Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
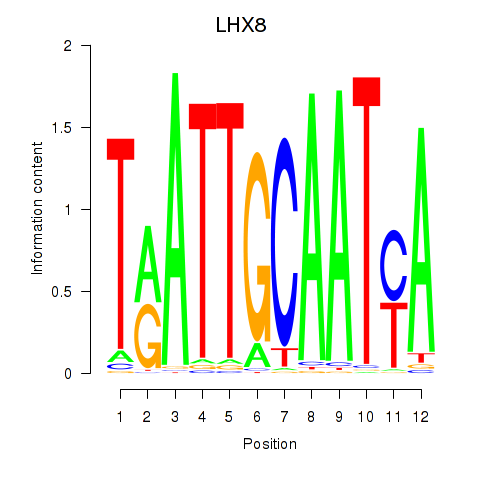
Results for LHX8
Z-value: 0.79
Transcription factors associated with LHX8
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
LHX8
|
ENSG00000162624.10 | LHX8 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| LHX8 | hg19_v2_chr1_+_75600567_75600567, hg19_v2_chr1_+_75594119_75594119 | -0.67 | 6.8e-02 | Click! |
Activity profile of LHX8 motif
Sorted Z-values of LHX8 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of LHX8
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_+_33022885 | 0.72 |
ENST00000322805.4 |
GREM1 |
gremlin 1, DAN family BMP antagonist |
| chr12_-_15038779 | 0.62 |
ENST00000228938.5 ENST00000539261.1 |
MGP |
matrix Gla protein |
| chr7_-_92777606 | 0.44 |
ENST00000437805.1 ENST00000446959.1 ENST00000439952.1 ENST00000414791.1 ENST00000446033.1 ENST00000411955.1 ENST00000318238.4 |
SAMD9L |
sterile alpha motif domain containing 9-like |
| chr9_-_20622478 | 0.44 |
ENST00000355930.6 ENST00000380338.4 |
MLLT3 |
myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 3 |
| chr3_-_151102529 | 0.41 |
ENST00000302632.3 |
P2RY12 |
purinergic receptor P2Y, G-protein coupled, 12 |
| chr8_+_17354617 | 0.40 |
ENST00000470360.1 |
SLC7A2 |
solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 |
| chr12_+_59989918 | 0.38 |
ENST00000547379.1 ENST00000549465.1 |
SLC16A7 |
solute carrier family 16 (monocarboxylate transporter), member 7 |
| chr3_+_119187785 | 0.35 |
ENST00000295588.4 ENST00000476573.1 |
POGLUT1 |
protein O-glucosyltransferase 1 |
| chr8_+_27168988 | 0.32 |
ENST00000397501.1 ENST00000338238.4 ENST00000544172.1 |
PTK2B |
protein tyrosine kinase 2 beta |
| chr6_+_160221293 | 0.32 |
ENST00000610273.1 ENST00000392167.3 |
PNLDC1 |
poly(A)-specific ribonuclease (PARN)-like domain containing 1 |
| chr4_-_100242549 | 0.30 |
ENST00000305046.8 ENST00000394887.3 |
ADH1B |
alcohol dehydrogenase 1B (class I), beta polypeptide |
| chr8_+_17354587 | 0.28 |
ENST00000494857.1 ENST00000522656.1 |
SLC7A2 |
solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 |
| chr3_+_42977846 | 0.27 |
ENST00000383748.4 |
KRBOX1 |
KRAB box domain containing 1 |
| chrX_+_103031421 | 0.27 |
ENST00000433491.1 ENST00000418604.1 ENST00000443502.1 |
PLP1 |
proteolipid protein 1 |
| chr4_+_166300084 | 0.27 |
ENST00000402744.4 |
CPE |
carboxypeptidase E |
| chrX_+_70798261 | 0.27 |
ENST00000373696.3 |
ACRC |
acidic repeat containing |
| chr7_-_14026063 | 0.26 |
ENST00000443608.1 ENST00000438956.1 |
ETV1 |
ets variant 1 |
| chr2_-_145275228 | 0.25 |
ENST00000427902.1 ENST00000409487.3 ENST00000470879.1 ENST00000435831.1 |
ZEB2 |
zinc finger E-box binding homeobox 2 |
| chr2_-_158345462 | 0.25 |
ENST00000439355.1 ENST00000540637.1 |
CYTIP |
cytohesin 1 interacting protein |
| chr1_+_144989309 | 0.24 |
ENST00000596396.1 |
AL590452.1 |
Uncharacterized protein |
| chr10_+_90672113 | 0.24 |
ENST00000371922.1 |
STAMBPL1 |
STAM binding protein-like 1 |
| chr10_+_81892347 | 0.22 |
ENST00000372267.2 |
PLAC9 |
placenta-specific 9 |
| chr1_-_48937838 | 0.22 |
ENST00000371847.3 |
SPATA6 |
spermatogenesis associated 6 |
| chr7_-_14026123 | 0.21 |
ENST00000420159.2 ENST00000399357.3 ENST00000403527.1 |
ETV1 |
ets variant 1 |
| chr8_+_22424551 | 0.21 |
ENST00000523348.1 |
SORBS3 |
sorbin and SH3 domain containing 3 |
| chr7_-_120498357 | 0.20 |
ENST00000415871.1 ENST00000222747.3 ENST00000430985.1 |
TSPAN12 |
tetraspanin 12 |
| chr4_-_83812248 | 0.20 |
ENST00000514326.1 ENST00000505434.1 ENST00000503058.1 ENST00000348405.4 ENST00000505984.1 ENST00000513858.1 ENST00000508479.1 ENST00000443462.2 ENST00000508502.1 ENST00000509142.1 ENST00000432794.1 ENST00000448323.1 ENST00000326950.5 ENST00000311785.7 |
SEC31A |
SEC31 homolog A (S. cerevisiae) |
| chr1_-_48937821 | 0.20 |
ENST00000396199.3 |
SPATA6 |
spermatogenesis associated 6 |
| chr4_+_74718906 | 0.19 |
ENST00000226524.3 |
PF4V1 |
platelet factor 4 variant 1 |
| chr4_-_83812402 | 0.19 |
ENST00000395310.2 |
SEC31A |
SEC31 homolog A (S. cerevisiae) |
| chr12_-_53893399 | 0.19 |
ENST00000267079.2 |
MAP3K12 |
mitogen-activated protein kinase kinase kinase 12 |
| chr16_-_11922665 | 0.19 |
ENST00000573319.1 ENST00000577041.1 ENST00000574028.1 ENST00000571259.1 ENST00000573037.1 ENST00000571158.1 |
BCAR4 |
breast cancer anti-estrogen resistance 4 (non-protein coding) |
| chrX_+_102883620 | 0.18 |
ENST00000372626.3 |
TCEAL1 |
transcription elongation factor A (SII)-like 1 |
| chr10_-_28571015 | 0.18 |
ENST00000375719.3 ENST00000375732.1 |
MPP7 |
membrane protein, palmitoylated 7 (MAGUK p55 subfamily member 7) |
| chr3_-_164913777 | 0.17 |
ENST00000475390.1 |
SLITRK3 |
SLIT and NTRK-like family, member 3 |
| chr2_+_158114051 | 0.17 |
ENST00000259056.4 |
GALNT5 |
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 5 (GalNAc-T5) |
| chr2_+_109237717 | 0.17 |
ENST00000409441.1 |
LIMS1 |
LIM and senescent cell antigen-like domains 1 |
| chr2_+_201994042 | 0.16 |
ENST00000417748.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr6_-_24489842 | 0.16 |
ENST00000230036.1 |
GPLD1 |
glycosylphosphatidylinositol specific phospholipase D1 |
| chr12_+_15475331 | 0.15 |
ENST00000281171.4 |
PTPRO |
protein tyrosine phosphatase, receptor type, O |
| chr18_-_11908329 | 0.14 |
ENST00000344987.7 ENST00000588103.1 ENST00000588191.1 ENST00000317235.7 ENST00000309976.9 ENST00000588186.1 ENST00000589267.1 |
MPPE1 |
metallophosphoesterase 1 |
| chr4_+_88754113 | 0.14 |
ENST00000560249.1 ENST00000540395.1 ENST00000511670.1 ENST00000361056.3 |
MEPE |
matrix extracellular phosphoglycoprotein |
| chr20_-_30458491 | 0.14 |
ENST00000339738.5 |
DUSP15 |
dual specificity phosphatase 15 |
| chr4_+_174089904 | 0.14 |
ENST00000265000.4 |
GALNT7 |
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 7 (GalNAc-T7) |
| chr19_+_44716678 | 0.14 |
ENST00000586228.1 ENST00000588219.1 ENST00000313040.7 ENST00000589707.1 ENST00000588394.1 ENST00000589005.1 |
ZNF227 |
zinc finger protein 227 |
| chr1_+_145293371 | 0.13 |
ENST00000342960.5 |
NBPF10 |
neuroblastoma breakpoint family, member 10 |
| chr5_-_150473127 | 0.13 |
ENST00000521001.1 |
TNIP1 |
TNFAIP3 interacting protein 1 |
| chr8_-_7243080 | 0.13 |
ENST00000400156.4 |
ZNF705G |
zinc finger protein 705G |
| chr8_-_27168737 | 0.13 |
ENST00000521253.1 ENST00000305364.4 |
TRIM35 |
tripartite motif containing 35 |
| chr12_-_71551652 | 0.13 |
ENST00000546561.1 |
TSPAN8 |
tetraspanin 8 |
| chr14_+_57046530 | 0.13 |
ENST00000536419.1 ENST00000538838.1 |
TMEM260 |
transmembrane protein 260 |
| chr17_+_45000483 | 0.13 |
ENST00000576910.2 ENST00000439730.2 ENST00000393456.2 ENST00000415811.2 ENST00000575949.1 ENST00000225567.4 ENST00000572403.1 ENST00000570879.1 |
GOSR2 |
golgi SNAP receptor complex member 2 |
| chr2_-_225266743 | 0.13 |
ENST00000409685.3 |
FAM124B |
family with sequence similarity 124B |
| chr15_-_43785274 | 0.12 |
ENST00000413546.1 |
TP53BP1 |
tumor protein p53 binding protein 1 |
| chr2_-_225266711 | 0.12 |
ENST00000389874.3 |
FAM124B |
family with sequence similarity 124B |
| chr4_+_71108300 | 0.12 |
ENST00000304954.3 |
CSN3 |
casein kappa |
| chr14_+_23067146 | 0.12 |
ENST00000428304.2 |
ABHD4 |
abhydrolase domain containing 4 |
| chr19_+_21688366 | 0.12 |
ENST00000358491.4 ENST00000597078.1 |
ZNF429 |
zinc finger protein 429 |
| chr12_-_71551868 | 0.12 |
ENST00000247829.3 |
TSPAN8 |
tetraspanin 8 |
| chr12_-_92536433 | 0.12 |
ENST00000551563.2 ENST00000546975.1 ENST00000549802.1 |
C12orf79 |
chromosome 12 open reading frame 79 |
| chr19_-_9006766 | 0.11 |
ENST00000599436.1 |
MUC16 |
mucin 16, cell surface associated |
| chr7_+_142498725 | 0.11 |
ENST00000466254.1 |
TRBC2 |
T cell receptor beta constant 2 |
| chr7_+_55980331 | 0.11 |
ENST00000429591.2 |
ZNF713 |
zinc finger protein 713 |
| chr13_-_24895566 | 0.11 |
ENST00000422229.2 |
AL359736.1 |
protein PCOTH isoform 1 |
| chr15_+_36338242 | 0.11 |
ENST00000560056.1 |
RP11-684B21.1 |
RP11-684B21.1 |
| chr9_-_13165457 | 0.11 |
ENST00000542239.1 ENST00000538841.1 ENST00000433359.2 |
MPDZ |
multiple PDZ domain protein |
| chr14_+_57046500 | 0.11 |
ENST00000261556.6 |
TMEM260 |
transmembrane protein 260 |
| chr1_-_146040968 | 0.11 |
ENST00000401010.3 |
NBPF11 |
neuroblastoma breakpoint family, member 11 |
| chr11_-_68780824 | 0.11 |
ENST00000441623.1 ENST00000309099.6 |
MRGPRF |
MAS-related GPR, member F |
| chr3_+_148545586 | 0.11 |
ENST00000282957.4 ENST00000468341.1 |
CPB1 |
carboxypeptidase B1 (tissue) |
| chr2_+_201994208 | 0.10 |
ENST00000440180.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr6_+_116691001 | 0.10 |
ENST00000537543.1 |
DSE |
dermatan sulfate epimerase |
| chr8_+_11961898 | 0.10 |
ENST00000400085.3 |
ZNF705D |
zinc finger protein 705D |
| chr19_-_4535233 | 0.10 |
ENST00000381848.3 ENST00000588887.1 ENST00000586133.1 |
PLIN5 |
perilipin 5 |
| chr7_+_12727250 | 0.10 |
ENST00000404894.1 |
ARL4A |
ADP-ribosylation factor-like 4A |
| chrX_-_19689106 | 0.10 |
ENST00000379716.1 |
SH3KBP1 |
SH3-domain kinase binding protein 1 |
| chr5_+_140480083 | 0.10 |
ENST00000231130.2 |
PCDHB3 |
protocadherin beta 3 |
| chr12_+_26205496 | 0.09 |
ENST00000537946.1 ENST00000541218.1 ENST00000282884.9 ENST00000545413.1 |
RASSF8 |
Ras association (RalGDS/AF-6) domain family (N-terminal) member 8 |
| chr15_-_34880646 | 0.09 |
ENST00000543376.1 |
GOLGA8A |
golgin A8 family, member A |
| chr2_-_74618964 | 0.09 |
ENST00000417090.1 ENST00000409868.1 |
DCTN1 |
dynactin 1 |
| chr12_-_10766184 | 0.09 |
ENST00000539554.1 ENST00000381881.2 ENST00000320756.2 |
MAGOHB |
mago-nashi homolog B (Drosophila) |
| chr14_+_102829300 | 0.09 |
ENST00000359520.7 |
TECPR2 |
tectonin beta-propeller repeat containing 2 |
| chr4_-_163085107 | 0.09 |
ENST00000379164.4 |
FSTL5 |
follistatin-like 5 |
| chr2_-_37068530 | 0.09 |
ENST00000593798.1 |
AC007382.1 |
Uncharacterized protein |
| chr20_+_56964169 | 0.09 |
ENST00000475243.1 |
VAPB |
VAMP (vesicle-associated membrane protein)-associated protein B and C |
| chr2_-_152118352 | 0.09 |
ENST00000331426.5 |
RBM43 |
RNA binding motif protein 43 |
| chr1_+_146373546 | 0.09 |
ENST00000446760.2 |
NBPF12 |
neuroblastoma breakpoint family, member 12 |
| chr9_+_96846740 | 0.09 |
ENST00000288976.3 |
PTPDC1 |
protein tyrosine phosphatase domain containing 1 |
| chr1_+_33722080 | 0.08 |
ENST00000483388.1 ENST00000539719.1 |
ZNF362 |
zinc finger protein 362 |
| chr4_+_189060573 | 0.08 |
ENST00000332517.3 |
TRIML1 |
tripartite motif family-like 1 |
| chr14_+_22433675 | 0.08 |
ENST00000390442.3 |
TRAV12-3 |
T cell receptor alpha variable 12-3 |
| chr7_-_14880892 | 0.08 |
ENST00000406247.3 ENST00000399322.3 ENST00000258767.5 |
DGKB |
diacylglycerol kinase, beta 90kDa |
| chr3_-_187455680 | 0.08 |
ENST00000438077.1 |
BCL6 |
B-cell CLL/lymphoma 6 |
| chr10_+_94352956 | 0.08 |
ENST00000260731.3 |
KIF11 |
kinesin family member 11 |
| chr4_-_186733363 | 0.08 |
ENST00000393523.2 ENST00000393528.3 ENST00000449407.2 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr15_-_43785303 | 0.08 |
ENST00000382039.3 ENST00000450115.2 ENST00000382044.4 |
TP53BP1 |
tumor protein p53 binding protein 1 |
| chr19_+_21264980 | 0.08 |
ENST00000596053.1 ENST00000597086.1 ENST00000596143.1 ENST00000596367.1 ENST00000601416.1 |
ZNF714 |
zinc finger protein 714 |
| chr3_+_107096188 | 0.08 |
ENST00000261058.1 |
CCDC54 |
coiled-coil domain containing 54 |
| chrX_+_108779870 | 0.07 |
ENST00000372107.1 |
NXT2 |
nuclear transport factor 2-like export factor 2 |
| chr4_-_95264008 | 0.07 |
ENST00000295256.5 |
HPGDS |
hematopoietic prostaglandin D synthase |
| chr7_+_7811992 | 0.07 |
ENST00000406829.1 |
RPA3-AS1 |
RPA3 antisense RNA 1 |
| chr13_+_24883703 | 0.07 |
ENST00000332018.4 |
C1QTNF9 |
C1q and tumor necrosis factor related protein 9 |
| chr2_+_114195268 | 0.07 |
ENST00000259199.4 ENST00000416503.2 ENST00000433343.2 |
CBWD2 |
COBW domain containing 2 |
| chr1_+_145883868 | 0.07 |
ENST00000447947.2 |
GPR89C |
G protein-coupled receptor 89C |
| chr8_-_62602327 | 0.07 |
ENST00000445642.3 ENST00000517847.2 ENST00000389204.4 ENST00000517661.1 ENST00000517903.1 ENST00000522603.1 ENST00000522349.1 ENST00000522835.1 ENST00000541428.1 ENST00000518306.1 |
ASPH |
aspartate beta-hydroxylase |
| chr1_-_110933663 | 0.07 |
ENST00000369781.4 ENST00000541986.1 ENST00000369779.4 |
SLC16A4 |
solute carrier family 16, member 4 |
| chr5_+_140593509 | 0.07 |
ENST00000341948.4 |
PCDHB13 |
protocadherin beta 13 |
| chr9_-_179018 | 0.07 |
ENST00000431099.2 ENST00000382447.4 ENST00000382389.1 ENST00000377447.3 ENST00000314367.10 ENST00000356521.4 ENST00000382393.1 ENST00000377400.4 |
CBWD1 |
COBW domain containing 1 |
| chr1_-_236445251 | 0.07 |
ENST00000354619.5 ENST00000327333.8 |
ERO1LB |
ERO1-like beta (S. cerevisiae) |
| chr12_-_88974236 | 0.07 |
ENST00000228280.5 ENST00000552044.1 ENST00000357116.4 |
KITLG |
KIT ligand |
| chr6_+_43543942 | 0.07 |
ENST00000372226.1 ENST00000443535.1 |
POLH |
polymerase (DNA directed), eta |
| chr2_-_74619152 | 0.07 |
ENST00000440727.1 ENST00000409240.1 |
DCTN1 |
dynactin 1 |
| chr17_-_76719807 | 0.06 |
ENST00000589297.1 |
CYTH1 |
cytohesin 1 |
| chr8_+_7801144 | 0.06 |
ENST00000443676.1 |
ZNF705B |
zinc finger protein 705B |
| chrX_+_108779004 | 0.06 |
ENST00000218004.1 |
NXT2 |
nuclear transport factor 2-like export factor 2 |
| chr9_-_69262509 | 0.06 |
ENST00000377449.1 ENST00000382399.4 ENST00000377439.1 ENST00000377441.1 ENST00000377457.5 |
CBWD6 |
COBW domain containing 6 |
| chr5_+_175511859 | 0.06 |
ENST00000503724.2 ENST00000253490.4 |
FAM153B |
family with sequence similarity 153, member B |
| chr7_-_28220354 | 0.06 |
ENST00000283928.5 |
JAZF1 |
JAZF zinc finger 1 |
| chr9_+_108424738 | 0.06 |
ENST00000334077.3 |
TAL2 |
T-cell acute lymphocytic leukemia 2 |
| chr18_-_14132422 | 0.06 |
ENST00000589498.1 ENST00000590202.1 |
ZNF519 |
zinc finger protein 519 |
| chr2_-_183291741 | 0.06 |
ENST00000351439.5 ENST00000409365.1 |
PDE1A |
phosphodiesterase 1A, calmodulin-dependent |
| chr1_+_247670415 | 0.06 |
ENST00000366491.2 ENST00000366489.1 ENST00000526896.1 |
GCSAML |
germinal center-associated, signaling and motility-like |
| chr7_-_140482926 | 0.06 |
ENST00000496384.2 |
BRAF |
v-raf murine sarcoma viral oncogene homolog B |
| chr19_+_12175504 | 0.06 |
ENST00000439326.3 |
ZNF844 |
zinc finger protein 844 |
| chr11_-_66964638 | 0.06 |
ENST00000444002.2 |
AP001885.1 |
AP001885.1 |
| chr6_+_29426230 | 0.06 |
ENST00000442615.1 |
OR2H1 |
olfactory receptor, family 2, subfamily H, member 1 |
| chr11_-_3400442 | 0.06 |
ENST00000429541.2 ENST00000532539.1 |
ZNF195 |
zinc finger protein 195 |
| chr17_-_3301704 | 0.06 |
ENST00000322608.2 |
OR1E1 |
olfactory receptor, family 1, subfamily E, member 1 |
| chr2_-_136288113 | 0.06 |
ENST00000401392.1 |
ZRANB3 |
zinc finger, RAN-binding domain containing 3 |
| chr9_+_70856397 | 0.06 |
ENST00000360171.6 |
CBWD3 |
COBW domain containing 3 |
| chr7_-_137028498 | 0.05 |
ENST00000393083.2 |
PTN |
pleiotrophin |
| chr7_-_7575477 | 0.05 |
ENST00000399429.3 |
COL28A1 |
collagen, type XXVIII, alpha 1 |
| chr5_+_135170331 | 0.05 |
ENST00000425402.1 ENST00000274513.5 ENST00000420621.1 ENST00000433282.2 ENST00000412661.2 |
SLC25A48 |
solute carrier family 25, member 48 |
| chr1_+_95616933 | 0.05 |
ENST00000604203.1 |
RP11-57H12.6 |
TMEM56-RWDD3 readthrough |
| chr15_-_35047166 | 0.05 |
ENST00000290374.4 |
GJD2 |
gap junction protein, delta 2, 36kDa |
| chr9_+_70856899 | 0.05 |
ENST00000377342.5 ENST00000478048.1 |
CBWD3 |
COBW domain containing 3 |
| chr9_-_21217310 | 0.05 |
ENST00000380216.1 |
IFNA16 |
interferon, alpha 16 |
| chr17_-_26220366 | 0.05 |
ENST00000460380.2 ENST00000508862.1 ENST00000379102.3 ENST00000582441.1 |
LYRM9 RP1-66C13.4 |
LYR motif containing 9 Uncharacterized protein |
| chrX_+_73164167 | 0.05 |
ENST00000414209.1 ENST00000602895.1 ENST00000453317.1 ENST00000602546.1 ENST00000602985.1 ENST00000415215.1 |
JPX |
JPX transcript, XIST activator (non-protein coding) |
| chr12_-_54982420 | 0.05 |
ENST00000257905.8 |
PPP1R1A |
protein phosphatase 1, regulatory (inhibitor) subunit 1A |
| chr12_+_26348246 | 0.05 |
ENST00000422622.2 |
SSPN |
sarcospan |
| chr2_-_178128250 | 0.05 |
ENST00000448782.1 ENST00000446151.2 |
NFE2L2 |
nuclear factor, erythroid 2-like 2 |
| chr22_+_22516550 | 0.05 |
ENST00000390284.2 |
IGLV4-60 |
immunoglobulin lambda variable 4-60 |
| chr14_-_81425828 | 0.04 |
ENST00000555529.1 ENST00000556042.1 ENST00000556981.1 |
CEP128 |
centrosomal protein 128kDa |
| chr5_+_79615790 | 0.04 |
ENST00000296739.4 |
SPZ1 |
spermatogenic leucine zipper 1 |
| chr12_+_48866448 | 0.04 |
ENST00000266594.1 |
ANP32D |
acidic (leucine-rich) nuclear phosphoprotein 32 family, member D |
| chr2_+_185463093 | 0.04 |
ENST00000302277.6 |
ZNF804A |
zinc finger protein 804A |
| chr3_+_195447738 | 0.04 |
ENST00000447234.2 ENST00000320736.6 ENST00000436408.1 |
MUC20 |
mucin 20, cell surface associated |
| chr19_+_21203426 | 0.04 |
ENST00000261560.5 ENST00000599548.1 ENST00000594110.1 |
ZNF430 |
zinc finger protein 430 |
| chr2_+_87135076 | 0.04 |
ENST00000409776.2 |
RGPD1 |
RANBP2-like and GRIP domain containing 1 |
| chr2_+_108145913 | 0.04 |
ENST00000443205.1 |
AC096669.3 |
AC096669.3 |
| chr16_-_15149828 | 0.04 |
ENST00000566419.1 ENST00000568320.1 |
NTAN1 |
N-terminal asparagine amidase |
| chr3_-_48956818 | 0.04 |
ENST00000408959.2 |
ARIH2OS |
ariadne homolog 2 opposite strand |
| chr5_+_157158205 | 0.04 |
ENST00000231198.7 |
THG1L |
tRNA-histidine guanylyltransferase 1-like (S. cerevisiae) |
| chr9_-_70490107 | 0.04 |
ENST00000377395.4 ENST00000429800.2 ENST00000430059.2 ENST00000377384.1 ENST00000382405.3 |
CBWD5 |
COBW domain containing 5 |
| chr9_-_139372141 | 0.04 |
ENST00000313050.7 |
SEC16A |
SEC16 homolog A (S. cerevisiae) |
| chr2_-_152830441 | 0.04 |
ENST00000534999.1 ENST00000397327.2 |
CACNB4 |
calcium channel, voltage-dependent, beta 4 subunit |
| chr7_-_144435985 | 0.04 |
ENST00000549981.1 |
TPK1 |
thiamin pyrophosphokinase 1 |
| chr1_-_110933611 | 0.04 |
ENST00000472422.2 ENST00000437429.2 |
SLC16A4 |
solute carrier family 16, member 4 |
| chr2_-_178128528 | 0.04 |
ENST00000397063.4 ENST00000421929.1 |
NFE2L2 |
nuclear factor, erythroid 2-like 2 |
| chr17_+_48823975 | 0.04 |
ENST00000513969.1 ENST00000503728.1 |
LUC7L3 |
LUC7-like 3 (S. cerevisiae) |
| chr7_-_137028534 | 0.04 |
ENST00000348225.2 |
PTN |
pleiotrophin |
| chr14_-_101295407 | 0.04 |
ENST00000596284.1 |
AL117190.2 |
AL117190.2 |
| chr2_+_87144738 | 0.03 |
ENST00000559485.1 |
RGPD1 |
RANBP2-like and GRIP domain containing 1 |
| chr2_-_88285309 | 0.03 |
ENST00000420840.2 |
RGPD2 |
RANBP2-like and GRIP domain containing 2 |
| chr12_+_94071341 | 0.03 |
ENST00000542893.2 |
CRADD |
CASP2 and RIPK1 domain containing adaptor with death domain |
| chr21_-_34863998 | 0.03 |
ENST00000402202.1 ENST00000381947.3 |
DNAJC28 |
DnaJ (Hsp40) homolog, subfamily C, member 28 |
| chr19_+_15838834 | 0.03 |
ENST00000305899.3 |
OR10H2 |
olfactory receptor, family 10, subfamily H, member 2 |
| chr11_+_125757556 | 0.03 |
ENST00000526028.1 |
HYLS1 |
hydrolethalus syndrome 1 |
| chr15_+_65843130 | 0.03 |
ENST00000569894.1 |
PTPLAD1 |
protein tyrosine phosphatase-like A domain containing 1 |
| chr4_-_109684120 | 0.03 |
ENST00000512646.1 ENST00000411864.2 ENST00000296486.3 ENST00000510706.1 |
ETNPPL |
ethanolamine-phosphate phospho-lyase |
| chr5_-_147162263 | 0.03 |
ENST00000333010.6 ENST00000265272.5 |
JAKMIP2 |
janus kinase and microtubule interacting protein 2 |
| chr8_+_119294456 | 0.03 |
ENST00000366457.2 |
AC023590.1 |
Uncharacterized protein |
| chr20_+_30458431 | 0.03 |
ENST00000375938.4 ENST00000535842.1 ENST00000310998.4 ENST00000375921.2 |
TTLL9 |
tubulin tyrosine ligase-like family, member 9 |
| chr1_-_247242048 | 0.03 |
ENST00000366503.2 |
ZNF670 |
zinc finger protein 670 |
| chr19_-_48547294 | 0.03 |
ENST00000293255.2 |
CABP5 |
calcium binding protein 5 |
| chr2_+_200775971 | 0.03 |
ENST00000319974.5 |
C2orf69 |
chromosome 2 open reading frame 69 |
| chr10_-_29923893 | 0.03 |
ENST00000355867.4 |
SVIL |
supervillin |
| chr3_-_119182523 | 0.03 |
ENST00000319172.5 |
TMEM39A |
transmembrane protein 39A |
| chr4_-_164534657 | 0.02 |
ENST00000339875.5 |
MARCH1 |
membrane-associated ring finger (C3HC4) 1, E3 ubiquitin protein ligase |
| chr16_-_53737722 | 0.02 |
ENST00000569716.1 ENST00000562588.1 ENST00000562230.1 ENST00000379925.3 ENST00000563746.1 ENST00000568653.3 |
RPGRIP1L |
RPGRIP1-like |
| chr3_-_149470229 | 0.02 |
ENST00000473414.1 |
COMMD2 |
COMM domain containing 2 |
| chr9_-_21974820 | 0.02 |
ENST00000579122.1 ENST00000498124.1 |
CDKN2A |
cyclin-dependent kinase inhibitor 2A |
| chr16_-_53737795 | 0.02 |
ENST00000262135.4 ENST00000564374.1 ENST00000566096.1 |
RPGRIP1L |
RPGRIP1-like |
| chr1_-_95391315 | 0.02 |
ENST00000545882.1 ENST00000415017.1 |
CNN3 |
calponin 3, acidic |
| chr19_+_47852538 | 0.02 |
ENST00000328771.4 |
DHX34 |
DEAH (Asp-Glu-Ala-His) box polypeptide 34 |
| chr14_+_22446680 | 0.02 |
ENST00000390443.3 |
TRAV8-6 |
T cell receptor alpha variable 8-6 |
| chr1_+_47603109 | 0.02 |
ENST00000371890.3 ENST00000294337.3 ENST00000371891.3 |
CYP4A22 |
cytochrome P450, family 4, subfamily A, polypeptide 22 |
| chr5_-_39270725 | 0.02 |
ENST00000512138.1 ENST00000512982.1 ENST00000540520.1 |
FYB |
FYN binding protein |
| chr11_+_60048129 | 0.02 |
ENST00000355131.3 |
MS4A4A |
membrane-spanning 4-domains, subfamily A, member 4A |
| chr11_+_72975559 | 0.02 |
ENST00000349767.2 |
P2RY6 |
pyrimidinergic receptor P2Y, G-protein coupled, 6 |
| chr16_-_15149917 | 0.02 |
ENST00000287706.3 |
NTAN1 |
N-terminal asparagine amidase |
| chr9_-_123676827 | 0.02 |
ENST00000546084.1 |
TRAF1 |
TNF receptor-associated factor 1 |
| chr6_+_32006042 | 0.02 |
ENST00000418967.2 |
CYP21A2 |
cytochrome P450, family 21, subfamily A, polypeptide 2 |
| chr8_+_75262612 | 0.02 |
ENST00000220822.7 |
GDAP1 |
ganglioside induced differentiation associated protein 1 |
| chr3_+_186915274 | 0.02 |
ENST00000312295.4 |
RTP1 |
receptor (chemosensory) transporter protein 1 |
| chr9_+_26956371 | 0.02 |
ENST00000380062.5 ENST00000518614.1 |
IFT74 |
intraflagellar transport 74 homolog (Chlamydomonas) |
| chr13_+_38923959 | 0.02 |
ENST00000379649.1 ENST00000239878.4 ENST00000437952.1 ENST00000379641.1 |
UFM1 |
ubiquitin-fold modifier 1 |
| chr1_+_28696111 | 0.02 |
ENST00000373839.3 |
PHACTR4 |
phosphatase and actin regulator 4 |
| chr11_-_31531121 | 0.01 |
ENST00000532287.1 ENST00000526776.1 ENST00000534812.1 ENST00000529749.1 ENST00000278200.1 ENST00000530023.1 ENST00000533642.1 |
IMMP1L |
IMP1 inner mitochondrial membrane peptidase-like (S. cerevisiae) |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:1900158 | negative regulation of osteoclast proliferation(GO:0090291) negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 0.2 | 0.7 | GO:1903410 | lysine import(GO:0034226) L-lysine import(GO:0061461) L-lysine import into cell(GO:1903410) |
| 0.1 | 0.3 | GO:0018242 | protein O-linked glycosylation via serine(GO:0018242) |
| 0.1 | 0.3 | GO:2000537 | regulation of cGMP-mediated signaling(GO:0010752) regulation of B cell chemotaxis(GO:2000537) positive regulation of B cell chemotaxis(GO:2000538) |
| 0.1 | 0.3 | GO:0030070 | insulin processing(GO:0030070) |
| 0.1 | 0.4 | GO:1904124 | microglial cell migration(GO:1904124) regulation of microglial cell migration(GO:1904139) |
| 0.0 | 0.1 | GO:0036058 | filtration diaphragm assembly(GO:0036058) slit diaphragm assembly(GO:0036060) |
| 0.0 | 0.4 | GO:0006848 | pyruvate transport(GO:0006848) pyruvate transmembrane transport(GO:1901475) |
| 0.0 | 0.2 | GO:2000672 | negative regulation of motor neuron apoptotic process(GO:2000672) |
| 0.0 | 0.3 | GO:0010897 | negative regulation of triglyceride catabolic process(GO:0010897) |
| 0.0 | 0.3 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.0 | 0.4 | GO:0034058 | endosomal vesicle fusion(GO:0034058) |
| 0.0 | 0.1 | GO:0052027 | modulation of signal transduction in other organism(GO:0044501) modulation by symbiont of host signal transduction pathway(GO:0052027) modulation of signal transduction in other organism involved in symbiotic interaction(GO:0052250) modulation by symbiont of host I-kappaB kinase/NF-kappaB cascade(GO:0085032) |
| 0.0 | 0.4 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.0 | 0.1 | GO:1904395 | positive regulation of skeletal muscle acetylcholine-gated channel clustering(GO:1904395) negative regulation of neuromuscular junction development(GO:1904397) |
| 0.0 | 0.1 | GO:0032764 | negative regulation of mast cell cytokine production(GO:0032764) |
| 0.0 | 0.3 | GO:1903943 | regulation of hepatocyte apoptotic process(GO:1903943) negative regulation of hepatocyte apoptotic process(GO:1903944) |
| 0.0 | 0.6 | GO:0001502 | cartilage condensation(GO:0001502) |
| 0.0 | 0.1 | GO:0033023 | mast cell homeostasis(GO:0033023) mast cell apoptotic process(GO:0033024) regulation of mast cell apoptotic process(GO:0033025) regulation of mast cell proliferation(GO:0070666) positive regulation of mast cell proliferation(GO:0070668) |
| 0.0 | 0.2 | GO:0090063 | positive regulation of microtubule nucleation(GO:0090063) |
| 0.0 | 0.2 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 0.0 | 0.1 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.0 | 0.4 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.0 | 0.3 | GO:0006069 | ethanol oxidation(GO:0006069) |
| 0.0 | 0.1 | GO:0042264 | peptidyl-aspartic acid hydroxylation(GO:0042264) |
| 0.0 | 0.1 | GO:1903788 | mycotoxin catabolic process(GO:0043387) aflatoxin catabolic process(GO:0046223) organic heteropentacyclic compound catabolic process(GO:1901377) regulation of glutathione biosynthetic process(GO:1903786) positive regulation of glutathione biosynthetic process(GO:1903788) |
| 0.0 | 0.2 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination(GO:2000042) |
| 0.0 | 0.3 | GO:0042759 | long-chain fatty acid biosynthetic process(GO:0042759) |
| 0.0 | 0.1 | GO:0051037 | regulation of transcription involved in meiotic cell cycle(GO:0051037) |
| 0.0 | 0.1 | GO:2000255 | negative regulation of male germ cell proliferation(GO:2000255) |
| 0.0 | 0.1 | GO:0036292 | DNA rewinding(GO:0036292) |
| 0.0 | 0.1 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.0 | 0.4 | GO:0044458 | motile cilium assembly(GO:0044458) |
| 0.0 | 0.1 | GO:2000301 | negative regulation of synaptic vesicle exocytosis(GO:2000301) |
| 0.0 | 0.0 | GO:0048213 | Golgi vesicle prefusion complex stabilization(GO:0048213) |
| 0.0 | 0.3 | GO:0007638 | mechanosensory behavior(GO:0007638) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0097224 | sperm connecting piece(GO:0097224) |
| 0.0 | 0.6 | GO:0070971 | endoplasmic reticulum exit site(GO:0070971) |
| 0.0 | 0.2 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 0.3 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) ripoptosome(GO:0097342) |
| 0.0 | 0.2 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.0 | 0.2 | GO:0005869 | dynactin complex(GO:0005869) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0005287 | high-affinity basic amino acid transmembrane transporter activity(GO:0005287) high-affinity arginine transmembrane transporter activity(GO:0005289) high-affinity lysine transmembrane transporter activity(GO:0005292) |
| 0.1 | 0.3 | GO:0030158 | protein xylosyltransferase activity(GO:0030158) |
| 0.1 | 0.3 | GO:0043423 | 3-phosphoinositide-dependent protein kinase binding(GO:0043423) |
| 0.1 | 0.4 | GO:0005477 | pyruvate secondary active transmembrane transporter activity(GO:0005477) |
| 0.1 | 0.7 | GO:0016015 | morphogen activity(GO:0016015) |
| 0.0 | 0.4 | GO:0001609 | G-protein coupled adenosine receptor activity(GO:0001609) |
| 0.0 | 0.3 | GO:0004024 | alcohol dehydrogenase activity, zinc-dependent(GO:0004024) |
| 0.0 | 0.4 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 0.0 | 0.1 | GO:0033149 | FFAT motif binding(GO:0033149) |
| 0.0 | 0.2 | GO:0061649 | ubiquitinated histone binding(GO:0061649) |
| 0.0 | 0.1 | GO:0004597 | peptide-aspartate beta-dioxygenase activity(GO:0004597) |
| 0.0 | 0.1 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.0 | 0.1 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.0 | 0.1 | GO:0035373 | chondroitin sulfate proteoglycan binding(GO:0035373) |
| 0.0 | 0.1 | GO:0047757 | chondroitin-glucuronate 5-epimerase activity(GO:0047757) |
| 0.0 | 0.2 | GO:0004630 | phospholipase D activity(GO:0004630) |
| 0.0 | 0.1 | GO:0004667 | prostaglandin-D synthase activity(GO:0004667) |
| 0.0 | 0.1 | GO:0034235 | GPI anchor binding(GO:0034235) |
| 0.0 | 0.0 | GO:0031862 | prostanoid receptor binding(GO:0031862) |
| 0.0 | 0.3 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.0 | 0.2 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.0 | 0.3 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.1 | GO:0035473 | lipase binding(GO:0035473) |
| 0.0 | 0.0 | GO:0008192 | RNA guanylyltransferase activity(GO:0008192) |
| 0.0 | 0.3 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.0 | 0.2 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.0 | 0.0 | GO:0016838 | carbon-oxygen lyase activity, acting on phosphates(GO:0016838) |
| 0.0 | 0.1 | GO:0000064 | L-ornithine transmembrane transporter activity(GO:0000064) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 0.3 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.0 | 0.4 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 0.7 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.0 | 0.4 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |