Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
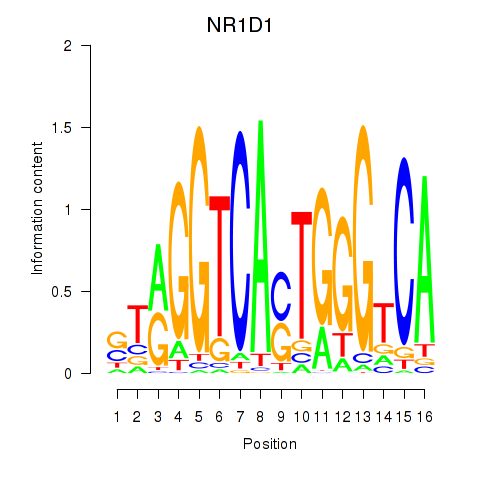
Results for NR1D1
Z-value: 0.48
Transcription factors associated with NR1D1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NR1D1
|
ENSG00000126368.5 | NR1D1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NR1D1 | hg19_v2_chr17_-_38256973_38256990 | 0.91 | 1.9e-03 | Click! |
Activity profile of NR1D1 motif
Sorted Z-values of NR1D1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of NR1D1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_44398943 | 0.47 |
ENST00000372359.5 ENST00000414809.3 |
ARTN |
artemin |
| chr16_+_67233412 | 0.36 |
ENST00000477898.1 |
ELMO3 |
engulfment and cell motility 3 |
| chr16_+_67233007 | 0.36 |
ENST00000360833.1 ENST00000393997.2 |
ELMO3 |
engulfment and cell motility 3 |
| chr17_-_3599327 | 0.32 |
ENST00000551178.1 ENST00000552276.1 ENST00000547178.1 |
P2RX5 |
purinergic receptor P2X, ligand-gated ion channel, 5 |
| chr17_-_3599492 | 0.31 |
ENST00000435558.1 ENST00000345901.3 |
P2RX5 |
purinergic receptor P2X, ligand-gated ion channel, 5 |
| chr15_+_26360970 | 0.26 |
ENST00000556159.1 ENST00000557523.1 |
LINC00929 |
long intergenic non-protein coding RNA 929 |
| chr3_+_169755715 | 0.23 |
ENST00000355897.5 |
GPR160 |
G protein-coupled receptor 160 |
| chr1_-_6557156 | 0.22 |
ENST00000537245.1 |
PLEKHG5 |
pleckstrin homology domain containing, family G (with RhoGef domain) member 5 |
| chr17_-_74528128 | 0.21 |
ENST00000590175.1 |
CYGB |
cytoglobin |
| chr1_+_20915409 | 0.21 |
ENST00000375071.3 |
CDA |
cytidine deaminase |
| chr17_-_76128488 | 0.20 |
ENST00000322914.3 |
TMC6 |
transmembrane channel-like 6 |
| chr2_-_64371546 | 0.19 |
ENST00000358912.4 |
PELI1 |
pellino E3 ubiquitin protein ligase 1 |
| chrX_-_49089771 | 0.18 |
ENST00000376251.1 ENST00000323022.5 ENST00000376265.2 |
CACNA1F |
calcium channel, voltage-dependent, L type, alpha 1F subunit |
| chr1_-_41131326 | 0.18 |
ENST00000372684.3 |
RIMS3 |
regulating synaptic membrane exocytosis 3 |
| chr19_-_52035044 | 0.15 |
ENST00000359982.4 ENST00000436458.1 ENST00000425629.3 ENST00000391797.3 ENST00000343300.4 |
SIGLEC6 |
sialic acid binding Ig-like lectin 6 |
| chr17_+_39405939 | 0.14 |
ENST00000334109.2 |
KRTAP9-4 |
keratin associated protein 9-4 |
| chr17_-_38074842 | 0.14 |
ENST00000309481.7 |
GSDMB |
gasdermin B |
| chr17_-_38074859 | 0.13 |
ENST00000520542.1 ENST00000418519.1 ENST00000394179.1 |
GSDMB |
gasdermin B |
| chr10_-_103880209 | 0.13 |
ENST00000425280.1 |
LDB1 |
LIM domain binding 1 |
| chr2_+_103353367 | 0.13 |
ENST00000454536.1 ENST00000409528.1 ENST00000409173.1 |
TMEM182 |
transmembrane protein 182 |
| chr19_-_42927251 | 0.13 |
ENST00000597001.1 |
LIPE |
lipase, hormone-sensitive |
| chr1_-_111991850 | 0.11 |
ENST00000411751.2 |
WDR77 |
WD repeat domain 77 |
| chr1_+_2398876 | 0.10 |
ENST00000449969.1 |
PLCH2 |
phospholipase C, eta 2 |
| chr17_-_56358287 | 0.10 |
ENST00000225275.3 ENST00000340482.3 |
MPO |
myeloperoxidase |
| chr10_-_8095412 | 0.10 |
ENST00000458727.1 ENST00000355358.1 |
RP11-379F12.3 GATA3-AS1 |
RP11-379F12.3 GATA3 antisense RNA 1 |
| chr2_+_220306745 | 0.10 |
ENST00000431523.1 ENST00000396698.1 ENST00000396695.2 |
SPEG |
SPEG complex locus |
| chr2_-_73053126 | 0.09 |
ENST00000272427.6 ENST00000410104.1 |
EXOC6B |
exocyst complex component 6B |
| chr17_+_39394250 | 0.09 |
ENST00000254072.6 |
KRTAP9-8 |
keratin associated protein 9-8 |
| chr8_+_105235572 | 0.08 |
ENST00000523362.1 |
RIMS2 |
regulating synaptic membrane exocytosis 2 |
| chr8_+_101162812 | 0.08 |
ENST00000353107.3 ENST00000522439.1 |
POLR2K |
polymerase (RNA) II (DNA directed) polypeptide K, 7.0kDa |
| chr17_-_38256973 | 0.08 |
ENST00000246672.3 |
NR1D1 |
nuclear receptor subfamily 1, group D, member 1 |
| chr1_-_11865982 | 0.07 |
ENST00000418034.1 |
MTHFR |
methylenetetrahydrofolate reductase (NAD(P)H) |
| chr15_-_65903574 | 0.07 |
ENST00000420799.2 ENST00000313182.2 ENST00000431261.2 ENST00000442903.3 |
VWA9 |
von Willebrand factor A domain containing 9 |
| chr3_-_182703688 | 0.07 |
ENST00000466812.1 ENST00000487822.1 ENST00000460412.1 ENST00000469954.1 |
DCUN1D1 |
DCN1, defective in cullin neddylation 1, domain containing 1 |
| chr7_+_97736197 | 0.07 |
ENST00000297293.5 |
LMTK2 |
lemur tyrosine kinase 2 |
| chr2_+_113342011 | 0.06 |
ENST00000324913.5 |
CHCHD5 |
coiled-coil-helix-coiled-coil-helix domain containing 5 |
| chr11_+_46740730 | 0.06 |
ENST00000311907.5 ENST00000530231.1 ENST00000442468.1 |
F2 |
coagulation factor II (thrombin) |
| chr17_+_6899366 | 0.06 |
ENST00000251535.6 |
ALOX12 |
arachidonate 12-lipoxygenase |
| chr9_+_101569944 | 0.06 |
ENST00000375011.3 |
GALNT12 |
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 12 (GalNAc-T12) |
| chr2_+_113342163 | 0.06 |
ENST00000409719.1 |
CHCHD5 |
coiled-coil-helix-coiled-coil-helix domain containing 5 |
| chr4_+_1873100 | 0.06 |
ENST00000508803.1 |
WHSC1 |
Wolf-Hirschhorn syndrome candidate 1 |
| chr22_+_37415700 | 0.06 |
ENST00000397129.1 |
MPST |
mercaptopyruvate sulfurtransferase |
| chr15_-_65903407 | 0.06 |
ENST00000395644.4 ENST00000567744.1 ENST00000568573.1 ENST00000562830.1 ENST00000569491.1 ENST00000561769.1 |
VWA9 |
von Willebrand factor A domain containing 9 |
| chr13_+_113622810 | 0.05 |
ENST00000397030.1 |
MCF2L |
MCF.2 cell line derived transforming sequence-like |
| chr1_-_111991908 | 0.05 |
ENST00000235090.5 |
WDR77 |
WD repeat domain 77 |
| chr22_+_37415776 | 0.05 |
ENST00000341116.3 ENST00000429360.2 ENST00000404393.1 |
MPST |
mercaptopyruvate sulfurtransferase |
| chr14_+_24540731 | 0.05 |
ENST00000558859.1 ENST00000559197.1 ENST00000560828.1 ENST00000216775.2 ENST00000560884.1 |
CPNE6 |
copine VI (neuronal) |
| chr22_+_37415728 | 0.05 |
ENST00000404802.3 |
MPST |
mercaptopyruvate sulfurtransferase |
| chr2_+_63816295 | 0.05 |
ENST00000539945.1 ENST00000544381.1 |
MDH1 |
malate dehydrogenase 1, NAD (soluble) |
| chr7_+_45039783 | 0.05 |
ENST00000258781.6 ENST00000541586.1 ENST00000544363.1 |
CCM2 |
cerebral cavernous malformation 2 |
| chr1_+_203764742 | 0.05 |
ENST00000432282.1 ENST00000453771.1 ENST00000367214.1 ENST00000367212.3 ENST00000332127.4 |
ZC3H11A |
zinc finger CCCH-type containing 11A |
| chr2_-_89513402 | 0.05 |
ENST00000498435.1 |
IGKV1-27 |
immunoglobulin kappa variable 1-27 |
| chr6_+_63921351 | 0.05 |
ENST00000370659.1 |
FKBP1C |
FK506 binding protein 1C |
| chr21_-_28820892 | 0.05 |
ENST00000420186.2 |
AP001604.3 |
AP001604.3 |
| chr11_+_125365110 | 0.05 |
ENST00000527818.1 |
AP000708.1 |
AP000708.1 |
| chr1_-_147245484 | 0.05 |
ENST00000271348.2 |
GJA5 |
gap junction protein, alpha 5, 40kDa |
| chr2_+_63816087 | 0.05 |
ENST00000409908.1 ENST00000442225.1 ENST00000409476.1 ENST00000436321.1 |
MDH1 |
malate dehydrogenase 1, NAD (soluble) |
| chr7_-_142630820 | 0.05 |
ENST00000442623.1 ENST00000265310.1 |
TRPV5 |
transient receptor potential cation channel, subfamily V, member 5 |
| chr1_-_156918806 | 0.05 |
ENST00000315174.8 |
ARHGEF11 |
Rho guanine nucleotide exchange factor (GEF) 11 |
| chr10_+_23217006 | 0.04 |
ENST00000376528.4 ENST00000447081.1 |
ARMC3 |
armadillo repeat containing 3 |
| chr22_-_37880543 | 0.04 |
ENST00000442496.1 |
MFNG |
MFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| chr4_-_147043058 | 0.04 |
ENST00000512063.1 ENST00000507726.1 |
RP11-6L6.3 |
long intergenic non-protein coding RNA 1095 |
| chr19_-_14629224 | 0.04 |
ENST00000254322.2 |
DNAJB1 |
DnaJ (Hsp40) homolog, subfamily B, member 1 |
| chr2_+_127656486 | 0.04 |
ENST00000568484.1 ENST00000450035.1 |
AC114783.1 |
Protein LOC339760 |
| chr4_+_88529681 | 0.04 |
ENST00000399271.1 |
DSPP |
dentin sialophosphoprotein |
| chr1_-_11865351 | 0.04 |
ENST00000413656.1 ENST00000376585.1 ENST00000423400.1 ENST00000431243.1 |
MTHFR |
methylenetetrahydrofolate reductase (NAD(P)H) |
| chr14_+_24540046 | 0.04 |
ENST00000397016.2 ENST00000537691.1 ENST00000560356.1 ENST00000558450.1 |
CPNE6 |
copine VI (neuronal) |
| chr1_-_197169672 | 0.04 |
ENST00000367405.4 |
ZBTB41 |
zinc finger and BTB domain containing 41 |
| chr10_+_23216944 | 0.04 |
ENST00000298032.5 ENST00000409983.3 ENST00000409049.3 |
ARMC3 |
armadillo repeat containing 3 |
| chr19_-_4455290 | 0.04 |
ENST00000394765.3 ENST00000592515.1 |
UBXN6 |
UBX domain protein 6 |
| chr11_+_46366918 | 0.04 |
ENST00000528615.1 ENST00000395574.3 |
DGKZ |
diacylglycerol kinase, zeta |
| chrY_+_2803322 | 0.03 |
ENST00000383052.1 ENST00000155093.3 ENST00000449237.1 ENST00000443793.1 |
ZFY |
zinc finger protein, Y-linked |
| chr6_+_31543334 | 0.03 |
ENST00000449264.2 |
TNF |
tumor necrosis factor |
| chr17_+_39411636 | 0.03 |
ENST00000394008.1 |
KRTAP9-9 |
keratin associated protein 9-9 |
| chr17_+_43922241 | 0.03 |
ENST00000329196.5 |
SPPL2C |
signal peptide peptidase like 2C |
| chr10_-_43762329 | 0.03 |
ENST00000395810.1 |
RASGEF1A |
RasGEF domain family, member 1A |
| chr1_-_1690014 | 0.03 |
ENST00000400922.2 ENST00000342348.5 |
NADK |
NAD kinase |
| chrX_+_68725084 | 0.03 |
ENST00000252338.4 |
FAM155B |
family with sequence similarity 155, member B |
| chrX_+_24167746 | 0.03 |
ENST00000428571.1 ENST00000539115.1 |
ZFX |
zinc finger protein, X-linked |
| chrX_+_24167828 | 0.03 |
ENST00000379188.3 ENST00000419690.1 ENST00000379177.1 ENST00000304543.5 |
ZFX |
zinc finger protein, X-linked |
| chr13_+_27844464 | 0.03 |
ENST00000241463.4 |
RASL11A |
RAS-like, family 11, member A |
| chr1_+_159796534 | 0.03 |
ENST00000289707.5 |
SLAMF8 |
SLAM family member 8 |
| chr19_+_21203481 | 0.03 |
ENST00000595401.1 |
ZNF430 |
zinc finger protein 430 |
| chr6_+_167525277 | 0.03 |
ENST00000400926.2 |
CCR6 |
chemokine (C-C motif) receptor 6 |
| chr1_+_111992064 | 0.02 |
ENST00000483994.1 |
ATP5F1 |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit B1 |
| chr19_+_21324863 | 0.02 |
ENST00000598331.1 |
ZNF431 |
zinc finger protein 431 |
| chr2_-_89442621 | 0.02 |
ENST00000492167.1 |
IGKV3-20 |
immunoglobulin kappa variable 3-20 |
| chr4_+_169842707 | 0.02 |
ENST00000503290.1 |
PALLD |
palladin, cytoskeletal associated protein |
| chr2_+_90153696 | 0.02 |
ENST00000417279.2 |
IGKV3D-15 |
immunoglobulin kappa variable 3D-15 (gene/pseudogene) |
| chr12_-_123755639 | 0.02 |
ENST00000535979.1 |
CDK2AP1 |
cyclin-dependent kinase 2 associated protein 1 |
| chr9_-_37592561 | 0.02 |
ENST00000544379.1 ENST00000377773.5 ENST00000401811.3 ENST00000321301.6 |
TOMM5 |
translocase of outer mitochondrial membrane 5 homolog (yeast) |
| chr3_-_45957088 | 0.02 |
ENST00000539217.1 |
LZTFL1 |
leucine zipper transcription factor-like 1 |
| chr2_-_103353277 | 0.02 |
ENST00000258436.5 |
MFSD9 |
major facilitator superfamily domain containing 9 |
| chr11_-_64013663 | 0.02 |
ENST00000392210.2 |
PPP1R14B |
protein phosphatase 1, regulatory (inhibitor) subunit 14B |
| chr1_-_11866034 | 0.02 |
ENST00000376590.3 |
MTHFR |
methylenetetrahydrofolate reductase (NAD(P)H) |
| chr6_-_31938700 | 0.02 |
ENST00000495340.1 |
DXO |
decapping exoribonuclease |
| chr1_+_197382957 | 0.02 |
ENST00000367397.1 |
CRB1 |
crumbs homolog 1 (Drosophila) |
| chr7_+_64126503 | 0.02 |
ENST00000360117.3 |
ZNF107 |
zinc finger protein 107 |
| chr8_+_11561660 | 0.02 |
ENST00000526716.1 ENST00000335135.4 ENST00000528027.1 |
GATA4 |
GATA binding protein 4 |
| chr10_+_90346519 | 0.01 |
ENST00000371939.3 |
LIPJ |
lipase, family member J |
| chr7_-_128695147 | 0.01 |
ENST00000482320.1 ENST00000393245.1 ENST00000471234.1 |
TNPO3 |
transportin 3 |
| chr19_-_19932501 | 0.01 |
ENST00000540806.2 ENST00000590766.1 ENST00000587452.1 ENST00000545006.1 ENST00000590319.1 ENST00000587461.1 ENST00000450683.2 ENST00000443905.2 ENST00000590274.1 |
ZNF506 CTC-559E9.4 |
zinc finger protein 506 CTC-559E9.4 |
| chr11_+_7534999 | 0.01 |
ENST00000528947.1 ENST00000299492.4 |
PPFIBP2 |
PTPRF interacting protein, binding protein 2 (liprin beta 2) |
| chr19_-_43702231 | 0.01 |
ENST00000597374.1 ENST00000599371.1 |
PSG4 |
pregnancy specific beta-1-glycoprotein 4 |
| chr11_-_327537 | 0.01 |
ENST00000602735.1 |
IFITM3 |
interferon induced transmembrane protein 3 |
| chr3_+_58223228 | 0.00 |
ENST00000478253.1 ENST00000295962.4 |
ABHD6 |
abhydrolase domain containing 6 |
| chr5_-_43313574 | 0.00 |
ENST00000325110.6 ENST00000433297.2 |
HMGCS1 |
3-hydroxy-3-methylglutaryl-CoA synthase 1 (soluble) |
| chr19_-_12833164 | 0.00 |
ENST00000356861.5 |
TNPO2 |
transportin 2 |
| chr10_-_105156198 | 0.00 |
ENST00000369815.1 ENST00000309579.3 ENST00000337003.4 |
USMG5 |
up-regulated during skeletal muscle growth 5 homolog (mouse) |
| chr1_-_11007927 | 0.00 |
ENST00000468348.1 |
C1orf127 |
chromosome 1 open reading frame 127 |
| chr19_-_12833361 | 0.00 |
ENST00000592287.1 |
TNPO2 |
transportin 2 |
| chr16_+_77225071 | 0.00 |
ENST00000439557.2 ENST00000545553.1 |
MON1B |
MON1 secretory trafficking family member B |
| chr3_-_48723268 | 0.00 |
ENST00000439518.1 ENST00000416649.2 ENST00000341520.4 ENST00000294129.2 |
NCKIPSD |
NCK interacting protein with SH3 domain |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0097021 | Peyer's patch morphogenesis(GO:0061146) lymphocyte migration into lymphoid organs(GO:0097021) |
| 0.1 | 0.2 | GO:0019858 | cytosine metabolic process(GO:0019858) |
| 0.0 | 0.2 | GO:2000490 | negative regulation of hepatic stellate cell activation(GO:2000490) |
| 0.0 | 0.2 | GO:0008063 | Toll signaling pathway(GO:0008063) |
| 0.0 | 0.2 | GO:0009439 | cyanate metabolic process(GO:0009439) cyanate catabolic process(GO:0009440) |
| 0.0 | 0.0 | GO:0045994 | positive regulation of translational initiation by iron(GO:0045994) |
| 0.0 | 0.2 | GO:0060528 | secretory columnal luminar epithelial cell differentiation involved in prostate glandular acinus development(GO:0060528) |
| 0.0 | 0.1 | GO:0070829 | response to vitamin B2(GO:0033274) heterochromatin maintenance(GO:0070829) |
| 0.0 | 0.1 | GO:0070859 | circadian temperature homeostasis(GO:0060086) positive regulation of bile acid biosynthetic process(GO:0070859) positive regulation of bile acid metabolic process(GO:1904253) |
| 0.0 | 0.1 | GO:0046340 | diacylglycerol catabolic process(GO:0046340) |
| 0.0 | 0.1 | GO:0046985 | positive regulation of hemoglobin biosynthetic process(GO:0046985) |
| 0.0 | 0.1 | GO:2001303 | lipoxin biosynthetic process(GO:2001301) lipoxin A4 metabolic process(GO:2001302) lipoxin A4 biosynthetic process(GO:2001303) |
| 0.0 | 0.6 | GO:0035590 | purinergic nucleotide receptor signaling pathway(GO:0035590) |
| 0.0 | 0.0 | GO:0010652 | pulmonary valve formation(GO:0003193) regulation of cell communication by chemical coupling(GO:0010645) positive regulation of cell communication by chemical coupling(GO:0010652) foramen ovale closure(GO:0035922) regulation of bundle of His cell action potential(GO:0098905) |
| 0.0 | 0.1 | GO:1900738 | cytolysis in other organism involved in symbiotic interaction(GO:0051801) positive regulation of phospholipase C-activating G-protein coupled receptor signaling pathway(GO:1900738) |
| 0.0 | 0.1 | GO:0001878 | response to yeast(GO:0001878) |
| 0.0 | 0.1 | GO:0098814 | spontaneous neurotransmitter secretion(GO:0061669) spontaneous synaptic transmission(GO:0098814) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | GO:0031229 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0004931 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.1 | 0.2 | GO:0047888 | fatty acid peroxidase activity(GO:0047888) |
| 0.0 | 0.1 | GO:0033878 | hormone-sensitive lipase activity(GO:0033878) |
| 0.0 | 0.2 | GO:0016784 | 3-mercaptopyruvate sulfurtransferase activity(GO:0016784) |
| 0.0 | 0.2 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.0 | 0.1 | GO:0004489 | methylenetetrahydrofolate reductase (NAD(P)H) activity(GO:0004489) |
| 0.0 | 0.1 | GO:0016165 | linoleate 13S-lipoxygenase activity(GO:0016165) hepoxilin-epoxide hydrolase activity(GO:0047977) |
| 0.0 | 0.1 | GO:0030060 | L-malate dehydrogenase activity(GO:0030060) |
| 0.0 | 0.1 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.0 | 0.0 | GO:0086020 | gap junction channel activity involved in SA node cell-atrial cardiac muscle cell electrical coupling(GO:0086020) |
| 0.0 | 0.0 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | REACTOME IRAK1 RECRUITS IKK COMPLEX | Genes involved in IRAK1 recruits IKK complex |