Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
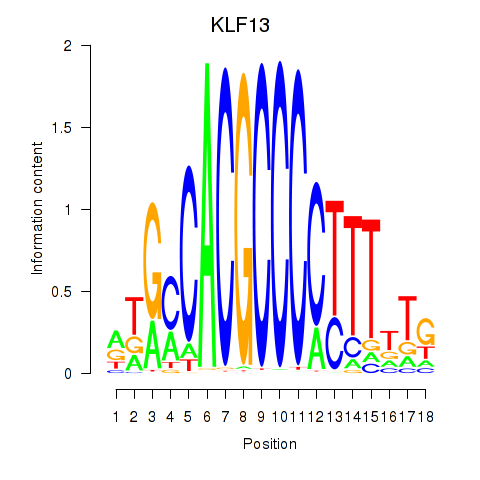
Results for KLF13
Z-value: 0.33
Transcription factors associated with KLF13
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
KLF13
|
ENSG00000169926.5 | KLF13 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| KLF13 | hg19_v2_chr15_+_31658349_31658358 | 0.22 | 6.0e-01 | Click! |
Activity profile of KLF13 motif
Sorted Z-values of KLF13 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of KLF13
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_-_7936344 | 0.48 |
ENST00000599142.1 |
CTD-3193O13.9 |
Protein FLJ22184 |
| chr6_-_31697563 | 0.30 |
ENST00000375789.2 ENST00000416410.1 |
DDAH2 |
dimethylarginine dimethylaminohydrolase 2 |
| chr6_-_31697977 | 0.29 |
ENST00000375787.2 |
DDAH2 |
dimethylarginine dimethylaminohydrolase 2 |
| chr5_+_178450753 | 0.27 |
ENST00000444149.2 ENST00000519896.1 ENST00000522442.1 |
ZNF879 |
zinc finger protein 879 |
| chr18_-_56940611 | 0.27 |
ENST00000256852.7 ENST00000334889.3 |
RAX |
retina and anterior neural fold homeobox |
| chr20_+_34129770 | 0.22 |
ENST00000348547.2 ENST00000357394.4 ENST00000447986.1 ENST00000279052.6 ENST00000416206.1 ENST00000411577.1 ENST00000413587.1 |
ERGIC3 |
ERGIC and golgi 3 |
| chr1_-_16939976 | 0.20 |
ENST00000430580.2 |
NBPF1 |
neuroblastoma breakpoint family, member 1 |
| chr19_-_12941163 | 0.19 |
ENST00000589567.1 |
RTBDN |
retbindin |
| chr6_-_169364429 | 0.19 |
ENST00000444586.1 |
RP3-495K2.3 |
RP3-495K2.3 |
| chr19_-_50311896 | 0.18 |
ENST00000529634.2 |
FUZ |
fuzzy planar cell polarity protein |
| chr15_+_41056218 | 0.18 |
ENST00000260447.4 |
GCHFR |
GTP cyclohydrolase I feedback regulator |
| chr15_+_41056255 | 0.17 |
ENST00000561160.1 ENST00000559445.1 |
GCHFR |
GTP cyclohydrolase I feedback regulator |
| chr12_+_58013693 | 0.16 |
ENST00000320442.4 ENST00000379218.2 |
SLC26A10 |
solute carrier family 26, member 10 |
| chr11_+_44587141 | 0.16 |
ENST00000227155.4 ENST00000342935.3 ENST00000532544.1 |
CD82 |
CD82 molecule |
| chr2_+_71295733 | 0.16 |
ENST00000443938.2 ENST00000244204.6 |
NAGK |
N-acetylglucosamine kinase |
| chr6_+_132129151 | 0.15 |
ENST00000360971.2 |
ENPP1 |
ectonucleotide pyrophosphatase/phosphodiesterase 1 |
| chr2_+_71295717 | 0.15 |
ENST00000418807.3 ENST00000443872.2 |
NAGK |
N-acetylglucosamine kinase |
| chr2_-_97405775 | 0.15 |
ENST00000264963.4 ENST00000537039.1 ENST00000377079.4 ENST00000426463.2 ENST00000534882.1 |
LMAN2L |
lectin, mannose-binding 2-like |
| chr14_-_67981870 | 0.15 |
ENST00000555994.1 |
TMEM229B |
transmembrane protein 229B |
| chr15_+_28623784 | 0.15 |
ENST00000526619.2 ENST00000337838.7 ENST00000532622.2 |
GOLGA8F |
golgin A8 family, member F |
| chr7_+_33169142 | 0.15 |
ENST00000242067.6 ENST00000350941.3 ENST00000396127.2 ENST00000355070.2 ENST00000354265.4 ENST00000425508.2 |
BBS9 |
Bardet-Biedl syndrome 9 |
| chr19_+_49866851 | 0.14 |
ENST00000221498.2 ENST00000596402.1 |
DKKL1 |
dickkopf-like 1 |
| chr17_+_42264395 | 0.13 |
ENST00000587989.1 ENST00000590235.1 |
TMUB2 |
transmembrane and ubiquitin-like domain containing 2 |
| chr17_+_42264322 | 0.13 |
ENST00000446571.3 ENST00000357984.3 ENST00000538716.2 |
TMUB2 |
transmembrane and ubiquitin-like domain containing 2 |
| chr6_-_31697255 | 0.13 |
ENST00000436437.1 |
DDAH2 |
dimethylarginine dimethylaminohydrolase 2 |
| chr17_+_27055798 | 0.13 |
ENST00000268766.6 |
NEK8 |
NIMA-related kinase 8 |
| chr4_+_55524085 | 0.12 |
ENST00000412167.2 ENST00000288135.5 |
KIT |
v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog |
| chr19_-_12941469 | 0.12 |
ENST00000586969.1 ENST00000589681.1 ENST00000585384.1 ENST00000589808.1 |
RTBDN |
retbindin |
| chr19_+_5455421 | 0.12 |
ENST00000222033.4 |
ZNRF4 |
zinc and ring finger 4 |
| chr19_+_45349432 | 0.12 |
ENST00000252485.4 |
PVRL2 |
poliovirus receptor-related 2 (herpesvirus entry mediator B) |
| chr1_+_45805342 | 0.12 |
ENST00000372090.5 |
TOE1 |
target of EGR1, member 1 (nuclear) |
| chr9_+_35605274 | 0.12 |
ENST00000336395.5 |
TESK1 |
testis-specific kinase 1 |
| chr20_-_8000426 | 0.12 |
ENST00000527925.1 ENST00000246024.2 |
TMX4 |
thioredoxin-related transmembrane protein 4 |
| chr10_-_13344341 | 0.12 |
ENST00000396920.3 |
PHYH |
phytanoyl-CoA 2-hydroxylase |
| chr9_-_116163400 | 0.12 |
ENST00000277315.5 ENST00000448137.1 ENST00000409155.3 |
ALAD |
aminolevulinate dehydratase |
| chr17_+_42264556 | 0.12 |
ENST00000319511.6 ENST00000589785.1 ENST00000592825.1 ENST00000589184.1 |
TMUB2 |
transmembrane and ubiquitin-like domain containing 2 |
| chr21_-_34100341 | 0.12 |
ENST00000382499.2 ENST00000433931.2 |
SYNJ1 |
synaptojanin 1 |
| chr16_-_28936493 | 0.12 |
ENST00000544477.1 ENST00000357573.6 |
RABEP2 |
rabaptin, RAB GTPase binding effector protein 2 |
| chr21_-_34100244 | 0.12 |
ENST00000382491.3 ENST00000357345.3 ENST00000429236.1 |
SYNJ1 |
synaptojanin 1 |
| chr15_+_23255242 | 0.11 |
ENST00000450802.3 |
GOLGA8I |
golgin A8 family, member I |
| chr8_-_37756972 | 0.11 |
ENST00000330843.4 ENST00000522727.1 ENST00000287263.4 |
RAB11FIP1 |
RAB11 family interacting protein 1 (class I) |
| chr17_-_6915646 | 0.11 |
ENST00000574377.1 ENST00000399541.2 ENST00000399540.2 ENST00000575727.1 ENST00000573939.1 |
AC027763.2 |
Uncharacterized protein |
| chr12_-_6580094 | 0.11 |
ENST00000361716.3 |
VAMP1 |
vesicle-associated membrane protein 1 (synaptobrevin 1) |
| chr5_+_170288856 | 0.11 |
ENST00000523189.1 |
RANBP17 |
RAN binding protein 17 |
| chrX_+_15756382 | 0.11 |
ENST00000318636.3 |
CA5B |
carbonic anhydrase VB, mitochondrial |
| chr2_+_179316163 | 0.10 |
ENST00000409117.3 |
DFNB59 |
deafness, autosomal recessive 59 |
| chr10_+_120789223 | 0.10 |
ENST00000425699.1 |
NANOS1 |
nanos homolog 1 (Drosophila) |
| chr9_+_72658490 | 0.10 |
ENST00000377182.4 |
MAMDC2 |
MAM domain containing 2 |
| chr1_+_33352036 | 0.10 |
ENST00000373467.3 |
HPCA |
hippocalcin |
| chrX_-_140271249 | 0.10 |
ENST00000370526.2 |
LDOC1 |
leucine zipper, down-regulated in cancer 1 |
| chrX_+_54835493 | 0.10 |
ENST00000396224.1 |
MAGED2 |
melanoma antigen family D, 2 |
| chr20_+_48599506 | 0.10 |
ENST00000244050.2 |
SNAI1 |
snail family zinc finger 1 |
| chr1_+_45805728 | 0.10 |
ENST00000539779.1 |
TOE1 |
target of EGR1, member 1 (nuclear) |
| chr3_+_158519654 | 0.09 |
ENST00000415822.2 ENST00000392813.4 ENST00000264266.8 |
MFSD1 |
major facilitator superfamily domain containing 1 |
| chr19_-_12941215 | 0.09 |
ENST00000458671.2 |
RTBDN |
retbindin |
| chr17_+_19281034 | 0.09 |
ENST00000308406.5 ENST00000299612.7 |
MAPK7 |
mitogen-activated protein kinase 7 |
| chr12_-_124457257 | 0.09 |
ENST00000545891.1 |
CCDC92 |
coiled-coil domain containing 92 |
| chr6_+_31126291 | 0.09 |
ENST00000376257.3 ENST00000376255.4 |
TCF19 |
transcription factor 19 |
| chr19_+_36249057 | 0.09 |
ENST00000301165.5 ENST00000536950.1 ENST00000537459.1 ENST00000421853.2 |
C19orf55 |
chromosome 19 open reading frame 55 |
| chr19_+_49867181 | 0.09 |
ENST00000597546.1 |
DKKL1 |
dickkopf-like 1 |
| chr19_-_19843900 | 0.09 |
ENST00000344099.3 |
ZNF14 |
zinc finger protein 14 |
| chr6_-_31125850 | 0.08 |
ENST00000507751.1 ENST00000448162.2 ENST00000502557.1 ENST00000503420.1 ENST00000507892.1 ENST00000507226.1 ENST00000513222.1 ENST00000503934.1 ENST00000396263.2 ENST00000508683.1 ENST00000428174.1 ENST00000448141.2 ENST00000507829.1 ENST00000455279.2 ENST00000376266.5 |
CCHCR1 |
coiled-coil alpha-helical rod protein 1 |
| chr19_-_12886327 | 0.08 |
ENST00000397668.3 ENST00000587178.1 ENST00000264827.5 |
HOOK2 |
hook microtubule-tethering protein 2 |
| chr15_-_28778117 | 0.08 |
ENST00000525590.2 ENST00000329523.6 |
GOLGA8G |
golgin A8 family, member G |
| chr1_-_146082633 | 0.08 |
ENST00000605317.1 ENST00000604938.1 ENST00000339388.5 |
NBPF11 |
neuroblastoma breakpoint family, member 11 |
| chr17_+_74733744 | 0.08 |
ENST00000586689.1 ENST00000587661.1 ENST00000593181.1 ENST00000336509.4 ENST00000355954.3 |
MFSD11 |
major facilitator superfamily domain containing 11 |
| chr7_+_100450328 | 0.08 |
ENST00000540482.1 ENST00000418037.1 ENST00000428758.1 ENST00000275729.3 ENST00000415287.1 ENST00000354161.3 ENST00000416675.1 |
SLC12A9 |
solute carrier family 12, member 9 |
| chr12_-_124457371 | 0.08 |
ENST00000238156.3 ENST00000545037.1 |
CCDC92 |
coiled-coil domain containing 92 |
| chr12_-_6579808 | 0.08 |
ENST00000535180.1 ENST00000400911.3 |
VAMP1 |
vesicle-associated membrane protein 1 (synaptobrevin 1) |
| chr14_+_23067146 | 0.08 |
ENST00000428304.2 |
ABHD4 |
abhydrolase domain containing 4 |
| chr6_+_30594619 | 0.08 |
ENST00000318999.7 ENST00000376485.4 ENST00000376478.2 ENST00000319027.5 ENST00000376483.4 ENST00000329992.8 ENST00000330083.5 |
ATAT1 |
alpha tubulin acetyltransferase 1 |
| chr10_+_104678032 | 0.08 |
ENST00000369878.4 ENST00000369875.3 |
CNNM2 |
cyclin M2 |
| chr12_+_121124921 | 0.07 |
ENST00000412616.2 |
MLEC |
malectin |
| chr10_+_102672712 | 0.07 |
ENST00000370271.3 ENST00000370269.3 ENST00000609386.1 |
FAM178A |
family with sequence similarity 178, member A |
| chr19_+_49713991 | 0.07 |
ENST00000597316.1 |
TRPM4 |
transient receptor potential cation channel, subfamily M, member 4 |
| chr15_+_82722225 | 0.07 |
ENST00000300515.8 |
GOLGA6L9 |
golgin A6 family-like 9 |
| chr5_+_154237778 | 0.07 |
ENST00000523698.1 ENST00000517876.1 ENST00000520472.1 |
CNOT8 |
CCR4-NOT transcription complex, subunit 8 |
| chr3_+_50606577 | 0.07 |
ENST00000434410.1 ENST00000232854.4 |
HEMK1 |
HemK methyltransferase family member 1 |
| chr2_-_8723918 | 0.07 |
ENST00000454224.1 |
AC011747.4 |
AC011747.4 |
| chr1_+_65775204 | 0.07 |
ENST00000371069.4 |
DNAJC6 |
DnaJ (Hsp40) homolog, subfamily C, member 6 |
| chr19_+_50979753 | 0.07 |
ENST00000597426.1 ENST00000334976.6 ENST00000376918.3 ENST00000598585.1 |
EMC10 |
ER membrane protein complex subunit 10 |
| chr2_-_8977714 | 0.07 |
ENST00000319688.5 ENST00000489024.1 ENST00000256707.3 ENST00000427284.1 ENST00000418530.1 ENST00000473731.1 |
KIDINS220 |
kinase D-interacting substrate, 220kDa |
| chr19_+_49403562 | 0.07 |
ENST00000407032.1 ENST00000452087.1 ENST00000411700.1 |
NUCB1 |
nucleobindin 1 |
| chr9_-_130533615 | 0.07 |
ENST00000373277.4 |
SH2D3C |
SH2 domain containing 3C |
| chr3_+_132136331 | 0.07 |
ENST00000260818.6 |
DNAJC13 |
DnaJ (Hsp40) homolog, subfamily C, member 13 |
| chr11_-_82612727 | 0.07 |
ENST00000531128.1 ENST00000535099.1 ENST00000527444.1 |
PRCP |
prolylcarboxypeptidase (angiotensinase C) |
| chr15_+_84904525 | 0.07 |
ENST00000510439.2 |
GOLGA6L4 |
golgin A6 family-like 4 |
| chr19_-_19729725 | 0.07 |
ENST00000251203.9 |
PBX4 |
pre-B-cell leukemia homeobox 4 |
| chr19_+_49866331 | 0.06 |
ENST00000597873.1 |
DKKL1 |
dickkopf-like 1 |
| chr15_-_101817616 | 0.06 |
ENST00000526049.1 ENST00000398226.3 ENST00000537379.1 |
VIMP |
VCP-interacting membrane protein |
| chr7_-_99756293 | 0.06 |
ENST00000316937.3 ENST00000456769.1 |
C7orf43 |
chromosome 7 open reading frame 43 |
| chr15_+_83098710 | 0.06 |
ENST00000561062.1 ENST00000358583.3 |
GOLGA6L9 |
golgin A6 family-like 20 |
| chr12_+_6982725 | 0.06 |
ENST00000433346.1 |
LRRC23 |
leucine rich repeat containing 23 |
| chr6_+_33378738 | 0.06 |
ENST00000374512.3 ENST00000374516.3 |
PHF1 |
PHD finger protein 1 |
| chr5_+_111496631 | 0.06 |
ENST00000508590.1 |
EPB41L4A-AS1 |
EPB41L4A antisense RNA 1 |
| chr11_+_7041677 | 0.06 |
ENST00000299481.4 |
NLRP14 |
NLR family, pyrin domain containing 14 |
| chr14_-_94970559 | 0.06 |
ENST00000556881.1 |
SERPINA12 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 12 |
| chr13_-_95953589 | 0.06 |
ENST00000538287.1 ENST00000376887.4 ENST00000412704.1 ENST00000536256.1 ENST00000431522.1 |
ABCC4 |
ATP-binding cassette, sub-family C (CFTR/MRP), member 4 |
| chr5_+_154238042 | 0.06 |
ENST00000519211.1 ENST00000522458.1 ENST00000519903.1 ENST00000521450.1 ENST00000403027.2 |
CNOT8 |
CCR4-NOT transcription complex, subunit 8 |
| chr1_+_172502336 | 0.06 |
ENST00000263688.3 |
SUCO |
SUN domain containing ossification factor |
| chr19_-_9420249 | 0.06 |
ENST00000591998.1 |
ZNF699 |
zinc finger protein 699 |
| chr19_+_45458503 | 0.05 |
ENST00000337392.5 ENST00000591304.1 |
CLPTM1 |
cleft lip and palate associated transmembrane protein 1 |
| chr14_-_74551096 | 0.05 |
ENST00000350259.4 |
ALDH6A1 |
aldehyde dehydrogenase 6 family, member A1 |
| chr14_-_74551172 | 0.05 |
ENST00000553458.1 |
ALDH6A1 |
aldehyde dehydrogenase 6 family, member A1 |
| chr5_+_154238096 | 0.05 |
ENST00000517568.1 ENST00000524105.1 ENST00000285896.6 |
CNOT8 |
CCR4-NOT transcription complex, subunit 8 |
| chr19_+_36236514 | 0.05 |
ENST00000222266.2 |
PSENEN |
presenilin enhancer gamma secretase subunit |
| chr17_+_74734052 | 0.05 |
ENST00000590514.1 |
MFSD11 |
major facilitator superfamily domain containing 11 |
| chr13_+_27131798 | 0.05 |
ENST00000361042.4 |
WASF3 |
WAS protein family, member 3 |
| chr7_-_111846435 | 0.05 |
ENST00000437633.1 ENST00000428084.1 |
DOCK4 |
dedicator of cytokinesis 4 |
| chr12_+_56521840 | 0.05 |
ENST00000394048.5 |
ESYT1 |
extended synaptotagmin-like protein 1 |
| chr19_+_36236491 | 0.05 |
ENST00000591949.1 |
PSENEN |
presenilin enhancer gamma secretase subunit |
| chr14_+_93260642 | 0.05 |
ENST00000355976.2 |
GOLGA5 |
golgin A5 |
| chr19_-_15236562 | 0.05 |
ENST00000263383.3 |
ILVBL |
ilvB (bacterial acetolactate synthase)-like |
| chr11_+_121163466 | 0.05 |
ENST00000527762.1 ENST00000534230.1 ENST00000392789.2 |
SC5D |
sterol-C5-desaturase |
| chr1_-_224517823 | 0.05 |
ENST00000469968.1 ENST00000436927.1 ENST00000469075.1 ENST00000488718.1 ENST00000482491.1 ENST00000340871.4 ENST00000492281.1 ENST00000361463.3 ENST00000391875.2 ENST00000461546.1 |
NVL |
nuclear VCP-like |
| chr1_+_162467595 | 0.04 |
ENST00000538489.1 ENST00000489294.1 |
UHMK1 |
U2AF homology motif (UHM) kinase 1 |
| chr7_+_99102267 | 0.04 |
ENST00000326775.5 ENST00000451158.1 |
ZKSCAN5 |
zinc finger with KRAB and SCAN domains 5 |
| chr3_+_184529929 | 0.04 |
ENST00000287546.4 ENST00000437079.3 |
VPS8 |
vacuolar protein sorting 8 homolog (S. cerevisiae) |
| chr21_+_48055527 | 0.04 |
ENST00000397638.2 ENST00000458387.2 ENST00000451211.2 ENST00000291705.6 ENST00000397637.1 ENST00000334494.4 ENST00000397628.1 ENST00000440086.1 |
PRMT2 |
protein arginine methyltransferase 2 |
| chr5_+_111496554 | 0.04 |
ENST00000442823.2 |
EPB41L4A-AS1 |
EPB41L4A antisense RNA 1 |
| chr7_+_99102573 | 0.04 |
ENST00000394170.2 |
ZKSCAN5 |
zinc finger with KRAB and SCAN domains 5 |
| chr4_+_4291924 | 0.04 |
ENST00000355834.3 ENST00000337872.4 ENST00000538529.1 ENST00000502918.1 |
ZBTB49 |
zinc finger and BTB domain containing 49 |
| chr17_+_7338737 | 0.04 |
ENST00000323206.1 ENST00000396568.1 |
TMEM102 |
transmembrane protein 102 |
| chr11_-_7041466 | 0.04 |
ENST00000536068.1 ENST00000278314.4 |
ZNF214 |
zinc finger protein 214 |
| chr14_+_54976546 | 0.04 |
ENST00000216420.7 |
CGRRF1 |
cell growth regulator with ring finger domain 1 |
| chr11_-_6426635 | 0.04 |
ENST00000608645.1 ENST00000608394.1 ENST00000529519.1 |
APBB1 |
amyloid beta (A4) precursor protein-binding, family B, member 1 (Fe65) |
| chr7_-_155089251 | 0.04 |
ENST00000609974.1 |
AC144652.1 |
AC144652.1 |
| chr3_+_184529948 | 0.04 |
ENST00000436792.2 ENST00000446204.2 ENST00000422105.1 |
VPS8 |
vacuolar protein sorting 8 homolog (S. cerevisiae) |
| chr12_-_29534074 | 0.04 |
ENST00000546839.1 ENST00000360150.4 ENST00000552155.1 ENST00000550353.1 ENST00000548441.1 ENST00000552132.1 |
ERGIC2 |
ERGIC and golgi 2 |
| chr8_-_145013711 | 0.04 |
ENST00000345136.3 |
PLEC |
plectin |
| chr15_-_74374891 | 0.04 |
ENST00000290438.3 |
GOLGA6A |
golgin A6 family, member A |
| chr6_-_2962331 | 0.04 |
ENST00000380524.1 |
SERPINB6 |
serpin peptidase inhibitor, clade B (ovalbumin), member 6 |
| chr7_-_91763822 | 0.04 |
ENST00000003100.8 |
CYP51A1 |
cytochrome P450, family 51, subfamily A, polypeptide 1 |
| chr2_-_160143158 | 0.04 |
ENST00000409124.1 ENST00000358147.4 |
WDSUB1 |
WD repeat, sterile alpha motif and U-box domain containing 1 |
| chr4_+_56719782 | 0.04 |
ENST00000381295.2 ENST00000346134.7 ENST00000349598.6 |
EXOC1 |
exocyst complex component 1 |
| chr22_-_41985865 | 0.04 |
ENST00000216259.7 |
PMM1 |
phosphomannomutase 1 |
| chr9_-_69262509 | 0.04 |
ENST00000377449.1 ENST00000382399.4 ENST00000377439.1 ENST00000377441.1 ENST00000377457.5 |
CBWD6 |
COBW domain containing 6 |
| chrX_+_72062617 | 0.04 |
ENST00000440247.1 |
DMRTC1B |
DMRT-like family C1B |
| chr13_-_30424821 | 0.04 |
ENST00000380680.4 |
UBL3 |
ubiquitin-like 3 |
| chr21_+_15588499 | 0.04 |
ENST00000400577.3 |
RBM11 |
RNA binding motif protein 11 |
| chr9_-_35111420 | 0.03 |
ENST00000378557.1 |
FAM214B |
family with sequence similarity 214, member B |
| chr21_+_44394620 | 0.03 |
ENST00000291547.5 |
PKNOX1 |
PBX/knotted 1 homeobox 1 |
| chr12_-_48152853 | 0.03 |
ENST00000171000.4 |
RAPGEF3 |
Rap guanine nucleotide exchange factor (GEF) 3 |
| chr9_-_179018 | 0.03 |
ENST00000431099.2 ENST00000382447.4 ENST00000382389.1 ENST00000377447.3 ENST00000314367.10 ENST00000356521.4 ENST00000382393.1 ENST00000377400.4 |
CBWD1 |
COBW domain containing 1 |
| chr10_+_104678102 | 0.03 |
ENST00000433628.2 |
CNNM2 |
cyclin M2 |
| chr17_-_5487277 | 0.03 |
ENST00000572272.1 ENST00000354411.3 ENST00000577119.1 |
NLRP1 |
NLR family, pyrin domain containing 1 |
| chr13_+_27131887 | 0.03 |
ENST00000335327.5 |
WASF3 |
WAS protein family, member 3 |
| chr15_-_30685752 | 0.03 |
ENST00000299847.2 ENST00000397827.3 |
CHRFAM7A |
CHRNA7 (cholinergic receptor, nicotinic, alpha 7, exons 5-10) and FAM7A (family with sequence similarity 7A, exons A-E) fusion |
| chr1_-_25558984 | 0.03 |
ENST00000236273.4 |
SYF2 |
SYF2 pre-mRNA-splicing factor |
| chr4_-_13546632 | 0.03 |
ENST00000382438.5 |
NKX3-2 |
NK3 homeobox 2 |
| chr6_+_52535878 | 0.03 |
ENST00000211314.4 |
TMEM14A |
transmembrane protein 14A |
| chr3_+_50606901 | 0.03 |
ENST00000455834.1 |
HEMK1 |
HemK methyltransferase family member 1 |
| chr19_+_36632056 | 0.03 |
ENST00000586851.1 ENST00000590211.1 |
CAPNS1 |
calpain, small subunit 1 |
| chr20_-_23402028 | 0.03 |
ENST00000398425.3 ENST00000432543.2 ENST00000377026.4 |
NAPB |
N-ethylmaleimide-sensitive factor attachment protein, beta |
| chr14_+_93260569 | 0.03 |
ENST00000163416.2 |
GOLGA5 |
golgin A5 |
| chr2_-_160143242 | 0.03 |
ENST00000359774.4 |
WDSUB1 |
WD repeat, sterile alpha motif and U-box domain containing 1 |
| chr2_-_160143084 | 0.03 |
ENST00000409990.3 |
WDSUB1 |
WD repeat, sterile alpha motif and U-box domain containing 1 |
| chr7_+_111846741 | 0.03 |
ENST00000421043.1 ENST00000425229.1 ENST00000450657.1 |
ZNF277 |
zinc finger protein 277 |
| chr9_-_35111570 | 0.03 |
ENST00000378561.1 ENST00000603301.1 |
FAM214B |
family with sequence similarity 214, member B |
| chr13_+_100153665 | 0.03 |
ENST00000376387.4 |
TM9SF2 |
transmembrane 9 superfamily member 2 |
| chr10_+_101292684 | 0.03 |
ENST00000344586.7 |
NKX2-3 |
NK2 homeobox 3 |
| chr18_-_51750948 | 0.03 |
ENST00000583046.1 ENST00000398398.2 |
MBD2 |
methyl-CpG binding domain protein 2 |
| chr1_+_100436065 | 0.03 |
ENST00000370153.1 |
SLC35A3 |
solute carrier family 35 (UDP-N-acetylglucosamine (UDP-GlcNAc) transporter), member A3 |
| chr19_-_2151523 | 0.03 |
ENST00000350812.6 ENST00000355272.6 ENST00000356926.4 ENST00000345016.5 |
AP3D1 |
adaptor-related protein complex 3, delta 1 subunit |
| chr15_+_41851211 | 0.03 |
ENST00000263798.3 |
TYRO3 |
TYRO3 protein tyrosine kinase |
| chr4_-_76439483 | 0.03 |
ENST00000380840.2 ENST00000513257.1 ENST00000507014.1 |
RCHY1 |
ring finger and CHY zinc finger domain containing 1, E3 ubiquitin protein ligase |
| chr15_+_90792760 | 0.03 |
ENST00000339615.5 ENST00000438251.1 |
TTLL13 |
tubulin tyrosine ligase-like family, member 13 |
| chr10_-_31320860 | 0.03 |
ENST00000436087.2 ENST00000442986.1 ENST00000413025.1 ENST00000452305.1 |
ZNF438 |
zinc finger protein 438 |
| chr5_+_154238149 | 0.03 |
ENST00000519430.1 ENST00000520671.1 ENST00000521583.1 ENST00000518028.1 ENST00000519404.1 ENST00000519394.1 ENST00000518775.1 |
CNOT8 |
CCR4-NOT transcription complex, subunit 8 |
| chr19_+_45349630 | 0.02 |
ENST00000252483.5 |
PVRL2 |
poliovirus receptor-related 2 (herpesvirus entry mediator B) |
| chr2_-_160143059 | 0.02 |
ENST00000392796.3 |
WDSUB1 |
WD repeat, sterile alpha motif and U-box domain containing 1 |
| chrX_+_153170189 | 0.02 |
ENST00000358927.2 |
AVPR2 |
arginine vasopressin receptor 2 |
| chr17_+_7184986 | 0.02 |
ENST00000317370.8 ENST00000571308.1 |
SLC2A4 |
solute carrier family 2 (facilitated glucose transporter), member 4 |
| chr19_+_7985198 | 0.02 |
ENST00000221573.6 ENST00000595637.1 |
SNAPC2 |
small nuclear RNA activating complex, polypeptide 2, 45kDa |
| chr1_+_15573757 | 0.02 |
ENST00000358897.4 ENST00000417793.1 ENST00000375999.3 ENST00000433640.2 |
FHAD1 |
forkhead-associated (FHA) phosphopeptide binding domain 1 |
| chr2_+_96257766 | 0.02 |
ENST00000272395.2 |
TRIM43 |
tripartite motif containing 43 |
| chr15_+_75575176 | 0.02 |
ENST00000434739.3 |
GOLGA6D |
golgin A6 family, member D |
| chr19_+_36631867 | 0.02 |
ENST00000588780.1 |
CAPNS1 |
calpain, small subunit 1 |
| chr12_+_121124599 | 0.02 |
ENST00000228506.3 |
MLEC |
malectin |
| chr19_-_47164386 | 0.02 |
ENST00000391916.2 ENST00000410105.2 |
DACT3 |
dishevelled-binding antagonist of beta-catenin 3 |
| chr17_-_27054952 | 0.02 |
ENST00000580518.1 |
TLCD1 |
TLC domain containing 1 |
| chr10_-_31320840 | 0.02 |
ENST00000375311.1 |
ZNF438 |
zinc finger protein 438 |
| chr20_+_2361545 | 0.02 |
ENST00000202625.2 ENST00000381423.1 |
TGM6 |
transglutaminase 6 |
| chr20_-_33264886 | 0.02 |
ENST00000217446.3 ENST00000452740.2 ENST00000374820.2 |
PIGU |
phosphatidylinositol glycan anchor biosynthesis, class U |
| chr6_+_28193037 | 0.02 |
ENST00000531981.1 ENST00000425468.2 ENST00000252207.5 ENST00000531979.1 ENST00000527436.1 |
ZSCAN9 |
zinc finger and SCAN domain containing 9 |
| chr2_+_200775971 | 0.02 |
ENST00000319974.5 |
C2orf69 |
chromosome 2 open reading frame 69 |
| chr8_-_62627057 | 0.02 |
ENST00000519234.1 ENST00000379449.6 ENST00000379454.4 ENST00000518068.1 ENST00000517856.1 ENST00000356457.5 |
ASPH |
aspartate beta-hydroxylase |
| chr20_-_44600810 | 0.02 |
ENST00000322927.2 ENST00000426788.1 |
ZNF335 |
zinc finger protein 335 |
| chr1_-_25558963 | 0.02 |
ENST00000354361.3 |
SYF2 |
SYF2 pre-mRNA-splicing factor |
| chr9_-_70490107 | 0.02 |
ENST00000377395.4 ENST00000429800.2 ENST00000430059.2 ENST00000377384.1 ENST00000382405.3 |
CBWD5 |
COBW domain containing 5 |
| chr16_-_30905263 | 0.02 |
ENST00000572628.1 |
BCL7C |
B-cell CLL/lymphoma 7C |
| chr19_-_36236292 | 0.02 |
ENST00000378975.3 ENST00000412391.2 ENST00000292879.5 |
U2AF1L4 |
U2 small nuclear RNA auxiliary factor 1-like 4 |
| chr9_+_70856899 | 0.02 |
ENST00000377342.5 ENST00000478048.1 |
CBWD3 |
COBW domain containing 3 |
| chr21_+_31768348 | 0.02 |
ENST00000355459.2 |
KRTAP13-1 |
keratin associated protein 13-1 |
| chrX_-_103401649 | 0.01 |
ENST00000357421.4 |
SLC25A53 |
solute carrier family 25, member 53 |
| chr7_+_29234028 | 0.01 |
ENST00000222792.6 |
CHN2 |
chimerin 2 |
| chr22_+_21771656 | 0.01 |
ENST00000407464.2 |
HIC2 |
hypermethylated in cancer 2 |
| chr7_+_111846643 | 0.01 |
ENST00000361822.3 |
ZNF277 |
zinc finger protein 277 |
| chr10_-_111683308 | 0.01 |
ENST00000502935.1 ENST00000322238.8 ENST00000369680.4 |
XPNPEP1 |
X-prolyl aminopeptidase (aminopeptidase P) 1, soluble |
| chr14_-_20929624 | 0.01 |
ENST00000398020.4 ENST00000250489.4 |
TMEM55B |
transmembrane protein 55B |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0043095 | regulation of GTP cyclohydrolase I activity(GO:0043095) negative regulation of GTP cyclohydrolase I activity(GO:0043105) |
| 0.1 | 0.3 | GO:0006050 | mannosamine metabolic process(GO:0006050) N-acetylmannosamine metabolic process(GO:0006051) |
| 0.1 | 0.2 | GO:1904978 | regulation of endosome organization(GO:1904978) |
| 0.1 | 0.2 | GO:0030505 | inorganic diphosphate transport(GO:0030505) |
| 0.0 | 0.2 | GO:0090301 | regulation of neural crest formation(GO:0090299) negative regulation of neural crest formation(GO:0090301) negative regulation of fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:2000314) |
| 0.0 | 0.1 | GO:0038162 | response to stem cell factor(GO:0036215) cellular response to stem cell factor stimulus(GO:0036216) Kit signaling pathway(GO:0038109) erythropoietin-mediated signaling pathway(GO:0038162) |
| 0.0 | 0.1 | GO:0070541 | response to platinum ion(GO:0070541) cellular response to lead ion(GO:0071284) |
| 0.0 | 0.7 | GO:0006527 | arginine catabolic process(GO:0006527) |
| 0.0 | 0.1 | GO:0097089 | methyl-branched fatty acid metabolic process(GO:0097089) |
| 0.0 | 0.1 | GO:0019859 | pyrimidine nucleobase catabolic process(GO:0006208) thymine catabolic process(GO:0006210) thymine metabolic process(GO:0019859) |
| 0.0 | 0.1 | GO:0030805 | regulation of cyclic nucleotide catabolic process(GO:0030805) regulation of cAMP catabolic process(GO:0030820) regulation of purine nucleotide catabolic process(GO:0033121) |
| 0.0 | 0.1 | GO:1902075 | cellular response to salt(GO:1902075) |
| 0.0 | 0.1 | GO:0060800 | regulation of cell differentiation involved in embryonic placenta development(GO:0060800) |
| 0.0 | 0.1 | GO:1990736 | positive regulation of regulation of vascular smooth muscle cell membrane depolarization(GO:1904199) regulation of vascular smooth muscle cell membrane depolarization(GO:1990736) |
| 0.0 | 0.1 | GO:1902954 | regulation of early endosome to recycling endosome transport(GO:1902954) |
| 0.0 | 0.1 | GO:0002155 | regulation of thyroid hormone mediated signaling pathway(GO:0002155) kinin cascade(GO:0002254) plasma kallikrein-kinin cascade(GO:0002353) |
| 0.0 | 0.1 | GO:0070164 | negative regulation of adiponectin secretion(GO:0070164) |
| 0.0 | 0.2 | GO:0034472 | snRNA 3'-end processing(GO:0034472) |
| 0.0 | 0.2 | GO:0019532 | oxalate transport(GO:0019532) |
| 0.0 | 0.0 | GO:0050760 | negative regulation of thymidylate synthase biosynthetic process(GO:0050760) |
| 0.0 | 0.0 | GO:1904694 | negative regulation of vascular smooth muscle contraction(GO:1904694) |
| 0.0 | 0.1 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.0 | 0.1 | GO:0061086 | negative regulation of histone H3-K27 methylation(GO:0061086) |
| 0.0 | 0.1 | GO:0016191 | synaptic vesicle uncoating(GO:0016191) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0033165 | interphotoreceptor matrix(GO:0033165) |
| 0.0 | 0.2 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.0 | 0.0 | GO:0035189 | Rb-E2F complex(GO:0035189) |
| 0.0 | 0.1 | GO:0090498 | extrinsic component of Golgi membrane(GO:0090498) |
| 0.0 | 0.1 | GO:0033263 | CORVET complex(GO:0033263) |
| 0.0 | 0.1 | GO:0070695 | FHF complex(GO:0070695) |
| 0.0 | 0.1 | GO:0034464 | BBSome(GO:0034464) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0044549 | GTP cyclohydrolase binding(GO:0044549) |
| 0.1 | 0.3 | GO:0045127 | N-acetylglucosamine kinase activity(GO:0045127) |
| 0.1 | 0.7 | GO:0016403 | dimethylargininase activity(GO:0016403) |
| 0.1 | 0.4 | GO:1902444 | riboflavin binding(GO:1902444) |
| 0.1 | 0.2 | GO:0043812 | phosphatidylinositol-4-phosphate phosphatase activity(GO:0043812) |
| 0.0 | 0.2 | GO:0035529 | NADH pyrophosphatase activity(GO:0035529) |
| 0.0 | 0.4 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.2 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.0 | 0.1 | GO:0016404 | 15-hydroxyprostaglandin dehydrogenase (NAD+) activity(GO:0016404) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |