Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
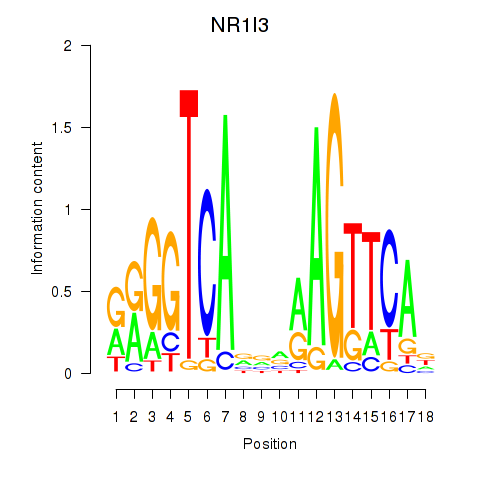
Results for NR1I3
Z-value: 1.14
Transcription factors associated with NR1I3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NR1I3
|
ENSG00000143257.7 | NR1I3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NR1I3 | hg19_v2_chr1_-_161207986_161208000 | 0.61 | 1.1e-01 | Click! |
Activity profile of NR1I3 motif
Sorted Z-values of NR1I3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of NR1I3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_44401479 | 1.27 |
ENST00000438616.3 |
ARTN |
artemin |
| chr6_+_36097992 | 1.17 |
ENST00000211287.4 |
MAPK13 |
mitogen-activated protein kinase 13 |
| chr15_+_43885252 | 1.01 |
ENST00000453782.1 ENST00000300283.6 ENST00000437924.1 ENST00000450086.2 |
CKMT1B |
creatine kinase, mitochondrial 1B |
| chr14_+_21525981 | 1.00 |
ENST00000308227.2 |
RNASE8 |
ribonuclease, RNase A family, 8 |
| chr15_+_43985084 | 0.98 |
ENST00000434505.1 ENST00000411750.1 |
CKMT1A |
creatine kinase, mitochondrial 1A |
| chr1_+_153330322 | 0.97 |
ENST00000368738.3 |
S100A9 |
S100 calcium binding protein A9 |
| chr13_-_20806440 | 0.96 |
ENST00000400066.3 ENST00000400065.3 ENST00000356192.6 |
GJB6 |
gap junction protein, beta 6, 30kDa |
| chr1_+_15250596 | 0.68 |
ENST00000361144.5 |
KAZN |
kazrin, periplakin interacting protein |
| chr1_-_205290865 | 0.56 |
ENST00000367157.3 |
NUAK2 |
NUAK family, SNF1-like kinase, 2 |
| chr12_-_54779511 | 0.51 |
ENST00000551109.1 ENST00000546970.1 |
ZNF385A |
zinc finger protein 385A |
| chr3_-_116163830 | 0.35 |
ENST00000333617.4 |
LSAMP |
limbic system-associated membrane protein |
| chr14_-_22005062 | 0.32 |
ENST00000317492.5 |
SALL2 |
spalt-like transcription factor 2 |
| chr2_-_69180083 | 0.32 |
ENST00000328895.4 |
GKN2 |
gastrokine 2 |
| chr1_-_23886285 | 0.28 |
ENST00000374561.5 |
ID3 |
inhibitor of DNA binding 3, dominant negative helix-loop-helix protein |
| chr14_+_22977587 | 0.27 |
ENST00000390504.1 |
TRAJ33 |
T cell receptor alpha joining 33 |
| chr2_-_69098566 | 0.26 |
ENST00000295379.1 |
BMP10 |
bone morphogenetic protein 10 |
| chr2_-_69180012 | 0.25 |
ENST00000481498.1 |
GKN2 |
gastrokine 2 |
| chr17_-_34270668 | 0.22 |
ENST00000293274.4 |
LYZL6 |
lysozyme-like 6 |
| chr17_-_34270697 | 0.22 |
ENST00000585556.1 |
LYZL6 |
lysozyme-like 6 |
| chr3_-_71802760 | 0.21 |
ENST00000295612.3 |
EIF4E3 |
eukaryotic translation initiation factor 4E family member 3 |
| chr16_+_577697 | 0.21 |
ENST00000562370.1 ENST00000568988.1 ENST00000219611.2 |
CAPN15 |
calpain 15 |
| chr11_+_63137251 | 0.18 |
ENST00000310969.4 ENST00000279178.3 |
SLC22A9 |
solute carrier family 22 (organic anion transporter), member 9 |
| chr19_-_49140609 | 0.17 |
ENST00000601104.1 |
DBP |
D site of albumin promoter (albumin D-box) binding protein |
| chr19_-_44306590 | 0.16 |
ENST00000377950.3 |
LYPD5 |
LY6/PLAUR domain containing 5 |
| chr11_-_59950519 | 0.16 |
ENST00000528851.1 |
MS4A6A |
membrane-spanning 4-domains, subfamily A, member 6A |
| chr7_-_31380502 | 0.15 |
ENST00000297142.3 |
NEUROD6 |
neuronal differentiation 6 |
| chr14_-_22005018 | 0.15 |
ENST00000546363.1 |
SALL2 |
spalt-like transcription factor 2 |
| chr20_+_57766075 | 0.15 |
ENST00000371030.2 |
ZNF831 |
zinc finger protein 831 |
| chr9_+_5450503 | 0.14 |
ENST00000381573.4 ENST00000381577.3 |
CD274 |
CD274 molecule |
| chrX_+_69674943 | 0.14 |
ENST00000542398.1 |
DLG3 |
discs, large homolog 3 (Drosophila) |
| chr8_-_109799793 | 0.13 |
ENST00000297459.3 |
TMEM74 |
transmembrane protein 74 |
| chr4_-_82136114 | 0.13 |
ENST00000395578.1 ENST00000418486.2 |
PRKG2 |
protein kinase, cGMP-dependent, type II |
| chr11_-_57089671 | 0.11 |
ENST00000532437.1 |
TNKS1BP1 |
tankyrase 1 binding protein 1, 182kDa |
| chr16_-_75018968 | 0.11 |
ENST00000262144.6 |
WDR59 |
WD repeat domain 59 |
| chr10_-_50396425 | 0.11 |
ENST00000374148.1 |
C10orf128 |
chromosome 10 open reading frame 128 |
| chr18_-_19283649 | 0.11 |
ENST00000584464.1 ENST00000578270.1 |
ABHD3 |
abhydrolase domain containing 3 |
| chr20_-_44007014 | 0.11 |
ENST00000372726.3 ENST00000537995.1 |
TP53TG5 |
TP53 target 5 |
| chr14_-_77787198 | 0.11 |
ENST00000261534.4 |
POMT2 |
protein-O-mannosyltransferase 2 |
| chr16_-_4465886 | 0.10 |
ENST00000539968.1 |
CORO7 |
coronin 7 |
| chr3_-_116164306 | 0.10 |
ENST00000490035.2 |
LSAMP |
limbic system-associated membrane protein |
| chr11_+_105480740 | 0.10 |
ENST00000393125.2 |
GRIA4 |
glutamate receptor, ionotropic, AMPA 4 |
| chr20_-_45142154 | 0.10 |
ENST00000347606.4 ENST00000457685.2 |
ZNF334 |
zinc finger protein 334 |
| chrX_+_128872998 | 0.09 |
ENST00000371106.3 |
XPNPEP2 |
X-prolyl aminopeptidase (aminopeptidase P) 2, membrane-bound |
| chr12_-_53729525 | 0.09 |
ENST00000303846.3 |
SP7 |
Sp7 transcription factor |
| chr3_+_122103014 | 0.09 |
ENST00000232125.5 ENST00000477892.1 ENST00000469967.1 |
FAM162A |
family with sequence similarity 162, member A |
| chr12_-_118810688 | 0.08 |
ENST00000542532.1 ENST00000392533.3 |
TAOK3 |
TAO kinase 3 |
| chr18_-_19284724 | 0.08 |
ENST00000580981.1 ENST00000289119.2 |
ABHD3 |
abhydrolase domain containing 3 |
| chr19_+_676385 | 0.08 |
ENST00000166139.4 |
FSTL3 |
follistatin-like 3 (secreted glycoprotein) |
| chrX_-_110655391 | 0.08 |
ENST00000356915.2 ENST00000356220.3 |
DCX |
doublecortin |
| chr6_+_32407619 | 0.08 |
ENST00000395388.2 ENST00000374982.5 |
HLA-DRA |
major histocompatibility complex, class II, DR alpha |
| chr9_+_37486005 | 0.08 |
ENST00000377792.3 |
POLR1E |
polymerase (RNA) I polypeptide E, 53kDa |
| chr6_-_43496605 | 0.07 |
ENST00000455285.2 |
XPO5 |
exportin 5 |
| chr18_-_33077556 | 0.07 |
ENST00000589273.1 ENST00000586489.1 |
INO80C |
INO80 complex subunit C |
| chr12_+_100594557 | 0.07 |
ENST00000546902.1 ENST00000552376.1 ENST00000551617.1 |
ACTR6 |
ARP6 actin-related protein 6 homolog (yeast) |
| chr3_-_50396978 | 0.06 |
ENST00000266025.3 |
TMEM115 |
transmembrane protein 115 |
| chr6_-_31509714 | 0.06 |
ENST00000456662.1 ENST00000431908.1 ENST00000456976.1 ENST00000428450.1 ENST00000453105.2 ENST00000418897.1 ENST00000415382.2 ENST00000449074.2 ENST00000419020.1 ENST00000428098.1 |
DDX39B |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 39B |
| chrX_-_15619076 | 0.06 |
ENST00000252519.3 |
ACE2 |
angiotensin I converting enzyme 2 |
| chr2_+_220462560 | 0.06 |
ENST00000456909.1 ENST00000295641.10 |
STK11IP |
serine/threonine kinase 11 interacting protein |
| chr6_+_33422343 | 0.06 |
ENST00000395064.2 |
ZBTB9 |
zinc finger and BTB domain containing 9 |
| chr6_-_34113856 | 0.05 |
ENST00000538487.2 |
GRM4 |
glutamate receptor, metabotropic 4 |
| chr13_-_95131923 | 0.05 |
ENST00000377028.5 ENST00000446125.1 |
DCT |
dopachrome tautomerase |
| chr8_-_53626974 | 0.05 |
ENST00000435644.2 ENST00000518710.1 ENST00000025008.5 ENST00000517963.1 |
RB1CC1 |
RB1-inducible coiled-coil 1 |
| chr6_+_71276596 | 0.05 |
ENST00000370474.3 |
C6orf57 |
chromosome 6 open reading frame 57 |
| chr1_+_43803475 | 0.05 |
ENST00000372470.3 ENST00000413998.2 |
MPL |
myeloproliferative leukemia virus oncogene |
| chr16_-_66968055 | 0.04 |
ENST00000568572.1 |
FAM96B |
family with sequence similarity 96, member B |
| chr2_-_86790593 | 0.04 |
ENST00000263856.4 ENST00000409225.2 |
CHMP3 |
charged multivesicular body protein 3 |
| chr16_+_30710462 | 0.04 |
ENST00000262518.4 ENST00000395059.2 ENST00000344771.4 |
SRCAP |
Snf2-related CREBBP activator protein |
| chr7_-_99381798 | 0.04 |
ENST00000415003.1 ENST00000354593.2 |
CYP3A4 |
cytochrome P450, family 3, subfamily A, polypeptide 4 |
| chr1_+_113217043 | 0.04 |
ENST00000413052.2 |
MOV10 |
Mov10, Moloney leukemia virus 10, homolog (mouse) |
| chr1_+_57320437 | 0.04 |
ENST00000361249.3 |
C8A |
complement component 8, alpha polypeptide |
| chr14_+_77787227 | 0.03 |
ENST00000216465.5 ENST00000361389.4 ENST00000554279.1 ENST00000557639.1 ENST00000349555.3 ENST00000556627.1 ENST00000557053.1 |
GSTZ1 |
glutathione S-transferase zeta 1 |
| chr19_-_49140692 | 0.03 |
ENST00000222122.5 |
DBP |
D site of albumin promoter (albumin D-box) binding protein |
| chr1_-_33336414 | 0.03 |
ENST00000373471.3 ENST00000609187.1 |
FNDC5 |
fibronectin type III domain containing 5 |
| chr7_-_99381884 | 0.03 |
ENST00000336411.2 |
CYP3A4 |
cytochrome P450, family 3, subfamily A, polypeptide 4 |
| chr16_-_16317321 | 0.03 |
ENST00000205557.7 ENST00000575728.1 |
ABCC6 |
ATP-binding cassette, sub-family C (CFTR/MRP), member 6 |
| chr2_+_177015950 | 0.03 |
ENST00000306324.3 |
HOXD4 |
homeobox D4 |
| chr17_-_62502639 | 0.02 |
ENST00000225792.5 ENST00000581697.1 ENST00000584279.1 ENST00000577922.1 |
DDX5 |
DEAD (Asp-Glu-Ala-Asp) box helicase 5 |
| chr6_+_108487245 | 0.02 |
ENST00000368986.4 |
NR2E1 |
nuclear receptor subfamily 2, group E, member 1 |
| chr2_-_128642434 | 0.02 |
ENST00000393001.1 |
AMMECR1L |
AMMECR1-like |
| chr3_-_58419537 | 0.02 |
ENST00000474765.1 ENST00000485460.1 ENST00000302746.6 ENST00000383714.4 |
PDHB |
pyruvate dehydrogenase (lipoamide) beta |
| chr19_-_46296011 | 0.02 |
ENST00000377735.3 ENST00000270223.6 |
DMWD |
dystrophia myotonica, WD repeat containing |
| chr15_+_85523671 | 0.02 |
ENST00000310298.4 ENST00000557957.1 |
PDE8A |
phosphodiesterase 8A |
| chr10_+_88414338 | 0.02 |
ENST00000241891.5 ENST00000443292.1 |
OPN4 |
opsin 4 |
| chr16_+_67906919 | 0.02 |
ENST00000358933.5 |
EDC4 |
enhancer of mRNA decapping 4 |
| chr19_+_58919992 | 0.02 |
ENST00000306910.4 ENST00000598901.1 ENST00000593920.1 ENST00000596281.1 |
ZNF584 |
zinc finger protein 584 |
| chr1_-_151119087 | 0.01 |
ENST00000341697.3 ENST00000368914.3 |
SEMA6C |
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6C |
| chr19_+_12203069 | 0.01 |
ENST00000430298.2 ENST00000339302.4 |
ZNF788 ZNF788 |
zinc finger family member 788 Zinc finger protein 788 |
| chr6_+_96969672 | 0.01 |
ENST00000369278.4 |
UFL1 |
UFM1-specific ligase 1 |
| chr16_-_66968265 | 0.00 |
ENST00000567511.1 ENST00000422424.2 |
FAM96B |
family with sequence similarity 96, member B |
| chr10_-_50396407 | 0.00 |
ENST00000374153.2 ENST00000374151.3 |
C10orf128 |
chromosome 10 open reading frame 128 |
| chr13_-_21635631 | 0.00 |
ENST00000382592.4 |
LATS2 |
large tumor suppressor kinase 2 |
| chr7_+_101460882 | 0.00 |
ENST00000292535.7 ENST00000549414.2 ENST00000550008.2 ENST00000546411.2 ENST00000556210.1 |
CUX1 |
cut-like homeobox 1 |
| chr9_-_80646374 | 0.00 |
ENST00000286548.4 |
GNAQ |
guanine nucleotide binding protein (G protein), q polypeptide |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.0 | GO:0035606 | peptidyl-cysteine S-trans-nitrosylation(GO:0035606) |
| 0.3 | 1.3 | GO:0061146 | Peyer's patch morphogenesis(GO:0061146) lymphocyte migration into lymphoid organs(GO:0097021) |
| 0.2 | 1.0 | GO:0061760 | antifungal innate immune response(GO:0061760) |
| 0.1 | 0.5 | GO:1902162 | mRNA localization resulting in posttranscriptional regulation of gene expression(GO:0010609) regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902162) positive regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902164) |
| 0.1 | 1.0 | GO:0006600 | creatine metabolic process(GO:0006600) |
| 0.0 | 0.1 | GO:2001226 | negative regulation of chloride transport(GO:2001226) |
| 0.0 | 0.3 | GO:0060298 | positive regulation of sarcomere organization(GO:0060298) |
| 0.0 | 0.4 | GO:0016998 | cell wall macromolecule catabolic process(GO:0016998) |
| 0.0 | 0.1 | GO:0035281 | pre-miRNA export from nucleus(GO:0035281) |
| 0.0 | 0.4 | GO:0035641 | locomotory exploration behavior(GO:0035641) |
| 0.0 | 0.1 | GO:1904100 | regulation of protein O-linked glycosylation(GO:1904098) positive regulation of protein O-linked glycosylation(GO:1904100) |
| 0.0 | 0.1 | GO:2001181 | positive regulation of interleukin-10 secretion(GO:2001181) |
| 0.0 | 0.1 | GO:0002503 | peptide antigen assembly with MHC class II protein complex(GO:0002503) |
| 0.0 | 0.1 | GO:0003051 | angiotensin-mediated drinking behavior(GO:0003051) |
| 0.0 | 0.1 | GO:0001189 | RNA polymerase I transcriptional preinitiation complex assembly(GO:0001188) RNA polymerase I transcriptional preinitiation complex assembly at the promoter for the nuclear large rRNA transcript(GO:0001189) |
| 0.0 | 0.1 | GO:0036378 | alkaloid catabolic process(GO:0009822) calcitriol biosynthetic process from calciol(GO:0036378) |
| 0.0 | 0.0 | GO:0038163 | thrombopoietin-mediated signaling pathway(GO:0038163) |
| 0.0 | 1.0 | GO:0051602 | response to electrical stimulus(GO:0051602) |
| 0.0 | 0.2 | GO:0015747 | urate transport(GO:0015747) |
| 0.0 | 0.3 | GO:0030903 | notochord development(GO:0030903) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.0 | GO:0005922 | connexon complex(GO:0005922) |
| 0.0 | 0.4 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 0.0 | 0.1 | GO:0042565 | RNA nuclear export complex(GO:0042565) |
| 0.0 | 0.7 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 0.5 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.0 | 0.2 | GO:0005845 | mRNA cap binding complex(GO:0005845) RNA cap binding complex(GO:0034518) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.0 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.1 | 1.0 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.1 | 1.0 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 0.0 | 1.2 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.0 | 0.2 | GO:0052740 | 1-acyl-2-lysophosphatidylserine acylhydrolase activity(GO:0052740) |
| 0.0 | 0.4 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.0 | 0.1 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.0 | 0.1 | GO:0090631 | pre-miRNA transporter activity(GO:0090631) |
| 0.0 | 0.3 | GO:0031433 | telethonin binding(GO:0031433) |
| 0.0 | 0.1 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.0 | 0.1 | GO:0070643 | vitamin D3 25-hydroxylase activity(GO:0030343) vitamin D 25-hydroxylase activity(GO:0070643) |
| 0.0 | 0.2 | GO:1901702 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.0 | 0.1 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.0 | 0.1 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.2 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 1.0 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.2 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.0 | 1.0 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.0 | 1.3 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 0.2 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |