000368089.3 |
KCNJ10 |
potassium inwardly-rectifying channel, subfamily J, member 10 |
| chr12_+_7341759 | 0.05 |
ENST00000455147.2 ENST00000540398.1 |
PEX5 |
peroxisomal biogenesis factor 5 |
| chr10_-_103603677 | 0.05 |
ENST00000358038.3 |
KCNIP2 |
Kv channel interacting protein 2 |
| chr15_-_65426174 | 0.05 |
ENST00000204549.4 |
PDCD7 |
programmed cell death 7 |
| chr5_+_60933634 | 0.04 |
ENST00000505642.1 |
C5orf64 |
chromosome 5 open reading frame 64 |
| chr14_+_31343951 | 0.04 |
ENST00000556908.1 ENST00000555881.1 ENST00000460581.2 |
COCH |
cochlin |
| chr12_-_50101003 | 0.04 |
ENST00000550488.1 |
FMNL3 |
formin-like 3 |
| chr11_+_10772534 | 0.04 |
ENST00000361367.2 |
CTR9 |
CTR9, Paf1/RNA polymerase II complex component |
| chr9_+_115983808 | 0.04 |
ENST00000374210.6 ENST00000374212.4 |
SLC31A1 |
solute carrier family 31 (copper transporter), member 1 |
| chr13_-_36871886 | 0.04 |
ENST00000491049.2 ENST00000503173.1 ENST00000239860.6 ENST00000379862.2 ENST00000239859.7 ENST00000379864.2 ENST00000510088.1 ENST00000554962.1 ENST00000511166.1 |
CCDC169 SOHLH2 CCDC169-SOHLH2 |
coiled-coil domain containing 169 spermatogenesis and oogenesis specific basic helix-loop-helix 2 CCDC169-SOHLH2 readthrough |
| chr6_-_29595779 | 0.04 |
ENST00000355973.3 ENST00000377012.4 |
GABBR1 |
gamma-aminobutyric acid (GABA) B receptor, 1 |
| chr15_-_77924689 | 0.04 |
ENST00000355300.6 |
LINGO1 |
leucine rich repeat and Ig domain containing 1 |
| chr12_-_52715179 | 0.04 |
ENST00000293670.3 |
KRT83 |
keratin 83 |
| chr20_+_48807351 | 0.03 |
ENST00000303004.3 |
CEBPB |
CCAAT/enhancer binding protein (C/EBP), beta |
| chr6_-_97345689 | 0.03 |
ENST00000316149.7 |
NDUFAF4 |
NADH dehydrogenase (ubiquinone) complex I, assembly factor 4 |
| chr14_+_96152754 | 0.03 |
ENST00000340722.7 |
TCL1B |
T-cell leukemia/lymphoma 1B |
| chr19_-_19626351 | 0.03 |
ENST00000585580.3 |
TSSK6 |
testis-specific serine kinase 6 |
| chr1_+_2398876 | 0.03 |
ENST00000449969.1 |
PLCH2 |
phospholipase C, eta 2 |
| chr7_-_79082867 | 0.03 |
ENST00000419488.1 ENST00000354212.4 |
MAGI2 |
membrane associated guanylate kinase, WW and PDZ domain containing 2 |
| chr12_-_123756781 | 0.03 |
ENST00000544658.1 |
CDK2AP1 |
cyclin-dependent kinase 2 associated protein 1 |
| chr8_-_101571933 | 0.03 |
ENST00000520311.1 |
ANKRD46 |
ankyrin repeat domain 46 |
| chr12_-_123756687 | 0.03 |
ENST00000261692.2 |
CDK2AP1 |
cyclin-dependent kinase 2 associated protein 1 |
| chr9_+_140125209 | 0.03 |
ENST00000538474.1 |
SLC34A3 |
solute carrier family 34 (type II sodium/phosphate contransporter), member 3 |
| chr7_-_128694927 | 0.03 |
ENST00000471166.1 ENST00000265388.5 |
TNPO3 |
transportin 3 |
| chr5_-_11904152 | 0.03 |
ENST00000304623.8 ENST00000458100.2 |
CTNND2 |
catenin (cadherin-associated protein), delta 2 |
| chr17_-_42441204 | 0.03 |
ENST00000293443.7 |
FAM171A2 |
family with sequence similarity 171, member A2 |
| chr16_+_88704978 | 0.02 |
ENST00000244241.4 |
IL17C |
interleukin 17C |
| chr1_+_156863470 | 0.02 |
ENST00000338302.3 ENST00000455314.1 ENST00000292357.7 |
PEAR1 |
platelet endothelial aggregation receptor 1 |
| chr2_+_152266392 | 0.02 |
ENST00000444746.2 ENST00000453091.2 ENST00000428287.2 ENST00000433166.2 ENST00000420714.3 ENST00000243326.5 ENST00000414861.2 |
RIF1 |
RAP1 interacting factor homolog (yeast) |
| chr11_+_64073699 | 0.02 |
ENST00000405666.1 ENST00000468670.1 |
ESRRA |
estrogen-related receptor alpha |
| chr12_+_51985001 | 0.02 |
ENST00000354534.6 |
SCN8A |
sodium channel, voltage gated, type VIII, alpha subunit |
| chr1_+_54411715 | 0.02 |
ENST00000371370.3 ENST00000371368.1 |
LRRC42 |
leucine rich repeat containing 42 |
| chr18_+_46065483 | 0.02 |
ENST00000382998.4 |
CTIF |
CBP80/20-dependent translation initiation factor |
| chr2_+_27435179 | 0.02 |
ENST00000606999.1 ENST00000405489.3 |
ATRAID |
all-trans retinoic acid-induced differentiation factor |
| chr11_-_57417405 | 0.02 |
ENST00000524669.1 ENST00000300022.3 |
YPEL4 |
yippee-like 4 (Drosophila) |
| chr17_-_62493131 | 0.02 |
ENST00000539111.2 |
POLG2 |
polymerase (DNA directed), gamma 2, accessory subunit |
| chr15_-_52263937 | 0.02 |
ENST00000315141.5 ENST00000299601.5 |
LEO1 |
Leo1, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) |
| chr17_+_46184911 | 0.02 |
ENST00000580219.1 ENST00000452859.2 ENST00000393405.2 ENST00000439357.2 ENST00000359238.2 |
SNX11 |
sorting nexin 11 |
| chr1_-_205326161 | 0.02 |
ENST00000367156.3 ENST00000606887.1 ENST00000607173.1 |
KLHDC8A |
kelch domain containing 8A |
| chr1_+_220863187 | 0.02 |
ENST00000294889.5 |
C1orf115 |
chromosome 1 open reading frame 115 |
| chr1_+_53098862 | 0.01 |
ENST00000517870.1 |
FAM159A |
family with sequence similarity 159, member A |
| chr17_-_36762095 | 0.01 |
ENST00000578925.1 ENST00000264659.7 |
SRCIN1 |
SRC kinase signaling inhibitor 1 |
| chr9_-_123691439 | 0.01 |
ENST00000540010.1 |
TRAF1 |
TNF receptor-associated factor 1 |
| chr17_-_8868991 | 0.01 |
ENST00000447110.1 |
PIK3R5 |
phosphoinositide-3-kinase, regulatory subunit 5 |
| chr6_+_155054459 | 0.01 |
ENST00000367178.3 ENST00000417268.1 ENST00000367186.4 |
SCAF8 |
SR-related CTD-associated factor 8 |
| chr20_+_35202909 | 0.01 |
ENST00000558530.1 ENST00000558028.1 ENST00000560025.1 |
TGIF2-C20orf24 TGIF2 |
TGIF2-C20orf24 readthrough TGFB-induced factor homeobox 2 |
| chr14_-_58332774 | 0.01 |
ENST00000556826.1 |
SLC35F4 |
solute carrier family 35, member F4 |
| chr9_-_98269481 | 0.01 |
ENST00000418258.1 ENST00000553011.1 ENST00000551845.1 |
PTCH1 |
patched 1 |
| chr9_-_136242956 | 0.01 |
ENST00000371989.3 ENST00000485435.2 |
SURF4 |
surfeit 4 |
| chr3_-_48601206 | 0.01 |
ENST00000273610.3 |
UCN2 |
urocortin 2 |
| chr9_+_140125385 | 0.01 |
ENST00000361134.2 |
SLC34A3 |
solute carrier family 34 (type II sodium/phosphate contransporter), member 3 |
| chr7_+_114055052 | 0.01 |
ENST00000462331.1 ENST00000408937.3 ENST00000403559.4 ENST00000350908.4 ENST00000393498.2 ENST00000393495.3 ENST00000378237.3 ENST00000393489.3 |
FOXP2 |
forkhead box P2 |
| chr1_-_236445251 | 0.01 |
ENST00000354619.5 ENST00000327333.8 |
ERO1LB |
ERO1-like beta (S. cerevisiae) |
| chr4_+_56814968 | 0.01 |
ENST00000422247.2 |
CEP135 |
centrosomal protein 135kDa |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.5 | GO:0051710 | cytolysis by symbiont of host cells(GO:0001897) regulation of cytolysis in other organism(GO:0051710) |
| 0.3 | 0.8 | GO:0060489 | orthogonal dichotomous subdivision of terminal units involved in lung branching morphogenesis(GO:0060488) planar dichotomous subdivision of terminal units involved in lung branching morphogenesis(GO:0060489) lateral sprouting involved in lung morphogenesis(GO:0060490) non-canonical Wnt signaling pathway involved in heart development(GO:0061341) planar cell polarity pathway involved in heart morphogenesis(GO:0061346) |
| 0.2 | 1.9 | GO:0035290 | trunk segmentation(GO:0035290) trunk neural crest cell migration(GO:0036484) ventral trunk neural crest cell migration(GO:0036486) |
| 0.2 | 0.7 | GO:1903028 | positive regulation of opsonization(GO:1903028) |
| 0.2 | 2.0 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.2 | 0.5 | GO:0061183 | Spemann organizer formation(GO:0060061) dermatome development(GO:0061054) regulation of dermatome development(GO:0061183) |
| 0.2 | 0.8 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.1 | 1.3 | GO:0034625 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.1 | 1.8 | GO:0007196 | adenylate cyclase-inhibiting G-protein coupled glutamate receptor signaling pathway(GO:0007196) |
| 0.1 | 0.9 | GO:0046940 | nucleoside monophosphate phosphorylation(GO:0046940) |
| 0.1 | 0.5 | GO:0032185 | septin cytoskeleton organization(GO:0032185) |
| 0.1 | 0.2 | GO:0000117 | regulation of transcription involved in G2/M transition of mitotic cell cycle(GO:0000117) |
| 0.1 | 0.4 | GO:1902162 | mRNA localization resulting in posttranscriptional regulation of gene expression(GO:0010609) regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902162) positive regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902164) |
| 0.1 | 0.2 | GO:2001037 | tongue muscle cell differentiation(GO:0035981) positive regulation of skeletal muscle fiber differentiation(GO:1902811) regulation of tongue muscle cell differentiation(GO:2001035) positive regulation of tongue muscle cell differentiation(GO:2001037) |
| 0.1 | 0.3 | GO:1901093 | regulation of protein tetramerization(GO:1901090) negative regulation of protein tetramerization(GO:1901091) regulation of protein homotetramerization(GO:1901093) negative regulation of protein homotetramerization(GO:1901094) |
| 0.1 | 0.1 | GO:1904933 | regulation of cell proliferation in midbrain(GO:1904933) |
| 0.0 | 0.2 | GO:0015692 | lead ion transport(GO:0015692) |
| 0.0 | 0.4 | GO:1903751 | regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:1903750) negative regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:1903751) |
| 0.0 | 0.4 | GO:0021546 | rhombomere development(GO:0021546) |
| 0.0 | 0.1 | GO:0006424 | glutamyl-tRNA aminoacylation(GO:0006424) |
| 0.0 | 0.3 | GO:0045163 | clustering of voltage-gated potassium channels(GO:0045163) |
| 0.0 | 0.3 | GO:0060282 | positive regulation of oocyte development(GO:0060282) |
| 0.0 | 0.2 | GO:1903659 | regulation of complement-dependent cytotoxicity(GO:1903659) |
| 0.0 | 0.2 | GO:2000301 | regulation of intracellular transport of viral material(GO:1901252) negative regulation of synaptic vesicle exocytosis(GO:2000301) |
| 0.0 | 0.2 | GO:0030382 | sperm mitochondrion organization(GO:0030382) |
| 0.0 | 0.1 | GO:0046832 | negative regulation of nucleobase-containing compound transport(GO:0032240) negative regulation of RNA export from nucleus(GO:0046832) |
| 0.0 | 0.2 | GO:0051182 | coenzyme transport(GO:0051182) |
| 0.0 | 0.2 | GO:0030174 | regulation of DNA-dependent DNA replication initiation(GO:0030174) |
| 0.0 | 0.4 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.0 | 0.1 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
| 0.0 | 0.1 | GO:0048377 | lateral mesodermal cell fate commitment(GO:0048372) lateral mesodermal cell fate specification(GO:0048377) regulation of lateral mesodermal cell fate specification(GO:0048378) |
| 0.0 | 0.2 | GO:0001886 | endothelial cell morphogenesis(GO:0001886) |
| 0.0 | 0.1 | GO:0006452 | translational frameshifting(GO:0006452) positive regulation of translational termination(GO:0045905) |
| 0.0 | 0.3 | GO:0015812 | gamma-aminobutyric acid transport(GO:0015812) |
| 0.0 | 0.1 | GO:0018242 | protein O-linked glycosylation via serine(GO:0018242) |
| 0.0 | 0.2 | GO:0036155 | acylglycerol acyl-chain remodeling(GO:0036155) |
| 0.0 | 0.0 | GO:0003402 | planar cell polarity pathway involved in axis elongation(GO:0003402) |
| 0.0 | 0.6 | GO:0007202 | activation of phospholipase C activity(GO:0007202) |
| 0.0 | 0.2 | GO:0038003 | opioid receptor signaling pathway(GO:0038003) |
| 0.0 | 0.1 | GO:0006701 | progesterone biosynthetic process(GO:0006701) |
| 0.0 | 0.0 | GO:2001162 | regulation of histone H3-K79 methylation(GO:2001160) positive regulation of histone H3-K79 methylation(GO:2001162) |
| 0.0 | 0.1 | GO:0070459 | prolactin secretion(GO:0070459) |
| 0.0 | 0.3 | GO:0030903 | notochord development(GO:0030903) |
| 0.0 | 0.1 | GO:0019348 | dolichol metabolic process(GO:0019348) |
| 0.0 | 0.1 | GO:0061343 | cell adhesion involved in heart morphogenesis(GO:0061343) |
| 0.0 | 0.2 | GO:0051964 | negative regulation of collateral sprouting(GO:0048671) negative regulation of synapse assembly(GO:0051964) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.8 | GO:0060187 | cell pole(GO:0060187) |
| 0.1 | 0.2 | GO:0019034 | viral replication complex(GO:0019034) |
| 0.0 | 0.2 | GO:0070826 | paraferritin complex(GO:0070826) |
| 0.0 | 0.2 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 0.3 | GO:0032039 | integrator complex(GO:0032039) |
| 0.0 | 0.1 | GO:0071256 | Sec61 translocon complex(GO:0005784) translocon complex(GO:0071256) |
| 0.0 | 3.6 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 1.3 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 0.4 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.0 | 1.7 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 0.1 | GO:0005642 | annulate lamellae(GO:0005642) |
| 0.0 | 0.2 | GO:0030285 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.0 | 0.8 | GO:0015030 | Cajal body(GO:0015030) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.8 | GO:0001641 | group II metabotropic glutamate receptor activity(GO:0001641) |
| 0.3 | 2.0 | GO:0015315 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.3 | 0.8 | GO:0008859 | exoribonuclease II activity(GO:0008859) |
| 0.2 | 0.9 | GO:0046899 | nucleoside triphosphate adenylate kinase activity(GO:0046899) |
| 0.1 | 1.3 | GO:0102336 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.1 | 2.1 | GO:0045499 | chemorepellent activity(GO:0045499) |
| 0.1 | 0.2 | GO:0008523 | sodium-dependent multivitamin transmembrane transporter activity(GO:0008523) |
| 0.1 | 0.2 | GO:0050252 | retinol O-fatty-acyltransferase activity(GO:0050252) |
| 0.1 | 0.5 | GO:0016015 | morphogen activity(GO:0016015) |
| 0.1 | 0.3 | GO:0005332 | gamma-aminobutyric acid:sodium symporter activity(GO:0005332) |
| 0.0 | 0.2 | GO:0015086 | cadmium ion transmembrane transporter activity(GO:0015086) cobalt ion transmembrane transporter activity(GO:0015087) lead ion transmembrane transporter activity(GO:0015094) ferrous iron uptake transmembrane transporter activity(GO:0015639) |
| 0.0 | 0.3 | GO:0017161 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) inositol-1,3,4-trisphosphate 4-phosphatase activity(GO:0017161) inositol-3,4-bisphosphate 4-phosphatase activity(GO:0052828) |
| 0.0 | 0.1 | GO:0004818 | glutamate-tRNA ligase activity(GO:0004818) |
| 0.0 | 0.1 | GO:0034511 | U3 snoRNA binding(GO:0034511) |
| 0.0 | 0.2 | GO:0008401 | retinoic acid 4-hydroxylase activity(GO:0008401) |
| 0.0 | 0.3 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.0 | 0.3 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.0 | 0.4 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.0 | 0.1 | GO:0016262 | protein N-acetylglucosaminyltransferase activity(GO:0016262) |
| 0.0 | 0.6 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.0 | 0.2 | GO:0034046 | poly(G) binding(GO:0034046) |
| 0.0 | 0.1 | GO:0004991 | parathyroid hormone receptor activity(GO:0004991) |
| 0.0 | 0.2 | GO:0015288 | porin activity(GO:0015288) |
| 0.0 | 0.5 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 0.2 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.0 | 0.1 | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity(GO:0003976) |
| 0.0 | 0.3 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.0 | 0.2 | GO:0004985 | opioid receptor activity(GO:0004985) |
| 0.0 | 0.2 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.0 | 0.2 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.0 | 0.4 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.0 | 0.7 | GO:0043531 | ADP binding(GO:0043531) |
| 0.0 | 0.1 | GO:0098821 | BMP receptor activity(GO:0098821) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.8 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.1 | 1.3 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 0.2 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.0 | 0.6 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.0 | 0.3 | REACTOME SYNTHESIS OF PIPS AT THE EARLY ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.0 | 0.6 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.0 | 0.3 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.0 | 0.9 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |