Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
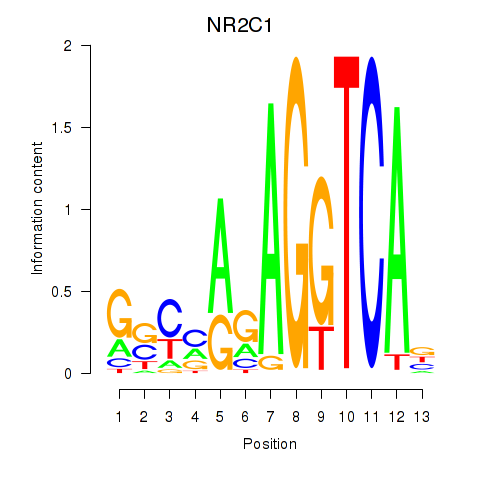
Results for NR2C1
Z-value: 0.60
Transcription factors associated with NR2C1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NR2C1
|
ENSG00000120798.12 | NR2C1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NR2C1 | hg19_v2_chr12_-_95467267_95467350 | -0.79 | 1.9e-02 | Click! |
Activity profile of NR2C1 motif
Sorted Z-values of NR2C1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of NR2C1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_+_70404996 | 0.50 |
ENST00000402687.4 ENST00000419716.3 |
SULF1 |
sulfatase 1 |
| chr13_-_33760216 | 0.48 |
ENST00000255486.4 |
STARD13 |
StAR-related lipid transfer (START) domain containing 13 |
| chr2_+_202316392 | 0.40 |
ENST00000194530.3 ENST00000392249.2 |
STRADB |
STE20-related kinase adaptor beta |
| chr2_-_38303218 | 0.39 |
ENST00000407341.1 ENST00000260630.3 |
CYP1B1 |
cytochrome P450, family 1, subfamily B, polypeptide 1 |
| chr9_-_35685452 | 0.29 |
ENST00000607559.1 |
TPM2 |
tropomyosin 2 (beta) |
| chr2_-_202316260 | 0.28 |
ENST00000332624.3 |
TRAK2 |
trafficking protein, kinesin binding 2 |
| chr12_-_6665200 | 0.25 |
ENST00000336604.4 ENST00000396840.2 ENST00000356896.4 |
IFFO1 |
intermediate filament family orphan 1 |
| chr1_+_159750776 | 0.22 |
ENST00000368107.1 |
DUSP23 |
dual specificity phosphatase 23 |
| chr15_+_75335604 | 0.22 |
ENST00000563393.1 |
PPCDC |
phosphopantothenoylcysteine decarboxylase |
| chr1_+_145727681 | 0.22 |
ENST00000417171.1 ENST00000451928.2 |
PDZK1 |
PDZ domain containing 1 |
| chr1_+_159750720 | 0.22 |
ENST00000368109.1 ENST00000368108.3 |
DUSP23 |
dual specificity phosphatase 23 |
| chr14_+_74035763 | 0.21 |
ENST00000238651.5 |
ACOT2 |
acyl-CoA thioesterase 2 |
| chr14_+_74003818 | 0.20 |
ENST00000311148.4 |
ACOT1 |
acyl-CoA thioesterase 1 |
| chr2_+_109237717 | 0.19 |
ENST00000409441.1 |
LIMS1 |
LIM and senescent cell antigen-like domains 1 |
| chr20_-_44485835 | 0.19 |
ENST00000457981.1 ENST00000426915.1 ENST00000217455.4 |
ACOT8 |
acyl-CoA thioesterase 8 |
| chr12_-_48500085 | 0.19 |
ENST00000549518.1 |
SENP1 |
SUMO1/sentrin specific peptidase 1 |
| chr2_-_219433014 | 0.18 |
ENST00000418019.1 ENST00000454775.1 ENST00000338465.5 ENST00000415516.1 ENST00000258399.3 |
USP37 |
ubiquitin specific peptidase 37 |
| chr15_-_83474806 | 0.18 |
ENST00000541889.1 ENST00000334574.8 ENST00000561368.1 |
FSD2 |
fibronectin type III and SPRY domain containing 2 |
| chr11_-_67275542 | 0.18 |
ENST00000531506.1 |
CDK2AP2 |
cyclin-dependent kinase 2 associated protein 2 |
| chr22_-_50699701 | 0.18 |
ENST00000395780.1 |
MAPK12 |
mitogen-activated protein kinase 12 |
| chr3_-_50605077 | 0.17 |
ENST00000426034.1 ENST00000441239.1 |
C3orf18 |
chromosome 3 open reading frame 18 |
| chr12_-_54070098 | 0.17 |
ENST00000394349.3 ENST00000549164.1 |
ATP5G2 |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit C2 (subunit 9) |
| chr12_-_54069856 | 0.17 |
ENST00000602871.1 |
ATP5G2 |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit C2 (subunit 9) |
| chr13_+_100741269 | 0.17 |
ENST00000376286.4 ENST00000376279.3 ENST00000376285.1 |
PCCA |
propionyl CoA carboxylase, alpha polypeptide |
| chr7_+_100770328 | 0.16 |
ENST00000223095.4 ENST00000445463.2 |
SERPINE1 |
serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 |
| chr9_+_33795533 | 0.16 |
ENST00000379405.3 |
PRSS3 |
protease, serine, 3 |
| chr3_-_58613323 | 0.16 |
ENST00000474531.1 ENST00000465970.1 |
FAM107A |
family with sequence similarity 107, member A |
| chr6_+_57037089 | 0.16 |
ENST00000370693.5 |
BAG2 |
BCL2-associated athanogene 2 |
| chr22_+_47158518 | 0.14 |
ENST00000337137.4 ENST00000380995.1 ENST00000407381.3 |
TBC1D22A |
TBC1 domain family, member 22A |
| chr19_-_49371711 | 0.14 |
ENST00000355496.5 ENST00000263265.6 |
PLEKHA4 |
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 4 |
| chr2_+_46926048 | 0.14 |
ENST00000306503.5 |
SOCS5 |
suppressor of cytokine signaling 5 |
| chr5_+_1801503 | 0.13 |
ENST00000274137.5 ENST00000469176.1 |
NDUFS6 |
NADH dehydrogenase (ubiquinone) Fe-S protein 6, 13kDa (NADH-coenzyme Q reductase) |
| chr17_+_4853442 | 0.13 |
ENST00000522301.1 |
ENO3 |
enolase 3 (beta, muscle) |
| chr12_-_48499591 | 0.13 |
ENST00000551330.1 ENST00000004980.5 ENST00000339976.6 ENST00000448372.1 |
SENP1 |
SUMO1/sentrin specific peptidase 1 |
| chrX_-_40036520 | 0.13 |
ENST00000406200.2 ENST00000378455.4 ENST00000342274.4 |
BCOR |
BCL6 corepressor |
| chr22_+_47158578 | 0.12 |
ENST00000355704.3 |
TBC1D22A |
TBC1 domain family, member 22A |
| chr14_+_105212297 | 0.12 |
ENST00000556623.1 ENST00000555674.1 |
ADSSL1 |
adenylosuccinate synthase like 1 |
| chr1_-_55089191 | 0.12 |
ENST00000302250.2 ENST00000371304.2 |
FAM151A |
family with sequence similarity 151, member A |
| chr2_+_201936458 | 0.12 |
ENST00000237889.4 |
NDUFB3 |
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 3, 12kDa |
| chr2_+_46926326 | 0.12 |
ENST00000394861.2 |
SOCS5 |
suppressor of cytokine signaling 5 |
| chr13_+_21714653 | 0.12 |
ENST00000382533.4 |
SAP18 |
Sin3A-associated protein, 18kDa |
| chr22_+_30163340 | 0.12 |
ENST00000330029.6 ENST00000401406.3 |
UQCR10 |
ubiquinol-cytochrome c reductase, complex III subunit X |
| chr1_-_229569834 | 0.11 |
ENST00000366684.3 ENST00000366683.2 |
ACTA1 |
actin, alpha 1, skeletal muscle |
| chr19_+_10197463 | 0.11 |
ENST00000590378.1 ENST00000397881.3 |
C19orf66 |
chromosome 19 open reading frame 66 |
| chr6_-_160148356 | 0.11 |
ENST00000401980.3 ENST00000545162.1 |
SOD2 |
superoxide dismutase 2, mitochondrial |
| chr11_-_66206260 | 0.11 |
ENST00000329819.4 ENST00000310999.7 ENST00000430466.2 |
MRPL11 |
mitochondrial ribosomal protein L11 |
| chr20_-_57607347 | 0.11 |
ENST00000395663.1 ENST00000395659.1 ENST00000243997.3 |
ATP5E |
ATP synthase, H+ transporting, mitochondrial F1 complex, epsilon subunit |
| chr13_+_21714711 | 0.11 |
ENST00000607003.1 ENST00000492245.1 |
SAP18 |
Sin3A-associated protein, 18kDa |
| chr2_+_120124497 | 0.10 |
ENST00000355857.3 ENST00000535617.1 ENST00000535757.1 ENST00000409094.1 ENST00000311521.4 |
DBI |
diazepam binding inhibitor (GABA receptor modulator, acyl-CoA binding protein) |
| chr8_+_96037255 | 0.10 |
ENST00000286687.4 |
NDUFAF6 |
NADH dehydrogenase (ubiquinone) complex I, assembly factor 6 |
| chr2_+_120125245 | 0.10 |
ENST00000393103.2 |
DBI |
diazepam binding inhibitor (GABA receptor modulator, acyl-CoA binding protein) |
| chr9_+_135037334 | 0.10 |
ENST00000393229.3 ENST00000360670.3 ENST00000393228.4 ENST00000372179.3 |
NTNG2 |
netrin G2 |
| chr11_+_67798363 | 0.10 |
ENST00000525628.1 |
NDUFS8 |
NADH dehydrogenase (ubiquinone) Fe-S protein 8, 23kDa (NADH-coenzyme Q reductase) |
| chr2_-_42721110 | 0.10 |
ENST00000394973.4 ENST00000306078.1 |
KCNG3 |
potassium voltage-gated channel, subfamily G, member 3 |
| chr1_-_45140074 | 0.10 |
ENST00000420706.1 ENST00000372235.3 ENST00000372242.3 ENST00000372243.3 ENST00000372244.3 |
TMEM53 |
transmembrane protein 53 |
| chr8_+_96037205 | 0.10 |
ENST00000396124.4 |
NDUFAF6 |
NADH dehydrogenase (ubiquinone) complex I, assembly factor 6 |
| chr2_-_163100045 | 0.09 |
ENST00000188790.4 |
FAP |
fibroblast activation protein, alpha |
| chr12_-_42631529 | 0.09 |
ENST00000548917.1 |
YAF2 |
YY1 associated factor 2 |
| chr7_-_38948774 | 0.09 |
ENST00000395969.2 ENST00000414632.1 ENST00000310301.4 |
VPS41 |
vacuolar protein sorting 41 homolog (S. cerevisiae) |
| chr11_-_73687997 | 0.09 |
ENST00000545212.1 |
UCP2 |
uncoupling protein 2 (mitochondrial, proton carrier) |
| chr2_-_163099885 | 0.09 |
ENST00000443424.1 |
FAP |
fibroblast activation protein, alpha |
| chr2_-_88427568 | 0.09 |
ENST00000393750.3 ENST00000295834.3 |
FABP1 |
fatty acid binding protein 1, liver |
| chr4_+_26321284 | 0.09 |
ENST00000506956.1 ENST00000512671.1 ENST00000345843.3 ENST00000342295.1 |
RBPJ |
recombination signal binding protein for immunoglobulin kappa J region |
| chr3_-_50605150 | 0.09 |
ENST00000357203.3 |
C3orf18 |
chromosome 3 open reading frame 18 |
| chr6_-_160147925 | 0.09 |
ENST00000535561.1 |
SOD2 |
superoxide dismutase 2, mitochondrial |
| chr20_+_30946106 | 0.09 |
ENST00000375687.4 ENST00000542461.1 |
ASXL1 |
additional sex combs like 1 (Drosophila) |
| chr19_+_10196981 | 0.09 |
ENST00000591813.1 |
C19orf66 |
chromosome 19 open reading frame 66 |
| chr12_+_96252706 | 0.08 |
ENST00000266735.5 ENST00000553192.1 ENST00000552085.1 |
SNRPF |
small nuclear ribonucleoprotein polypeptide F |
| chr11_+_67798114 | 0.08 |
ENST00000453471.2 ENST00000528492.1 ENST00000526339.1 ENST00000525419.1 |
NDUFS8 |
NADH dehydrogenase (ubiquinone) Fe-S protein 8, 23kDa (NADH-coenzyme Q reductase) |
| chr4_+_159593418 | 0.08 |
ENST00000507475.1 ENST00000307738.5 |
ETFDH |
electron-transferring-flavoprotein dehydrogenase |
| chr19_+_44100632 | 0.08 |
ENST00000533118.1 |
ZNF576 |
zinc finger protein 576 |
| chr1_+_25071848 | 0.08 |
ENST00000374379.4 |
CLIC4 |
chloride intracellular channel 4 |
| chr3_+_159557637 | 0.08 |
ENST00000445224.2 |
SCHIP1 |
schwannomin interacting protein 1 |
| chr12_-_42632016 | 0.08 |
ENST00000442791.3 ENST00000327791.4 ENST00000534854.2 ENST00000380788.3 ENST00000380790.4 |
YAF2 |
YY1 associated factor 2 |
| chr1_-_155881156 | 0.08 |
ENST00000539040.1 ENST00000368323.3 |
RIT1 |
Ras-like without CAAX 1 |
| chr13_+_103046954 | 0.08 |
ENST00000606448.1 |
FGF14-AS2 |
FGF14 antisense RNA 2 |
| chr1_-_173886491 | 0.07 |
ENST00000367698.3 |
SERPINC1 |
serpin peptidase inhibitor, clade C (antithrombin), member 1 |
| chr19_-_38806390 | 0.07 |
ENST00000589247.1 ENST00000329420.8 ENST00000591784.1 |
YIF1B |
Yip1 interacting factor homolog B (S. cerevisiae) |
| chr19_+_48828582 | 0.07 |
ENST00000270221.6 ENST00000596315.1 |
EMP3 |
epithelial membrane protein 3 |
| chr6_+_30594619 | 0.07 |
ENST00000318999.7 ENST00000376485.4 ENST00000376478.2 ENST00000319027.5 ENST00000376483.4 ENST00000329992.8 ENST00000330083.5 |
ATAT1 |
alpha tubulin acetyltransferase 1 |
| chr11_+_131781290 | 0.07 |
ENST00000425719.2 ENST00000374784.1 |
NTM |
neurotrimin |
| chr19_+_48828788 | 0.07 |
ENST00000594198.1 ENST00000597279.1 ENST00000593437.1 |
EMP3 |
epithelial membrane protein 3 |
| chr1_-_31381528 | 0.07 |
ENST00000339394.6 |
SDC3 |
syndecan 3 |
| chr4_-_99064387 | 0.07 |
ENST00000295268.3 |
STPG2 |
sperm-tail PG-rich repeat containing 2 |
| chr20_+_37590942 | 0.07 |
ENST00000373325.2 ENST00000252011.3 ENST00000373323.4 |
DHX35 |
DEAH (Asp-Glu-Ala-His) box polypeptide 35 |
| chr16_+_2255841 | 0.07 |
ENST00000301725.7 |
MLST8 |
MTOR associated protein, LST8 homolog (S. cerevisiae) |
| chr17_+_80674559 | 0.07 |
ENST00000269373.6 ENST00000535965.1 ENST00000577128.1 ENST00000573158.1 |
FN3KRP |
fructosamine 3 kinase related protein |
| chr19_-_38806540 | 0.06 |
ENST00000592694.1 |
YIF1B |
Yip1 interacting factor homolog B (S. cerevisiae) |
| chr1_-_45253377 | 0.06 |
ENST00000372207.3 |
BEST4 |
bestrophin 4 |
| chr17_-_15902951 | 0.06 |
ENST00000472495.1 |
ZSWIM7 |
zinc finger, SWIM-type containing 7 |
| chr12_-_63544718 | 0.06 |
ENST00000299178.2 |
AVPR1A |
arginine vasopressin receptor 1A |
| chr1_-_45140227 | 0.06 |
ENST00000372237.3 |
TMEM53 |
transmembrane protein 53 |
| chr11_+_67798090 | 0.06 |
ENST00000313468.5 |
NDUFS8 |
NADH dehydrogenase (ubiquinone) Fe-S protein 8, 23kDa (NADH-coenzyme Q reductase) |
| chr1_+_40713573 | 0.06 |
ENST00000372766.3 |
TMCO2 |
transmembrane and coiled-coil domains 2 |
| chr17_+_37821593 | 0.06 |
ENST00000578283.1 |
TCAP |
titin-cap |
| chr6_+_54172653 | 0.06 |
ENST00000370869.3 |
TINAG |
tubulointerstitial nephritis antigen |
| chr19_-_38806560 | 0.06 |
ENST00000591755.1 ENST00000337679.8 ENST00000339413.6 |
YIF1B |
Yip1 interacting factor homolog B (S. cerevisiae) |
| chr22_-_38851205 | 0.06 |
ENST00000303592.3 |
KCNJ4 |
potassium inwardly-rectifying channel, subfamily J, member 4 |
| chr6_+_31515337 | 0.06 |
ENST00000376148.4 ENST00000376145.4 |
NFKBIL1 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor-like 1 |
| chr12_+_109577202 | 0.05 |
ENST00000377848.3 ENST00000377854.5 |
ACACB |
acetyl-CoA carboxylase beta |
| chr17_-_15903002 | 0.05 |
ENST00000399277.1 |
ZSWIM7 |
zinc finger, SWIM-type containing 7 |
| chr11_-_64013663 | 0.05 |
ENST00000392210.2 |
PPP1R14B |
protein phosphatase 1, regulatory (inhibitor) subunit 14B |
| chr16_+_2255710 | 0.05 |
ENST00000397124.1 ENST00000565250.1 |
MLST8 |
MTOR associated protein, LST8 homolog (S. cerevisiae) |
| chrX_-_128657457 | 0.05 |
ENST00000371121.3 ENST00000371123.1 ENST00000371122.4 |
SMARCA1 |
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 1 |
| chr17_-_15902903 | 0.05 |
ENST00000486655.1 |
ZSWIM7 |
zinc finger, SWIM-type containing 7 |
| chr1_-_155880672 | 0.05 |
ENST00000609492.1 ENST00000368322.3 |
RIT1 |
Ras-like without CAAX 1 |
| chr19_-_41256207 | 0.05 |
ENST00000598485.2 ENST00000470681.1 ENST00000339153.3 ENST00000598729.1 |
C19orf54 |
chromosome 19 open reading frame 54 |
| chr7_-_229557 | 0.05 |
ENST00000514988.1 |
AC145676.2 |
Uncharacterized protein |
| chr8_+_77318769 | 0.05 |
ENST00000518732.1 |
RP11-706J10.1 |
long intergenic non-protein coding RNA 1111 |
| chr12_-_22487618 | 0.05 |
ENST00000404299.3 ENST00000396037.4 |
ST8SIA1 |
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 1 |
| chr3_-_113464906 | 0.05 |
ENST00000477813.1 |
NAA50 |
N(alpha)-acetyltransferase 50, NatE catalytic subunit |
| chr12_-_90049828 | 0.04 |
ENST00000261173.2 ENST00000348959.3 |
ATP2B1 |
ATPase, Ca++ transporting, plasma membrane 1 |
| chr11_-_64013288 | 0.04 |
ENST00000542235.1 |
PPP1R14B |
protein phosphatase 1, regulatory (inhibitor) subunit 14B |
| chr19_-_10679697 | 0.04 |
ENST00000335766.2 |
CDKN2D |
cyclin-dependent kinase inhibitor 2D (p19, inhibits CDK4) |
| chr1_+_160097462 | 0.04 |
ENST00000447527.1 |
ATP1A2 |
ATPase, Na+/K+ transporting, alpha 2 polypeptide |
| chr4_-_76598296 | 0.04 |
ENST00000395719.3 |
G3BP2 |
GTPase activating protein (SH3 domain) binding protein 2 |
| chr3_+_183903811 | 0.04 |
ENST00000429586.2 ENST00000292808.5 |
ABCF3 |
ATP-binding cassette, sub-family F (GCN20), member 3 |
| chr5_-_42811986 | 0.04 |
ENST00000511224.1 ENST00000507920.1 ENST00000510965.1 |
SEPP1 |
selenoprotein P, plasma, 1 |
| chr1_-_101491319 | 0.04 |
ENST00000342173.7 ENST00000488176.1 ENST00000370109.3 |
DPH5 |
diphthamide biosynthesis 5 |
| chr15_-_43802769 | 0.04 |
ENST00000263801.3 |
TP53BP1 |
tumor protein p53 binding protein 1 |
| chr12_-_90049878 | 0.04 |
ENST00000359142.3 |
ATP2B1 |
ATPase, Ca++ transporting, plasma membrane 1 |
| chr2_-_164592497 | 0.03 |
ENST00000333129.3 ENST00000409634.1 |
FIGN |
fidgetin |
| chr15_-_65715401 | 0.03 |
ENST00000352385.2 |
IGDCC4 |
immunoglobulin superfamily, DCC subclass, member 4 |
| chr2_-_154335300 | 0.03 |
ENST00000325926.3 |
RPRM |
reprimo, TP53 dependent G2 arrest mediator candidate |
| chr6_+_10585979 | 0.03 |
ENST00000265012.4 |
GCNT2 |
glucosaminyl (N-acetyl) transferase 2, I-branching enzyme (I blood group) |
| chr6_+_160148593 | 0.03 |
ENST00000337387.4 |
WTAP |
Wilms tumor 1 associated protein |
| chr2_-_25194963 | 0.03 |
ENST00000264711.2 |
DNAJC27 |
DnaJ (Hsp40) homolog, subfamily C, member 27 |
| chr10_+_63808970 | 0.03 |
ENST00000309334.5 |
ARID5B |
AT rich interactive domain 5B (MRF1-like) |
| chr20_-_30458432 | 0.03 |
ENST00000375966.4 ENST00000278979.3 |
DUSP15 |
dual specificity phosphatase 15 |
| chr17_+_73997419 | 0.02 |
ENST00000425876.2 |
CDK3 |
cyclin-dependent kinase 3 |
| chr1_-_226374373 | 0.02 |
ENST00000366812.5 |
ACBD3 |
acyl-CoA binding domain containing 3 |
| chr11_-_116663127 | 0.02 |
ENST00000433069.1 ENST00000542499.1 |
APOA5 |
apolipoprotein A-V |
| chr16_-_4292071 | 0.02 |
ENST00000399609.3 |
SRL |
sarcalumenin |
| chr9_-_139839064 | 0.02 |
ENST00000325285.3 ENST00000428398.1 |
FBXW5 |
F-box and WD repeat domain containing 5 |
| chr14_+_42077552 | 0.02 |
ENST00000554120.1 |
LRFN5 |
leucine rich repeat and fibronectin type III domain containing 5 |
| chr16_+_83932684 | 0.02 |
ENST00000262430.4 |
MLYCD |
malonyl-CoA decarboxylase |
| chr2_+_210636697 | 0.02 |
ENST00000439458.1 ENST00000272845.6 |
UNC80 |
unc-80 homolog (C. elegans) |
| chr11_-_26743546 | 0.02 |
ENST00000280467.6 ENST00000396005.3 |
SLC5A12 |
solute carrier family 5 (sodium/monocarboxylate cotransporter), member 12 |
| chr13_+_21714913 | 0.02 |
ENST00000450573.1 ENST00000467636.1 |
SAP18 |
Sin3A-associated protein, 18kDa |
| chr19_+_4153598 | 0.02 |
ENST00000078445.2 ENST00000252587.3 ENST00000595923.1 ENST00000602257.1 ENST00000602147.1 |
CREB3L3 |
cAMP responsive element binding protein 3-like 3 |
| chr20_+_44486246 | 0.01 |
ENST00000255152.2 ENST00000454862.2 |
ZSWIM3 |
zinc finger, SWIM-type containing 3 |
| chr3_-_150264272 | 0.01 |
ENST00000491660.1 ENST00000487153.1 ENST00000239944.2 |
SERP1 |
stress-associated endoplasmic reticulum protein 1 |
| chr2_+_201936707 | 0.01 |
ENST00000433898.1 ENST00000454214.1 |
NDUFB3 |
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 3, 12kDa |
| chr16_-_70729496 | 0.01 |
ENST00000567648.1 |
VAC14 |
Vac14 homolog (S. cerevisiae) |
| chr3_+_46538981 | 0.01 |
ENST00000296142.3 |
RTP3 |
receptor (chemosensory) transporter protein 3 |
| chr5_-_42812143 | 0.01 |
ENST00000514985.1 |
SEPP1 |
selenoprotein P, plasma, 1 |
| chr2_-_219134343 | 0.01 |
ENST00000447885.1 ENST00000420660.1 |
AAMP |
angio-associated, migratory cell protein |
| chr7_+_120969045 | 0.01 |
ENST00000222462.2 |
WNT16 |
wingless-type MMTV integration site family, member 16 |
| chr3_+_113465866 | 0.01 |
ENST00000273398.3 ENST00000538620.1 ENST00000496747.1 ENST00000475322.1 |
ATP6V1A |
ATPase, H+ transporting, lysosomal 70kDa, V1 subunit A |
| chr8_-_30706608 | 0.01 |
ENST00000256246.2 |
TEX15 |
testis expressed 15 |
| chr3_+_184055240 | 0.01 |
ENST00000383847.2 |
FAM131A |
family with sequence similarity 131, member A |
| chr19_+_44100544 | 0.01 |
ENST00000391965.2 ENST00000525771.1 |
ZNF576 |
zinc finger protein 576 |
| chr21_-_32931290 | 0.01 |
ENST00000286827.3 |
TIAM1 |
T-cell lymphoma invasion and metastasis 1 |
| chr3_+_37284824 | 0.01 |
ENST00000431105.1 |
GOLGA4 |
golgin A4 |
| chrX_-_132095419 | 0.00 |
ENST00000370836.2 ENST00000521489.1 |
HS6ST2 |
heparan sulfate 6-O-sulfotransferase 2 |
| chr10_-_23633720 | 0.00 |
ENST00000323327.4 |
C10orf67 |
chromosome 10 open reading frame 67 |
| chr17_-_39203519 | 0.00 |
ENST00000542137.1 ENST00000391419.3 |
KRTAP2-1 |
keratin associated protein 2-1 |
| chr16_-_88752889 | 0.00 |
ENST00000332281.5 |
SNAI3 |
snail family zinc finger 3 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0014846 | esophagus smooth muscle contraction(GO:0014846) |
| 0.1 | 0.4 | GO:0061304 | retinal blood vessel morphogenesis(GO:0061304) |
| 0.1 | 0.3 | GO:0098937 | dendritic transport(GO:0098935) anterograde dendritic transport(GO:0098937) |
| 0.1 | 0.5 | GO:0097498 | endothelial tube lumen extension(GO:0097498) |
| 0.1 | 0.2 | GO:2000097 | chronological cell aging(GO:0001300) regulation of smooth muscle cell-matrix adhesion(GO:2000097) |
| 0.0 | 0.2 | GO:0097325 | melanocyte proliferation(GO:0097325) |
| 0.0 | 0.1 | GO:0000415 | negative regulation of histone H3-K36 methylation(GO:0000415) |
| 0.0 | 0.2 | GO:0003069 | age-dependent response to oxidative stress(GO:0001306) age-dependent response to reactive oxygen species(GO:0001315) regulation of systemic arterial blood pressure by acetylcholine(GO:0003068) vasodilation by acetylcholine involved in regulation of systemic arterial blood pressure(GO:0003069) regulation of systemic arterial blood pressure by neurotransmitter(GO:0003070) age-dependent general metabolic decline(GO:0007571) |
| 0.0 | 0.3 | GO:0010724 | regulation of definitive erythrocyte differentiation(GO:0010724) |
| 0.0 | 0.3 | GO:0045629 | negative regulation of T-helper 2 cell differentiation(GO:0045629) negative regulation of monocyte chemotactic protein-1 production(GO:0071638) |
| 0.0 | 0.1 | GO:1905068 | positive regulation of ephrin receptor signaling pathway(GO:1901189) positive regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901297) positive regulation of canonical Wnt signaling pathway involved in heart development(GO:1905068) |
| 0.0 | 0.1 | GO:0035359 | negative regulation of peroxisome proliferator activated receptor signaling pathway(GO:0035359) |
| 0.0 | 0.2 | GO:0019626 | short-chain fatty acid catabolic process(GO:0019626) |
| 0.0 | 0.2 | GO:2000035 | regulation of stem cell division(GO:2000035) |
| 0.0 | 0.1 | GO:0044208 | 'de novo' AMP biosynthetic process(GO:0044208) |
| 0.0 | 0.1 | GO:0031394 | maternal aggressive behavior(GO:0002125) positive regulation of prostaglandin biosynthetic process(GO:0031394) positive regulation of unsaturated fatty acid biosynthetic process(GO:2001280) |
| 0.0 | 0.1 | GO:0090131 | mesenchyme migration(GO:0090131) |
| 0.0 | 0.2 | GO:0035871 | protein K11-linked deubiquitination(GO:0035871) |
| 0.0 | 0.1 | GO:2001293 | malonyl-CoA metabolic process(GO:2001293) |
| 0.0 | 0.4 | GO:0042776 | mitochondrial ATP synthesis coupled proton transport(GO:0042776) |
| 0.0 | 0.1 | GO:1902774 | late endosome to lysosome transport(GO:1902774) |
| 0.0 | 0.1 | GO:0017062 | respiratory chain complex III assembly(GO:0017062) mitochondrial respiratory chain complex III assembly(GO:0034551) mitochondrial respiratory chain complex III biogenesis(GO:0097033) |
| 0.0 | 0.1 | GO:0030241 | skeletal muscle myosin thick filament assembly(GO:0030241) |
| 0.0 | 0.2 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.0 | 0.1 | GO:0032000 | positive regulation of fatty acid beta-oxidation(GO:0032000) |
| 0.0 | 0.2 | GO:0006975 | DNA damage induced protein phosphorylation(GO:0006975) |
| 0.0 | 0.2 | GO:0016559 | peroxisome fission(GO:0016559) |
| 0.0 | 0.1 | GO:1990034 | cellular response to corticosterone stimulus(GO:0071386) calcium ion export from cell(GO:1990034) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | GO:0097196 | Shu complex(GO:0097196) |
| 0.0 | 0.2 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.0 | 0.3 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.0 | 0.1 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.0 | 0.1 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.0 | 0.2 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.0 | 0.1 | GO:0033263 | CORVET complex(GO:0033263) |
| 0.0 | 0.1 | GO:0000275 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) |
| 0.0 | 0.3 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.0 | 0.1 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.0 | 0.1 | GO:0034715 | pICln-Sm protein complex(GO:0034715) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.1 | 0.2 | GO:0005124 | scavenger receptor binding(GO:0005124) |
| 0.1 | 0.3 | GO:0070137 | ubiquitin-like protein-specific endopeptidase activity(GO:0070137) SUMO-specific endopeptidase activity(GO:0070139) |
| 0.0 | 0.2 | GO:0003986 | acetyl-CoA hydrolase activity(GO:0003986) |
| 0.0 | 0.1 | GO:0004019 | adenylosuccinate synthase activity(GO:0004019) |
| 0.0 | 0.4 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.0 | 0.2 | GO:0036042 | long-chain fatty acyl-CoA binding(GO:0036042) |
| 0.0 | 0.1 | GO:0004174 | electron-transferring-flavoprotein dehydrogenase activity(GO:0004174) oxidoreductase activity, acting on the CH-NH group of donors, quinone or similar compound as acceptor(GO:0016649) |
| 0.0 | 0.2 | GO:0004784 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.0 | 0.2 | GO:0004075 | biotin carboxylase activity(GO:0004075) biotin binding(GO:0009374) |
| 0.0 | 0.1 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.0 | 0.4 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.0 | 0.1 | GO:0016681 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors(GO:0016679) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.0 | 0.4 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.0 | 0.1 | GO:0017077 | oxidative phosphorylation uncoupler activity(GO:0017077) |
| 0.0 | 0.1 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.0 | 0.1 | GO:0031893 | vasopressin receptor binding(GO:0031893) |
| 0.0 | 0.2 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.0 | 0.3 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.0 | 0.1 | GO:0051373 | FATZ binding(GO:0051373) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | REACTOME ENDOGENOUS STEROLS | Genes involved in Endogenous sterols |
| 0.0 | 0.4 | REACTOME REGULATION OF AMPK ACTIVITY VIA LKB1 | Genes involved in Regulation of AMPK activity via LKB1 |
| 0.0 | 0.2 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |