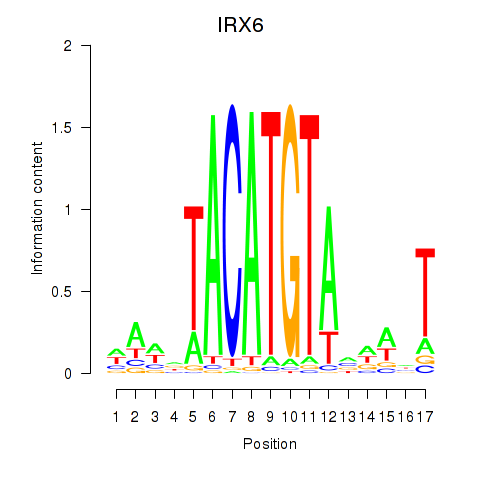
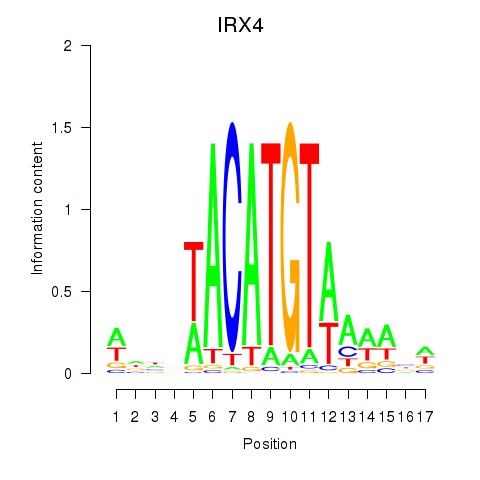
Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
Results for IRX6_IRX4
Z-value: 0.38
Transcription factors associated with IRX6_IRX4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
IRX6
|
ENSG00000159387.7 | IRX6 |
|
IRX4
|
ENSG00000113430.5 | IRX4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| IRX4 | hg19_v2_chr5_-_1882858_1883003 | 0.82 | 1.3e-02 | Click! |
Activity profile of IRX6_IRX4 motif
Sorted Z-values of IRX6_IRX4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of IRX6_IRX4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr18_-_47376197 | 0.36 |
ENST00000592688.1 |
MYO5B |
myosin VB |
| chr14_-_23624511 | 0.20 |
ENST00000529705.2 |
SLC7A8 |
solute carrier family 7 (amino acid transporter light chain, L system), member 8 |
| chr18_-_28622774 | 0.17 |
ENST00000434452.1 |
DSC3 |
desmocollin 3 |
| chr14_-_23623577 | 0.17 |
ENST00000422941.2 ENST00000453702.1 |
SLC7A8 |
solute carrier family 7 (amino acid transporter light chain, L system), member 8 |
| chr13_-_46716969 | 0.16 |
ENST00000435666.2 |
LCP1 |
lymphocyte cytosolic protein 1 (L-plastin) |
| chr1_-_201438282 | 0.15 |
ENST00000367311.3 ENST00000367309.1 |
PHLDA3 |
pleckstrin homology-like domain, family A, member 3 |
| chr3_-_27498235 | 0.13 |
ENST00000295736.5 ENST00000428386.1 ENST00000428179.1 |
SLC4A7 |
solute carrier family 4, sodium bicarbonate cotransporter, member 7 |
| chr11_+_6897856 | 0.13 |
ENST00000379829.2 |
OR10A4 |
olfactory receptor, family 10, subfamily A, member 4 |
| chr8_-_17555164 | 0.11 |
ENST00000297488.6 |
MTUS1 |
microtubule associated tumor suppressor 1 |
| chr21_-_34186006 | 0.10 |
ENST00000490358.1 |
C21orf62 |
chromosome 21 open reading frame 62 |
| chr21_-_34185989 | 0.09 |
ENST00000487113.1 ENST00000382373.4 |
C21orf62 |
chromosome 21 open reading frame 62 |
| chr12_+_32655110 | 0.09 |
ENST00000546442.1 ENST00000583694.1 |
FGD4 |
FYVE, RhoGEF and PH domain containing 4 |
| chr21_-_34185944 | 0.08 |
ENST00000479548.1 |
C21orf62 |
chromosome 21 open reading frame 62 |
| chr7_-_16844611 | 0.08 |
ENST00000401412.1 ENST00000419304.2 |
AGR2 |
anterior gradient 2 |
| chr4_+_71458012 | 0.07 |
ENST00000449493.2 |
AMBN |
ameloblastin (enamel matrix protein) |
| chr17_-_42327236 | 0.07 |
ENST00000399246.2 |
AC003102.1 |
AC003102.1 |
| chr1_+_220267429 | 0.06 |
ENST00000366922.1 ENST00000302637.5 |
IARS2 |
isoleucyl-tRNA synthetase 2, mitochondrial |
| chr18_+_616672 | 0.06 |
ENST00000338387.7 |
CLUL1 |
clusterin-like 1 (retinal) |
| chr16_+_777246 | 0.06 |
ENST00000561546.1 ENST00000564545.1 ENST00000389703.3 ENST00000567414.1 ENST00000568141.1 |
HAGHL |
hydroxyacylglutathione hydrolase-like |
| chr4_-_153303658 | 0.05 |
ENST00000296555.5 |
FBXW7 |
F-box and WD repeat domain containing 7, E3 ubiquitin protein ligase |
| chr2_+_113829895 | 0.05 |
ENST00000393197.2 |
IL1F10 |
interleukin 1 family, member 10 (theta) |
| chr16_+_776936 | 0.05 |
ENST00000549114.1 ENST00000341413.4 ENST00000562187.1 ENST00000564537.1 |
HAGHL |
hydroxyacylglutathione hydrolase-like |
| chr16_+_777118 | 0.05 |
ENST00000562141.1 |
HAGHL |
hydroxyacylglutathione hydrolase-like |
| chr4_-_143481822 | 0.04 |
ENST00000510812.1 |
INPP4B |
inositol polyphosphate-4-phosphatase, type II, 105kDa |
| chr9_-_27573651 | 0.03 |
ENST00000379995.1 ENST00000379997.3 |
C9orf72 |
chromosome 9 open reading frame 72 |
| chr2_-_231989808 | 0.03 |
ENST00000258400.3 |
HTR2B |
5-hydroxytryptamine (serotonin) receptor 2B, G protein-coupled |
| chr4_+_41361616 | 0.03 |
ENST00000513024.1 |
LIMCH1 |
LIM and calponin homology domains 1 |
| chr14_+_56127989 | 0.02 |
ENST00000555573.1 |
KTN1 |
kinectin 1 (kinesin receptor) |
| chr15_-_64665911 | 0.02 |
ENST00000606793.1 ENST00000561349.1 ENST00000560278.1 |
CTD-2116N17.1 |
Uncharacterized protein |
| chr5_+_147443534 | 0.02 |
ENST00000398454.1 ENST00000359874.3 ENST00000508733.1 ENST00000256084.7 |
SPINK5 |
serine peptidase inhibitor, Kazal type 5 |
| chr14_-_106845789 | 0.02 |
ENST00000390617.2 |
IGHV3-35 |
immunoglobulin heavy variable 3-35 (non-functional) |
| chr3_+_111630451 | 0.02 |
ENST00000495180.1 |
PHLDB2 |
pleckstrin homology-like domain, family B, member 2 |
| chr3_+_171561127 | 0.02 |
ENST00000334567.5 ENST00000450693.1 |
TMEM212 |
transmembrane protein 212 |
| chr2_+_58655461 | 0.02 |
ENST00000429095.1 ENST00000429664.1 ENST00000452840.1 |
AC007092.1 |
long intergenic non-protein coding RNA 1122 |
| chr4_+_141264597 | 0.02 |
ENST00000338517.4 ENST00000394203.3 ENST00000506322.1 |
SCOC |
short coiled-coil protein |
| chr14_-_21737551 | 0.02 |
ENST00000554891.1 ENST00000555883.1 ENST00000553753.1 ENST00000555914.1 ENST00000557336.1 ENST00000555215.1 ENST00000556628.1 ENST00000555137.1 ENST00000556226.1 ENST00000555309.1 ENST00000556142.1 ENST00000554969.1 ENST00000554455.1 ENST00000556513.1 ENST00000557201.1 ENST00000420743.2 ENST00000557768.1 ENST00000553300.1 ENST00000554383.1 ENST00000554539.1 |
HNRNPC |
heterogeneous nuclear ribonucleoprotein C (C1/C2) |
| chr2_+_133874577 | 0.02 |
ENST00000596384.1 |
AC011755.1 |
HCG2006742; Protein LOC100996685 |
| chr6_-_39290744 | 0.01 |
ENST00000507712.1 |
KCNK16 |
potassium channel, subfamily K, member 16 |
| chrY_-_6742068 | 0.01 |
ENST00000215479.5 |
AMELY |
amelogenin, Y-linked |
| chr8_+_84824920 | 0.01 |
ENST00000523678.1 |
RP11-120I21.2 |
RP11-120I21.2 |
| chr9_-_37465396 | 0.01 |
ENST00000307750.4 |
ZBTB5 |
zinc finger and BTB domain containing 5 |
| chr15_+_66585879 | 0.01 |
ENST00000319212.4 |
DIS3L |
DIS3 mitotic control homolog (S. cerevisiae)-like |
| chrX_+_99899180 | 0.01 |
ENST00000373004.3 |
SRPX2 |
sushi-repeat containing protein, X-linked 2 |
| chr6_-_108145499 | 0.01 |
ENST00000369020.3 ENST00000369022.2 |
SCML4 |
sex comb on midleg-like 4 (Drosophila) |
| chr19_-_48059113 | 0.01 |
ENST00000391901.3 ENST00000314121.4 ENST00000448976.1 |
ZNF541 |
zinc finger protein 541 |
| chr15_+_66585555 | 0.01 |
ENST00000319194.5 ENST00000525134.2 ENST00000441424.2 |
DIS3L |
DIS3 mitotic control homolog (S. cerevisiae)-like |
| chr5_+_54320078 | 0.01 |
ENST00000231009.2 |
GZMK |
granzyme K (granzyme 3; tryptase II) |
| chr6_-_52705641 | 0.01 |
ENST00000370989.2 |
GSTA5 |
glutathione S-transferase alpha 5 |
| chr20_+_11008408 | 0.01 |
ENST00000378252.1 |
C20orf187 |
chromosome 20 open reading frame 187 |
| chr2_+_160590469 | 0.01 |
ENST00000409591.1 |
MARCH7 |
membrane-associated ring finger (C3HC4) 7, E3 ubiquitin protein ligase |
| chr12_+_5541267 | 0.01 |
ENST00000423158.3 |
NTF3 |
neurotrophin 3 |
| chr11_+_61891445 | 0.01 |
ENST00000394818.3 ENST00000533896.1 ENST00000278849.4 |
INCENP |
inner centromere protein antigens 135/155kDa |
| chr4_-_177190364 | 0.01 |
ENST00000296525.3 |
ASB5 |
ankyrin repeat and SOCS box containing 5 |
| chrX_+_15767971 | 0.01 |
ENST00000479740.1 ENST00000454127.2 |
CA5B |
carbonic anhydrase VB, mitochondrial |
| chr6_-_49834209 | 0.01 |
ENST00000507853.1 |
CRISP1 |
cysteine-rich secretory protein 1 |
| chr2_-_183106641 | 0.01 |
ENST00000346717.4 |
PDE1A |
phosphodiesterase 1A, calmodulin-dependent |
| chr1_+_53308398 | 0.01 |
ENST00000371528.1 |
ZYG11A |
zyg-11 family member A, cell cycle regulator |
| chr1_+_170904612 | 0.00 |
ENST00000367759.4 ENST00000367758.3 |
MROH9 |
maestro heat-like repeat family member 9 |
| chr6_-_49834240 | 0.00 |
ENST00000335847.4 |
CRISP1 |
cysteine-rich secretory protein 1 |
| chr4_+_130017268 | 0.00 |
ENST00000425929.1 ENST00000508673.1 ENST00000508622.1 |
C4orf33 |
chromosome 4 open reading frame 33 |
| chr15_-_51630772 | 0.00 |
ENST00000557858.1 ENST00000558328.1 ENST00000396404.4 ENST00000561075.1 ENST00000405011.2 ENST00000559980.1 ENST00000453807.2 ENST00000396402.1 |
CYP19A1 |
cytochrome P450, family 19, subfamily A, polypeptide 1 |
| chr8_+_27631903 | 0.00 |
ENST00000305188.8 |
ESCO2 |
establishment of sister chromatid cohesion N-acetyltransferase 2 |
| chr20_+_10199468 | 0.00 |
ENST00000254976.2 ENST00000304886.2 |
SNAP25 |
synaptosomal-associated protein, 25kDa |
| chr1_+_26644441 | 0.00 |
ENST00000374213.2 |
CD52 |
CD52 molecule |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | GO:0032439 | endosome localization(GO:0032439) |
| 0.0 | 0.1 | GO:0006428 | isoleucyl-tRNA aminoacylation(GO:0006428) |
| 0.0 | 0.1 | GO:1903899 | lung goblet cell differentiation(GO:0060480) positive regulation of PERK-mediated unfolded protein response(GO:1903899) |
| 0.0 | 0.4 | GO:0015695 | organic cation transport(GO:0015695) |
| 0.0 | 0.1 | GO:2000638 | regulation of SREBP signaling pathway(GO:2000638) negative regulation of SREBP signaling pathway(GO:2000639) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | GO:0045179 | apical cortex(GO:0045179) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | GO:0019534 | toxin transporter activity(GO:0019534) |
| 0.0 | 0.1 | GO:0004822 | isoleucine-tRNA ligase activity(GO:0004822) |