Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
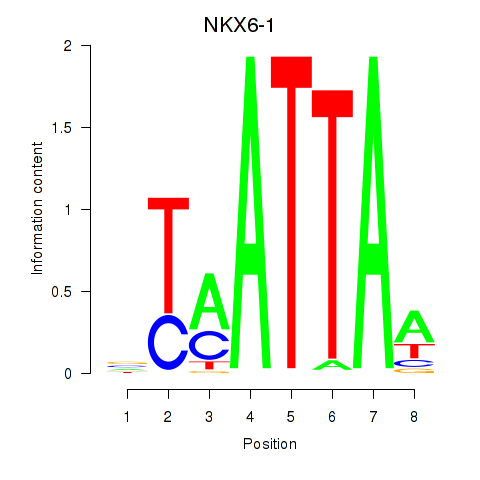
Results for NOTO_VSX2_DLX2_DLX6_NKX6-1
Z-value: 0.43
Transcription factors associated with NOTO_VSX2_DLX2_DLX6_NKX6-1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NOTO
|
ENSG00000214513.3 | NOTO |
|
VSX2
|
ENSG00000119614.2 | VSX2 |
|
DLX2
|
ENSG00000115844.6 | DLX2 |
|
DLX6
|
ENSG00000006377.9 | DLX6 |
|
NKX6-1
|
ENSG00000163623.5 | NKX6-1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NKX6-1 | hg19_v2_chr4_-_85418103_85418103 | 0.39 | 3.3e-01 | Click! |
| DLX2 | hg19_v2_chr2_-_172967621_172967637 | 0.34 | 4.1e-01 | Click! |
| DLX6 | hg19_v2_chr7_+_96634850_96634874, hg19_v2_chr7_+_96635277_96635301 | -0.20 | 6.3e-01 | Click! |
Activity profile of NOTO_VSX2_DLX2_DLX6_NKX6-1 motif
Sorted Z-values of NOTO_VSX2_DLX2_DLX6_NKX6-1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of NOTO_VSX2_DLX2_DLX6_NKX6-1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_-_28123206 | 0.71 |
ENST00000542963.1 ENST00000535992.1 |
PTHLH |
parathyroid hormone-like hormone |
| chr12_-_28122980 | 0.71 |
ENST00000395868.3 ENST00000534890.1 |
PTHLH |
parathyroid hormone-like hormone |
| chr5_+_66300446 | 0.64 |
ENST00000261569.7 |
MAST4 |
microtubule associated serine/threonine kinase family member 4 |
| chr1_+_62439037 | 0.47 |
ENST00000545929.1 |
INADL |
InaD-like (Drosophila) |
| chr4_-_74486217 | 0.46 |
ENST00000335049.5 ENST00000307439.5 |
RASSF6 |
Ras association (RalGDS/AF-6) domain family member 6 |
| chr4_-_25865159 | 0.44 |
ENST00000502949.1 ENST00000264868.5 ENST00000513691.1 ENST00000514872.1 |
SEL1L3 |
sel-1 suppressor of lin-12-like 3 (C. elegans) |
| chr3_-_151034734 | 0.43 |
ENST00000260843.4 |
GPR87 |
G protein-coupled receptor 87 |
| chr3_+_111718036 | 0.43 |
ENST00000455401.2 |
TAGLN3 |
transgelin 3 |
| chr3_+_111717600 | 0.41 |
ENST00000273368.4 |
TAGLN3 |
transgelin 3 |
| chr3_+_111717511 | 0.37 |
ENST00000478951.1 ENST00000393917.2 |
TAGLN3 |
transgelin 3 |
| chr5_-_1882858 | 0.37 |
ENST00000511126.1 ENST00000231357.2 |
IRX4 |
iroquois homeobox 4 |
| chr3_+_111718173 | 0.36 |
ENST00000494932.1 |
TAGLN3 |
transgelin 3 |
| chr13_-_46716969 | 0.36 |
ENST00000435666.2 |
LCP1 |
lymphocyte cytosolic protein 1 (L-plastin) |
| chr20_-_50418947 | 0.33 |
ENST00000371539.3 |
SALL4 |
spalt-like transcription factor 4 |
| chr19_-_51522955 | 0.29 |
ENST00000358789.3 |
KLK10 |
kallikrein-related peptidase 10 |
| chr12_+_41831485 | 0.28 |
ENST00000539469.2 ENST00000298919.7 |
PDZRN4 |
PDZ domain containing ring finger 4 |
| chr6_-_9933500 | 0.27 |
ENST00000492169.1 |
OFCC1 |
orofacial cleft 1 candidate 1 |
| chr20_-_50418972 | 0.27 |
ENST00000395997.3 |
SALL4 |
spalt-like transcription factor 4 |
| chr11_-_124190184 | 0.27 |
ENST00000357438.2 |
OR8D2 |
olfactory receptor, family 8, subfamily D, member 2 |
| chr20_+_58630972 | 0.25 |
ENST00000313426.1 |
C20orf197 |
chromosome 20 open reading frame 197 |
| chr20_-_50419055 | 0.25 |
ENST00000217086.4 |
SALL4 |
spalt-like transcription factor 4 |
| chr4_-_74486347 | 0.24 |
ENST00000342081.3 |
RASSF6 |
Ras association (RalGDS/AF-6) domain family member 6 |
| chr4_-_74486109 | 0.23 |
ENST00000395777.2 |
RASSF6 |
Ras association (RalGDS/AF-6) domain family member 6 |
| chr5_-_24645078 | 0.23 |
ENST00000264463.4 |
CDH10 |
cadherin 10, type 2 (T2-cadherin) |
| chr13_-_86373536 | 0.22 |
ENST00000400286.2 |
SLITRK6 |
SLIT and NTRK-like family, member 6 |
| chr4_+_77356248 | 0.22 |
ENST00000296043.6 |
SHROOM3 |
shroom family member 3 |
| chr10_+_24755416 | 0.21 |
ENST00000396446.1 ENST00000396445.1 ENST00000376451.2 |
KIAA1217 |
KIAA1217 |
| chr4_+_40198527 | 0.21 |
ENST00000381799.5 |
RHOH |
ras homolog family member H |
| chr8_-_42234745 | 0.21 |
ENST00000220812.2 |
DKK4 |
dickkopf WNT signaling pathway inhibitor 4 |
| chr7_-_71868354 | 0.19 |
ENST00000412588.1 |
CALN1 |
calneuron 1 |
| chr19_-_51523412 | 0.19 |
ENST00000391805.1 ENST00000599077.1 |
KLK10 |
kallikrein-related peptidase 10 |
| chr6_+_130339710 | 0.19 |
ENST00000526087.1 ENST00000533560.1 ENST00000361794.2 |
L3MBTL3 |
l(3)mbt-like 3 (Drosophila) |
| chr5_-_141249154 | 0.18 |
ENST00000357517.5 ENST00000536585.1 |
PCDH1 |
protocadherin 1 |
| chr18_-_53089723 | 0.18 |
ENST00000561992.1 ENST00000562512.2 |
TCF4 |
transcription factor 4 |
| chr21_+_17442799 | 0.18 |
ENST00000602580.1 ENST00000458468.1 ENST00000602935.1 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr19_-_51523275 | 0.18 |
ENST00000309958.3 |
KLK10 |
kallikrein-related peptidase 10 |
| chr8_-_139926236 | 0.18 |
ENST00000303045.6 ENST00000435777.1 |
COL22A1 |
collagen, type XXII, alpha 1 |
| chr14_-_38064198 | 0.18 |
ENST00000250448.2 |
FOXA1 |
forkhead box A1 |
| chr13_-_36050819 | 0.17 |
ENST00000379919.4 |
MAB21L1 |
mab-21-like 1 (C. elegans) |
| chr18_+_61445007 | 0.17 |
ENST00000447428.1 ENST00000546027.1 |
SERPINB7 |
serpin peptidase inhibitor, clade B (ovalbumin), member 7 |
| chr17_-_64225508 | 0.16 |
ENST00000205948.6 |
APOH |
apolipoprotein H (beta-2-glycoprotein I) |
| chr4_+_69313145 | 0.16 |
ENST00000305363.4 |
TMPRSS11E |
transmembrane protease, serine 11E |
| chr8_-_109799793 | 0.16 |
ENST00000297459.3 |
TMEM74 |
transmembrane protein 74 |
| chr1_-_242612779 | 0.16 |
ENST00000427495.1 |
PLD5 |
phospholipase D family, member 5 |
| chr2_-_70780572 | 0.16 |
ENST00000450929.1 |
TGFA |
transforming growth factor, alpha |
| chr14_-_54423529 | 0.16 |
ENST00000245451.4 ENST00000559087.1 |
BMP4 |
bone morphogenetic protein 4 |
| chr1_+_160160346 | 0.15 |
ENST00000368078.3 |
CASQ1 |
calsequestrin 1 (fast-twitch, skeletal muscle) |
| chr1_+_160160283 | 0.15 |
ENST00000368079.3 |
CASQ1 |
calsequestrin 1 (fast-twitch, skeletal muscle) |
| chr12_+_4385230 | 0.15 |
ENST00000536537.1 |
CCND2 |
cyclin D2 |
| chr2_-_216003127 | 0.15 |
ENST00000412081.1 ENST00000272895.7 |
ABCA12 |
ATP-binding cassette, sub-family A (ABC1), member 12 |
| chr21_-_42219065 | 0.14 |
ENST00000400454.1 |
DSCAM |
Down syndrome cell adhesion molecule |
| chr2_-_70780770 | 0.14 |
ENST00000444975.1 ENST00000445399.1 ENST00000418333.2 |
TGFA |
transforming growth factor, alpha |
| chr12_-_71031220 | 0.14 |
ENST00000334414.6 |
PTPRB |
protein tyrosine phosphatase, receptor type, B |
| chr17_-_7167279 | 0.14 |
ENST00000571932.2 |
CLDN7 |
claudin 7 |
| chr11_+_65554493 | 0.14 |
ENST00000335987.3 |
OVOL1 |
ovo-like zinc finger 1 |
| chr5_+_126984710 | 0.13 |
ENST00000379445.3 |
CTXN3 |
cortexin 3 |
| chr1_+_160370344 | 0.13 |
ENST00000368061.2 |
VANGL2 |
VANGL planar cell polarity protein 2 |
| chr12_-_71031185 | 0.13 |
ENST00000548122.1 ENST00000551525.1 ENST00000550358.1 |
PTPRB |
protein tyrosine phosphatase, receptor type, B |
| chr2_-_50201327 | 0.12 |
ENST00000412315.1 |
NRXN1 |
neurexin 1 |
| chr17_-_27418537 | 0.12 |
ENST00000408971.2 |
TIAF1 |
TGFB1-induced anti-apoptotic factor 1 |
| chr17_-_38911580 | 0.12 |
ENST00000312150.4 |
KRT25 |
keratin 25 |
| chr5_+_147582387 | 0.12 |
ENST00000325630.2 |
SPINK6 |
serine peptidase inhibitor, Kazal type 6 |
| chr16_+_76587314 | 0.12 |
ENST00000563764.1 |
RP11-58C22.1 |
Uncharacterized protein |
| chr12_-_28124903 | 0.12 |
ENST00000395872.1 ENST00000354417.3 ENST00000201015.4 |
PTHLH |
parathyroid hormone-like hormone |
| chr12_-_28125638 | 0.12 |
ENST00000545234.1 |
PTHLH |
parathyroid hormone-like hormone |
| chr9_+_12693336 | 0.11 |
ENST00000381137.2 ENST00000388918.5 |
TYRP1 |
tyrosinase-related protein 1 |
| chr14_+_61654271 | 0.11 |
ENST00000555185.1 ENST00000557294.1 ENST00000556778.1 |
PRKCH |
protein kinase C, eta |
| chr8_-_20040638 | 0.11 |
ENST00000519026.1 ENST00000276373.5 ENST00000440926.1 ENST00000437980.1 |
SLC18A1 |
solute carrier family 18 (vesicular monoamine transporter), member 1 |
| chr8_-_20040601 | 0.11 |
ENST00000265808.7 ENST00000522513.1 |
SLC18A1 |
solute carrier family 18 (vesicular monoamine transporter), member 1 |
| chr2_+_162272605 | 0.11 |
ENST00000389554.3 |
TBR1 |
T-box, brain, 1 |
| chr8_+_105235572 | 0.11 |
ENST00000523362.1 |
RIMS2 |
regulating synaptic membrane exocytosis 2 |
| chrX_+_105192423 | 0.10 |
ENST00000540278.1 |
NRK |
Nik related kinase |
| chr14_+_22966642 | 0.10 |
ENST00000390496.1 |
TRAJ41 |
T cell receptor alpha joining 41 |
| chr11_-_115375107 | 0.10 |
ENST00000545380.1 ENST00000452722.3 ENST00000537058.1 ENST00000536727.1 ENST00000542447.2 ENST00000331581.6 |
CADM1 |
cell adhesion molecule 1 |
| chr10_+_118083919 | 0.10 |
ENST00000333254.3 |
CCDC172 |
coiled-coil domain containing 172 |
| chr11_-_129062093 | 0.10 |
ENST00000310343.9 |
ARHGAP32 |
Rho GTPase activating protein 32 |
| chr11_+_5410607 | 0.10 |
ENST00000328611.3 |
OR51M1 |
olfactory receptor, family 51, subfamily M, member 1 |
| chr6_-_161695042 | 0.10 |
ENST00000366908.5 ENST00000366911.5 ENST00000366905.3 |
AGPAT4 |
1-acylglycerol-3-phosphate O-acyltransferase 4 |
| chr11_-_117747434 | 0.09 |
ENST00000529335.2 ENST00000530956.1 ENST00000260282.4 |
FXYD6 |
FXYD domain containing ion transport regulator 6 |
| chr11_+_20620946 | 0.09 |
ENST00000525748.1 |
SLC6A5 |
solute carrier family 6 (neurotransmitter transporter), member 5 |
| chr17_-_38956205 | 0.09 |
ENST00000306658.7 |
KRT28 |
keratin 28 |
| chr12_-_52828147 | 0.09 |
ENST00000252245.5 |
KRT75 |
keratin 75 |
| chrX_+_105937068 | 0.09 |
ENST00000324342.3 |
RNF128 |
ring finger protein 128, E3 ubiquitin protein ligase |
| chr3_+_130569429 | 0.09 |
ENST00000505330.1 ENST00000504381.1 ENST00000507488.2 ENST00000393221.4 |
ATP2C1 |
ATPase, Ca++ transporting, type 2C, member 1 |
| chr11_-_117747607 | 0.09 |
ENST00000540359.1 ENST00000539526.1 |
FXYD6 |
FXYD domain containing ion transport regulator 6 |
| chr1_+_107683436 | 0.09 |
ENST00000370068.1 |
NTNG1 |
netrin G1 |
| chr6_-_161695074 | 0.09 |
ENST00000457520.2 ENST00000366906.5 ENST00000320285.4 |
AGPAT4 |
1-acylglycerol-3-phosphate O-acyltransferase 4 |
| chr14_-_37051798 | 0.09 |
ENST00000258829.5 |
NKX2-8 |
NK2 homeobox 8 |
| chr6_-_155776966 | 0.08 |
ENST00000159060.2 |
NOX3 |
NADPH oxidase 3 |
| chr5_+_147582348 | 0.08 |
ENST00000514389.1 |
SPINK6 |
serine peptidase inhibitor, Kazal type 6 |
| chr13_-_103719196 | 0.08 |
ENST00000245312.3 |
SLC10A2 |
solute carrier family 10 (sodium/bile acid cotransporter), member 2 |
| chr1_+_107683644 | 0.08 |
ENST00000370067.1 |
NTNG1 |
netrin G1 |
| chrX_-_133792480 | 0.08 |
ENST00000359237.4 |
PLAC1 |
placenta-specific 1 |
| chr17_-_38938786 | 0.08 |
ENST00000301656.3 |
KRT27 |
keratin 27 |
| chr11_+_10476851 | 0.08 |
ENST00000396553.2 |
AMPD3 |
adenosine monophosphate deaminase 3 |
| chr11_-_117748138 | 0.08 |
ENST00000527717.1 |
FXYD6 |
FXYD domain containing ion transport regulator 6 |
| chr12_-_89746173 | 0.08 |
ENST00000308385.6 |
DUSP6 |
dual specificity phosphatase 6 |
| chr13_-_95131923 | 0.08 |
ENST00000377028.5 ENST00000446125.1 |
DCT |
dopachrome tautomerase |
| chr17_-_46716647 | 0.08 |
ENST00000608940.1 |
RP11-357H14.17 |
RP11-357H14.17 |
| chr18_+_28956740 | 0.08 |
ENST00000308128.4 ENST00000359747.4 |
DSG4 |
desmoglein 4 |
| chrX_-_117119243 | 0.08 |
ENST00000539496.1 ENST00000469946.1 |
KLHL13 |
kelch-like family member 13 |
| chrX_+_139791917 | 0.08 |
ENST00000607004.1 ENST00000370535.3 |
LINC00632 |
long intergenic non-protein coding RNA 632 |
| chr1_-_152386732 | 0.08 |
ENST00000271835.3 |
CRNN |
cornulin |
| chr1_+_180601139 | 0.07 |
ENST00000367590.4 ENST00000367589.3 |
XPR1 |
xenotropic and polytropic retrovirus receptor 1 |
| chr4_+_41614720 | 0.07 |
ENST00000509277.1 |
LIMCH1 |
LIM and calponin homology domains 1 |
| chr7_-_25268104 | 0.07 |
ENST00000222674.2 |
NPVF |
neuropeptide VF precursor |
| chr17_-_10421853 | 0.07 |
ENST00000226207.5 |
MYH1 |
myosin, heavy chain 1, skeletal muscle, adult |
| chr17_+_47448102 | 0.07 |
ENST00000576461.1 |
RP11-81K2.1 |
Uncharacterized protein |
| chr12_+_19358228 | 0.07 |
ENST00000424268.1 ENST00000543806.1 |
PLEKHA5 |
pleckstrin homology domain containing, family A member 5 |
| chr10_-_116444371 | 0.07 |
ENST00000533213.2 ENST00000369252.4 |
ABLIM1 |
actin binding LIM protein 1 |
| chr7_-_130080977 | 0.07 |
ENST00000223208.5 |
CEP41 |
centrosomal protein 41kDa |
| chr6_+_50681541 | 0.07 |
ENST00000008391.3 |
TFAP2D |
transcription factor AP-2 delta (activating enhancer binding protein 2 delta) |
| chr14_+_72052983 | 0.07 |
ENST00000358550.2 |
SIPA1L1 |
signal-induced proliferation-associated 1 like 1 |
| chrX_-_110655306 | 0.07 |
ENST00000371993.2 |
DCX |
doublecortin |
| chr4_-_123542224 | 0.07 |
ENST00000264497.3 |
IL21 |
interleukin 21 |
| chr18_-_53069419 | 0.07 |
ENST00000570177.2 |
TCF4 |
transcription factor 4 |
| chr3_-_27764190 | 0.07 |
ENST00000537516.1 |
EOMES |
eomesodermin |
| chr7_-_37026108 | 0.07 |
ENST00000396045.3 |
ELMO1 |
engulfment and cell motility 1 |
| chr11_+_34664014 | 0.07 |
ENST00000527935.1 |
EHF |
ets homologous factor |
| chr7_-_83278322 | 0.07 |
ENST00000307792.3 |
SEMA3E |
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3E |
| chr1_-_92952433 | 0.07 |
ENST00000294702.5 |
GFI1 |
growth factor independent 1 transcription repressor |
| chr7_+_116660246 | 0.07 |
ENST00000434836.1 ENST00000393443.1 ENST00000465133.1 ENST00000477742.1 ENST00000393447.4 ENST00000393444.3 |
ST7 |
suppression of tumorigenicity 7 |
| chr13_-_84456527 | 0.06 |
ENST00000377084.2 |
SLITRK1 |
SLIT and NTRK-like family, member 1 |
| chr3_+_120626919 | 0.06 |
ENST00000273666.6 ENST00000471454.1 ENST00000472879.1 ENST00000497029.1 ENST00000492541.1 |
STXBP5L |
syntaxin binding protein 5-like |
| chr2_-_99871570 | 0.06 |
ENST00000333017.2 ENST00000409679.1 ENST00000423306.1 |
LYG2 |
lysozyme G-like 2 |
| chr19_+_13049413 | 0.06 |
ENST00000316448.5 ENST00000588454.1 |
CALR |
calreticulin |
| chr2_+_128403720 | 0.06 |
ENST00000272644.3 |
GPR17 |
G protein-coupled receptor 17 |
| chr2_+_128403439 | 0.06 |
ENST00000544369.1 |
GPR17 |
G protein-coupled receptor 17 |
| chr11_+_34663913 | 0.06 |
ENST00000532302.1 |
EHF |
ets homologous factor |
| chrX_+_68835911 | 0.06 |
ENST00000525810.1 ENST00000527388.1 ENST00000374553.2 ENST00000374552.4 ENST00000338901.3 ENST00000524573.1 |
EDA |
ectodysplasin A |
| chr22_-_32860427 | 0.06 |
ENST00000534972.1 ENST00000397450.1 ENST00000397452.1 |
BPIFC |
BPI fold containing family C |
| chr4_+_71494461 | 0.06 |
ENST00000396073.3 |
ENAM |
enamelin |
| chr1_+_81771806 | 0.06 |
ENST00000370721.1 ENST00000370727.1 ENST00000370725.1 ENST00000370723.1 ENST00000370728.1 ENST00000370730.1 |
LPHN2 |
latrophilin 2 |
| chr9_+_470288 | 0.06 |
ENST00000382303.1 |
KANK1 |
KN motif and ankyrin repeat domains 1 |
| chr9_-_124989804 | 0.06 |
ENST00000373755.2 ENST00000373754.2 |
LHX6 |
LIM homeobox 6 |
| chr1_-_150738261 | 0.06 |
ENST00000448301.2 ENST00000368985.3 |
CTSS |
cathepsin S |
| chr14_+_22977587 | 0.06 |
ENST00000390504.1 |
TRAJ33 |
T cell receptor alpha joining 33 |
| chr3_-_79816965 | 0.06 |
ENST00000464233.1 |
ROBO1 |
roundabout, axon guidance receptor, homolog 1 (Drosophila) |
| chr1_+_153746683 | 0.06 |
ENST00000271857.2 |
SLC27A3 |
solute carrier family 27 (fatty acid transporter), member 3 |
| chr11_-_107729887 | 0.06 |
ENST00000525815.1 |
SLC35F2 |
solute carrier family 35, member F2 |
| chr1_-_234667504 | 0.06 |
ENST00000421207.1 ENST00000435574.1 |
RP5-855F14.1 |
RP5-855F14.1 |
| chrX_-_73512411 | 0.06 |
ENST00000602576.1 ENST00000429124.1 |
FTX |
FTX transcript, XIST regulator (non-protein coding) |
| chr4_+_88571429 | 0.05 |
ENST00000339673.6 ENST00000282479.7 |
DMP1 |
dentin matrix acidic phosphoprotein 1 |
| chr8_-_17579726 | 0.05 |
ENST00000381861.3 |
MTUS1 |
microtubule associated tumor suppressor 1 |
| chr21_-_31538971 | 0.05 |
ENST00000286808.3 |
CLDN17 |
claudin 17 |
| chr8_+_50824233 | 0.05 |
ENST00000522124.1 |
SNTG1 |
syntrophin, gamma 1 |
| chr7_+_141478242 | 0.05 |
ENST00000247881.2 |
TAS2R4 |
taste receptor, type 2, member 4 |
| chr17_+_48243352 | 0.05 |
ENST00000344627.6 ENST00000262018.3 ENST00000543315.1 ENST00000451235.2 ENST00000511303.1 |
SGCA |
sarcoglycan, alpha (50kDa dystrophin-associated glycoprotein) |
| chr4_+_114214125 | 0.05 |
ENST00000509550.1 |
ANK2 |
ankyrin 2, neuronal |
| chr12_+_20963647 | 0.05 |
ENST00000381545.3 |
SLCO1B3 |
solute carrier organic anion transporter family, member 1B3 |
| chr3_-_191000172 | 0.05 |
ENST00000427544.2 |
UTS2B |
urotensin 2B |
| chr12_-_51422017 | 0.05 |
ENST00000394904.3 |
SLC11A2 |
solute carrier family 11 (proton-coupled divalent metal ion transporter), member 2 |
| chr12_+_20963632 | 0.05 |
ENST00000540853.1 ENST00000261196.2 |
SLCO1B3 |
solute carrier organic anion transporter family, member 1B3 |
| chr17_-_26733179 | 0.05 |
ENST00000440501.1 ENST00000321666.5 |
SLC46A1 |
solute carrier family 46 (folate transporter), member 1 |
| chr4_+_155484155 | 0.05 |
ENST00000509493.1 |
FGB |
fibrinogen beta chain |
| chr3_-_157824292 | 0.05 |
ENST00000483851.2 |
SHOX2 |
short stature homeobox 2 |
| chr17_-_39093672 | 0.05 |
ENST00000209718.3 ENST00000436344.3 ENST00000485751.1 |
KRT23 |
keratin 23 (histone deacetylase inducible) |
| chr19_-_36304201 | 0.05 |
ENST00000301175.3 |
PRODH2 |
proline dehydrogenase (oxidase) 2 |
| chr3_-_33686925 | 0.05 |
ENST00000485378.2 ENST00000313350.6 ENST00000487200.1 |
CLASP2 |
cytoplasmic linker associated protein 2 |
| chr3_-_33686743 | 0.05 |
ENST00000333778.6 ENST00000539981.1 |
CLASP2 |
cytoplasmic linker associated protein 2 |
| chrX_-_44202857 | 0.05 |
ENST00000420999.1 |
EFHC2 |
EF-hand domain (C-terminal) containing 2 |
| chr1_+_28586006 | 0.05 |
ENST00000253063.3 |
SESN2 |
sestrin 2 |
| chr19_-_7968427 | 0.05 |
ENST00000539278.1 |
AC010336.1 |
Uncharacterized protein |
| chr20_-_50722183 | 0.05 |
ENST00000371523.4 |
ZFP64 |
ZFP64 zinc finger protein |
| chr3_-_108248169 | 0.05 |
ENST00000273353.3 |
MYH15 |
myosin, heavy chain 15 |
| chr18_-_53257027 | 0.05 |
ENST00000568740.1 ENST00000564403.2 ENST00000537578.1 |
TCF4 |
transcription factor 4 |
| chr12_-_95510743 | 0.05 |
ENST00000551521.1 |
FGD6 |
FYVE, RhoGEF and PH domain containing 6 |
| chr6_-_39693111 | 0.05 |
ENST00000373215.3 ENST00000538893.1 ENST00000287152.7 ENST00000373216.3 |
KIF6 |
kinesin family member 6 |
| chr6_+_135502501 | 0.05 |
ENST00000527615.1 ENST00000420123.2 ENST00000525369.1 ENST00000528774.1 ENST00000534121.1 ENST00000534044.1 ENST00000533624.1 |
MYB |
v-myb avian myeloblastosis viral oncogene homolog |
| chr2_-_136678123 | 0.05 |
ENST00000422708.1 |
DARS |
aspartyl-tRNA synthetase |
| chr6_+_151042224 | 0.04 |
ENST00000358517.2 |
PLEKHG1 |
pleckstrin homology domain containing, family G (with RhoGef domain) member 1 |
| chr11_-_7847519 | 0.04 |
ENST00000328375.1 |
OR5P3 |
olfactory receptor, family 5, subfamily P, member 3 |
| chr6_-_66417107 | 0.04 |
ENST00000370621.3 ENST00000370618.3 ENST00000503581.1 ENST00000393380.2 |
EYS |
eyes shut homolog (Drosophila) |
| chr18_-_33709268 | 0.04 |
ENST00000269187.5 ENST00000590986.1 ENST00000440549.2 |
SLC39A6 |
solute carrier family 39 (zinc transporter), member 6 |
| chr2_+_171571827 | 0.04 |
ENST00000375281.3 |
SP5 |
Sp5 transcription factor |
| chr4_-_41884582 | 0.04 |
ENST00000499082.2 |
LINC00682 |
long intergenic non-protein coding RNA 682 |
| chr4_-_111563076 | 0.04 |
ENST00000354925.2 ENST00000511990.1 |
PITX2 |
paired-like homeodomain 2 |
| chr3_-_72149553 | 0.04 |
ENST00000468646.2 ENST00000464271.1 |
LINC00877 |
long intergenic non-protein coding RNA 877 |
| chr14_-_78083112 | 0.04 |
ENST00000216484.2 |
SPTLC2 |
serine palmitoyltransferase, long chain base subunit 2 |
| chr21_-_32253874 | 0.04 |
ENST00000332378.4 |
KRTAP11-1 |
keratin associated protein 11-1 |
| chr2_+_163175394 | 0.04 |
ENST00000446271.1 ENST00000429691.2 |
GCA |
grancalcin, EF-hand calcium binding protein |
| chr9_-_131486367 | 0.04 |
ENST00000372663.4 ENST00000406904.2 ENST00000452105.1 ENST00000372672.2 ENST00000372667.5 |
ZDHHC12 |
zinc finger, DHHC-type containing 12 |
| chr14_+_22739823 | 0.04 |
ENST00000390464.2 |
TRAV38-1 |
T cell receptor alpha variable 38-1 |
| chr3_+_121774202 | 0.04 |
ENST00000469710.1 ENST00000493101.1 ENST00000330540.2 ENST00000264468.5 |
CD86 |
CD86 molecule |
| chr17_-_9694614 | 0.04 |
ENST00000330255.5 ENST00000571134.1 |
DHRS7C |
dehydrogenase/reductase (SDR family) member 7C |
| chr12_+_21207503 | 0.04 |
ENST00000545916.1 |
SLCO1B7 |
solute carrier organic anion transporter family, member 1B7 (non-functional) |
| chr17_-_39280419 | 0.04 |
ENST00000394014.1 |
KRTAP4-12 |
keratin associated protein 4-12 |
| chr2_-_80531399 | 0.04 |
ENST00000409148.1 ENST00000415098.1 ENST00000452811.1 |
LRRTM1 |
leucine rich repeat transmembrane neuronal 1 |
| chr3_-_123339343 | 0.04 |
ENST00000578202.1 |
MYLK |
myosin light chain kinase |
| chr7_-_81399411 | 0.04 |
ENST00000423064.2 |
HGF |
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr11_-_69633792 | 0.04 |
ENST00000334134.2 |
FGF3 |
fibroblast growth factor 3 |
| chr9_+_130026756 | 0.04 |
ENST00000314904.5 ENST00000373387.4 |
GARNL3 |
GTPase activating Rap/RanGAP domain-like 3 |
| chr16_-_30102547 | 0.04 |
ENST00000279386.2 |
TBX6 |
T-box 6 |
| chr12_+_28410128 | 0.04 |
ENST00000381259.1 ENST00000381256.1 |
CCDC91 |
coiled-coil domain containing 91 |
| chr13_-_99910673 | 0.04 |
ENST00000397473.2 ENST00000397470.2 |
GPR18 |
G protein-coupled receptor 18 |
| chr14_-_36988882 | 0.04 |
ENST00000498187.2 |
NKX2-1 |
NK2 homeobox 1 |
| chr6_-_51952418 | 0.04 |
ENST00000371117.3 |
PKHD1 |
polycystic kidney and hepatic disease 1 (autosomal recessive) |
| chr1_-_11024258 | 0.04 |
ENST00000418570.2 |
C1orf127 |
chromosome 1 open reading frame 127 |
| chr10_-_24770632 | 0.04 |
ENST00000596413.1 |
AL353583.1 |
AL353583.1 |
| chr3_-_27763803 | 0.03 |
ENST00000449599.1 |
EOMES |
eomesodermin |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0060738 | epithelial-mesenchymal signaling involved in prostate gland development(GO:0060738) |
| 0.1 | 0.9 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.1 | 0.1 | GO:0008217 | regulation of blood pressure(GO:0008217) |
| 0.1 | 1.7 | GO:0032331 | negative regulation of chondrocyte differentiation(GO:0032331) |
| 0.1 | 0.2 | GO:0061346 | non-canonical Wnt signaling pathway involved in heart development(GO:0061341) planar cell polarity pathway involved in heart morphogenesis(GO:0061346) |
| 0.1 | 0.1 | GO:0060809 | mesodermal to mesenchymal transition involved in gastrulation(GO:0060809) |
| 0.0 | 0.2 | GO:0090214 | spongiotrophoblast layer developmental growth(GO:0090214) |
| 0.0 | 0.1 | GO:1901994 | meiotic cell cycle phase transition(GO:0044771) regulation of meiotic cell cycle phase transition(GO:1901993) negative regulation of meiotic cell cycle phase transition(GO:1901994) |
| 0.0 | 0.2 | GO:0090362 | positive regulation of platelet-derived growth factor production(GO:0090362) |
| 0.0 | 0.2 | GO:0015842 | aminergic neurotransmitter loading into synaptic vesicle(GO:0015842) |
| 0.0 | 0.1 | GO:0035627 | ceramide transport(GO:0035627) |
| 0.0 | 0.2 | GO:0060005 | vestibular reflex(GO:0060005) |
| 0.0 | 0.4 | GO:0014877 | response to muscle inactivity involved in regulation of muscle adaptation(GO:0014877) response to denervation involved in regulation of muscle adaptation(GO:0014894) |
| 0.0 | 0.2 | GO:0051799 | negative regulation of hair follicle development(GO:0051799) |
| 0.0 | 0.1 | GO:0042663 | regulation of endodermal cell fate specification(GO:0042663) |
| 0.0 | 0.1 | GO:0042271 | susceptibility to natural killer cell mediated cytotoxicity(GO:0042271) |
| 0.0 | 0.4 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.0 | 0.1 | GO:0070105 | positive regulation of interleukin-6-mediated signaling pathway(GO:0070105) |
| 0.0 | 0.1 | GO:2000393 | negative regulation of lamellipodium morphogenesis(GO:2000393) |
| 0.0 | 0.1 | GO:0021823 | cerebral cortex tangential migration using cell-cell interactions(GO:0021823) postnatal olfactory bulb interneuron migration(GO:0021827) chemorepulsion involved in postnatal olfactory bulb interneuron migration(GO:0021836) |
| 0.0 | 0.1 | GO:1904502 | regulation of lipophagy(GO:1904502) positive regulation of lipophagy(GO:1904504) |
| 0.0 | 0.1 | GO:0021764 | amygdala development(GO:0021764) |
| 0.0 | 0.4 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.0 | 0.1 | GO:0036371 | protein localization to T-tubule(GO:0036371) |
| 0.0 | 0.3 | GO:0045741 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) |
| 0.0 | 0.0 | GO:0086073 | bundle of His cell-Purkinje myocyte adhesion involved in cell communication(GO:0086073) |
| 0.0 | 0.0 | GO:1904899 | regulation of hepatic stellate cell proliferation(GO:1904897) positive regulation of hepatic stellate cell proliferation(GO:1904899) hepatic stellate cell proliferation(GO:1990922) |
| 0.0 | 0.0 | GO:0006422 | aspartyl-tRNA aminoacylation(GO:0006422) |
| 0.0 | 0.1 | GO:0032468 | Golgi calcium ion homeostasis(GO:0032468) |
| 0.0 | 0.2 | GO:0045176 | apical protein localization(GO:0045176) |
| 0.0 | 0.2 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.0 | 0.0 | GO:0060127 | subthalamic nucleus development(GO:0021763) prolactin secreting cell differentiation(GO:0060127) superior vena cava morphogenesis(GO:0060578) |
| 0.0 | 0.0 | GO:0002644 | negative regulation of tolerance induction(GO:0002644) positive regulation of interleukin-4 biosynthetic process(GO:0045404) |
| 0.0 | 0.1 | GO:0097116 | gephyrin clustering involved in postsynaptic density assembly(GO:0097116) |
| 0.0 | 0.1 | GO:0002074 | extraocular skeletal muscle development(GO:0002074) |
| 0.0 | 0.2 | GO:0034196 | acylglycerol transport(GO:0034196) triglyceride transport(GO:0034197) |
| 0.0 | 0.1 | GO:0098814 | spontaneous neurotransmitter secretion(GO:0061669) spontaneous synaptic transmission(GO:0098814) |
| 0.0 | 0.1 | GO:0006196 | AMP catabolic process(GO:0006196) |
| 0.0 | 0.2 | GO:0043010 | camera-type eye development(GO:0043010) |
| 0.0 | 0.1 | GO:0051958 | methotrexate transport(GO:0051958) |
| 0.0 | 0.1 | GO:0002501 | peptide antigen assembly with MHC protein complex(GO:0002501) |
| 0.0 | 0.1 | GO:0061153 | trachea submucosa development(GO:0061152) trachea gland development(GO:0061153) |
| 0.0 | 0.0 | GO:0046013 | regulation of T cell homeostatic proliferation(GO:0046013) |
| 0.0 | 0.1 | GO:0034351 | regulation of glial cell apoptotic process(GO:0034350) negative regulation of glial cell apoptotic process(GO:0034351) |
| 0.0 | 0.0 | GO:2000854 | positive regulation of corticosterone secretion(GO:2000854) |
| 0.0 | 0.1 | GO:0021847 | ventricular zone neuroblast division(GO:0021847) |
| 0.0 | 0.1 | GO:0015692 | lead ion transport(GO:0015692) |
| 0.0 | 0.0 | GO:0050883 | musculoskeletal movement, spinal reflex action(GO:0050883) |
| 0.0 | 0.1 | GO:0042488 | positive regulation of odontogenesis of dentin-containing tooth(GO:0042488) |
| 0.0 | 0.1 | GO:0060012 | synaptic transmission, glycinergic(GO:0060012) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | GO:0014802 | terminal cisterna(GO:0014802) |
| 0.0 | 0.1 | GO:0060187 | cell pole(GO:0060187) |
| 0.0 | 0.2 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 0.0 | 0.2 | GO:0070081 | clathrin-sculpted monoamine transport vesicle(GO:0070081) clathrin-sculpted monoamine transport vesicle membrane(GO:0070083) |
| 0.0 | 0.1 | GO:0097209 | epidermal lamellar body(GO:0097209) |
| 0.0 | 0.4 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.0 | 0.1 | GO:0070826 | paraferritin complex(GO:0070826) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0015222 | serotonin transmembrane transporter activity(GO:0015222) |
| 0.1 | 1.7 | GO:0051428 | peptide hormone receptor binding(GO:0051428) |
| 0.0 | 0.1 | GO:0034040 | lipid-transporting ATPase activity(GO:0034040) |
| 0.0 | 0.3 | GO:0001087 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.0 | 0.2 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 0.0 | 0.1 | GO:0015375 | glycine:sodium symporter activity(GO:0015375) |
| 0.0 | 0.1 | GO:0004699 | calcium-independent protein kinase C activity(GO:0004699) |
| 0.0 | 0.3 | GO:0005549 | odorant binding(GO:0005549) |
| 0.0 | 0.1 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.0 | 0.1 | GO:0000822 | inositol hexakisphosphate binding(GO:0000822) |
| 0.0 | 0.4 | GO:0001608 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.0 | 0.0 | GO:0036134 | thromboxane-A synthase activity(GO:0004796) 12-hydroxyheptadecatrienoic acid synthase activity(GO:0036134) |
| 0.0 | 0.1 | GO:0004167 | dopachrome isomerase activity(GO:0004167) |
| 0.0 | 0.1 | GO:0015350 | methotrexate transporter activity(GO:0015350) |
| 0.0 | 0.1 | GO:0004657 | proline dehydrogenase activity(GO:0004657) |
| 0.0 | 0.2 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.0 | 0.1 | GO:0015094 | cadmium ion transmembrane transporter activity(GO:0015086) cobalt ion transmembrane transporter activity(GO:0015087) lead ion transmembrane transporter activity(GO:0015094) ferrous iron uptake transmembrane transporter activity(GO:0015639) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.6 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.0 | 1.7 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.3 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.0 | 0.2 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |