Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
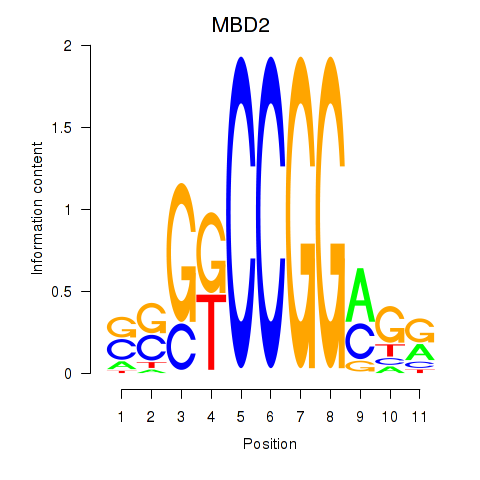
Results for MBD2
Z-value: 1.36
Transcription factors associated with MBD2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
MBD2
|
ENSG00000134046.7 | MBD2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| MBD2 | hg19_v2_chr18_-_51751132_51751158 | 0.79 | 1.9e-02 | Click! |
Activity profile of MBD2 motif
Sorted Z-values of MBD2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of MBD2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_+_54671047 | 1.59 |
ENST00000332822.4 |
NOG |
noggin |
| chr6_+_125474992 | 0.97 |
ENST00000528193.1 |
TPD52L1 |
tumor protein D52-like 1 |
| chr6_+_125475335 | 0.95 |
ENST00000532429.1 ENST00000534199.1 |
TPD52L1 |
tumor protein D52-like 1 |
| chr10_-_123357598 | 0.84 |
ENST00000358487.5 ENST00000369058.3 ENST00000369060.4 ENST00000359354.2 |
FGFR2 |
fibroblast growth factor receptor 2 |
| chr14_-_21566731 | 0.78 |
ENST00000360947.3 |
ZNF219 |
zinc finger protein 219 |
| chr6_-_99797522 | 0.77 |
ENST00000389677.5 |
FAXC |
failed axon connections homolog (Drosophila) |
| chr20_-_46415297 | 0.69 |
ENST00000467815.1 ENST00000359930.4 |
SULF2 |
sulfatase 2 |
| chr20_-_46415341 | 0.68 |
ENST00000484875.1 ENST00000361612.4 |
SULF2 |
sulfatase 2 |
| chr2_+_173292390 | 0.67 |
ENST00000442250.1 ENST00000458358.1 ENST00000409080.1 |
ITGA6 |
integrin, alpha 6 |
| chr10_-_123357910 | 0.66 |
ENST00000336553.6 ENST00000457416.2 ENST00000360144.3 ENST00000369059.1 ENST00000356226.4 ENST00000351936.6 |
FGFR2 |
fibroblast growth factor receptor 2 |
| chr1_+_82266053 | 0.65 |
ENST00000370715.1 ENST00000370713.1 ENST00000319517.6 ENST00000370717.2 ENST00000394879.1 ENST00000271029.4 ENST00000335786.5 |
LPHN2 |
latrophilin 2 |
| chr7_+_69064300 | 0.61 |
ENST00000342771.4 |
AUTS2 |
autism susceptibility candidate 2 |
| chr2_-_46769694 | 0.60 |
ENST00000522587.1 |
ATP6V1E2 |
ATPase, H+ transporting, lysosomal 31kDa, V1 subunit E2 |
| chr14_-_99737822 | 0.55 |
ENST00000345514.2 ENST00000443726.2 |
BCL11B |
B-cell CLL/lymphoma 11B (zinc finger protein) |
| chrX_+_152953505 | 0.55 |
ENST00000253122.5 |
SLC6A8 |
solute carrier family 6 (neurotransmitter transporter), member 8 |
| chr6_+_90142884 | 0.53 |
ENST00000369408.5 ENST00000339746.4 ENST00000447838.2 |
ANKRD6 |
ankyrin repeat domain 6 |
| chr6_+_17281573 | 0.50 |
ENST00000379052.5 |
RBM24 |
RNA binding motif protein 24 |
| chr16_+_2059872 | 0.48 |
ENST00000567649.1 |
NPW |
neuropeptide W |
| chr2_+_173292301 | 0.46 |
ENST00000264106.6 ENST00000375221.2 ENST00000343713.4 |
ITGA6 |
integrin, alpha 6 |
| chr9_-_135996537 | 0.43 |
ENST00000372050.3 ENST00000372047.3 |
RALGDS |
ral guanine nucleotide dissociation stimulator |
| chr5_+_167718604 | 0.42 |
ENST00000265293.4 |
WWC1 |
WW and C2 domain containing 1 |
| chr1_+_156252708 | 0.42 |
ENST00000295694.5 ENST00000357501.2 |
TMEM79 |
transmembrane protein 79 |
| chr9_+_124413873 | 0.41 |
ENST00000408936.3 |
DAB2IP |
DAB2 interacting protein |
| chr4_-_153456153 | 0.41 |
ENST00000603548.1 |
FBXW7 |
F-box and WD repeat domain containing 7, E3 ubiquitin protein ligase |
| chr3_+_112930306 | 0.41 |
ENST00000495514.1 |
BOC |
BOC cell adhesion associated, oncogene regulated |
| chr22_+_38035459 | 0.38 |
ENST00000357436.4 |
SH3BP1 |
SH3-domain binding protein 1 |
| chr10_-_92617671 | 0.37 |
ENST00000371721.3 |
HTR7 |
5-hydroxytryptamine (serotonin) receptor 7, adenylate cyclase-coupled |
| chr19_-_41222775 | 0.36 |
ENST00000324464.3 ENST00000450541.1 ENST00000594720.1 |
ADCK4 |
aarF domain containing kinase 4 |
| chr12_+_122064673 | 0.36 |
ENST00000537188.1 |
ORAI1 |
ORAI calcium release-activated calcium modulator 1 |
| chr1_-_6545502 | 0.35 |
ENST00000535355.1 |
PLEKHG5 |
pleckstrin homology domain containing, family G (with RhoGef domain) member 5 |
| chr2_+_26915584 | 0.34 |
ENST00000302909.3 |
KCNK3 |
potassium channel, subfamily K, member 3 |
| chr16_-_2059797 | 0.34 |
ENST00000563630.1 |
ZNF598 |
zinc finger protein 598 |
| chr20_-_50808236 | 0.33 |
ENST00000361387.2 |
ZFP64 |
ZFP64 zinc finger protein |
| chr1_-_54872059 | 0.33 |
ENST00000371320.3 |
SSBP3 |
single stranded DNA binding protein 3 |
| chr11_-_75236867 | 0.32 |
ENST00000376282.3 ENST00000336898.3 |
GDPD5 |
glycerophosphodiester phosphodiesterase domain containing 5 |
| chr6_+_4776580 | 0.32 |
ENST00000397588.3 |
CDYL |
chromodomain protein, Y-like |
| chr7_-_139876812 | 0.32 |
ENST00000397560.2 |
JHDM1D |
lysine (K)-specific demethylase 7A |
| chr17_-_41623075 | 0.31 |
ENST00000545089.1 |
ETV4 |
ets variant 4 |
| chr17_-_41623259 | 0.31 |
ENST00000538265.1 ENST00000591713.1 |
ETV4 |
ets variant 4 |
| chr10_-_92617437 | 0.31 |
ENST00000336152.3 ENST00000277874.6 ENST00000371719.2 |
HTR7 |
5-hydroxytryptamine (serotonin) receptor 7, adenylate cyclase-coupled |
| chr3_-_185542817 | 0.31 |
ENST00000382199.2 |
IGF2BP2 |
insulin-like growth factor 2 mRNA binding protein 2 |
| chr10_-_105615164 | 0.30 |
ENST00000355946.2 ENST00000369774.4 |
SH3PXD2A |
SH3 and PX domains 2A |
| chr10_+_129845823 | 0.30 |
ENST00000306042.5 |
PTPRE |
protein tyrosine phosphatase, receptor type, E |
| chr3_-_185542761 | 0.29 |
ENST00000457616.2 ENST00000346192.3 |
IGF2BP2 |
insulin-like growth factor 2 mRNA binding protein 2 |
| chr8_-_145688231 | 0.28 |
ENST00000530374.1 |
CYHR1 |
cysteine/histidine-rich 1 |
| chr1_-_32801825 | 0.28 |
ENST00000329421.7 |
MARCKSL1 |
MARCKS-like 1 |
| chr14_-_100070363 | 0.28 |
ENST00000380243.4 |
CCDC85C |
coiled-coil domain containing 85C |
| chr17_-_41623009 | 0.28 |
ENST00000393664.2 |
ETV4 |
ets variant 4 |
| chr11_-_72353451 | 0.28 |
ENST00000376450.3 |
PDE2A |
phosphodiesterase 2A, cGMP-stimulated |
| chr1_-_161008697 | 0.27 |
ENST00000318289.10 ENST00000368023.3 ENST00000368024.1 ENST00000423014.2 |
TSTD1 |
thiosulfate sulfurtransferase (rhodanese)-like domain containing 1 |
| chr12_+_50478977 | 0.27 |
ENST00000381513.4 |
SMARCD1 |
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 1 |
| chr10_-_103880209 | 0.27 |
ENST00000425280.1 |
LDB1 |
LIM domain binding 1 |
| chr11_-_115375107 | 0.27 |
ENST00000545380.1 ENST00000452722.3 ENST00000537058.1 ENST00000536727.1 ENST00000542447.2 ENST00000331581.6 |
CADM1 |
cell adhesion molecule 1 |
| chr3_+_112930387 | 0.27 |
ENST00000485230.1 |
BOC |
BOC cell adhesion associated, oncogene regulated |
| chr16_-_2059748 | 0.27 |
ENST00000562103.1 ENST00000431526.1 |
ZNF598 |
zinc finger protein 598 |
| chr1_+_4714792 | 0.27 |
ENST00000378190.3 |
AJAP1 |
adherens junctions associated protein 1 |
| chr12_-_57400227 | 0.26 |
ENST00000300101.2 |
ZBTB39 |
zinc finger and BTB domain containing 39 |
| chr8_+_30241934 | 0.24 |
ENST00000538486.1 |
RBPMS |
RNA binding protein with multiple splicing |
| chr19_-_17137625 | 0.24 |
ENST00000443236.1 ENST00000388925.4 |
CPAMD8 |
C3 and PZP-like, alpha-2-macroglobulin domain containing 8 |
| chr15_-_65067773 | 0.24 |
ENST00000300069.4 |
RBPMS2 |
RNA binding protein with multiple splicing 2 |
| chr11_-_79151695 | 0.24 |
ENST00000278550.7 |
TENM4 |
teneurin transmembrane protein 4 |
| chr3_-_52090461 | 0.23 |
ENST00000296483.6 ENST00000495880.1 |
DUSP7 |
dual specificity phosphatase 7 |
| chr3_-_71802760 | 0.23 |
ENST00000295612.3 |
EIF4E3 |
eukaryotic translation initiation factor 4E family member 3 |
| chr12_+_110152033 | 0.23 |
ENST00000538780.1 |
FAM222A |
family with sequence similarity 222, member A |
| chr2_-_129076151 | 0.23 |
ENST00000259241.6 |
HS6ST1 |
heparan sulfate 6-O-sulfotransferase 1 |
| chr9_+_129376722 | 0.23 |
ENST00000526117.1 ENST00000373474.4 ENST00000355497.5 ENST00000425646.2 ENST00000561065.1 |
LMX1B |
LIM homeobox transcription factor 1, beta |
| chr19_-_42759300 | 0.23 |
ENST00000222329.4 |
ERF |
Ets2 repressor factor |
| chr9_-_112260531 | 0.22 |
ENST00000374541.2 ENST00000262539.3 |
PTPN3 |
protein tyrosine phosphatase, non-receptor type 3 |
| chr18_-_53255766 | 0.22 |
ENST00000566286.1 ENST00000564999.1 ENST00000566279.1 ENST00000354452.3 ENST00000356073.4 |
TCF4 |
transcription factor 4 |
| chr14_+_93799556 | 0.22 |
ENST00000256339.4 |
UNC79 |
unc-79 homolog (C. elegans) |
| chr18_+_56530794 | 0.22 |
ENST00000590285.1 ENST00000586085.1 ENST00000589288.1 |
ZNF532 |
zinc finger protein 532 |
| chr16_+_75033210 | 0.21 |
ENST00000566250.1 ENST00000567962.1 |
ZNRF1 |
zinc and ring finger 1, E3 ubiquitin protein ligase |
| chr3_-_48470838 | 0.21 |
ENST00000358459.4 ENST00000358536.4 |
PLXNB1 |
plexin B1 |
| chr11_+_74459876 | 0.21 |
ENST00000299563.4 |
RNF169 |
ring finger protein 169 |
| chr3_-_128206759 | 0.21 |
ENST00000430265.2 |
GATA2 |
GATA binding protein 2 |
| chr4_+_6988884 | 0.20 |
ENST00000451522.2 |
TBC1D14 |
TBC1 domain family, member 14 |
| chr9_+_103189405 | 0.20 |
ENST00000395067.2 |
MSANTD3 |
Myb/SANT-like DNA-binding domain containing 3 |
| chr3_+_184279566 | 0.20 |
ENST00000330394.2 |
EPHB3 |
EPH receptor B3 |
| chr15_+_40650408 | 0.20 |
ENST00000267889.3 |
DISP2 |
dispatched homolog 2 (Drosophila) |
| chr19_+_35491174 | 0.20 |
ENST00000317991.5 ENST00000504615.2 |
GRAMD1A |
GRAM domain containing 1A |
| chr11_-_46142948 | 0.20 |
ENST00000257821.4 |
PHF21A |
PHD finger protein 21A |
| chr19_+_18496957 | 0.19 |
ENST00000252809.3 |
GDF15 |
growth differentiation factor 15 |
| chr14_+_103851712 | 0.19 |
ENST00000440884.3 ENST00000416682.2 ENST00000429436.2 ENST00000303622.9 |
MARK3 |
MAP/microtubule affinity-regulating kinase 3 |
| chr2_+_220306745 | 0.19 |
ENST00000431523.1 ENST00000396698.1 ENST00000396695.2 |
SPEG |
SPEG complex locus |
| chr2_-_220408430 | 0.19 |
ENST00000243776.6 |
CHPF |
chondroitin polymerizing factor |
| chr2_+_42275153 | 0.19 |
ENST00000294964.5 |
PKDCC |
protein kinase domain containing, cytoplasmic |
| chr6_-_10415470 | 0.19 |
ENST00000379604.2 ENST00000379613.3 |
TFAP2A |
transcription factor AP-2 alpha (activating enhancer binding protein 2 alpha) |
| chr6_-_43596899 | 0.19 |
ENST00000307126.5 ENST00000452781.1 |
GTPBP2 |
GTP binding protein 2 |
| chr17_+_65374075 | 0.18 |
ENST00000581322.1 |
PITPNC1 |
phosphatidylinositol transfer protein, cytoplasmic 1 |
| chr4_+_38665810 | 0.18 |
ENST00000261438.5 ENST00000514033.1 |
KLF3 |
Kruppel-like factor 3 (basic) |
| chr3_+_14989186 | 0.18 |
ENST00000435454.1 ENST00000323373.6 |
NR2C2 |
nuclear receptor subfamily 2, group C, member 2 |
| chr19_-_46148820 | 0.18 |
ENST00000587152.1 |
EML2 |
echinoderm microtubule associated protein like 2 |
| chr14_-_93581615 | 0.18 |
ENST00000555553.1 ENST00000555495.1 ENST00000554999.1 |
ITPK1 |
inositol-tetrakisphosphate 1-kinase |
| chr11_+_67033881 | 0.18 |
ENST00000308595.5 ENST00000526285.1 |
ADRBK1 |
adrenergic, beta, receptor kinase 1 |
| chr16_+_57769635 | 0.18 |
ENST00000379661.3 ENST00000562592.1 ENST00000566726.1 |
KATNB1 |
katanin p80 (WD repeat containing) subunit B 1 |
| chr2_-_54197915 | 0.18 |
ENST00000404125.1 |
PSME4 |
proteasome (prosome, macropain) activator subunit 4 |
| chr1_+_84543821 | 0.18 |
ENST00000370688.3 |
PRKACB |
protein kinase, cAMP-dependent, catalytic, beta |
| chr13_-_21476900 | 0.18 |
ENST00000400602.2 ENST00000255305.6 |
XPO4 |
exportin 4 |
| chr9_+_138628365 | 0.18 |
ENST00000491806.2 ENST00000488444.2 ENST00000490355.2 ENST00000263604.3 |
KCNT1 |
potassium channel, subfamily T, member 1 |
| chr2_+_113033164 | 0.17 |
ENST00000409871.1 ENST00000343936.4 |
ZC3H6 |
zinc finger CCCH-type containing 6 |
| chr11_+_100558384 | 0.17 |
ENST00000524892.2 ENST00000298815.8 |
ARHGAP42 |
Rho GTPase activating protein 42 |
| chr17_+_80477571 | 0.17 |
ENST00000335255.5 |
FOXK2 |
forkhead box K2 |
| chr20_+_20348740 | 0.17 |
ENST00000310227.1 |
INSM1 |
insulinoma-associated 1 |
| chr6_+_139456226 | 0.17 |
ENST00000367658.2 |
HECA |
headcase homolog (Drosophila) |
| chr19_+_35491330 | 0.16 |
ENST00000411896.2 ENST00000424536.2 |
GRAMD1A |
GRAM domain containing 1A |
| chr3_-_48471454 | 0.16 |
ENST00000296440.6 ENST00000448774.2 |
PLXNB1 |
plexin B1 |
| chr1_+_156698234 | 0.16 |
ENST00000368218.4 ENST00000368216.4 |
RRNAD1 |
ribosomal RNA adenine dimethylase domain containing 1 |
| chr11_+_46354455 | 0.16 |
ENST00000343674.6 |
DGKZ |
diacylglycerol kinase, zeta |
| chr19_-_1568057 | 0.16 |
ENST00000402693.4 ENST00000388824.6 |
MEX3D |
mex-3 RNA binding family member D |
| chr3_-_38691119 | 0.15 |
ENST00000333535.4 ENST00000413689.1 ENST00000443581.1 ENST00000425664.1 ENST00000451551.2 |
SCN5A |
sodium channel, voltage-gated, type V, alpha subunit |
| chr14_+_105886150 | 0.15 |
ENST00000331320.7 ENST00000406191.1 |
MTA1 |
metastasis associated 1 |
| chr17_+_17942684 | 0.15 |
ENST00000376345.3 |
GID4 |
GID complex subunit 4 |
| chr2_-_38604398 | 0.15 |
ENST00000443098.1 ENST00000449130.1 ENST00000378954.4 ENST00000539122.1 ENST00000419554.2 ENST00000451483.1 ENST00000406122.1 |
ATL2 |
atlastin GTPase 2 |
| chr17_-_61920280 | 0.15 |
ENST00000448276.2 ENST00000577990.1 |
SMARCD2 |
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 2 |
| chr6_+_29795595 | 0.15 |
ENST00000360323.6 ENST00000376818.3 ENST00000376815.3 |
HLA-G |
major histocompatibility complex, class I, G |
| chr6_+_42981922 | 0.15 |
ENST00000326974.4 ENST00000244670.8 |
KLHDC3 |
kelch domain containing 3 |
| chr19_-_17356697 | 0.15 |
ENST00000291442.3 |
NR2F6 |
nuclear receptor subfamily 2, group F, member 6 |
| chr19_-_46476791 | 0.14 |
ENST00000263257.5 |
NOVA2 |
neuro-oncological ventral antigen 2 |
| chr20_+_55204351 | 0.14 |
ENST00000201031.2 |
TFAP2C |
transcription factor AP-2 gamma (activating enhancer binding protein 2 gamma) |
| chr7_-_129251459 | 0.14 |
ENST00000608694.1 |
RP11-448A19.1 |
RP11-448A19.1 |
| chr7_+_94285637 | 0.14 |
ENST00000482108.1 ENST00000488574.1 |
PEG10 |
paternally expressed 10 |
| chr18_+_8609402 | 0.14 |
ENST00000329286.6 |
RAB12 |
RAB12, member RAS oncogene family |
| chr1_-_9884011 | 0.14 |
ENST00000361311.4 |
CLSTN1 |
calsyntenin 1 |
| chr11_+_66886717 | 0.14 |
ENST00000398645.2 |
KDM2A |
lysine (K)-specific demethylase 2A |
| chr1_-_6546001 | 0.14 |
ENST00000400913.1 |
PLEKHG5 |
pleckstrin homology domain containing, family G (with RhoGef domain) member 5 |
| chr3_-_27525826 | 0.14 |
ENST00000454389.1 ENST00000440156.1 ENST00000437179.1 ENST00000446700.1 ENST00000455077.1 ENST00000435667.2 ENST00000388777.4 ENST00000425128.2 |
SLC4A7 |
solute carrier family 4, sodium bicarbonate cotransporter, member 7 |
| chr17_-_42441204 | 0.14 |
ENST00000293443.7 |
FAM171A2 |
family with sequence similarity 171, member A2 |
| chr6_-_10415218 | 0.14 |
ENST00000466073.1 ENST00000498450.1 |
TFAP2A |
transcription factor AP-2 alpha (activating enhancer binding protein 2 alpha) |
| chrX_-_40594755 | 0.14 |
ENST00000324817.1 |
MED14 |
mediator complex subunit 14 |
| chr16_-_23160591 | 0.14 |
ENST00000219689.7 |
USP31 |
ubiquitin specific peptidase 31 |
| chr3_-_53080047 | 0.14 |
ENST00000482396.1 ENST00000358080.2 ENST00000296295.6 ENST00000394752.3 |
SFMBT1 |
Scm-like with four mbt domains 1 |
| chr1_-_156698181 | 0.14 |
ENST00000313146.6 |
ISG20L2 |
interferon stimulated exonuclease gene 20kDa-like 2 |
| chr19_-_19051103 | 0.14 |
ENST00000542541.2 ENST00000433218.2 |
HOMER3 |
homer homolog 3 (Drosophila) |
| chr1_-_200379129 | 0.13 |
ENST00000367353.1 |
ZNF281 |
zinc finger protein 281 |
| chr2_+_9346892 | 0.13 |
ENST00000281419.3 ENST00000315273.4 |
ASAP2 |
ArfGAP with SH3 domain, ankyrin repeat and PH domain 2 |
| chr1_+_9599540 | 0.13 |
ENST00000302692.6 |
SLC25A33 |
solute carrier family 25 (pyrimidine nucleotide carrier), member 33 |
| chr18_+_77155942 | 0.13 |
ENST00000397790.2 |
NFATC1 |
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 1 |
| chr2_+_48541776 | 0.13 |
ENST00000413569.1 ENST00000340553.3 |
FOXN2 |
forkhead box N2 |
| chr19_-_17445613 | 0.13 |
ENST00000159087.4 |
ANO8 |
anoctamin 8 |
| chr17_+_17942594 | 0.13 |
ENST00000268719.4 |
GID4 |
GID complex subunit 4 |
| chrY_+_2803322 | 0.13 |
ENST00000383052.1 ENST00000155093.3 ENST00000449237.1 ENST00000443793.1 |
ZFY |
zinc finger protein, Y-linked |
| chr14_-_103523745 | 0.13 |
ENST00000361246.2 |
CDC42BPB |
CDC42 binding protein kinase beta (DMPK-like) |
| chr3_+_14989076 | 0.13 |
ENST00000413118.1 ENST00000425241.1 |
NR2C2 |
nuclear receptor subfamily 2, group C, member 2 |
| chr3_+_133292759 | 0.13 |
ENST00000431519.2 |
CDV3 |
CDV3 homolog (mouse) |
| chr3_-_197024394 | 0.13 |
ENST00000434148.1 ENST00000412364.2 ENST00000436682.1 ENST00000456699.2 ENST00000392380.2 |
DLG1 |
discs, large homolog 1 (Drosophila) |
| chr2_-_208031542 | 0.13 |
ENST00000423015.1 |
KLF7 |
Kruppel-like factor 7 (ubiquitous) |
| chr4_-_16228083 | 0.13 |
ENST00000399920.3 |
TAPT1 |
transmembrane anterior posterior transformation 1 |
| chr13_+_98795505 | 0.13 |
ENST00000319562.6 |
FARP1 |
FERM, RhoGEF (ARHGEF) and pleckstrin domain protein 1 (chondrocyte-derived) |
| chr18_-_812231 | 0.12 |
ENST00000314574.4 |
YES1 |
v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 |
| chr13_+_98795434 | 0.12 |
ENST00000376586.2 |
FARP1 |
FERM, RhoGEF (ARHGEF) and pleckstrin domain protein 1 (chondrocyte-derived) |
| chr7_+_40174565 | 0.12 |
ENST00000309930.5 ENST00000401647.2 ENST00000335693.4 ENST00000413931.1 ENST00000416370.1 ENST00000540834.1 |
C7orf10 |
succinylCoA:glutarate-CoA transferase |
| chr17_-_48227877 | 0.12 |
ENST00000316878.6 |
PPP1R9B |
protein phosphatase 1, regulatory subunit 9B |
| chrX_+_24167828 | 0.12 |
ENST00000379188.3 ENST00000419690.1 ENST00000379177.1 ENST00000304543.5 |
ZFX |
zinc finger protein, X-linked |
| chr18_-_812517 | 0.12 |
ENST00000584307.1 |
YES1 |
v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 |
| chr1_+_153606513 | 0.12 |
ENST00000368694.3 |
CHTOP |
chromatin target of PRMT1 |
| chr6_-_31763721 | 0.12 |
ENST00000375663.3 |
VARS |
valyl-tRNA synthetase |
| chr19_+_38397839 | 0.12 |
ENST00000222345.6 |
SIPA1L3 |
signal-induced proliferation-associated 1 like 3 |
| chr15_+_90544532 | 0.12 |
ENST00000268154.4 |
ZNF710 |
zinc finger protein 710 |
| chr5_+_122847908 | 0.12 |
ENST00000511130.2 ENST00000512718.3 |
CSNK1G3 |
casein kinase 1, gamma 3 |
| chr19_+_42388437 | 0.12 |
ENST00000378152.4 ENST00000337665.4 |
ARHGEF1 |
Rho guanine nucleotide exchange factor (GEF) 1 |
| chr2_+_9696028 | 0.12 |
ENST00000607241.1 |
RP11-214N9.1 |
RP11-214N9.1 |
| chr3_-_133969437 | 0.12 |
ENST00000460933.1 ENST00000296084.4 |
RYK |
receptor-like tyrosine kinase |
| chr1_-_154531095 | 0.11 |
ENST00000292211.4 |
UBE2Q1 |
ubiquitin-conjugating enzyme E2Q family member 1 |
| chrX_+_68725084 | 0.11 |
ENST00000252338.4 |
FAM155B |
family with sequence similarity 155, member B |
| chr20_-_60640866 | 0.11 |
ENST00000252996.4 |
TAF4 |
TAF4 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 135kDa |
| chr16_+_2022036 | 0.11 |
ENST00000568546.1 |
TBL3 |
transducin (beta)-like 3 |
| chr6_-_45983549 | 0.11 |
ENST00000544153.1 |
CLIC5 |
chloride intracellular channel 5 |
| chr1_+_28844778 | 0.11 |
ENST00000411533.1 |
RCC1 |
regulator of chromosome condensation 1 |
| chr6_+_139349817 | 0.11 |
ENST00000367660.3 |
ABRACL |
ABRA C-terminal like |
| chr10_+_106400859 | 0.11 |
ENST00000369701.3 |
SORCS3 |
sortilin-related VPS10 domain containing receptor 3 |
| chr6_+_41514078 | 0.11 |
ENST00000373063.3 ENST00000373060.1 |
FOXP4 |
forkhead box P4 |
| chr17_+_6544356 | 0.10 |
ENST00000574838.1 |
TXNDC17 |
thioredoxin domain containing 17 |
| chr8_-_119124045 | 0.10 |
ENST00000378204.2 |
EXT1 |
exostosin glycosyltransferase 1 |
| chr1_-_156698591 | 0.10 |
ENST00000368219.1 |
ISG20L2 |
interferon stimulated exonuclease gene 20kDa-like 2 |
| chr19_+_36208877 | 0.10 |
ENST00000420124.1 ENST00000222270.7 ENST00000341701.1 |
KMT2B |
Histone-lysine N-methyltransferase 2B |
| chr6_+_1610681 | 0.10 |
ENST00000380874.2 |
FOXC1 |
forkhead box C1 |
| chr14_+_103243813 | 0.10 |
ENST00000560371.1 ENST00000347662.4 ENST00000392745.2 ENST00000539721.1 ENST00000560463.1 |
TRAF3 |
TNF receptor-associated factor 3 |
| chr1_+_110881945 | 0.10 |
ENST00000602849.1 ENST00000487146.2 |
RBM15 |
RNA binding motif protein 15 |
| chr15_+_92397051 | 0.10 |
ENST00000424469.2 |
SLCO3A1 |
solute carrier organic anion transporter family, member 3A1 |
| chr8_+_30241995 | 0.10 |
ENST00000397323.4 ENST00000339877.4 ENST00000320203.4 ENST00000287771.5 |
RBPMS |
RNA binding protein with multiple splicing |
| chr1_+_3689325 | 0.10 |
ENST00000444870.2 ENST00000452264.1 |
SMIM1 |
small integral membrane protein 1 (Vel blood group) |
| chr3_+_196295482 | 0.10 |
ENST00000440469.1 ENST00000311630.6 |
FBXO45 |
F-box protein 45 |
| chr17_+_17685422 | 0.10 |
ENST00000395774.1 |
RAI1 |
retinoic acid induced 1 |
| chr16_+_2732476 | 0.10 |
ENST00000301738.4 ENST00000564195.1 |
KCTD5 |
potassium channel tetramerization domain containing 5 |
| chr6_-_91296737 | 0.10 |
ENST00000369332.3 ENST00000369329.3 |
MAP3K7 |
mitogen-activated protein kinase kinase kinase 7 |
| chr2_+_176972000 | 0.10 |
ENST00000249504.5 |
HOXD11 |
homeobox D11 |
| chr18_+_77155856 | 0.10 |
ENST00000253506.5 ENST00000591814.1 |
NFATC1 |
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 1 |
| chr1_-_156252590 | 0.10 |
ENST00000361813.5 ENST00000368267.5 |
SMG5 |
SMG5 nonsense mediated mRNA decay factor |
| chr12_-_117319236 | 0.10 |
ENST00000257572.5 |
HRK |
harakiri, BCL2 interacting protein (contains only BH3 domain) |
| chr12_+_19282713 | 0.10 |
ENST00000299275.6 ENST00000539256.1 ENST00000538714.1 |
PLEKHA5 |
pleckstrin homology domain containing, family A member 5 |
| chr17_+_65373531 | 0.10 |
ENST00000580974.1 |
PITPNC1 |
phosphatidylinositol transfer protein, cytoplasmic 1 |
| chr11_-_73471655 | 0.10 |
ENST00000400470.2 |
RAB6A |
RAB6A, member RAS oncogene family |
| chr1_-_244013384 | 0.10 |
ENST00000366539.1 |
AKT3 |
v-akt murine thymoma viral oncogene homolog 3 |
| chr1_-_179851611 | 0.10 |
ENST00000610272.1 |
RP11-533E19.7 |
RP11-533E19.7 |
| chr8_+_145137489 | 0.10 |
ENST00000355091.4 ENST00000525087.1 ENST00000361036.6 ENST00000524418.1 |
GPAA1 |
glycosylphosphatidylinositol anchor attachment 1 |
| chr17_+_27920486 | 0.10 |
ENST00000394859.3 |
ANKRD13B |
ankyrin repeat domain 13B |
| chr12_+_112856690 | 0.10 |
ENST00000392597.1 ENST00000351677.2 |
PTPN11 |
protein tyrosine phosphatase, non-receptor type 11 |
| chr17_-_29151794 | 0.10 |
ENST00000324238.6 |
CRLF3 |
cytokine receptor-like factor 3 |
| chr5_-_180076613 | 0.10 |
ENST00000261937.6 ENST00000393347.3 |
FLT4 |
fms-related tyrosine kinase 4 |
| chr11_-_89224638 | 0.10 |
ENST00000535633.1 ENST00000263317.4 |
NOX4 |
NADPH oxidase 4 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.5 | GO:0035604 | fibroblast growth factor receptor signaling pathway involved in negative regulation of apoptotic process in bone marrow(GO:0035602) fibroblast growth factor receptor signaling pathway involved in hemopoiesis(GO:0035603) fibroblast growth factor receptor signaling pathway involved in positive regulation of cell proliferation in bone marrow(GO:0035604) |
| 0.3 | 1.4 | GO:0014846 | esophagus smooth muscle contraction(GO:0014846) |
| 0.3 | 1.6 | GO:2000313 | fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:0060825) regulation of fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:2000313) |
| 0.2 | 0.6 | GO:0098582 | innate vocalization behavior(GO:0098582) |
| 0.2 | 0.7 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.2 | 0.6 | GO:0072344 | rescue of stalled ribosome(GO:0072344) |
| 0.1 | 0.4 | GO:1900220 | semaphorin-plexin signaling pathway involved in bone trabecula morphogenesis(GO:1900220) |
| 0.1 | 0.6 | GO:0021773 | striatal medium spiny neuron differentiation(GO:0021773) |
| 0.1 | 0.3 | GO:0030860 | regulation of polarized epithelial cell differentiation(GO:0030860) |
| 0.1 | 0.4 | GO:0043553 | vascular endothelial growth factor receptor-2 signaling pathway(GO:0036324) negative regulation of phosphatidylinositol 3-kinase activity(GO:0043553) |
| 0.1 | 0.4 | GO:2000639 | regulation of SREBP signaling pathway(GO:2000638) negative regulation of SREBP signaling pathway(GO:2000639) |
| 0.1 | 0.3 | GO:0003409 | optic cup structural organization(GO:0003409) |
| 0.1 | 1.1 | GO:0010668 | ectodermal cell differentiation(GO:0010668) |
| 0.1 | 0.2 | GO:0032289 | central nervous system myelin formation(GO:0032289) |
| 0.1 | 0.3 | GO:0033159 | negative regulation of protein import into nucleus, translocation(GO:0033159) |
| 0.1 | 0.2 | GO:0035854 | regulation of primitive erythrocyte differentiation(GO:0010725) eosinophil fate commitment(GO:0035854) |
| 0.1 | 0.4 | GO:0042335 | cuticle development(GO:0042335) |
| 0.1 | 0.3 | GO:0042271 | susceptibility to natural killer cell mediated cytotoxicity(GO:0042271) |
| 0.1 | 0.3 | GO:0021999 | neural plate anterior/posterior regionalization(GO:0021999) |
| 0.1 | 0.2 | GO:0003358 | noradrenergic neuron development(GO:0003358) |
| 0.1 | 0.6 | GO:0006600 | creatine metabolic process(GO:0006600) |
| 0.1 | 0.2 | GO:0071955 | recycling endosome to Golgi transport(GO:0071955) |
| 0.1 | 0.3 | GO:0072675 | multinuclear osteoclast differentiation(GO:0072674) osteoclast fusion(GO:0072675) |
| 0.1 | 0.6 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 0.0 | 0.2 | GO:0036518 | chemorepulsion of dopaminergic neuron axon(GO:0036518) |
| 0.0 | 0.2 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.0 | 0.3 | GO:0046985 | positive regulation of hemoglobin biosynthetic process(GO:0046985) |
| 0.0 | 0.2 | GO:0031117 | positive regulation of microtubule depolymerization(GO:0031117) |
| 0.0 | 0.1 | GO:0006864 | pyrimidine nucleotide transport(GO:0006864) mitochondrial pyrimidine nucleotide import(GO:1990519) |
| 0.0 | 0.2 | GO:1904694 | negative regulation of vascular smooth muscle contraction(GO:1904694) |
| 0.0 | 0.2 | GO:1902498 | regulation of protein autoubiquitination(GO:1902498) |
| 0.0 | 0.2 | GO:2001021 | negative regulation of DNA repair(GO:0045738) negative regulation of double-strand break repair(GO:2000780) negative regulation of response to DNA damage stimulus(GO:2001021) |
| 0.0 | 0.1 | GO:0001560 | regulation of cell growth by extracellular stimulus(GO:0001560) |
| 0.0 | 0.1 | GO:0003275 | apoptotic process involved in outflow tract morphogenesis(GO:0003275) negative regulation of lymphangiogenesis(GO:1901491) regulation of apoptotic process involved in outflow tract morphogenesis(GO:1902256) |
| 0.0 | 0.1 | GO:0061582 | intestinal epithelial cell migration(GO:0061582) |
| 0.0 | 0.0 | GO:0010512 | negative regulation of phosphatidylinositol biosynthetic process(GO:0010512) |
| 0.0 | 0.1 | GO:0033023 | mast cell homeostasis(GO:0033023) mast cell apoptotic process(GO:0033024) regulation of mast cell apoptotic process(GO:0033025) |
| 0.0 | 0.2 | GO:0010609 | mRNA localization resulting in posttranscriptional regulation of gene expression(GO:0010609) |
| 0.0 | 0.1 | GO:0021919 | BMP signaling pathway involved in spinal cord dorsal/ventral patterning(GO:0021919) |
| 0.0 | 0.1 | GO:0003032 | detection of oxygen(GO:0003032) |
| 0.0 | 0.3 | GO:0040034 | regulation of development, heterochronic(GO:0040034) regulation of timing of cell differentiation(GO:0048505) |
| 0.0 | 0.1 | GO:0006535 | cysteine biosynthetic process from serine(GO:0006535) |
| 0.0 | 0.2 | GO:0035093 | spermatogenesis, exchange of chromosomal proteins(GO:0035093) |
| 0.0 | 0.1 | GO:0007221 | positive regulation of transcription of Notch receptor target(GO:0007221) |
| 0.0 | 0.2 | GO:0060373 | regulation of ventricular cardiac muscle cell membrane depolarization(GO:0060373) |
| 0.0 | 0.1 | GO:0035526 | retrograde transport, plasma membrane to Golgi(GO:0035526) |
| 0.0 | 0.7 | GO:0045663 | positive regulation of myoblast differentiation(GO:0045663) |
| 0.0 | 0.1 | GO:0006438 | valyl-tRNA aminoacylation(GO:0006438) |
| 0.0 | 0.2 | GO:0042997 | negative regulation of Golgi to plasma membrane protein transport(GO:0042997) |
| 0.0 | 0.1 | GO:0044314 | protein K27-linked ubiquitination(GO:0044314) |
| 0.0 | 0.3 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.0 | 0.4 | GO:0002115 | store-operated calcium entry(GO:0002115) |
| 0.0 | 0.1 | GO:0071921 | establishment of sister chromatid cohesion(GO:0034085) cohesin loading(GO:0071921) regulation of cohesin loading(GO:0071922) |
| 0.0 | 0.2 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.0 | 0.4 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.0 | 0.4 | GO:0060391 | positive regulation of SMAD protein import into nucleus(GO:0060391) |
| 0.0 | 0.1 | GO:0021722 | superior olivary nucleus development(GO:0021718) superior olivary nucleus maturation(GO:0021722) |
| 0.0 | 0.1 | GO:0042137 | sequestering of neurotransmitter(GO:0042137) |
| 0.0 | 0.1 | GO:0018125 | peptidyl-cysteine methylation(GO:0018125) |
| 0.0 | 0.1 | GO:0018076 | N-terminal peptidyl-lysine acetylation(GO:0018076) |
| 0.0 | 0.1 | GO:0046952 | ketone body catabolic process(GO:0046952) |
| 0.0 | 0.1 | GO:0090118 | receptor-mediated endocytosis of low-density lipoprotein particle involved in cholesterol transport(GO:0090118) |
| 0.0 | 0.1 | GO:0010748 | regulation of plasma membrane long-chain fatty acid transport(GO:0010746) negative regulation of plasma membrane long-chain fatty acid transport(GO:0010748) |
| 0.0 | 0.1 | GO:0008063 | Toll signaling pathway(GO:0008063) |
| 0.0 | 0.1 | GO:0097338 | response to clozapine(GO:0097338) |
| 0.0 | 0.1 | GO:1902683 | positive regulation of long-term neuronal synaptic plasticity(GO:0048170) regulation of receptor localization to synapse(GO:1902683) |
| 0.0 | 0.3 | GO:0030322 | stabilization of membrane potential(GO:0030322) |
| 0.0 | 0.2 | GO:0051533 | positive regulation of NFAT protein import into nucleus(GO:0051533) |
| 0.0 | 0.1 | GO:0021957 | corticospinal tract morphogenesis(GO:0021957) |
| 0.0 | 0.1 | GO:0061357 | positive regulation of Wnt protein secretion(GO:0061357) |
| 0.0 | 0.0 | GO:0071930 | negative regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0071930) |
| 0.0 | 0.0 | GO:0070634 | transepithelial ammonium transport(GO:0070634) |
| 0.0 | 0.2 | GO:0051151 | negative regulation of smooth muscle cell differentiation(GO:0051151) |
| 0.0 | 0.1 | GO:2000051 | Wnt receptor catabolic process(GO:0038018) negative regulation of non-canonical Wnt signaling pathway(GO:2000051) |
| 0.0 | 0.1 | GO:1903764 | regulation of potassium ion export across plasma membrane(GO:1903764) |
| 0.0 | 0.1 | GO:0070544 | histone H3-K36 demethylation(GO:0070544) |
| 0.0 | 0.0 | GO:1903233 | regulation of calcium ion-dependent exocytosis of neurotransmitter(GO:1903233) |
| 0.0 | 0.6 | GO:0090383 | phagosome acidification(GO:0090383) |
| 0.0 | 0.2 | GO:0010968 | regulation of microtubule nucleation(GO:0010968) |
| 0.0 | 0.1 | GO:0002480 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-independent(GO:0002480) |
| 0.0 | 0.1 | GO:2000052 | positive regulation of non-canonical Wnt signaling pathway(GO:2000052) |
| 0.0 | 0.1 | GO:1902174 | positive regulation of keratinocyte apoptotic process(GO:1902174) |
| 0.0 | 0.3 | GO:0048149 | behavioral response to ethanol(GO:0048149) |
| 0.0 | 0.2 | GO:1901741 | positive regulation of myoblast fusion(GO:1901741) |
| 0.0 | 0.1 | GO:1904098 | regulation of protein O-linked glycosylation(GO:1904098) positive regulation of protein O-linked glycosylation(GO:1904100) |
| 0.0 | 0.1 | GO:0051096 | positive regulation of helicase activity(GO:0051096) |
| 0.0 | 0.0 | GO:0002838 | negative regulation of response to tumor cell(GO:0002835) negative regulation of immune response to tumor cell(GO:0002838) negative regulation of natural killer cell mediated immune response to tumor cell(GO:0002856) negative regulation of natural killer cell mediated cytotoxicity directed against tumor cell target(GO:0002859) |
| 0.0 | 0.1 | GO:0090625 | siRNA loading onto RISC involved in RNA interference(GO:0035087) mRNA cleavage involved in gene silencing by siRNA(GO:0090625) |
| 0.0 | 0.2 | GO:0090037 | regulation of blood vessel remodeling(GO:0060312) positive regulation of protein kinase C signaling(GO:0090037) |
| 0.0 | 0.0 | GO:0021571 | rhombomere 5 development(GO:0021571) central nervous system segmentation(GO:0035283) brain segmentation(GO:0035284) |
| 0.0 | 0.1 | GO:2000382 | positive regulation of mesoderm development(GO:2000382) |
| 0.0 | 0.0 | GO:0002071 | glandular epithelial cell maturation(GO:0002071) type B pancreatic cell maturation(GO:0072560) |
| 0.0 | 0.1 | GO:1900738 | positive regulation of phospholipase C-activating G-protein coupled receptor signaling pathway(GO:1900738) |
| 0.0 | 0.1 | GO:0016480 | negative regulation of transcription from RNA polymerase III promoter(GO:0016480) |
| 0.0 | 0.1 | GO:0006572 | tyrosine catabolic process(GO:0006572) |
| 0.0 | 0.5 | GO:0035767 | endothelial cell chemotaxis(GO:0035767) |
| 0.0 | 0.1 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.0 | 0.1 | GO:0000389 | mRNA 3'-splice site recognition(GO:0000389) |
| 0.0 | 0.1 | GO:1900452 | regulation of long term synaptic depression(GO:1900452) |
| 0.0 | 0.1 | GO:0051987 | positive regulation of attachment of spindle microtubules to kinetochore(GO:0051987) |
| 0.0 | 0.1 | GO:0090170 | regulation of Golgi inheritance(GO:0090170) |
| 0.0 | 0.1 | GO:0015014 | heparan sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0015014) |
| 0.0 | 0.1 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.0 | 0.0 | GO:0061054 | dermatome development(GO:0061054) regulation of dermatome development(GO:0061183) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.1 | GO:0034676 | integrin alpha6-beta4 complex(GO:0034676) |
| 0.1 | 0.4 | GO:1990032 | parallel fiber(GO:1990032) |
| 0.1 | 0.4 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
| 0.1 | 0.3 | GO:0044214 | spanning component of plasma membrane(GO:0044214) spanning component of membrane(GO:0089717) |
| 0.0 | 0.4 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.0 | 0.1 | GO:0070985 | TFIIK complex(GO:0070985) |
| 0.0 | 0.6 | GO:0033178 | proton-transporting two-sector ATPase complex, catalytic domain(GO:0033178) |
| 0.0 | 0.6 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.0 | 0.1 | GO:1990742 | microvesicle(GO:1990742) |
| 0.0 | 0.2 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 0.0 | 0.1 | GO:0070847 | core mediator complex(GO:0070847) |
| 0.0 | 0.1 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.0 | 0.1 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.0 | 0.3 | GO:0005845 | mRNA cap binding complex(GO:0005845) RNA cap binding complex(GO:0034518) |
| 0.0 | 0.1 | GO:0070381 | endosome to plasma membrane transport vesicle(GO:0070381) |
| 0.0 | 0.0 | GO:0031372 | UBC13-MMS2 complex(GO:0031372) |
| 0.0 | 0.1 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.0 | 0.4 | GO:0016514 | SWI/SNF complex(GO:0016514) |
| 0.0 | 0.1 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.0 | 0.3 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.0 | 0.1 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.0 | 0.3 | GO:0005770 | late endosome(GO:0005770) |
| 0.0 | 0.2 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.0 | 0.1 | GO:0043219 | lateral loop(GO:0043219) |
| 0.0 | 0.1 | GO:0030121 | AP-1 adaptor complex(GO:0030121) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.4 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.1 | 1.5 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 0.1 | 0.3 | GO:0047389 | glycerophosphocholine phosphodiesterase activity(GO:0047389) |
| 0.1 | 1.1 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.1 | 0.2 | GO:0017095 | heparan sulfate 6-O-sulfotransferase activity(GO:0017095) |
| 0.1 | 0.8 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.1 | 0.2 | GO:0052725 | inositol tetrakisphosphate 1-kinase activity(GO:0047325) inositol-1,3,4-trisphosphate 6-kinase activity(GO:0052725) inositol-1,3,4-trisphosphate 5-kinase activity(GO:0052726) inositol-1,3,4,5,6-pentakisphosphate 1-phosphatase activity(GO:0052825) inositol-1,3,4,6-tetrakisphosphate 6-phosphatase activity(GO:0052830) inositol-1,3,4,6-tetrakisphosphate 1-phosphatase activity(GO:0052831) inositol-3,4,6-trisphosphate 1-kinase activity(GO:0052835) |
| 0.1 | 0.6 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.1 | 0.4 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 0.1 | 0.7 | GO:0051378 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.1 | 0.4 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 0.1 | 0.2 | GO:0086062 | voltage-gated sodium channel activity involved in Purkinje myocyte action potential(GO:0086062) |
| 0.0 | 0.3 | GO:0035614 | snRNA stem-loop binding(GO:0035614) |
| 0.0 | 0.2 | GO:0050510 | N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase activity(GO:0050510) |
| 0.0 | 0.3 | GO:0004118 | cGMP-stimulated cyclic-nucleotide phosphodiesterase activity(GO:0004118) |
| 0.0 | 0.1 | GO:0015218 | pyrimidine nucleotide transmembrane transporter activity(GO:0015218) |
| 0.0 | 0.2 | GO:0000179 | rRNA (adenine-N6,N6-)-dimethyltransferase activity(GO:0000179) |
| 0.0 | 0.1 | GO:0015275 | stretch-activated, cation-selective, calcium channel activity(GO:0015275) |
| 0.0 | 0.1 | GO:0098808 | mRNA cap binding(GO:0098808) |
| 0.0 | 0.1 | GO:0047696 | beta-adrenergic receptor kinase activity(GO:0047696) |
| 0.0 | 0.3 | GO:0008525 | phosphatidylcholine transporter activity(GO:0008525) |
| 0.0 | 0.2 | GO:1904929 | coreceptor activity involved in Wnt signaling pathway, planar cell polarity pathway(GO:1904929) |
| 0.0 | 0.1 | GO:0070025 | cystathionine beta-synthase activity(GO:0004122) oxidoreductase activity, acting on other nitrogenous compounds as donors, cytochrome as acceptor(GO:0016662) nitrite reductase (NO-forming) activity(GO:0050421) carbon monoxide binding(GO:0070025) nitrite reductase activity(GO:0098809) |
| 0.0 | 0.4 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 0.1 | GO:0050509 | N-acetylglucosaminyl-proteoglycan 4-beta-glucuronosyltransferase activity(GO:0050509) |
| 0.0 | 0.2 | GO:0001225 | RNA polymerase II transcription coactivator binding(GO:0001225) |
| 0.0 | 0.4 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.0 | 0.1 | GO:0016520 | growth hormone-releasing hormone receptor activity(GO:0016520) |
| 0.0 | 0.1 | GO:0004832 | valine-tRNA ligase activity(GO:0004832) |
| 0.0 | 0.1 | GO:0008410 | CoA-transferase activity(GO:0008410) |
| 0.0 | 0.2 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.0 | 0.6 | GO:0005328 | neurotransmitter:sodium symporter activity(GO:0005328) |
| 0.0 | 0.3 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.0 | 0.1 | GO:0016534 | cyclin-dependent protein kinase 5 activator activity(GO:0016534) |
| 0.0 | 0.1 | GO:0015222 | serotonin transmembrane transporter activity(GO:0015222) |
| 0.0 | 0.1 | GO:0003923 | GPI-anchor transamidase activity(GO:0003923) |
| 0.0 | 0.3 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.0 | 0.4 | GO:0044548 | S100 protein binding(GO:0044548) |
| 0.0 | 0.2 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.0 | 0.2 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.0 | 0.0 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 0.0 | 0.1 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.0 | 0.6 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.0 | 0.1 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
| 0.0 | 0.1 | GO:0017002 | activin-activated receptor activity(GO:0017002) |
| 0.0 | 0.1 | GO:0005087 | Ran guanyl-nucleotide exchange factor activity(GO:0005087) |
| 0.0 | 0.0 | GO:0030272 | 5-formyltetrahydrofolate cyclo-ligase activity(GO:0030272) |
| 0.0 | 0.2 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.0 | 0.2 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.0 | 0.2 | GO:0047134 | protein-disulfide reductase activity(GO:0047134) |
| 0.0 | 0.2 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.0 | 0.2 | GO:0010314 | phosphatidylinositol-5-phosphate binding(GO:0010314) |
| 0.0 | 0.1 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.0 | 0.1 | GO:0097016 | L27 domain binding(GO:0097016) |
| 0.0 | 0.1 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.0 | 0.2 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.0 | 0.3 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.0 | 0.1 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 0.0 | 1.7 | GO:0019955 | cytokine binding(GO:0019955) |
| 0.0 | 0.1 | GO:0004887 | thyroid hormone receptor activity(GO:0004887) |
| 0.0 | 0.0 | GO:0034046 | poly(G) binding(GO:0034046) |
| 0.0 | 0.3 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.1 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.0 | 1.7 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 0.8 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 0.7 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.5 | REACTOME FGFR2C LIGAND BINDING AND ACTIVATION | Genes involved in FGFR2c ligand binding and activation |
| 0.0 | 0.7 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.0 | 1.6 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.0 | 0.4 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.0 | 0.3 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 0.0 | 0.4 | REACTOME P38MAPK EVENTS | Genes involved in p38MAPK events |
| 0.0 | 0.2 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.0 | 0.6 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.0 | 1.7 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 0.2 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.0 | 0.9 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 0.1 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 0.0 | 0.2 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.0 | 0.2 | REACTOME IRAK2 MEDIATED ACTIVATION OF TAK1 COMPLEX UPON TLR7 8 OR 9 STIMULATION | Genes involved in IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation |
| 0.0 | 0.1 | REACTOME RAF MAP KINASE CASCADE | Genes involved in RAF/MAP kinase cascade |
| 0.0 | 0.4 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 0.1 | REACTOME RECEPTOR LIGAND BINDING INITIATES THE SECOND PROTEOLYTIC CLEAVAGE OF NOTCH RECEPTOR | Genes involved in Receptor-ligand binding initiates the second proteolytic cleavage of Notch receptor |
| 0.0 | 0.4 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |