Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
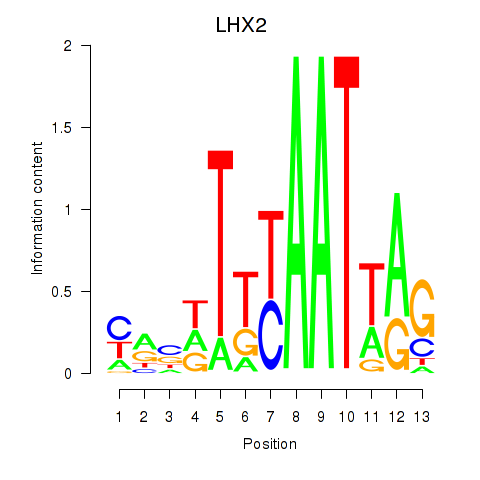
Results for LHX2
Z-value: 0.60
Transcription factors associated with LHX2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
LHX2
|
ENSG00000106689.6 | LHX2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| LHX2 | hg19_v2_chr9_+_126773880_126773895, hg19_v2_chr9_+_126777676_126777700 | -0.03 | 9.5e-01 | Click! |
Activity profile of LHX2 motif
Sorted Z-values of LHX2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of LHX2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr18_+_29027696 | 1.28 |
ENST00000257189.4 |
DSG3 |
desmoglein 3 |
| chr6_+_125524785 | 0.42 |
ENST00000392482.2 |
TPD52L1 |
tumor protein D52-like 1 |
| chr1_+_160370344 | 0.37 |
ENST00000368061.2 |
VANGL2 |
VANGL planar cell polarity protein 2 |
| chr1_+_62439037 | 0.29 |
ENST00000545929.1 |
INADL |
InaD-like (Drosophila) |
| chr2_+_68961934 | 0.28 |
ENST00000409202.3 |
ARHGAP25 |
Rho GTPase activating protein 25 |
| chr2_+_68961905 | 0.28 |
ENST00000295381.3 |
ARHGAP25 |
Rho GTPase activating protein 25 |
| chr18_+_34124507 | 0.28 |
ENST00000591635.1 |
FHOD3 |
formin homology 2 domain containing 3 |
| chrX_+_105937068 | 0.27 |
ENST00000324342.3 |
RNF128 |
ring finger protein 128, E3 ubiquitin protein ligase |
| chr6_+_30848557 | 0.27 |
ENST00000460944.2 ENST00000324771.8 |
DDR1 |
discoidin domain receptor tyrosine kinase 1 |
| chr4_-_90756769 | 0.26 |
ENST00000345009.4 ENST00000505199.1 ENST00000502987.1 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr12_+_122688090 | 0.25 |
ENST00000324189.4 ENST00000546192.1 |
B3GNT4 |
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 4 |
| chr10_-_116286656 | 0.24 |
ENST00000428430.1 ENST00000369266.3 ENST00000392952.3 |
ABLIM1 |
actin binding LIM protein 1 |
| chr7_-_22234381 | 0.23 |
ENST00000458533.1 |
RAPGEF5 |
Rap guanine nucleotide exchange factor (GEF) 5 |
| chr5_-_35938674 | 0.19 |
ENST00000397366.1 ENST00000513623.1 ENST00000514524.1 ENST00000397367.2 |
CAPSL |
calcyphosine-like |
| chr4_+_69313145 | 0.18 |
ENST00000305363.4 |
TMPRSS11E |
transmembrane protease, serine 11E |
| chr2_+_68962014 | 0.18 |
ENST00000467265.1 |
ARHGAP25 |
Rho GTPase activating protein 25 |
| chr11_-_104972158 | 0.15 |
ENST00000598974.1 ENST00000593315.1 ENST00000594519.1 ENST00000415981.2 ENST00000525374.1 ENST00000375707.1 |
CASP1 CARD16 CARD17 |
caspase 1, apoptosis-related cysteine peptidase caspase recruitment domain family, member 16 caspase recruitment domain family, member 17 |
| chr1_-_183560011 | 0.15 |
ENST00000367536.1 |
NCF2 |
neutrophil cytosolic factor 2 |
| chr18_-_19994830 | 0.15 |
ENST00000525417.1 |
CTAGE1 |
cutaneous T-cell lymphoma-associated antigen 1 |
| chr14_-_22005343 | 0.14 |
ENST00000327430.3 |
SALL2 |
spalt-like transcription factor 2 |
| chr15_-_51535208 | 0.13 |
ENST00000405913.3 ENST00000559878.1 |
CYP19A1 |
cytochrome P450, family 19, subfamily A, polypeptide 1 |
| chr19_+_42212501 | 0.12 |
ENST00000398599.4 |
CEACAM5 |
carcinoembryonic antigen-related cell adhesion molecule 5 |
| chr4_-_159956333 | 0.12 |
ENST00000434826.2 |
C4orf45 |
chromosome 4 open reading frame 45 |
| chr10_+_95326416 | 0.12 |
ENST00000371481.4 ENST00000371483.4 ENST00000604414.1 |
FFAR4 |
free fatty acid receptor 4 |
| chr14_-_34420259 | 0.12 |
ENST00000250457.3 ENST00000547327.2 |
EGLN3 |
egl-9 family hypoxia-inducible factor 3 |
| chr5_-_78281775 | 0.11 |
ENST00000396151.3 ENST00000565165.1 |
ARSB |
arylsulfatase B |
| chr3_-_191000172 | 0.10 |
ENST00000427544.2 |
UTS2B |
urotensin 2B |
| chr1_+_160709076 | 0.10 |
ENST00000359331.4 ENST00000495334.1 |
SLAMF7 |
SLAM family member 7 |
| chrX_-_11284095 | 0.10 |
ENST00000303025.6 ENST00000534860.1 |
ARHGAP6 |
Rho GTPase activating protein 6 |
| chr4_-_153332886 | 0.09 |
ENST00000603841.1 |
FBXW7 |
F-box and WD repeat domain containing 7, E3 ubiquitin protein ligase |
| chr2_+_145780725 | 0.09 |
ENST00000451478.1 |
TEX41 |
testis expressed 41 (non-protein coding) |
| chr6_+_26199737 | 0.09 |
ENST00000359985.1 |
HIST1H2BF |
histone cluster 1, H2bf |
| chr4_+_147096837 | 0.09 |
ENST00000296581.5 ENST00000502781.1 |
LSM6 |
LSM6 homolog, U6 small nuclear RNA associated (S. cerevisiae) |
| chr6_-_26199499 | 0.09 |
ENST00000377831.5 |
HIST1H3D |
histone cluster 1, H3d |
| chr1_+_244515930 | 0.09 |
ENST00000366537.1 ENST00000308105.4 |
C1orf100 |
chromosome 1 open reading frame 100 |
| chr7_-_112758589 | 0.09 |
ENST00000413744.1 ENST00000439551.1 ENST00000441359.1 |
LINC00998 |
long intergenic non-protein coding RNA 998 |
| chr13_+_73632897 | 0.09 |
ENST00000377687.4 |
KLF5 |
Kruppel-like factor 5 (intestinal) |
| chr6_-_66417107 | 0.08 |
ENST00000370621.3 ENST00000370618.3 ENST00000503581.1 ENST00000393380.2 |
EYS |
eyes shut homolog (Drosophila) |
| chr1_+_160709055 | 0.08 |
ENST00000368043.3 ENST00000368042.3 ENST00000458602.2 ENST00000458104.2 |
SLAMF7 |
SLAM family member 7 |
| chr17_+_61086917 | 0.08 |
ENST00000424789.2 ENST00000389520.4 |
TANC2 |
tetratricopeptide repeat, ankyrin repeat and coiled-coil containing 2 |
| chr6_+_130339710 | 0.08 |
ENST00000526087.1 ENST00000533560.1 ENST00000361794.2 |
L3MBTL3 |
l(3)mbt-like 3 (Drosophila) |
| chr11_-_104827425 | 0.08 |
ENST00000393150.3 |
CASP4 |
caspase 4, apoptosis-related cysteine peptidase |
| chr4_+_88532028 | 0.08 |
ENST00000282478.7 |
DSPP |
dentin sialophosphoprotein |
| chr14_+_22694606 | 0.07 |
ENST00000390463.3 |
TRAV36DV7 |
T cell receptor alpha variable 36/delta variable 7 |
| chr6_-_26199471 | 0.07 |
ENST00000341023.1 |
HIST1H2AD |
histone cluster 1, H2ad |
| chr4_+_128554081 | 0.07 |
ENST00000335251.6 ENST00000296461.5 |
INTU |
inturned planar cell polarity protein |
| chr8_+_52730143 | 0.07 |
ENST00000415643.1 |
AC090186.1 |
Uncharacterized protein |
| chr17_-_7760457 | 0.07 |
ENST00000576384.1 |
LSMD1 |
LSM domain containing 1 |
| chr4_-_120243545 | 0.07 |
ENST00000274024.3 |
FABP2 |
fatty acid binding protein 2, intestinal |
| chr2_+_145780767 | 0.07 |
ENST00000599358.1 ENST00000596278.1 ENST00000596747.1 ENST00000608652.1 ENST00000609705.1 ENST00000608432.1 ENST00000596970.1 ENST00000602041.1 ENST00000601578.1 ENST00000596034.1 ENST00000414195.2 ENST00000594837.1 |
TEX41 |
testis expressed 41 (non-protein coding) |
| chr3_+_35722487 | 0.07 |
ENST00000441454.1 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chr12_-_100656134 | 0.06 |
ENST00000548313.1 |
DEPDC4 |
DEP domain containing 4 |
| chr9_-_139137648 | 0.06 |
ENST00000358701.5 |
QSOX2 |
quiescin Q6 sulfhydryl oxidase 2 |
| chr11_+_27076764 | 0.06 |
ENST00000525090.1 |
BBOX1 |
butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 |
| chr11_+_63606373 | 0.06 |
ENST00000402010.2 ENST00000315032.8 ENST00000377809.4 ENST00000413835.2 ENST00000377810.3 |
MARK2 |
MAP/microtubule affinity-regulating kinase 2 |
| chr1_+_152975488 | 0.06 |
ENST00000542696.1 |
SPRR3 |
small proline-rich protein 3 |
| chr2_+_102953608 | 0.06 |
ENST00000311734.2 ENST00000409584.1 |
IL1RL1 |
interleukin 1 receptor-like 1 |
| chr21_-_27423339 | 0.06 |
ENST00000415997.1 |
APP |
amyloid beta (A4) precursor protein |
| chr3_+_57881966 | 0.06 |
ENST00000495364.1 |
SLMAP |
sarcolemma associated protein |
| chr3_+_62936098 | 0.06 |
ENST00000475886.1 ENST00000465684.1 ENST00000465262.1 ENST00000468072.1 |
LINC00698 |
long intergenic non-protein coding RNA 698 |
| chr1_-_168464875 | 0.06 |
ENST00000422253.1 |
RP5-968D22.3 |
RP5-968D22.3 |
| chr4_-_100575781 | 0.06 |
ENST00000511828.1 |
RP11-766F14.2 |
Protein LOC285556 |
| chr3_+_147795932 | 0.06 |
ENST00000490465.1 |
RP11-639B1.1 |
RP11-639B1.1 |
| chr2_+_145780739 | 0.05 |
ENST00000597173.1 ENST00000602108.1 ENST00000420472.1 |
TEX41 |
testis expressed 41 (non-protein coding) |
| chr3_+_38017264 | 0.05 |
ENST00000436654.1 |
CTDSPL |
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase-like |
| chr14_+_56584414 | 0.05 |
ENST00000559044.1 |
PELI2 |
pellino E3 ubiquitin protein ligase family member 2 |
| chr6_+_123038689 | 0.05 |
ENST00000354275.2 ENST00000368446.1 |
PKIB |
protein kinase (cAMP-dependent, catalytic) inhibitor beta |
| chr10_-_61900762 | 0.05 |
ENST00000355288.2 |
ANK3 |
ankyrin 3, node of Ranvier (ankyrin G) |
| chr14_-_69864993 | 0.05 |
ENST00000555373.1 |
ERH |
enhancer of rudimentary homolog (Drosophila) |
| chr15_+_69854027 | 0.05 |
ENST00000498938.2 |
RP11-279F6.1 |
RP11-279F6.1 |
| chr9_-_77643307 | 0.04 |
ENST00000376834.3 ENST00000376830.3 |
C9orf41 |
chromosome 9 open reading frame 41 |
| chr6_-_27841289 | 0.04 |
ENST00000355981.2 |
HIST1H4L |
histone cluster 1, H4l |
| chr8_-_109260897 | 0.04 |
ENST00000521297.1 ENST00000519030.1 ENST00000521440.1 ENST00000518345.1 ENST00000519627.1 ENST00000220849.5 |
EIF3E |
eukaryotic translation initiation factor 3, subunit E |
| chr7_+_37960163 | 0.04 |
ENST00000199448.4 ENST00000559325.1 ENST00000423717.1 |
EPDR1 |
ependymin related 1 |
| chr9_+_116207007 | 0.04 |
ENST00000374140.2 |
RGS3 |
regulator of G-protein signaling 3 |
| chr2_+_120436733 | 0.04 |
ENST00000401466.1 ENST00000424086.1 ENST00000272521.6 |
TMEM177 |
transmembrane protein 177 |
| chr4_+_70796784 | 0.04 |
ENST00000246891.4 ENST00000444405.3 |
CSN1S1 |
casein alpha s1 |
| chr16_-_23652570 | 0.04 |
ENST00000261584.4 |
PALB2 |
partner and localizer of BRCA2 |
| chr13_+_78315466 | 0.04 |
ENST00000314070.5 ENST00000462234.1 |
SLAIN1 |
SLAIN motif family, member 1 |
| chr12_+_27932803 | 0.04 |
ENST00000381271.2 |
KLHL42 |
kelch-like family member 42 |
| chr21_+_35014706 | 0.04 |
ENST00000399353.1 ENST00000444491.1 ENST00000381318.3 |
ITSN1 |
intersectin 1 (SH3 domain protein) |
| chr2_-_112237835 | 0.04 |
ENST00000442293.1 ENST00000439494.1 |
MIR4435-1HG |
MIR4435-1 host gene (non-protein coding) |
| chr2_+_103035102 | 0.04 |
ENST00000264260.2 |
IL18RAP |
interleukin 18 receptor accessory protein |
| chr14_+_69865401 | 0.04 |
ENST00000556605.1 ENST00000336643.5 ENST00000031146.4 |
SLC39A9 |
solute carrier family 39, member 9 |
| chr12_+_128399965 | 0.04 |
ENST00000540882.1 ENST00000542089.1 |
LINC00507 |
long intergenic non-protein coding RNA 507 |
| chr11_+_64001962 | 0.04 |
ENST00000309422.2 |
VEGFB |
vascular endothelial growth factor B |
| chr2_+_234216454 | 0.03 |
ENST00000447536.1 ENST00000409110.1 |
SAG |
S-antigen; retina and pineal gland (arrestin) |
| chr1_-_89591749 | 0.03 |
ENST00000370466.3 |
GBP2 |
guanylate binding protein 2, interferon-inducible |
| chr1_+_160709029 | 0.03 |
ENST00000444090.2 ENST00000441662.2 |
SLAMF7 |
SLAM family member 7 |
| chr13_-_19755975 | 0.03 |
ENST00000400113.3 |
TUBA3C |
tubulin, alpha 3c |
| chr12_+_9980069 | 0.03 |
ENST00000354855.3 ENST00000324214.4 ENST00000279544.3 |
KLRF1 |
killer cell lectin-like receptor subfamily F, member 1 |
| chr16_-_66864806 | 0.03 |
ENST00000566336.1 ENST00000394074.2 ENST00000563185.2 ENST00000359087.4 ENST00000379463.2 ENST00000565535.1 ENST00000290810.3 |
NAE1 |
NEDD8 activating enzyme E1 subunit 1 |
| chr4_-_109541539 | 0.03 |
ENST00000509984.1 ENST00000507248.1 ENST00000506795.1 |
RPL34-AS1 |
RPL34 antisense RNA 1 (head to head) |
| chr3_-_180707306 | 0.03 |
ENST00000479269.1 |
DNAJC19 |
DnaJ (Hsp40) homolog, subfamily C, member 19 |
| chr8_+_76452097 | 0.03 |
ENST00000396423.2 |
HNF4G |
hepatocyte nuclear factor 4, gamma |
| chr17_-_39324424 | 0.03 |
ENST00000391356.2 |
KRTAP4-3 |
keratin associated protein 4-3 |
| chr16_+_31213206 | 0.03 |
ENST00000561916.2 |
C16orf98 |
chromosome 16 open reading frame 98 |
| chrX_-_20074895 | 0.03 |
ENST00000543767.1 |
MAP7D2 |
MAP7 domain containing 2 |
| chrX_+_30233668 | 0.03 |
ENST00000378988.4 |
MAGEB2 |
melanoma antigen family B, 2 |
| chr3_+_57882024 | 0.03 |
ENST00000494088.1 |
SLMAP |
sarcolemma associated protein |
| chr12_+_6309517 | 0.03 |
ENST00000382519.4 ENST00000009180.4 |
CD9 |
CD9 molecule |
| chr2_-_55496174 | 0.03 |
ENST00000417363.1 ENST00000412530.1 ENST00000394600.3 ENST00000366137.2 ENST00000420637.1 |
MTIF2 |
mitochondrial translational initiation factor 2 |
| chr17_-_39140549 | 0.03 |
ENST00000377755.4 |
KRT40 |
keratin 40 |
| chr5_-_156536126 | 0.03 |
ENST00000522593.1 |
HAVCR2 |
hepatitis A virus cellular receptor 2 |
| chr18_+_3466248 | 0.03 |
ENST00000581029.1 ENST00000581442.1 ENST00000579007.1 |
RP11-838N2.4 |
RP11-838N2.4 |
| chr12_+_6554021 | 0.03 |
ENST00000266557.3 |
CD27 |
CD27 molecule |
| chr16_+_24266874 | 0.03 |
ENST00000005284.3 |
CACNG3 |
calcium channel, voltage-dependent, gamma subunit 3 |
| chr14_-_78083112 | 0.02 |
ENST00000216484.2 |
SPTLC2 |
serine palmitoyltransferase, long chain base subunit 2 |
| chr4_-_105416039 | 0.02 |
ENST00000394767.2 |
CXXC4 |
CXXC finger protein 4 |
| chr21_+_34619079 | 0.02 |
ENST00000433395.2 |
AP000295.9 |
AP000295.9 |
| chrY_-_20935572 | 0.02 |
ENST00000382852.1 ENST00000344884.4 ENST00000304790.3 |
HSFY2 |
heat shock transcription factor, Y linked 2 |
| chr4_+_15471489 | 0.02 |
ENST00000424120.1 ENST00000413206.1 ENST00000438599.2 ENST00000511544.1 ENST00000512702.1 ENST00000507954.1 ENST00000515124.1 ENST00000503292.1 ENST00000503658.1 |
CC2D2A |
coiled-coil and C2 domain containing 2A |
| chr7_-_54826920 | 0.02 |
ENST00000395535.3 ENST00000352861.4 |
SEC61G |
Sec61 gamma subunit |
| chr16_+_31271274 | 0.02 |
ENST00000287497.8 ENST00000544665.3 |
ITGAM |
integrin, alpha M (complement component 3 receptor 3 subunit) |
| chr2_-_128284020 | 0.02 |
ENST00000295321.4 ENST00000455721.2 |
IWS1 |
IWS1 homolog (S. cerevisiae) |
| chr12_-_11548496 | 0.02 |
ENST00000389362.4 ENST00000565533.1 ENST00000546254.1 |
PRB2 PRB1 |
proline-rich protein BstNI subfamily 2 proline-rich protein BstNI subfamily 1 |
| chr7_-_5998714 | 0.02 |
ENST00000539903.1 |
RSPH10B |
radial spoke head 10 homolog B (Chlamydomonas) |
| chr13_+_19756173 | 0.02 |
ENST00000382988.2 |
RP11-408E5.4 |
RP11-408E5.4 |
| chr2_+_234627424 | 0.02 |
ENST00000373409.3 |
UGT1A4 |
UDP glucuronosyltransferase 1 family, polypeptide A4 |
| chr2_+_87769459 | 0.02 |
ENST00000414030.1 ENST00000437561.1 |
LINC00152 |
long intergenic non-protein coding RNA 152 |
| chr15_+_58430567 | 0.02 |
ENST00000536493.1 |
AQP9 |
aquaporin 9 |
| chrY_+_20708557 | 0.02 |
ENST00000307393.2 ENST00000309834.4 ENST00000382856.2 |
HSFY1 |
heat shock transcription factor, Y-linked 1 |
| chr15_+_58430368 | 0.02 |
ENST00000558772.1 ENST00000219919.4 |
AQP9 |
aquaporin 9 |
| chr17_-_57229155 | 0.02 |
ENST00000584089.1 |
SKA2 |
spindle and kinetochore associated complex subunit 2 |
| chr6_+_112408768 | 0.02 |
ENST00000368656.2 ENST00000604268.1 |
FAM229B |
family with sequence similarity 229, member B |
| chr12_-_74686314 | 0.02 |
ENST00000551210.1 ENST00000515416.2 ENST00000549905.1 |
RP11-81H3.2 |
RP11-81H3.2 |
| chr17_-_38859996 | 0.02 |
ENST00000264651.2 |
KRT24 |
keratin 24 |
| chr5_+_179921344 | 0.02 |
ENST00000261951.4 |
CNOT6 |
CCR4-NOT transcription complex, subunit 6 |
| chr20_+_30697298 | 0.02 |
ENST00000398022.2 |
TM9SF4 |
transmembrane 9 superfamily protein member 4 |
| chr12_-_113574028 | 0.02 |
ENST00000546530.1 ENST00000261729.5 |
RASAL1 |
RAS protein activator like 1 (GAP1 like) |
| chr2_+_173686303 | 0.02 |
ENST00000397087.3 |
RAPGEF4 |
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr9_+_21440440 | 0.02 |
ENST00000276927.1 |
IFNA1 |
interferon, alpha 1 |
| chr11_-_2323089 | 0.02 |
ENST00000456145.2 |
C11orf21 |
chromosome 11 open reading frame 21 |
| chr2_-_166930131 | 0.02 |
ENST00000303395.4 ENST00000409050.1 ENST00000423058.2 ENST00000375405.3 |
SCN1A |
sodium channel, voltage-gated, type I, alpha subunit |
| chr4_-_89951028 | 0.02 |
ENST00000506913.1 |
FAM13A |
family with sequence similarity 13, member A |
| chr11_-_40315640 | 0.02 |
ENST00000278198.2 |
LRRC4C |
leucine rich repeat containing 4C |
| chr11_+_94706973 | 0.01 |
ENST00000536741.1 |
KDM4D |
lysine (K)-specific demethylase 4D |
| chr10_-_69597828 | 0.01 |
ENST00000339758.7 |
DNAJC12 |
DnaJ (Hsp40) homolog, subfamily C, member 12 |
| chr12_+_12510352 | 0.01 |
ENST00000298571.6 |
LOH12CR1 |
loss of heterozygosity, 12, chromosomal region 1 |
| chr19_+_20188830 | 0.01 |
ENST00000418063.2 |
ZNF90 |
zinc finger protein 90 |
| chr7_-_54826869 | 0.01 |
ENST00000450622.1 |
SEC61G |
Sec61 gamma subunit |
| chr5_-_133702761 | 0.01 |
ENST00000521118.1 ENST00000265334.4 ENST00000435211.1 |
CDKL3 |
cyclin-dependent kinase-like 3 |
| chr6_+_3259148 | 0.01 |
ENST00000419065.2 ENST00000473000.2 ENST00000451246.2 ENST00000454610.2 |
PSMG4 |
proteasome (prosome, macropain) assembly chaperone 4 |
| chr3_-_160823040 | 0.01 |
ENST00000484127.1 ENST00000492353.1 ENST00000473142.1 ENST00000468268.1 ENST00000460353.1 ENST00000320474.4 ENST00000392781.2 |
B3GALNT1 |
beta-1,3-N-acetylgalactosaminyltransferase 1 (globoside blood group) |
| chr16_+_23652773 | 0.01 |
ENST00000563998.1 ENST00000568589.1 ENST00000568272.1 |
DCTN5 |
dynactin 5 (p25) |
| chr1_+_182419261 | 0.01 |
ENST00000294854.8 ENST00000542961.1 |
RGSL1 |
regulator of G-protein signaling like 1 |
| chr11_-_55703876 | 0.01 |
ENST00000301532.3 |
OR5I1 |
olfactory receptor, family 5, subfamily I, member 1 |
| chr3_-_160823158 | 0.01 |
ENST00000392779.2 ENST00000392780.1 ENST00000494173.1 |
B3GALNT1 |
beta-1,3-N-acetylgalactosaminyltransferase 1 (globoside blood group) |
| chr12_+_48722763 | 0.01 |
ENST00000335017.1 |
H1FNT |
H1 histone family, member N, testis-specific |
| chr1_-_28527152 | 0.01 |
ENST00000321830.5 |
AL353354.1 |
Uncharacterized protein |
| chr5_-_175388327 | 0.01 |
ENST00000432305.2 ENST00000505969.1 |
THOC3 |
THO complex 3 |
| chr2_-_208989225 | 0.01 |
ENST00000264376.4 |
CRYGD |
crystallin, gamma D |
| chr9_-_21202204 | 0.01 |
ENST00000239347.3 |
IFNA7 |
interferon, alpha 7 |
| chr1_-_198509804 | 0.01 |
ENST00000489986.1 ENST00000367382.1 |
ATP6V1G3 |
ATPase, H+ transporting, lysosomal 13kDa, V1 subunit G3 |
| chr8_+_55528627 | 0.01 |
ENST00000220676.1 |
RP1 |
retinitis pigmentosa 1 (autosomal dominant) |
| chr1_+_219347203 | 0.01 |
ENST00000366927.3 |
LYPLAL1 |
lysophospholipase-like 1 |
| chr15_-_72563585 | 0.01 |
ENST00000287196.9 ENST00000260376.7 |
PARP6 |
poly (ADP-ribose) polymerase family, member 6 |
| chr8_-_82395461 | 0.01 |
ENST00000256104.4 |
FABP4 |
fatty acid binding protein 4, adipocyte |
| chr14_+_22554680 | 0.01 |
ENST00000390451.2 |
TRAV23DV6 |
T cell receptor alpha variable 23/delta variable 6 |
| chr10_+_96443204 | 0.00 |
ENST00000339022.5 |
CYP2C18 |
cytochrome P450, family 2, subfamily C, polypeptide 18 |
| chr14_+_27342334 | 0.00 |
ENST00000548170.1 ENST00000552926.1 |
RP11-384J4.1 |
RP11-384J4.1 |
| chr12_-_118797475 | 0.00 |
ENST00000541786.1 ENST00000419821.2 ENST00000541878.1 |
TAOK3 |
TAO kinase 3 |
| chr3_+_35722424 | 0.00 |
ENST00000396481.2 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chr20_-_31989307 | 0.00 |
ENST00000473997.1 ENST00000544843.1 ENST00000346416.2 ENST00000357886.4 ENST00000339269.5 |
CDK5RAP1 |
CDK5 regulatory subunit associated protein 1 |
| chr7_-_151217166 | 0.00 |
ENST00000496004.1 |
RHEB |
Ras homolog enriched in brain |
| chr12_+_16064106 | 0.00 |
ENST00000428559.2 |
DERA |
deoxyribose-phosphate aldolase (putative) |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0061341 | orthogonal dichotomous subdivision of terminal units involved in lung branching morphogenesis(GO:0060488) planar dichotomous subdivision of terminal units involved in lung branching morphogenesis(GO:0060489) lateral sprouting involved in lung morphogenesis(GO:0060490) non-canonical Wnt signaling pathway involved in heart development(GO:0061341) planar cell polarity pathway involved in heart morphogenesis(GO:0061346) |
| 0.1 | 0.1 | GO:0008217 | regulation of blood pressure(GO:0008217) |
| 0.1 | 0.2 | GO:0030311 | poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 0.1 | 0.3 | GO:0051945 | negative regulation of dopamine uptake involved in synaptic transmission(GO:0051585) norepinephrine uptake(GO:0051620) regulation of norepinephrine uptake(GO:0051621) negative regulation of norepinephrine uptake(GO:0051622) negative regulation of catecholamine uptake involved in synaptic transmission(GO:0051945) regulation of glutathione peroxidase activity(GO:1903282) positive regulation of glutathione peroxidase activity(GO:1903284) positive regulation of hydrogen peroxide catabolic process(GO:1903285) positive regulation of peroxidase activity(GO:2000470) |
| 0.0 | 0.1 | GO:2000863 | positive regulation of estrogen secretion(GO:2000863) positive regulation of estradiol secretion(GO:2000866) |
| 0.0 | 0.2 | GO:0050717 | positive regulation of interleukin-1 alpha secretion(GO:0050717) |
| 0.0 | 0.3 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.0 | 0.3 | GO:1904352 | positive regulation of protein catabolic process in the vacuole(GO:1904352) |
| 0.0 | 0.1 | GO:2000639 | regulation of SREBP signaling pathway(GO:2000638) negative regulation of SREBP signaling pathway(GO:2000639) |
| 0.0 | 0.3 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.0 | 0.1 | GO:1905150 | regulation of voltage-gated sodium channel activity(GO:1905150) |
| 0.0 | 0.1 | GO:0038172 | interleukin-33-mediated signaling pathway(GO:0038172) |
| 0.0 | 0.1 | GO:0071874 | cellular response to norepinephrine stimulus(GO:0071874) |
| 0.0 | 0.0 | GO:0072658 | maintenance of protein location in membrane(GO:0072658) maintenance of protein location in plasma membrane(GO:0072660) positive regulation of membrane depolarization during cardiac muscle cell action potential(GO:1900827) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0060187 | cell pole(GO:0060187) |
| 0.0 | 1.3 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 0.2 | GO:0097169 | AIM2 inflammasome complex(GO:0097169) |
| 0.0 | 0.1 | GO:0032010 | phagolysosome(GO:0032010) |
| 0.0 | 0.1 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0008532 | N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase activity(GO:0008532) |
| 0.1 | 0.3 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 0.1 | 0.3 | GO:0060961 | phospholipase D inhibitor activity(GO:0060961) |
| 0.0 | 0.2 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.0 | 0.1 | GO:0008336 | gamma-butyrobetaine dioxygenase activity(GO:0008336) |
| 0.0 | 0.1 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.0 | 0.1 | GO:0002114 | interleukin-33 receptor activity(GO:0002114) |
| 0.0 | 0.2 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.0 | 0.1 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 0.0 | 0.1 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.0 | 0.0 | GO:0005275 | amine transmembrane transporter activity(GO:0005275) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.3 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.0 | 0.2 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |