Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
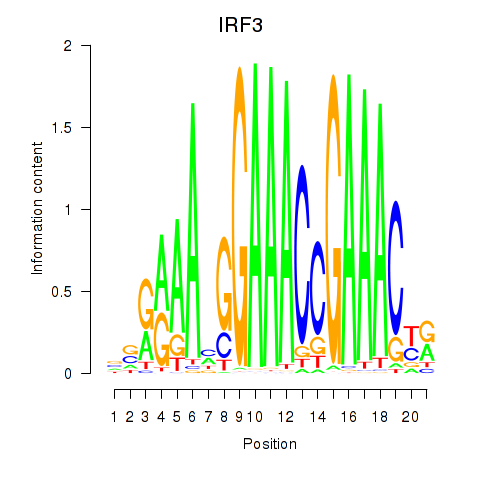
Results for IRF3
Z-value: 1.08
Transcription factors associated with IRF3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
IRF3
|
ENSG00000126456.11 | IRF3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| IRF3 | hg19_v2_chr19_-_50168861_50168873 | -0.78 | 2.2e-02 | Click! |
Activity profile of IRF3 motif
Sorted Z-values of IRF3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of IRF3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_-_95158644 | 1.11 |
ENST00000237858.6 |
GLRX |
glutaredoxin (thioltransferase) |
| chr1_+_221054584 | 1.10 |
ENST00000549319.1 |
HLX |
H2.0-like homeobox |
| chr5_-_95158375 | 1.05 |
ENST00000512469.2 ENST00000379979.4 ENST00000505427.1 ENST00000508780.1 |
GLRX |
glutaredoxin (thioltransferase) |
| chr2_-_190044480 | 0.84 |
ENST00000374866.3 |
COL5A2 |
collagen, type V, alpha 2 |
| chr1_+_79115503 | 0.82 |
ENST00000370747.4 ENST00000438486.1 ENST00000545124.1 |
IFI44 |
interferon-induced protein 44 |
| chr10_+_91087651 | 0.82 |
ENST00000371818.4 |
IFIT3 |
interferon-induced protein with tetratricopeptide repeats 3 |
| chr7_-_92777606 | 0.79 |
ENST00000437805.1 ENST00000446959.1 ENST00000439952.1 ENST00000414791.1 ENST00000446033.1 ENST00000411955.1 ENST00000318238.4 |
SAMD9L |
sterile alpha motif domain containing 9-like |
| chr10_+_91092241 | 0.74 |
ENST00000371811.4 |
IFIT3 |
interferon-induced protein with tetratricopeptide repeats 3 |
| chr6_+_72922590 | 0.72 |
ENST00000523963.1 |
RIMS1 |
regulating synaptic membrane exocytosis 1 |
| chr11_-_57335280 | 0.72 |
ENST00000287156.4 |
UBE2L6 |
ubiquitin-conjugating enzyme E2L 6 |
| chr4_+_124320665 | 0.71 |
ENST00000394339.2 |
SPRY1 |
sprouty homolog 1, antagonist of FGF signaling (Drosophila) |
| chr6_+_72922505 | 0.71 |
ENST00000401910.3 |
RIMS1 |
regulating synaptic membrane exocytosis 1 |
| chr10_-_49701686 | 0.70 |
ENST00000417247.2 |
ARHGAP22 |
Rho GTPase activating protein 22 |
| chr1_+_948803 | 0.64 |
ENST00000379389.4 |
ISG15 |
ISG15 ubiquitin-like modifier |
| chr3_-_122283079 | 0.59 |
ENST00000471785.1 ENST00000466126.1 |
PARP9 |
poly (ADP-ribose) polymerase family, member 9 |
| chr3_-_122283424 | 0.57 |
ENST00000477522.2 ENST00000360356.2 |
PARP9 |
poly (ADP-ribose) polymerase family, member 9 |
| chrX_+_80457442 | 0.57 |
ENST00000373212.5 |
SH3BGRL |
SH3 domain binding glutamic acid-rich protein like |
| chr7_-_122526799 | 0.52 |
ENST00000334010.7 ENST00000313070.7 |
CADPS2 |
Ca++-dependent secretion activator 2 |
| chr2_-_145275228 | 0.50 |
ENST00000427902.1 ENST00000409487.3 ENST00000470879.1 ENST00000435831.1 |
ZEB2 |
zinc finger E-box binding homeobox 2 |
| chr19_+_49977466 | 0.49 |
ENST00000596435.1 ENST00000344019.3 ENST00000597551.1 ENST00000204637.2 ENST00000600429.1 |
FLT3LG |
fms-related tyrosine kinase 3 ligand |
| chr1_+_210502238 | 0.49 |
ENST00000545154.1 ENST00000537898.1 ENST00000391905.3 ENST00000545781.1 ENST00000261458.3 ENST00000308852.6 |
HHAT |
hedgehog acyltransferase |
| chr17_+_6659153 | 0.48 |
ENST00000441631.1 ENST00000438512.1 ENST00000346752.4 ENST00000361842.3 |
XAF1 |
XIAP associated factor 1 |
| chr2_+_201980827 | 0.46 |
ENST00000309955.3 ENST00000443227.1 ENST00000341222.6 ENST00000355558.4 ENST00000340870.5 ENST00000341582.6 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr2_+_201980961 | 0.46 |
ENST00000342795.5 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr12_+_133614119 | 0.46 |
ENST00000327668.7 |
ZNF84 |
zinc finger protein 84 |
| chr19_-_17516449 | 0.45 |
ENST00000252593.6 |
BST2 |
bone marrow stromal cell antigen 2 |
| chr12_+_133613878 | 0.44 |
ENST00000392319.2 ENST00000543758.1 |
ZNF84 |
zinc finger protein 84 |
| chr19_+_49977818 | 0.44 |
ENST00000594009.1 ENST00000595510.1 |
FLT3LG |
fms-related tyrosine kinase 3 ligand |
| chr4_-_156875003 | 0.43 |
ENST00000433477.3 |
CTSO |
cathepsin O |
| chr6_-_3752222 | 0.42 |
ENST00000380283.4 |
PXDC1 |
PX domain containing 1 |
| chr20_-_47894569 | 0.42 |
ENST00000371744.1 ENST00000371752.1 ENST00000396105.1 |
ZNFX1 |
zinc finger, NFX1-type containing 1 |
| chr10_+_92980517 | 0.41 |
ENST00000336126.5 |
PCGF5 |
polycomb group ring finger 5 |
| chr14_+_24605389 | 0.40 |
ENST00000382708.3 ENST00000561435.1 |
PSME1 |
proteasome (prosome, macropain) activator subunit 1 (PA28 alpha) |
| chr19_+_10196781 | 0.39 |
ENST00000253110.11 |
C19orf66 |
chromosome 19 open reading frame 66 |
| chr16_-_67970990 | 0.38 |
ENST00000358514.4 |
PSMB10 |
proteasome (prosome, macropain) subunit, beta type, 10 |
| chr6_+_32821924 | 0.38 |
ENST00000374859.2 ENST00000453265.2 |
PSMB9 |
proteasome (prosome, macropain) subunit, beta type, 9 |
| chr1_-_146644036 | 0.38 |
ENST00000425272.2 |
PRKAB2 |
protein kinase, AMP-activated, beta 2 non-catalytic subunit |
| chr12_-_4758159 | 0.36 |
ENST00000545990.2 |
AKAP3 |
A kinase (PRKA) anchor protein 3 |
| chr3_+_187086120 | 0.36 |
ENST00000259030.2 |
RTP4 |
receptor (chemosensory) transporter protein 4 |
| chr20_+_388791 | 0.35 |
ENST00000441733.1 ENST00000353660.3 |
RBCK1 |
RanBP-type and C3HC4-type zinc finger containing 1 |
| chr7_+_100728720 | 0.35 |
ENST00000306085.6 ENST00000412507.1 |
TRIM56 |
tripartite motif containing 56 |
| chr20_-_33872518 | 0.35 |
ENST00000374436.3 |
EIF6 |
eukaryotic translation initiation factor 6 |
| chr2_+_201981527 | 0.34 |
ENST00000441224.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr9_+_95087766 | 0.33 |
ENST00000375587.3 |
CENPP |
centromere protein P |
| chr20_-_33872548 | 0.32 |
ENST00000374443.3 |
EIF6 |
eukaryotic translation initiation factor 6 |
| chr19_+_39786962 | 0.32 |
ENST00000333625.2 |
IFNL1 |
interferon, lambda 1 |
| chr2_-_119605253 | 0.30 |
ENST00000295206.6 |
EN1 |
engrailed homeobox 1 |
| chr5_+_140749803 | 0.30 |
ENST00000576222.1 |
PCDHGB3 |
protocadherin gamma subfamily B, 3 |
| chr12_+_133613937 | 0.30 |
ENST00000539354.1 ENST00000542874.1 ENST00000438628.2 |
ZNF84 |
zinc finger protein 84 |
| chr10_-_14996321 | 0.30 |
ENST00000378289.4 |
DCLRE1C |
DNA cross-link repair 1C |
| chr19_+_10196981 | 0.29 |
ENST00000591813.1 |
C19orf66 |
chromosome 19 open reading frame 66 |
| chr7_+_142829162 | 0.28 |
ENST00000291009.3 |
PIP |
prolactin-induced protein |
| chr6_+_3068606 | 0.28 |
ENST00000259808.4 |
RIPK1 |
receptor (TNFRSF)-interacting serine-threonine kinase 1 |
| chr10_+_35415978 | 0.28 |
ENST00000429130.3 ENST00000469949.2 ENST00000460270.1 |
CREM |
cAMP responsive element modulator |
| chr5_-_180288248 | 0.27 |
ENST00000512132.1 ENST00000506439.1 ENST00000502412.1 ENST00000359141.6 |
ZFP62 |
ZFP62 zinc finger protein |
| chr4_-_77134742 | 0.27 |
ENST00000452464.2 |
SCARB2 |
scavenger receptor class B, member 2 |
| chr14_+_24605361 | 0.26 |
ENST00000206451.6 ENST00000559123.1 |
PSME1 |
proteasome (prosome, macropain) activator subunit 1 (PA28 alpha) |
| chr10_+_91061712 | 0.26 |
ENST00000371826.3 |
IFIT2 |
interferon-induced protein with tetratricopeptide repeats 2 |
| chr4_-_74847800 | 0.26 |
ENST00000296029.3 |
PF4 |
platelet factor 4 |
| chr1_+_145292806 | 0.25 |
ENST00000448873.2 |
NBPF10 |
neuroblastoma breakpoint family, member 10 |
| chr12_+_6561190 | 0.25 |
ENST00000544021.1 ENST00000266556.7 |
TAPBPL |
TAP binding protein-like |
| chr4_-_77135046 | 0.25 |
ENST00000264896.2 |
SCARB2 |
scavenger receptor class B, member 2 |
| chr10_+_115439630 | 0.24 |
ENST00000369318.3 |
CASP7 |
caspase 7, apoptosis-related cysteine peptidase |
| chr6_+_26440700 | 0.24 |
ENST00000494393.1 ENST00000482451.1 ENST00000244519.2 ENST00000339789.4 ENST00000471353.1 ENST00000361232.3 ENST00000487627.1 ENST00000496719.1 ENST00000490254.1 ENST00000487272.1 |
BTN3A3 |
butyrophilin, subfamily 3, member A3 |
| chr15_-_62352570 | 0.24 |
ENST00000261517.5 ENST00000395896.4 ENST00000395898.3 |
VPS13C |
vacuolar protein sorting 13 homolog C (S. cerevisiae) |
| chr1_+_159931002 | 0.24 |
ENST00000443364.1 ENST00000423943.1 |
RP11-48O20.4 |
long intergenic non-protein coding RNA 1133 |
| chr2_-_152118352 | 0.23 |
ENST00000331426.5 |
RBM43 |
RNA binding motif protein 43 |
| chr2_-_231084617 | 0.23 |
ENST00000409815.2 |
SP110 |
SP110 nuclear body protein |
| chr22_-_36556821 | 0.23 |
ENST00000531095.1 ENST00000397293.2 ENST00000349314.2 |
APOL3 |
apolipoprotein L, 3 |
| chr19_+_17516531 | 0.23 |
ENST00000528911.1 ENST00000528604.1 ENST00000595892.1 ENST00000500836.2 ENST00000598546.1 ENST00000600369.1 ENST00000598356.1 ENST00000594426.1 |
MVB12A CTD-2521M24.9 |
multivesicular body subunit 12A CTD-2521M24.9 |
| chr10_+_115439282 | 0.23 |
ENST00000369321.2 ENST00000345633.4 |
CASP7 |
caspase 7, apoptosis-related cysteine peptidase |
| chr1_+_151129135 | 0.22 |
ENST00000602841.1 |
SCNM1 |
sodium channel modifier 1 |
| chr6_-_110964453 | 0.22 |
ENST00000413605.2 |
CDK19 |
cyclin-dependent kinase 19 |
| chr8_-_67525473 | 0.22 |
ENST00000522677.3 |
MYBL1 |
v-myb avian myeloblastosis viral oncogene homolog-like 1 |
| chr11_-_615570 | 0.22 |
ENST00000525445.1 ENST00000348655.6 ENST00000397566.1 |
IRF7 |
interferon regulatory factor 7 |
| chr7_-_138794081 | 0.22 |
ENST00000464606.1 |
ZC3HAV1 |
zinc finger CCCH-type, antiviral 1 |
| chr14_-_70883708 | 0.21 |
ENST00000256366.4 |
SYNJ2BP |
synaptojanin 2 binding protein |
| chr10_+_115439699 | 0.21 |
ENST00000369315.1 |
CASP7 |
caspase 7, apoptosis-related cysteine peptidase |
| chr14_+_73525265 | 0.20 |
ENST00000525161.1 |
RBM25 |
RNA binding motif protein 25 |
| chr10_-_102046417 | 0.20 |
ENST00000370372.2 |
BLOC1S2 |
biogenesis of lysosomal organelles complex-1, subunit 2 |
| chr17_-_40264692 | 0.20 |
ENST00000591220.1 ENST00000251642.3 |
DHX58 |
DEXH (Asp-Glu-X-His) box polypeptide 58 |
| chr1_-_182558374 | 0.20 |
ENST00000367559.3 ENST00000539397.1 |
RNASEL |
ribonuclease L (2',5'-oligoisoadenylate synthetase-dependent) |
| chr22_-_36635684 | 0.20 |
ENST00000358502.5 |
APOL2 |
apolipoprotein L, 2 |
| chr2_+_69240302 | 0.20 |
ENST00000303714.4 |
ANTXR1 |
anthrax toxin receptor 1 |
| chr10_-_22292675 | 0.19 |
ENST00000376946.1 |
DNAJC1 |
DnaJ (Hsp40) homolog, subfamily C, member 1 |
| chr22_+_36649056 | 0.19 |
ENST00000397278.3 ENST00000422706.1 ENST00000426053.1 ENST00000319136.4 |
APOL1 |
apolipoprotein L, 1 |
| chr10_-_102046098 | 0.19 |
ENST00000441611.1 |
BLOC1S2 |
biogenesis of lysosomal organelles complex-1, subunit 2 |
| chr2_-_231084820 | 0.18 |
ENST00000258382.5 ENST00000338556.3 |
SP110 |
SP110 nuclear body protein |
| chrX_-_48937531 | 0.17 |
ENST00000473974.1 ENST00000475880.1 ENST00000396681.4 ENST00000553851.1 ENST00000471338.1 ENST00000476728.1 ENST00000376368.2 ENST00000485908.1 ENST00000376372.3 ENST00000376358.3 |
WDR45 AF196779.12 |
WD repeat domain 45 WD repeat domain phosphoinositide-interacting protein 4 |
| chr2_+_201981119 | 0.17 |
ENST00000395148.2 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr2_+_113672770 | 0.17 |
ENST00000311328.2 |
IL37 |
interleukin 37 |
| chr14_-_20881579 | 0.16 |
ENST00000556935.1 ENST00000262715.5 ENST00000556549.1 |
TEP1 |
telomerase-associated protein 1 |
| chr2_+_239756671 | 0.16 |
ENST00000448943.2 |
TWIST2 |
twist family bHLH transcription factor 2 |
| chr1_-_161039753 | 0.16 |
ENST00000368015.1 |
ARHGAP30 |
Rho GTPase activating protein 30 |
| chr1_-_177939348 | 0.15 |
ENST00000464631.2 |
SEC16B |
SEC16 homolog B (S. cerevisiae) |
| chr1_+_25599018 | 0.15 |
ENST00000417538.2 ENST00000357542.4 ENST00000568195.1 ENST00000342055.5 ENST00000423810.2 |
RHD |
Rh blood group, D antigen |
| chr15_+_64428529 | 0.15 |
ENST00000560861.1 |
SNX1 |
sorting nexin 1 |
| chr7_-_92747269 | 0.15 |
ENST00000446617.1 ENST00000379958.2 |
SAMD9 |
sterile alpha motif domain containing 9 |
| chrX_+_77154935 | 0.15 |
ENST00000481445.1 |
COX7B |
cytochrome c oxidase subunit VIIb |
| chr5_+_140071178 | 0.14 |
ENST00000508522.1 ENST00000448069.2 |
HARS2 |
histidyl-tRNA synthetase 2, mitochondrial |
| chr6_-_28226984 | 0.14 |
ENST00000423974.2 |
ZKSCAN4 |
zinc finger with KRAB and SCAN domains 4 |
| chr2_-_175629135 | 0.14 |
ENST00000409542.1 ENST00000409219.1 |
CHRNA1 |
cholinergic receptor, nicotinic, alpha 1 (muscle) |
| chr1_-_161039647 | 0.14 |
ENST00000368013.3 |
ARHGAP30 |
Rho GTPase activating protein 30 |
| chr5_-_77656175 | 0.14 |
ENST00000513755.1 ENST00000421004.3 |
CTD-2037K23.2 |
CTD-2037K23.2 |
| chr16_+_75681650 | 0.14 |
ENST00000300086.4 |
TERF2IP |
telomeric repeat binding factor 2, interacting protein |
| chr19_+_50431959 | 0.14 |
ENST00000595125.1 |
ATF5 |
activating transcription factor 5 |
| chr3_+_122399444 | 0.13 |
ENST00000474629.2 |
PARP14 |
poly (ADP-ribose) polymerase family, member 14 |
| chr2_+_219125714 | 0.13 |
ENST00000522678.1 ENST00000519574.1 ENST00000521462.1 |
GPBAR1 |
G protein-coupled bile acid receptor 1 |
| chr6_-_36355513 | 0.13 |
ENST00000340181.4 ENST00000373737.4 |
ETV7 |
ets variant 7 |
| chr20_-_20033052 | 0.12 |
ENST00000536226.1 |
CRNKL1 |
crooked neck pre-mRNA splicing factor 1 |
| chr3_+_184529929 | 0.12 |
ENST00000287546.4 ENST00000437079.3 |
VPS8 |
vacuolar protein sorting 8 homolog (S. cerevisiae) |
| chr17_-_63052929 | 0.12 |
ENST00000439174.2 |
GNA13 |
guanine nucleotide binding protein (G protein), alpha 13 |
| chr20_+_61436146 | 0.12 |
ENST00000290291.6 |
OGFR |
opioid growth factor receptor |
| chr1_+_236557569 | 0.12 |
ENST00000334232.4 |
EDARADD |
EDAR-associated death domain |
| chr3_+_184529948 | 0.12 |
ENST00000436792.2 ENST00000446204.2 ENST00000422105.1 |
VPS8 |
vacuolar protein sorting 8 homolog (S. cerevisiae) |
| chr2_-_231084659 | 0.12 |
ENST00000258381.6 ENST00000358662.4 ENST00000455674.1 ENST00000392048.3 |
SP110 |
SP110 nuclear body protein |
| chr10_+_35416223 | 0.12 |
ENST00000489321.1 ENST00000427847.2 ENST00000345491.3 ENST00000395895.2 ENST00000374728.3 ENST00000487132.1 |
CREM |
cAMP responsive element modulator |
| chr6_-_121655593 | 0.12 |
ENST00000398212.2 |
TBC1D32 |
TBC1 domain family, member 32 |
| chr1_+_150480551 | 0.12 |
ENST00000369049.4 ENST00000369047.4 |
ECM1 |
extracellular matrix protein 1 |
| chr1_+_151129103 | 0.12 |
ENST00000368910.3 |
TNFAIP8L2 |
tumor necrosis factor, alpha-induced protein 8-like 2 |
| chr1_-_146644122 | 0.11 |
ENST00000254101.3 |
PRKAB2 |
protein kinase, AMP-activated, beta 2 non-catalytic subunit |
| chr12_+_129028500 | 0.11 |
ENST00000315208.8 |
TMEM132C |
transmembrane protein 132C |
| chr16_+_66968343 | 0.11 |
ENST00000417689.1 ENST00000561697.1 ENST00000317091.4 ENST00000566182.1 |
CES2 |
carboxylesterase 2 |
| chr7_-_138794394 | 0.10 |
ENST00000242351.5 ENST00000471652.1 |
ZC3HAV1 |
zinc finger CCCH-type, antiviral 1 |
| chr1_+_150480576 | 0.10 |
ENST00000346569.6 |
ECM1 |
extracellular matrix protein 1 |
| chr14_-_24615805 | 0.10 |
ENST00000560410.1 |
PSME2 |
proteasome (prosome, macropain) activator subunit 2 (PA28 beta) |
| chr22_+_18632666 | 0.10 |
ENST00000215794.7 |
USP18 |
ubiquitin specific peptidase 18 |
| chr6_+_28193037 | 0.10 |
ENST00000531981.1 ENST00000425468.2 ENST00000252207.5 ENST00000531979.1 ENST00000527436.1 |
ZSCAN9 |
zinc finger and SCAN domain containing 9 |
| chr10_-_135187193 | 0.09 |
ENST00000368547.3 |
ECHS1 |
enoyl CoA hydratase, short chain, 1, mitochondrial |
| chr14_-_24615523 | 0.08 |
ENST00000559056.1 |
PSME2 |
proteasome (prosome, macropain) activator subunit 2 (PA28 beta) |
| chr14_+_35452104 | 0.08 |
ENST00000216774.6 ENST00000546080.1 |
SRP54 |
signal recognition particle 54kDa |
| chr6_+_31540056 | 0.07 |
ENST00000418386.2 |
LTA |
lymphotoxin alpha |
| chr14_+_35452169 | 0.07 |
ENST00000555557.1 |
SRP54 |
signal recognition particle 54kDa |
| chr9_-_140082983 | 0.07 |
ENST00000323927.2 |
ANAPC2 |
anaphase promoting complex subunit 2 |
| chr6_+_42018614 | 0.07 |
ENST00000465926.1 ENST00000482432.1 |
TAF8 |
TAF8 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 43kDa |
| chr12_-_54582655 | 0.07 |
ENST00000504338.1 ENST00000514685.1 ENST00000504797.1 ENST00000513838.1 ENST00000505128.1 ENST00000337581.3 ENST00000503306.1 ENST00000243112.5 ENST00000514196.1 ENST00000506169.1 ENST00000507904.1 ENST00000508394.2 |
SMUG1 |
single-strand-selective monofunctional uracil-DNA glycosylase 1 |
| chr8_+_22446763 | 0.06 |
ENST00000450780.2 ENST00000430850.2 ENST00000447849.1 |
AC037459.4 |
Uncharacterized protein |
| chr19_+_17516909 | 0.06 |
ENST00000601007.1 ENST00000594913.1 ENST00000599975.1 ENST00000600514.1 |
CTD-2521M24.9 MVB12A |
CTD-2521M24.9 multivesicular body subunit 12A |
| chr17_+_21730180 | 0.06 |
ENST00000584398.1 |
UBBP4 |
ubiquitin B pseudogene 4 |
| chrX_+_154610428 | 0.06 |
ENST00000354514.4 |
H2AFB2 |
H2A histone family, member B2 |
| chr17_+_25958174 | 0.06 |
ENST00000313648.6 ENST00000577392.1 ENST00000584661.1 ENST00000413914.2 |
LGALS9 |
lectin, galactoside-binding, soluble, 9 |
| chr20_+_388679 | 0.05 |
ENST00000356286.5 ENST00000475269.1 |
RBCK1 |
RanBP-type and C3HC4-type zinc finger containing 1 |
| chr4_+_100495864 | 0.05 |
ENST00000265517.5 ENST00000422897.2 |
MTTP |
microsomal triglyceride transfer protein |
| chr6_+_26402517 | 0.05 |
ENST00000414912.2 |
BTN3A1 |
butyrophilin, subfamily 3, member A1 |
| chr14_+_77790901 | 0.05 |
ENST00000553586.1 ENST00000555583.1 |
GSTZ1 |
glutathione S-transferase zeta 1 |
| chr1_-_177939041 | 0.05 |
ENST00000308284.6 |
SEC16B |
SEC16 homolog B (S. cerevisiae) |
| chr11_-_615942 | 0.05 |
ENST00000397562.3 ENST00000330243.5 ENST00000397570.1 ENST00000397574.2 |
IRF7 |
interferon regulatory factor 7 |
| chr22_+_31892373 | 0.05 |
ENST00000443011.1 ENST00000400289.1 ENST00000444859.1 ENST00000400288.2 |
SFI1 |
Sfi1 homolog, spindle assembly associated (yeast) |
| chr6_-_30181156 | 0.05 |
ENST00000418026.1 ENST00000416596.1 ENST00000453195.1 |
TRIM26 |
tripartite motif containing 26 |
| chr2_+_127413704 | 0.05 |
ENST00000409836.3 |
GYPC |
glycophorin C (Gerbich blood group) |
| chr2_-_37384175 | 0.04 |
ENST00000411537.2 ENST00000233057.4 ENST00000395127.2 ENST00000390013.3 |
EIF2AK2 |
eukaryotic translation initiation factor 2-alpha kinase 2 |
| chr19_+_50432400 | 0.04 |
ENST00000423777.2 ENST00000600336.1 ENST00000597227.1 |
ATF5 |
activating transcription factor 5 |
| chr16_-_29874462 | 0.04 |
ENST00000566113.1 ENST00000569956.1 ENST00000570016.1 |
CDIPT |
CDP-diacylglycerol--inositol 3-phosphatidyltransferase |
| chrX_+_119384607 | 0.04 |
ENST00000326624.2 ENST00000557385.1 |
ZBTB33 |
zinc finger and BTB domain containing 33 |
| chr2_+_127413677 | 0.04 |
ENST00000356887.7 |
GYPC |
glycophorin C (Gerbich blood group) |
| chr4_+_118955500 | 0.04 |
ENST00000296499.5 |
NDST3 |
N-deacetylase/N-sulfotransferase (heparan glucosaminyl) 3 |
| chr19_+_17516624 | 0.04 |
ENST00000596322.1 ENST00000600008.1 ENST00000601885.1 |
CTD-2521M24.9 |
CTD-2521M24.9 |
| chr4_+_130017268 | 0.04 |
ENST00000425929.1 ENST00000508673.1 ENST00000508622.1 |
C4orf33 |
chromosome 4 open reading frame 33 |
| chr22_+_31892261 | 0.04 |
ENST00000432498.1 ENST00000540643.1 ENST00000443326.1 ENST00000414585.1 |
SFI1 |
Sfi1 homolog, spindle assembly associated (yeast) |
| chr5_-_96143796 | 0.04 |
ENST00000296754.3 |
ERAP1 |
endoplasmic reticulum aminopeptidase 1 |
| chr3_+_63898275 | 0.04 |
ENST00000538065.1 |
ATXN7 |
ataxin 7 |
| chr6_-_30181133 | 0.04 |
ENST00000454678.2 ENST00000434785.1 |
TRIM26 |
tripartite motif containing 26 |
| chr20_+_388935 | 0.03 |
ENST00000382181.2 ENST00000400247.3 |
RBCK1 |
RanBP-type and C3HC4-type zinc finger containing 1 |
| chr3_+_63897605 | 0.03 |
ENST00000487717.1 |
ATXN7 |
ataxin 7 |
| chrX_+_57313113 | 0.03 |
ENST00000374900.4 |
FAAH2 |
fatty acid amide hydrolase 2 |
| chr3_-_122283100 | 0.03 |
ENST00000492382.1 ENST00000462315.1 |
PARP9 |
poly (ADP-ribose) polymerase family, member 9 |
| chr6_+_26402465 | 0.03 |
ENST00000476549.2 ENST00000289361.6 ENST00000450085.2 ENST00000425234.2 ENST00000427334.1 ENST00000506698.1 |
BTN3A1 |
butyrophilin, subfamily 3, member A1 |
| chrX_+_10031499 | 0.02 |
ENST00000454666.1 |
WWC3 |
WWC family member 3 |
| chr20_+_44509857 | 0.02 |
ENST00000372523.1 ENST00000372520.1 |
ZSWIM1 |
zinc finger, SWIM-type containing 1 |
| chr8_+_145438870 | 0.02 |
ENST00000527931.1 |
FAM203B |
family with sequence similarity 203, member B |
| chr16_-_29875057 | 0.02 |
ENST00000219789.6 |
CDIPT |
CDP-diacylglycerol--inositol 3-phosphatidyltransferase |
| chr6_-_36355486 | 0.02 |
ENST00000538992.1 |
ETV7 |
ets variant 7 |
| chr2_+_69240415 | 0.02 |
ENST00000409829.3 |
ANTXR1 |
anthrax toxin receptor 1 |
| chr22_+_39436862 | 0.02 |
ENST00000381565.2 ENST00000452957.2 |
APOBEC3F APOBEC3G |
apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3F apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3G |
| chr17_+_21729899 | 0.02 |
ENST00000583708.1 |
UBBP4 |
ubiquitin B pseudogene 4 |
| chr9_+_140083099 | 0.01 |
ENST00000322310.5 |
SSNA1 |
Sjogren syndrome nuclear autoantigen 1 |
| chr2_-_175629164 | 0.01 |
ENST00000409323.1 ENST00000261007.5 ENST00000348749.5 |
CHRNA1 |
cholinergic receptor, nicotinic, alpha 1 (muscle) |
| chr12_-_127256772 | 0.01 |
ENST00000536517.1 |
LINC00944 |
long intergenic non-protein coding RNA 944 |
| chr19_+_17516494 | 0.01 |
ENST00000534306.1 |
CTD-2521M24.9 |
CTD-2521M24.9 |
| chr2_+_99771418 | 0.01 |
ENST00000393473.2 ENST00000393477.3 ENST00000393474.3 ENST00000340066.1 ENST00000393471.2 ENST00000449211.1 ENST00000434566.1 ENST00000410042.1 |
LIPT1 MRPL30 |
lipoyltransferase 1 39S ribosomal protein L30, mitochondrial |
| chr3_+_122283064 | 0.01 |
ENST00000296161.4 |
DTX3L |
deltex 3-like (Drosophila) |
| chr1_-_89664595 | 0.01 |
ENST00000355754.6 |
GBP4 |
guanylate binding protein 4 |
| chrX_+_52920336 | 0.00 |
ENST00000452154.2 |
FAM156B |
family with sequence similarity 156, member B |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.9 | GO:0090290 | positive regulation of osteoclast proliferation(GO:0090290) |
| 0.3 | 0.8 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.2 | 0.2 | GO:0002590 | regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002589) negative regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002590) |
| 0.2 | 0.7 | GO:0060940 | epithelial to mesenchymal transition involved in cardiac fibroblast development(GO:0060940) |
| 0.2 | 0.7 | GO:1902626 | assembly of large subunit precursor of preribosome(GO:1902626) |
| 0.2 | 1.5 | GO:0070212 | protein poly-ADP-ribosylation(GO:0070212) |
| 0.2 | 0.5 | GO:0002731 | negative regulation of dendritic cell cytokine production(GO:0002731) negative regulation of intracellular transport of viral material(GO:1901253) |
| 0.1 | 1.4 | GO:1903943 | regulation of hepatocyte apoptotic process(GO:1903943) negative regulation of hepatocyte apoptotic process(GO:1903944) |
| 0.1 | 1.8 | GO:0035457 | cellular response to interferon-alpha(GO:0035457) |
| 0.1 | 1.1 | GO:0045629 | negative regulation of T-helper 2 cell differentiation(GO:0045629) |
| 0.1 | 0.5 | GO:1905123 | regulation of endosome organization(GO:1904978) regulation of glucosylceramidase activity(GO:1905123) |
| 0.1 | 2.2 | GO:0045838 | positive regulation of membrane potential(GO:0045838) |
| 0.1 | 1.2 | GO:0034058 | endosomal vesicle fusion(GO:0034058) |
| 0.1 | 0.7 | GO:0072733 | response to staurosporine(GO:0072733) cellular response to staurosporine(GO:0072734) |
| 0.1 | 0.3 | GO:0032242 | regulation of nucleoside transport(GO:0032242) positive regulation of necroptotic process(GO:0060545) |
| 0.1 | 0.5 | GO:1990504 | dense core granule exocytosis(GO:1990504) |
| 0.1 | 0.5 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.1 | 0.2 | GO:1904924 | negative regulation of mitophagy in response to mitochondrial depolarization(GO:1904924) |
| 0.1 | 0.8 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 0.1 | 1.4 | GO:0098828 | positive regulation of inhibitory postsynaptic potential(GO:0097151) modulation of inhibitory postsynaptic potential(GO:0098828) |
| 0.1 | 0.3 | GO:0061743 | motor learning(GO:0061743) |
| 0.1 | 0.2 | GO:1901202 | negative regulation of extracellular matrix assembly(GO:1901202) |
| 0.1 | 0.3 | GO:0045918 | negative regulation of cytolysis(GO:0045918) |
| 0.1 | 0.3 | GO:0019075 | virus maturation(GO:0019075) |
| 0.1 | 0.4 | GO:0097039 | protein linear polyubiquitination(GO:0097039) |
| 0.1 | 0.3 | GO:2000110 | negative regulation of macrophage apoptotic process(GO:2000110) |
| 0.1 | 0.2 | GO:0039534 | negative regulation of MDA-5 signaling pathway(GO:0039534) |
| 0.0 | 0.1 | GO:0006617 | SRP-dependent cotranslational protein targeting to membrane, signal sequence recognition(GO:0006617) |
| 0.0 | 0.1 | GO:0006427 | histidyl-tRNA aminoacylation(GO:0006427) |
| 0.0 | 0.3 | GO:0032696 | negative regulation of interleukin-13 production(GO:0032696) |
| 0.0 | 0.1 | GO:0072695 | negative regulation of DNA recombination at telomere(GO:0048239) regulation of DNA recombination at telomere(GO:0072695) |
| 0.0 | 0.4 | GO:0060155 | platelet dense granule organization(GO:0060155) |
| 0.0 | 0.1 | GO:0038183 | bile acid signaling pathway(GO:0038183) |
| 0.0 | 0.1 | GO:0002874 | regulation of chronic inflammatory response to antigenic stimulus(GO:0002874) |
| 0.0 | 0.5 | GO:0006853 | carnitine shuttle(GO:0006853) |
| 0.0 | 0.2 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.0 | 0.1 | GO:0032831 | natural killer cell tolerance induction(GO:0002519) regulation of CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0032829) positive regulation of CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0032831) positive regulation of T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:2001190) |
| 0.0 | 0.6 | GO:0001580 | detection of chemical stimulus involved in sensory perception of bitter taste(GO:0001580) |
| 0.0 | 0.2 | GO:0048630 | skeletal muscle tissue growth(GO:0048630) |
| 0.0 | 0.2 | GO:0000722 | telomere maintenance via recombination(GO:0000722) |
| 0.0 | 1.6 | GO:0006521 | regulation of cellular amino acid metabolic process(GO:0006521) |
| 0.0 | 0.5 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.0 | 0.1 | GO:0003406 | retinal pigment epithelium development(GO:0003406) |
| 0.0 | 0.1 | GO:0031584 | activation of phospholipase D activity(GO:0031584) |
| 0.0 | 0.1 | GO:0031848 | protection from non-homologous end joining at telomere(GO:0031848) |
| 0.0 | 0.2 | GO:0016559 | peroxisome fission(GO:0016559) positive regulation of protein exit from endoplasmic reticulum(GO:0070863) |
| 0.0 | 0.5 | GO:0035456 | response to interferon-beta(GO:0035456) |
| 0.0 | 0.2 | GO:0021891 | olfactory bulb interneuron development(GO:0021891) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.1 | 0.8 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.1 | 0.3 | GO:0032002 | interleukin-28 receptor complex(GO:0032002) |
| 0.1 | 0.4 | GO:0071797 | LUBAC complex(GO:0071797) |
| 0.1 | 0.3 | GO:0097679 | symbiont-containing vacuole(GO:0020003) symbiont-containing vacuole membrane(GO:0020005) other organism cytoplasm(GO:0097679) |
| 0.1 | 0.8 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 0.1 | 1.7 | GO:0097342 | ripoptosome(GO:0097342) |
| 0.0 | 0.2 | GO:0033263 | CORVET complex(GO:0033263) |
| 0.0 | 0.4 | GO:0035686 | sperm fibrous sheath(GO:0035686) |
| 0.0 | 0.9 | GO:0031233 | intrinsic component of external side of plasma membrane(GO:0031233) |
| 0.0 | 1.4 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 0.5 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 0.3 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.0 | 0.1 | GO:0030905 | retromer, tubulation complex(GO:0030905) |
| 0.0 | 0.1 | GO:0071012 | catalytic step 1 spliceosome(GO:0071012) |
| 0.0 | 0.6 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.0 | 0.1 | GO:0030870 | Mre11 complex(GO:0030870) |
| 0.0 | 0.4 | GO:0031083 | BLOC-1 complex(GO:0031083) |
| 0.0 | 0.1 | GO:0005786 | signal recognition particle, endoplasmic reticulum targeting(GO:0005786) |
| 0.0 | 0.2 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.2 | GO:0015038 | glutathione disulfide oxidoreductase activity(GO:0015038) |
| 0.1 | 0.8 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.1 | 0.4 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.1 | 2.1 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.1 | 1.5 | GO:0070403 | NAD+ binding(GO:0070403) |
| 0.1 | 0.6 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.1 | 0.6 | GO:0031386 | protein tag(GO:0031386) |
| 0.1 | 0.3 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 0.0 | 0.1 | GO:0004821 | histidine-tRNA ligase activity(GO:0004821) |
| 0.0 | 0.5 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.0 | 0.1 | GO:1990460 | leptin receptor binding(GO:1990460) |
| 0.0 | 0.2 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 0.0 | 0.1 | GO:0038181 | bile acid receptor activity(GO:0038181) |
| 0.0 | 0.8 | GO:0004298 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.0 | 0.4 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
| 0.0 | 0.1 | GO:0047374 | methylumbelliferyl-acetate deacetylase activity(GO:0047374) |
| 0.0 | 0.1 | GO:0031752 | D5 dopamine receptor binding(GO:0031752) |
| 0.0 | 0.1 | GO:0030942 | endoplasmic reticulum signal peptide binding(GO:0030942) |
| 0.0 | 0.1 | GO:0000702 | oxidized base lesion DNA N-glycosylase activity(GO:0000702) |
| 0.0 | 0.5 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.3 | GO:0019864 | IgG binding(GO:0019864) |
| 0.0 | 0.3 | GO:0070513 | death domain binding(GO:0070513) |
| 0.0 | 0.1 | GO:0098505 | G-rich strand telomeric DNA binding(GO:0098505) |
| 0.0 | 0.1 | GO:0042296 | ISG15 transferase activity(GO:0042296) |
| 0.0 | 0.2 | GO:0003964 | telomerase activity(GO:0003720) RNA-directed DNA polymerase activity(GO:0003964) |
| 0.0 | 0.2 | GO:0008035 | high-density lipoprotein particle binding(GO:0008035) |
| 0.0 | 0.1 | GO:0035312 | 5'-3' exodeoxyribonuclease activity(GO:0035312) |
| 0.0 | 0.0 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 0.0 | 0.1 | GO:0048030 | disaccharide binding(GO:0048030) |
| 0.0 | 0.9 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.0 | 0.5 | GO:0016409 | palmitoyltransferase activity(GO:0016409) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.4 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 0.7 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.0 | 0.8 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 0.5 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.4 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.1 | 2.2 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.0 | 0.8 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 0.5 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.0 | 1.6 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 0.0 | 0.7 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 0.3 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 0.0 | 0.0 | REACTOME VIF MEDIATED DEGRADATION OF APOBEC3G | Genes involved in Vif-mediated degradation of APOBEC3G |
| 0.0 | 0.7 | REACTOME EGFR DOWNREGULATION | Genes involved in EGFR downregulation |
| 0.0 | 0.3 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.0 | 0.8 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 1.1 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |