Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
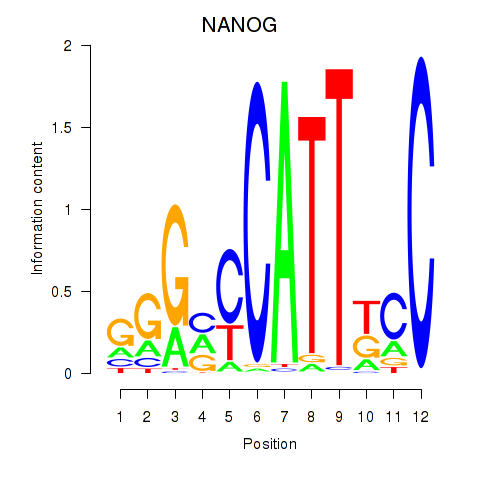
Results for NANOG
Z-value: 0.86
Transcription factors associated with NANOG
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NANOG
|
ENSG00000111704.6 | NANOG |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NANOG | hg19_v2_chr12_+_7941989_7942014 | 0.23 | 5.8e-01 | Click! |
Activity profile of NANOG motif
Sorted Z-values of NANOG motif
Network of associatons between targets according to the STRING database.
First level regulatory network of NANOG
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_+_156693091 | 0.90 |
ENST00000318218.6 ENST00000442283.2 ENST00000522463.1 ENST00000521420.1 |
CYFIP2 |
cytoplasmic FMR1 interacting protein 2 |
| chr5_+_156693159 | 0.90 |
ENST00000347377.6 |
CYFIP2 |
cytoplasmic FMR1 interacting protein 2 |
| chrX_-_107019181 | 0.47 |
ENST00000315660.4 ENST00000372384.2 ENST00000502650.1 ENST00000506724.1 |
TSC22D3 |
TSC22 domain family, member 3 |
| chr10_-_49812997 | 0.42 |
ENST00000417912.2 |
ARHGAP22 |
Rho GTPase activating protein 22 |
| chr10_+_64133934 | 0.36 |
ENST00000395254.3 ENST00000395255.3 ENST00000410046.3 |
ZNF365 |
zinc finger protein 365 |
| chr8_-_27469196 | 0.35 |
ENST00000546343.1 ENST00000560566.1 |
CLU |
clusterin |
| chr10_-_49813090 | 0.34 |
ENST00000249601.4 |
ARHGAP22 |
Rho GTPase activating protein 22 |
| chrX_-_118827333 | 0.33 |
ENST00000360156.7 ENST00000354228.4 ENST00000489216.1 ENST00000354416.3 ENST00000394610.1 ENST00000343984.5 |
SEPT6 |
septin 6 |
| chrX_+_85403487 | 0.31 |
ENST00000373125.4 |
DACH2 |
dachshund homolog 2 (Drosophila) |
| chr1_+_210406121 | 0.30 |
ENST00000367012.3 |
SERTAD4 |
SERTA domain containing 4 |
| chr8_-_27468945 | 0.29 |
ENST00000405140.3 |
CLU |
clusterin |
| chrX_+_85403445 | 0.28 |
ENST00000373131.1 |
DACH2 |
dachshund homolog 2 (Drosophila) |
| chr15_-_78526855 | 0.27 |
ENST00000541759.1 ENST00000558130.1 |
ACSBG1 |
acyl-CoA synthetase bubblegum family member 1 |
| chr6_+_155537771 | 0.27 |
ENST00000275246.7 |
TIAM2 |
T-cell lymphoma invasion and metastasis 2 |
| chr14_+_21156915 | 0.26 |
ENST00000397990.4 ENST00000555597.1 |
ANG RNASE4 |
angiogenin, ribonuclease, RNase A family, 5 ribonuclease, RNase A family, 4 |
| chr3_-_93692781 | 0.26 |
ENST00000394236.3 |
PROS1 |
protein S (alpha) |
| chr8_-_27468842 | 0.26 |
ENST00000523500.1 |
CLU |
clusterin |
| chr5_+_138089100 | 0.25 |
ENST00000520339.1 ENST00000355078.5 ENST00000302763.7 ENST00000518910.1 |
CTNNA1 |
catenin (cadherin-associated protein), alpha 1, 102kDa |
| chr6_+_30524663 | 0.25 |
ENST00000376560.3 |
PRR3 |
proline rich 3 |
| chr22_+_31090793 | 0.25 |
ENST00000332585.6 ENST00000382310.3 ENST00000446658.2 |
OSBP2 |
oxysterol binding protein 2 |
| chr6_+_151561085 | 0.24 |
ENST00000402676.2 |
AKAP12 |
A kinase (PRKA) anchor protein 12 |
| chr19_+_926000 | 0.24 |
ENST00000263620.3 |
ARID3A |
AT rich interactive domain 3A (BRIGHT-like) |
| chr3_-_49851313 | 0.24 |
ENST00000333486.3 |
UBA7 |
ubiquitin-like modifier activating enzyme 7 |
| chr6_+_30525051 | 0.23 |
ENST00000376557.3 |
PRR3 |
proline rich 3 |
| chr12_-_31743901 | 0.22 |
ENST00000354285.4 |
DENND5B |
DENN/MADD domain containing 5B |
| chr10_-_70092671 | 0.21 |
ENST00000358769.2 ENST00000432941.1 ENST00000495025.2 |
PBLD |
phenazine biosynthesis-like protein domain containing |
| chr9_+_124030338 | 0.21 |
ENST00000449773.1 ENST00000432226.1 ENST00000436847.1 ENST00000394353.2 ENST00000449733.1 ENST00000412819.1 ENST00000341272.2 ENST00000373808.2 |
GSN |
gelsolin |
| chrX_+_103173457 | 0.20 |
ENST00000419165.1 |
TMSB15B |
thymosin beta 15B |
| chrX_-_118827113 | 0.20 |
ENST00000394617.2 |
SEPT6 |
septin 6 |
| chr16_+_68573640 | 0.18 |
ENST00000398253.2 ENST00000573161.1 |
ZFP90 |
ZFP90 zinc finger protein |
| chr10_-_70092635 | 0.17 |
ENST00000309049.4 |
PBLD |
phenazine biosynthesis-like protein domain containing |
| chr15_+_41549105 | 0.17 |
ENST00000560965.1 |
CHP1 |
calcineurin-like EF-hand protein 1 |
| chr12_-_53901266 | 0.16 |
ENST00000609999.1 ENST00000267017.3 |
NPFF |
neuropeptide FF-amide peptide precursor |
| chr4_+_154074217 | 0.15 |
ENST00000437508.2 |
TRIM2 |
tripartite motif containing 2 |
| chr14_-_92198403 | 0.15 |
ENST00000553329.1 ENST00000256343.3 |
CATSPERB |
catsper channel auxiliary subunit beta |
| chr6_-_43021437 | 0.14 |
ENST00000265348.3 |
CUL7 |
cullin 7 |
| chr1_-_47655686 | 0.14 |
ENST00000294338.2 |
PDZK1IP1 |
PDZK1 interacting protein 1 |
| chr22_+_37447771 | 0.14 |
ENST00000402077.3 ENST00000403888.3 ENST00000456470.1 |
KCTD17 |
potassium channel tetramerization domain containing 17 |
| chr16_+_31494323 | 0.13 |
ENST00000569576.1 ENST00000330498.3 |
SLC5A2 |
solute carrier family 5 (sodium/glucose cotransporter), member 2 |
| chr6_-_43021612 | 0.13 |
ENST00000535468.1 |
CUL7 |
cullin 7 |
| chrX_+_47092314 | 0.13 |
ENST00000218348.3 |
USP11 |
ubiquitin specific peptidase 11 |
| chr6_-_31632962 | 0.12 |
ENST00000456540.1 ENST00000445768.1 |
GPANK1 |
G patch domain and ankyrin repeats 1 |
| chr20_-_21494654 | 0.12 |
ENST00000377142.4 |
NKX2-2 |
NK2 homeobox 2 |
| chr7_+_134576317 | 0.12 |
ENST00000424922.1 ENST00000495522.1 |
CALD1 |
caldesmon 1 |
| chr6_+_34204642 | 0.12 |
ENST00000347617.6 ENST00000401473.3 ENST00000311487.5 ENST00000447654.1 ENST00000395004.3 |
HMGA1 |
high mobility group AT-hook 1 |
| chr16_+_68573116 | 0.11 |
ENST00000570495.1 ENST00000563169.2 ENST00000564323.1 ENST00000562156.1 ENST00000573685.1 |
ZFP90 |
ZFP90 zinc finger protein |
| chr7_-_2883928 | 0.11 |
ENST00000275364.3 |
GNA12 |
guanine nucleotide binding protein (G protein) alpha 12 |
| chr19_-_42806919 | 0.11 |
ENST00000595530.1 ENST00000538771.1 ENST00000601865.1 |
PAFAH1B3 |
platelet-activating factor acetylhydrolase 1b, catalytic subunit 3 (29kDa) |
| chr19_-_42806723 | 0.11 |
ENST00000262890.3 |
PAFAH1B3 |
platelet-activating factor acetylhydrolase 1b, catalytic subunit 3 (29kDa) |
| chr1_+_27719148 | 0.10 |
ENST00000374024.3 |
GPR3 |
G protein-coupled receptor 3 |
| chr4_-_5891918 | 0.10 |
ENST00000512574.1 |
CRMP1 |
collapsin response mediator protein 1 |
| chr6_+_52226897 | 0.10 |
ENST00000442253.2 |
PAQR8 |
progestin and adipoQ receptor family member VIII |
| chr8_+_22022800 | 0.10 |
ENST00000397814.3 |
BMP1 |
bone morphogenetic protein 1 |
| chr2_+_45878790 | 0.10 |
ENST00000306156.3 |
PRKCE |
protein kinase C, epsilon |
| chr19_-_42806842 | 0.09 |
ENST00000596265.1 |
PAFAH1B3 |
platelet-activating factor acetylhydrolase 1b, catalytic subunit 3 (29kDa) |
| chr6_-_131277510 | 0.09 |
ENST00000525193.1 ENST00000527659.1 |
EPB41L2 |
erythrocyte membrane protein band 4.1-like 2 |
| chr8_-_42751820 | 0.09 |
ENST00000526349.1 ENST00000527424.1 ENST00000534961.1 ENST00000319073.4 |
RNF170 |
ring finger protein 170 |
| chr13_+_108921977 | 0.09 |
ENST00000430559.1 ENST00000375887.4 |
TNFSF13B |
tumor necrosis factor (ligand) superfamily, member 13b |
| chr5_-_55290773 | 0.09 |
ENST00000502326.3 ENST00000381298.2 |
IL6ST |
interleukin 6 signal transducer (gp130, oncostatin M receptor) |
| chr10_+_102891048 | 0.08 |
ENST00000467928.2 |
TLX1 |
T-cell leukemia homeobox 1 |
| chr7_-_28220354 | 0.08 |
ENST00000283928.5 |
JAZF1 |
JAZF zinc finger 1 |
| chr3_-_52567792 | 0.08 |
ENST00000307092.4 ENST00000422318.2 ENST00000459839.1 |
NT5DC2 |
5'-nucleotidase domain containing 2 |
| chr6_+_31540056 | 0.08 |
ENST00000418386.2 |
LTA |
lymphotoxin alpha |
| chrX_+_21392553 | 0.07 |
ENST00000279451.4 |
CNKSR2 |
connector enhancer of kinase suppressor of Ras 2 |
| chr15_+_44580955 | 0.07 |
ENST00000345795.2 ENST00000360824.3 |
CASC4 |
cancer susceptibility candidate 4 |
| chr17_+_7905912 | 0.07 |
ENST00000254854.4 |
GUCY2D |
guanylate cyclase 2D, membrane (retina-specific) |
| chr2_-_69664586 | 0.07 |
ENST00000303698.3 ENST00000394305.1 ENST00000410022.2 |
NFU1 |
NFU1 iron-sulfur cluster scaffold homolog (S. cerevisiae) |
| chr17_-_56065484 | 0.07 |
ENST00000581208.1 |
VEZF1 |
vascular endothelial zinc finger 1 |
| chrX_+_44703249 | 0.07 |
ENST00000339042.4 |
DUSP21 |
dual specificity phosphatase 21 |
| chr4_+_154073469 | 0.06 |
ENST00000441616.1 |
TRIM2 |
tripartite motif containing 2 |
| chr6_-_33714667 | 0.06 |
ENST00000293756.4 |
IP6K3 |
inositol hexakisphosphate kinase 3 |
| chr2_-_198650037 | 0.06 |
ENST00000392296.4 |
BOLL |
boule-like RNA-binding protein |
| chr1_-_202129704 | 0.06 |
ENST00000476061.1 ENST00000544762.1 ENST00000467283.1 ENST00000464870.1 ENST00000435759.2 ENST00000486116.1 ENST00000543735.1 ENST00000308986.5 ENST00000477625.1 |
PTPN7 |
protein tyrosine phosphatase, non-receptor type 7 |
| chr10_-_27443155 | 0.06 |
ENST00000427324.1 ENST00000326799.3 |
YME1L1 |
YME1-like 1 ATPase |
| chr12_-_52946923 | 0.06 |
ENST00000267119.5 |
KRT71 |
keratin 71 |
| chr1_+_161632937 | 0.06 |
ENST00000236937.9 ENST00000367961.4 ENST00000358671.5 |
FCGR2B |
Fc fragment of IgG, low affinity IIb, receptor (CD32) |
| chr15_+_78833071 | 0.06 |
ENST00000559365.1 |
PSMA4 |
proteasome (prosome, macropain) subunit, alpha type, 4 |
| chr6_+_31633011 | 0.06 |
ENST00000375885.4 |
CSNK2B |
casein kinase 2, beta polypeptide |
| chr3_+_75955817 | 0.06 |
ENST00000487694.3 ENST00000602589.1 |
ROBO2 |
roundabout, axon guidance receptor, homolog 2 (Drosophila) |
| chr15_+_44580899 | 0.06 |
ENST00000559222.1 ENST00000299957.6 |
CASC4 |
cancer susceptibility candidate 4 |
| chr18_-_23670546 | 0.06 |
ENST00000542743.1 ENST00000545952.1 ENST00000539849.1 ENST00000415083.2 |
SS18 |
synovial sarcoma translocation, chromosome 18 |
| chr15_-_50978965 | 0.06 |
ENST00000560955.1 ENST00000313478.7 |
TRPM7 |
transient receptor potential cation channel, subfamily M, member 7 |
| chr20_+_36974759 | 0.06 |
ENST00000217407.2 |
LBP |
lipopolysaccharide binding protein |
| chr1_+_35734562 | 0.06 |
ENST00000314607.6 ENST00000373297.2 |
ZMYM4 |
zinc finger, MYM-type 4 |
| chr1_+_145575980 | 0.06 |
ENST00000393045.2 |
PIAS3 |
protein inhibitor of activated STAT, 3 |
| chr1_+_145576007 | 0.05 |
ENST00000369298.1 |
PIAS3 |
protein inhibitor of activated STAT, 3 |
| chr5_+_139493665 | 0.05 |
ENST00000331327.3 |
PURA |
purine-rich element binding protein A |
| chr6_-_136610911 | 0.05 |
ENST00000530767.1 ENST00000527759.1 ENST00000527536.1 ENST00000529826.1 ENST00000531224.1 ENST00000353331.4 |
BCLAF1 |
BCL2-associated transcription factor 1 |
| chr14_-_98444461 | 0.05 |
ENST00000499006.2 |
C14orf64 |
chromosome 14 open reading frame 64 |
| chr1_+_161551101 | 0.05 |
ENST00000367962.4 ENST00000367960.5 ENST00000403078.3 ENST00000428605.2 |
FCGR2B |
Fc fragment of IgG, low affinity IIb, receptor (CD32) |
| chr19_+_6372444 | 0.05 |
ENST00000245812.3 |
ALKBH7 |
alkB, alkylation repair homolog 7 (E. coli) |
| chr5_+_68513557 | 0.05 |
ENST00000256441.4 |
MRPS36 |
mitochondrial ribosomal protein S36 |
| chr6_+_292051 | 0.05 |
ENST00000344450.5 |
DUSP22 |
dual specificity phosphatase 22 |
| chr18_-_23671139 | 0.05 |
ENST00000579061.1 ENST00000542420.2 |
SS18 |
synovial sarcoma translocation, chromosome 18 |
| chr6_-_119031228 | 0.05 |
ENST00000392500.3 ENST00000368488.5 ENST00000434604.1 |
CEP85L |
centrosomal protein 85kDa-like |
| chr3_-_149093499 | 0.05 |
ENST00000472441.1 |
TM4SF1 |
transmembrane 4 L six family member 1 |
| chr3_+_128968437 | 0.05 |
ENST00000314797.6 |
COPG1 |
coatomer protein complex, subunit gamma 1 |
| chr20_-_61557821 | 0.05 |
ENST00000354665.4 ENST00000370368.1 ENST00000395343.1 ENST00000395340.1 |
DIDO1 |
death inducer-obliterator 1 |
| chr6_-_33714752 | 0.05 |
ENST00000451316.1 |
IP6K3 |
inositol hexakisphosphate kinase 3 |
| chr14_-_98444386 | 0.05 |
ENST00000556462.1 ENST00000556138.1 |
C14orf64 |
chromosome 14 open reading frame 64 |
| chr8_+_9413410 | 0.04 |
ENST00000520408.1 ENST00000310430.6 ENST00000522110.1 |
TNKS |
tankyrase, TRF1-interacting ankyrin-related ADP-ribose polymerase |
| chr1_+_173837488 | 0.04 |
ENST00000427304.1 ENST00000432989.1 ENST00000367702.1 |
ZBTB37 |
zinc finger and BTB domain containing 37 |
| chr17_+_43224684 | 0.04 |
ENST00000332499.2 |
HEXIM1 |
hexamethylene bis-acetamide inducible 1 |
| chr1_-_151254362 | 0.04 |
ENST00000447795.2 |
RP11-126K1.2 |
Uncharacterized protein |
| chr9_+_71819927 | 0.04 |
ENST00000535702.1 |
TJP2 |
tight junction protein 2 |
| chr2_+_170590321 | 0.04 |
ENST00000392647.2 |
KLHL23 |
kelch-like family member 23 |
| chr17_-_43128943 | 0.04 |
ENST00000588499.1 ENST00000593094.1 |
DCAKD |
dephospho-CoA kinase domain containing |
| chr16_+_30759700 | 0.04 |
ENST00000328273.7 |
PHKG2 |
phosphorylase kinase, gamma 2 (testis) |
| chr4_-_16077741 | 0.04 |
ENST00000447510.2 ENST00000540805.1 ENST00000539194.1 |
PROM1 |
prominin 1 |
| chr3_+_107244229 | 0.04 |
ENST00000456419.1 ENST00000402163.2 |
BBX |
bobby sox homolog (Drosophila) |
| chr1_+_159409512 | 0.04 |
ENST00000423932.3 |
OR10J1 |
olfactory receptor, family 10, subfamily J, member 1 |
| chr10_+_69644404 | 0.04 |
ENST00000212015.6 |
SIRT1 |
sirtuin 1 |
| chr1_-_202129105 | 0.04 |
ENST00000367279.4 |
PTPN7 |
protein tyrosine phosphatase, non-receptor type 7 |
| chr2_-_214016314 | 0.04 |
ENST00000434687.1 ENST00000374319.4 |
IKZF2 |
IKAROS family zinc finger 2 (Helios) |
| chr22_-_32026810 | 0.04 |
ENST00000266095.5 ENST00000397500.1 |
PISD |
phosphatidylserine decarboxylase |
| chr19_-_39926268 | 0.04 |
ENST00000599705.1 |
RPS16 |
ribosomal protein S16 |
| chr6_-_30524951 | 0.04 |
ENST00000376621.3 |
GNL1 |
guanine nucleotide binding protein-like 1 |
| chr19_-_42806444 | 0.04 |
ENST00000594989.1 |
PAFAH1B3 |
platelet-activating factor acetylhydrolase 1b, catalytic subunit 3 (29kDa) |
| chr13_-_103389159 | 0.03 |
ENST00000322527.2 |
CCDC168 |
coiled-coil domain containing 168 |
| chr19_+_42381173 | 0.03 |
ENST00000221972.3 |
CD79A |
CD79a molecule, immunoglobulin-associated alpha |
| chr6_+_292459 | 0.03 |
ENST00000419235.2 ENST00000605035.1 ENST00000605863.1 |
DUSP22 |
dual specificity phosphatase 22 |
| chr19_-_39926555 | 0.03 |
ENST00000599539.1 ENST00000339471.4 ENST00000601655.1 ENST00000251453.3 |
RPS16 |
ribosomal protein S16 |
| chr9_+_71820057 | 0.03 |
ENST00000539225.1 |
TJP2 |
tight junction protein 2 |
| chr18_+_3262415 | 0.03 |
ENST00000581193.1 ENST00000400175.5 |
MYL12B |
myosin, light chain 12B, regulatory |
| chrX_-_133931164 | 0.03 |
ENST00000370790.1 ENST00000298090.6 |
FAM122B |
family with sequence similarity 122B |
| chr6_+_88032299 | 0.03 |
ENST00000608353.1 ENST00000392863.1 ENST00000229570.5 ENST00000608525.1 ENST00000608868.1 |
SMIM8 |
small integral membrane protein 8 |
| chr1_-_145589424 | 0.03 |
ENST00000334513.5 |
NUDT17 |
nudix (nucleoside diphosphate linked moiety X)-type motif 17 |
| chr15_+_78832747 | 0.03 |
ENST00000560217.1 ENST00000044462.7 ENST00000559082.1 ENST00000559948.1 ENST00000413382.2 ENST00000559146.1 ENST00000558281.1 |
PSMA4 |
proteasome (prosome, macropain) subunit, alpha type, 4 |
| chr3_-_15469006 | 0.03 |
ENST00000443029.1 ENST00000383790.3 ENST00000383789.5 |
METTL6 |
methyltransferase like 6 |
| chr1_+_159272111 | 0.03 |
ENST00000368114.1 |
FCER1A |
Fc fragment of IgE, high affinity I, receptor for; alpha polypeptide |
| chrX_+_47082408 | 0.03 |
ENST00000518022.1 ENST00000276052.6 |
CDK16 |
cyclin-dependent kinase 16 |
| chr11_-_67205538 | 0.03 |
ENST00000326294.3 |
PTPRCAP |
protein tyrosine phosphatase, receptor type, C-associated protein |
| chr10_-_1034237 | 0.02 |
ENST00000381466.1 |
AL359878.1 |
Uncharacterized protein |
| chr3_-_15469045 | 0.02 |
ENST00000450816.2 |
METTL6 |
methyltransferase like 6 |
| chr19_-_39342962 | 0.02 |
ENST00000600873.1 |
HNRNPL |
heterogeneous nuclear ribonucleoprotein L |
| chr7_+_131012605 | 0.02 |
ENST00000446815.1 ENST00000352689.6 |
MKLN1 |
muskelin 1, intracellular mediator containing kelch motifs |
| chr3_+_32147997 | 0.02 |
ENST00000282541.5 |
GPD1L |
glycerol-3-phosphate dehydrogenase 1-like |
| chr7_-_76829125 | 0.02 |
ENST00000248598.5 |
FGL2 |
fibrinogen-like 2 |
| chr16_-_2264779 | 0.02 |
ENST00000333503.7 |
PGP |
phosphoglycolate phosphatase |
| chr17_-_55911970 | 0.02 |
ENST00000581805.1 ENST00000580960.1 |
RP11-60A24.3 |
RP11-60A24.3 |
| chr2_+_87769459 | 0.02 |
ENST00000414030.1 ENST00000437561.1 |
LINC00152 |
long intergenic non-protein coding RNA 152 |
| chr18_+_3262954 | 0.02 |
ENST00000584539.1 |
MYL12B |
myosin, light chain 12B, regulatory |
| chr14_-_98444438 | 0.02 |
ENST00000512901.2 |
C14orf64 |
chromosome 14 open reading frame 64 |
| chr19_-_13213662 | 0.02 |
ENST00000264824.4 |
LYL1 |
lymphoblastic leukemia derived sequence 1 |
| chr8_+_42752053 | 0.02 |
ENST00000307602.4 |
HOOK3 |
hook microtubule-tethering protein 3 |
| chr7_-_102715263 | 0.02 |
ENST00000379305.3 |
FBXL13 |
F-box and leucine-rich repeat protein 13 |
| chr10_-_79686284 | 0.02 |
ENST00000372391.2 ENST00000372388.2 |
DLG5 |
discs, large homolog 5 (Drosophila) |
| chr5_-_169816638 | 0.02 |
ENST00000521859.1 ENST00000274629.4 |
KCNMB1 |
potassium large conductance calcium-activated channel, subfamily M, beta member 1 |
| chr1_+_20465805 | 0.02 |
ENST00000375102.3 |
PLA2G2F |
phospholipase A2, group IIF |
| chr7_-_99277610 | 0.02 |
ENST00000343703.5 ENST00000222982.4 ENST00000439761.1 ENST00000339843.2 |
CYP3A5 |
cytochrome P450, family 3, subfamily A, polypeptide 5 |
| chr11_-_14379997 | 0.02 |
ENST00000526063.1 ENST00000532814.1 |
RRAS2 |
related RAS viral (r-ras) oncogene homolog 2 |
| chr20_-_3762087 | 0.01 |
ENST00000379756.3 |
SPEF1 |
sperm flagellar 1 |
| chr17_+_75372165 | 0.01 |
ENST00000427674.2 |
SEPT9 |
septin 9 |
| chr2_-_97405775 | 0.01 |
ENST00000264963.4 ENST00000537039.1 ENST00000377079.4 ENST00000426463.2 ENST00000534882.1 |
LMAN2L |
lectin, mannose-binding 2-like |
| chr4_+_3344141 | 0.01 |
ENST00000306648.7 |
RGS12 |
regulator of G-protein signaling 12 |
| chr9_-_99381660 | 0.01 |
ENST00000375240.3 ENST00000463569.1 |
CDC14B |
cell division cycle 14B |
| chr2_-_175113088 | 0.01 |
ENST00000409546.1 ENST00000428402.2 |
OLA1 |
Obg-like ATPase 1 |
| chr15_+_78833105 | 0.01 |
ENST00000558341.1 ENST00000559437.1 |
PSMA4 |
proteasome (prosome, macropain) subunit, alpha type, 4 |
| chr14_-_51297837 | 0.01 |
ENST00000245441.5 ENST00000389868.3 ENST00000382041.3 ENST00000324330.9 ENST00000453196.1 ENST00000453401.2 |
NIN |
ninein (GSK3B interacting protein) |
| chr20_+_43343517 | 0.01 |
ENST00000372865.4 |
WISP2 |
WNT1 inducible signaling pathway protein 2 |
| chr2_-_112237835 | 0.01 |
ENST00000442293.1 ENST00000439494.1 |
MIR4435-1HG |
MIR4435-1 host gene (non-protein coding) |
| chr20_+_43343476 | 0.01 |
ENST00000372868.2 |
WISP2 |
WNT1 inducible signaling pathway protein 2 |
| chr6_-_31633402 | 0.01 |
ENST00000375893.2 |
GPANK1 |
G patch domain and ankyrin repeats 1 |
| chr19_+_4969116 | 0.01 |
ENST00000588337.1 ENST00000159111.4 ENST00000381759.4 |
KDM4B |
lysine (K)-specific demethylase 4B |
| chr16_+_6069072 | 0.01 |
ENST00000547605.1 ENST00000550418.1 ENST00000553186.1 |
RBFOX1 |
RNA binding protein, fox-1 homolog (C. elegans) 1 |
| chr15_-_78526942 | 0.01 |
ENST00000258873.4 |
ACSBG1 |
acyl-CoA synthetase bubblegum family member 1 |
| chr8_-_102181718 | 0.01 |
ENST00000565617.1 |
KB-1460A1.5 |
KB-1460A1.5 |
| chr3_-_134369853 | 0.01 |
ENST00000508956.1 ENST00000503669.1 ENST00000423778.2 |
KY |
kyphoscoliosis peptidase |
| chr5_-_138739739 | 0.01 |
ENST00000514983.1 ENST00000507779.2 ENST00000451821.2 ENST00000450845.2 ENST00000509959.1 ENST00000302091.5 |
SPATA24 |
spermatogenesis associated 24 |
| chr19_-_54327542 | 0.01 |
ENST00000391775.3 ENST00000324134.6 ENST00000535162.1 ENST00000351894.4 ENST00000354278.3 ENST00000391773.1 ENST00000345770.5 ENST00000391772.1 |
NLRP12 |
NLR family, pyrin domain containing 12 |
| chr1_+_203096831 | 0.00 |
ENST00000337894.4 |
ADORA1 |
adenosine A1 receptor |
| chr16_+_3508063 | 0.00 |
ENST00000576787.1 ENST00000572942.1 ENST00000576916.1 ENST00000575076.1 ENST00000572131.1 |
NAA60 |
N(alpha)-acetyltransferase 60, NatF catalytic subunit |
| chr21_-_34960930 | 0.00 |
ENST00000437395.1 |
DONSON |
downstream neighbor of SON |
| chr12_-_51477333 | 0.00 |
ENST00000228515.1 ENST00000548206.1 ENST00000546935.1 ENST00000548981.1 |
CSRNP2 |
cysteine-serine-rich nuclear protein 2 |
| chr17_-_34417479 | 0.00 |
ENST00000225245.5 |
CCL3 |
chemokine (C-C motif) ligand 3 |
| chr6_-_43484718 | 0.00 |
ENST00000372422.2 |
YIPF3 |
Yip1 domain family, member 3 |
| chr16_+_640055 | 0.00 |
ENST00000568586.1 ENST00000538492.1 ENST00000248139.3 |
RAB40C |
RAB40C, member RAS oncogene family |
| chr19_+_37825614 | 0.00 |
ENST00000591259.1 ENST00000590582.1 |
HKR1 |
HKR1, GLI-Kruppel zinc finger family member |
| chr6_+_107077471 | 0.00 |
ENST00000369044.1 |
QRSL1 |
glutaminyl-tRNA synthase (glutamine-hydrolyzing)-like 1 |
| chrX_-_120119638 | 0.00 |
ENST00000434414.1 ENST00000433030.1 ENST00000371280.3 |
CT47A1 |
cancer/testis antigen family 47, member A1 |
| chr22_-_30662828 | 0.00 |
ENST00000403463.1 ENST00000215781.2 |
OSM |
oncostatin M |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.8 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.1 | 0.9 | GO:1902998 | macrophage proliferation(GO:0061517) microglial cell proliferation(GO:0061518) regulation of neuronal signal transduction(GO:1902847) positive regulation of neurofibrillary tangle assembly(GO:1902998) |
| 0.1 | 0.4 | GO:0060392 | negative regulation of SMAD protein import into nucleus(GO:0060392) |
| 0.1 | 0.3 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.1 | 0.5 | GO:0070236 | negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.0 | 0.3 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 0.0 | 0.2 | GO:0044856 | plasma membrane raft distribution(GO:0044855) plasma membrane raft localization(GO:0044856) plasma membrane raft polarization(GO:0044858) regulation of plasma membrane raft polarization(GO:1903906) |
| 0.0 | 0.1 | GO:0021529 | spinal cord oligodendrocyte cell differentiation(GO:0021529) spinal cord oligodendrocyte cell fate specification(GO:0021530) |
| 0.0 | 0.2 | GO:0070253 | somatostatin secretion(GO:0070253) |
| 0.0 | 0.1 | GO:0043317 | negative regulation of dendritic cell antigen processing and presentation(GO:0002605) regulation of cytotoxic T cell degranulation(GO:0043317) negative regulation of cytotoxic T cell degranulation(GO:0043318) |
| 0.0 | 0.1 | GO:1904530 | negative regulation of actin filament binding(GO:1904530) negative regulation of actin binding(GO:1904617) |
| 0.0 | 0.1 | GO:0035668 | TRAM-dependent toll-like receptor signaling pathway(GO:0035668) TRAM-dependent toll-like receptor 4 signaling pathway(GO:0035669) |
| 0.0 | 0.1 | GO:0002874 | regulation of chronic inflammatory response to antigenic stimulus(GO:0002874) |
| 0.0 | 0.1 | GO:0090402 | oncogene-induced cell senescence(GO:0090402) |
| 0.0 | 0.1 | GO:0031296 | positive regulation of germinal center formation(GO:0002636) B cell costimulation(GO:0031296) |
| 0.0 | 0.2 | GO:0070885 | positive regulation of sodium:proton antiporter activity(GO:0032417) negative regulation of calcineurin-NFAT signaling cascade(GO:0070885) |
| 0.0 | 0.0 | GO:2000481 | positive regulation of cAMP-dependent protein kinase activity(GO:2000481) |
| 0.0 | 0.1 | GO:0016340 | calcium-dependent cell-matrix adhesion(GO:0016340) |
| 0.0 | 0.1 | GO:0015920 | lipopolysaccharide transport(GO:0015920) |
| 0.0 | 0.1 | GO:0048861 | leukemia inhibitory factor signaling pathway(GO:0048861) |
| 0.0 | 0.2 | GO:0010739 | positive regulation of protein kinase A signaling(GO:0010739) |
| 0.0 | 0.3 | GO:0018214 | peptidyl-glutamic acid carboxylation(GO:0017187) protein carboxylation(GO:0018214) |
| 0.0 | 0.1 | GO:0050925 | negative regulation of negative chemotaxis(GO:0050925) |
| 0.0 | 0.0 | GO:0070213 | negative regulation of sister chromatid cohesion(GO:0045875) protein auto-ADP-ribosylation(GO:0070213) |
| 0.0 | 0.0 | GO:2000698 | glomerular parietal epithelial cell differentiation(GO:0072139) positive regulation of epithelial cell differentiation involved in kidney development(GO:2000698) positive regulation of nephron tubule epithelial cell differentiation(GO:2000768) |
| 0.0 | 0.6 | GO:0051123 | RNA polymerase II transcriptional preinitiation complex assembly(GO:0051123) |
| 0.0 | 0.1 | GO:0033211 | adiponectin-activated signaling pathway(GO:0033211) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0005940 | septin ring(GO:0005940) septin collar(GO:0032173) |
| 0.1 | 0.9 | GO:0097418 | neurofibrillary tangle(GO:0097418) |
| 0.0 | 0.3 | GO:0032311 | angiogenin-PRI complex(GO:0032311) |
| 0.0 | 0.3 | GO:1990393 | 3M complex(GO:1990393) |
| 0.0 | 0.1 | GO:0005900 | oncostatin-M receptor complex(GO:0005900) |
| 0.0 | 0.3 | GO:0030478 | actin cap(GO:0030478) |
| 0.0 | 0.1 | GO:0036128 | CatSper complex(GO:0036128) |
| 0.0 | 0.3 | GO:0016342 | catenin complex(GO:0016342) |
| 0.0 | 0.2 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.0 | 0.4 | GO:0005815 | microtubule organizing center(GO:0005815) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0047179 | platelet-activating factor acetyltransferase activity(GO:0047179) |
| 0.1 | 0.5 | GO:0043426 | MRF binding(GO:0043426) |
| 0.0 | 0.9 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.0 | 0.2 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.0 | 0.3 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.0 | 0.6 | GO:0001075 | transcription factor activity, RNA polymerase II core promoter sequence-specific binding involved in preinitiation complex assembly(GO:0001075) |
| 0.0 | 0.1 | GO:0004915 | interleukin-6 receptor activity(GO:0004915) leukemia inhibitory factor receptor activity(GO:0004923) interleukin-6 binding(GO:0019981) |
| 0.0 | 0.1 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 0.0 | 0.1 | GO:0052836 | inositol 5-diphosphate pentakisphosphate 5-kinase activity(GO:0052836) inositol diphosphate tetrakisphosphate kinase activity(GO:0052839) |
| 0.0 | 0.1 | GO:0004699 | calcium-independent protein kinase C activity(GO:0004699) |
| 0.0 | 0.1 | GO:0031752 | D5 dopamine receptor binding(GO:0031752) |
| 0.0 | 0.1 | GO:0032422 | purine-rich negative regulatory element binding(GO:0032422) |
| 0.0 | 0.2 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.0 | 0.3 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.0 | 0.1 | GO:0005412 | glucose:sodium symporter activity(GO:0005412) |
| 0.0 | 0.2 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.0 | 0.3 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.0 | 0.0 | GO:0004609 | phosphatidylserine decarboxylase activity(GO:0004609) |
| 0.0 | 0.0 | GO:0004140 | dephospho-CoA kinase activity(GO:0004140) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 2.1 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.0 | 0.1 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |