Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
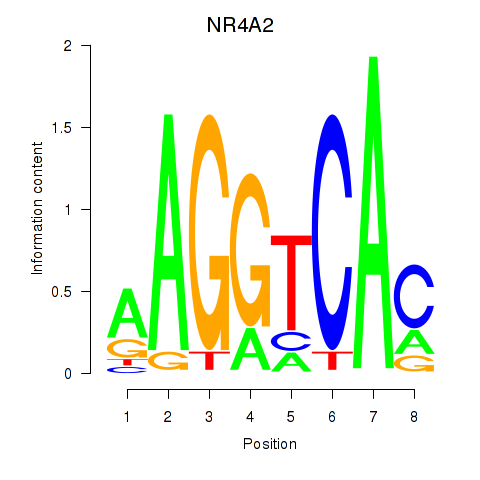
Results for NR4A2
Z-value: 0.52
Transcription factors associated with NR4A2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NR4A2
|
ENSG00000153234.9 | NR4A2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NR4A2 | hg19_v2_chr2_-_157189180_157189290 | -0.94 | 4.4e-04 | Click! |
Activity profile of NR4A2 motif
Sorted Z-values of NR4A2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of NR4A2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_+_43885252 | 0.68 |
ENST00000453782.1 ENST00000300283.6 ENST00000437924.1 ENST00000450086.2 |
CKMT1B |
creatine kinase, mitochondrial 1B |
| chr15_+_43985725 | 0.65 |
ENST00000413453.2 |
CKMT1A |
creatine kinase, mitochondrial 1A |
| chr15_+_43985084 | 0.65 |
ENST00000434505.1 ENST00000411750.1 |
CKMT1A |
creatine kinase, mitochondrial 1A |
| chr20_+_58179582 | 0.62 |
ENST00000371015.1 ENST00000395639.4 |
PHACTR3 |
phosphatase and actin regulator 3 |
| chr16_-_68269971 | 0.54 |
ENST00000565858.1 |
ESRP2 |
epithelial splicing regulatory protein 2 |
| chr5_+_66124590 | 0.42 |
ENST00000490016.2 ENST00000403666.1 ENST00000450827.1 |
MAST4 |
microtubule associated serine/threonine kinase family member 4 |
| chr3_-_151034734 | 0.38 |
ENST00000260843.4 |
GPR87 |
G protein-coupled receptor 87 |
| chr10_+_81107271 | 0.30 |
ENST00000448165.1 |
PPIF |
peptidylprolyl isomerase F |
| chr20_-_46415297 | 0.29 |
ENST00000467815.1 ENST00000359930.4 |
SULF2 |
sulfatase 2 |
| chr10_-_76995675 | 0.27 |
ENST00000469299.1 |
COMTD1 |
catechol-O-methyltransferase domain containing 1 |
| chr17_-_2614927 | 0.23 |
ENST00000435359.1 |
CLUH |
clustered mitochondria (cluA/CLU1) homolog |
| chr10_-_76995769 | 0.23 |
ENST00000372538.3 |
COMTD1 |
catechol-O-methyltransferase domain containing 1 |
| chr2_+_220491973 | 0.23 |
ENST00000358055.3 |
SLC4A3 |
solute carrier family 4 (anion exchanger), member 3 |
| chr11_-_66139199 | 0.21 |
ENST00000357440.2 |
SLC29A2 |
solute carrier family 29 (equilibrative nucleoside transporter), member 2 |
| chr10_+_81107216 | 0.21 |
ENST00000394579.3 ENST00000225174.3 |
PPIF |
peptidylprolyl isomerase F |
| chr19_+_45394477 | 0.19 |
ENST00000252487.5 ENST00000405636.2 ENST00000592434.1 ENST00000426677.2 ENST00000589649.1 |
TOMM40 |
translocase of outer mitochondrial membrane 40 homolog (yeast) |
| chr3_+_14989186 | 0.19 |
ENST00000435454.1 ENST00000323373.6 |
NR2C2 |
nuclear receptor subfamily 2, group C, member 2 |
| chr11_+_76571911 | 0.18 |
ENST00000534206.1 ENST00000532485.1 ENST00000526597.1 ENST00000533873.1 ENST00000538157.1 |
ACER3 |
alkaline ceramidase 3 |
| chr4_-_40517984 | 0.18 |
ENST00000381795.6 |
RBM47 |
RNA binding motif protein 47 |
| chr1_-_116311402 | 0.18 |
ENST00000261448.5 |
CASQ2 |
calsequestrin 2 (cardiac muscle) |
| chr17_-_27503770 | 0.18 |
ENST00000533112.1 |
MYO18A |
myosin XVIIIA |
| chr8_+_21911054 | 0.17 |
ENST00000519850.1 ENST00000381470.3 |
DMTN |
dematin actin binding protein |
| chr16_+_85942594 | 0.17 |
ENST00000566369.1 |
IRF8 |
interferon regulatory factor 8 |
| chr8_-_131028869 | 0.17 |
ENST00000518283.1 ENST00000519110.1 |
FAM49B |
family with sequence similarity 49, member B |
| chr4_+_20255123 | 0.16 |
ENST00000504154.1 ENST00000273739.5 |
SLIT2 |
slit homolog 2 (Drosophila) |
| chr15_-_75017711 | 0.16 |
ENST00000567032.1 ENST00000564596.1 ENST00000566503.1 ENST00000395049.4 ENST00000395048.2 ENST00000379727.3 |
CYP1A1 |
cytochrome P450, family 1, subfamily A, polypeptide 1 |
| chr15_+_59730348 | 0.16 |
ENST00000288228.5 ENST00000559628.1 ENST00000557914.1 ENST00000560474.1 |
FAM81A |
family with sequence similarity 81, member A |
| chrX_-_101098171 | 0.16 |
ENST00000473265.2 |
NXF5 |
nuclear RNA export factor 5 |
| chr19_+_1407733 | 0.15 |
ENST00000592453.1 |
DAZAP1 |
DAZ associated protein 1 |
| chr7_-_72936608 | 0.15 |
ENST00000404251.1 |
BAZ1B |
bromodomain adjacent to zinc finger domain, 1B |
| chr4_-_139163491 | 0.14 |
ENST00000280612.5 |
SLC7A11 |
solute carrier family 7 (anionic amino acid transporter light chain, xc- system), member 11 |
| chr14_-_21492113 | 0.13 |
ENST00000554094.1 |
NDRG2 |
NDRG family member 2 |
| chr1_+_174417095 | 0.13 |
ENST00000367685.2 |
GPR52 |
G protein-coupled receptor 52 |
| chr4_+_41614909 | 0.13 |
ENST00000509454.1 ENST00000396595.3 ENST00000381753.4 |
LIMCH1 |
LIM and calponin homology domains 1 |
| chr14_-_21492251 | 0.13 |
ENST00000554398.1 |
NDRG2 |
NDRG family member 2 |
| chr16_-_4466565 | 0.13 |
ENST00000572467.1 ENST00000423908.2 ENST00000572044.1 ENST00000571052.1 |
CORO7-PAM16 CORO7 |
CORO7-PAM16 readthrough coronin 7 |
| chr7_-_72936531 | 0.12 |
ENST00000339594.4 |
BAZ1B |
bromodomain adjacent to zinc finger domain, 1B |
| chr17_-_56350797 | 0.12 |
ENST00000577220.1 |
MPO |
myeloperoxidase |
| chr8_-_131028641 | 0.12 |
ENST00000523509.1 |
FAM49B |
family with sequence similarity 49, member B |
| chrX_+_153639856 | 0.12 |
ENST00000426834.1 ENST00000369790.4 ENST00000454722.1 ENST00000350743.4 ENST00000299328.5 ENST00000351413.4 |
TAZ |
tafazzin |
| chr7_+_73245193 | 0.12 |
ENST00000340958.2 |
CLDN4 |
claudin 4 |
| chr22_-_37880543 | 0.12 |
ENST00000442496.1 |
MFNG |
MFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| chr12_+_6309963 | 0.12 |
ENST00000382515.2 |
CD9 |
CD9 molecule |
| chr12_+_10163231 | 0.11 |
ENST00000396502.1 ENST00000338896.5 |
CLEC12B |
C-type lectin domain family 12, member B |
| chr7_-_151433393 | 0.11 |
ENST00000492843.1 |
PRKAG2 |
protein kinase, AMP-activated, gamma 2 non-catalytic subunit |
| chr14_-_55369525 | 0.11 |
ENST00000543643.2 ENST00000536224.2 ENST00000395514.1 ENST00000491895.2 |
GCH1 |
GTP cyclohydrolase 1 |
| chr7_-_151433342 | 0.11 |
ENST00000433631.2 |
PRKAG2 |
protein kinase, AMP-activated, gamma 2 non-catalytic subunit |
| chr2_+_120189422 | 0.11 |
ENST00000306406.4 |
TMEM37 |
transmembrane protein 37 |
| chr11_-_9336234 | 0.11 |
ENST00000528080.1 |
TMEM41B |
transmembrane protein 41B |
| chr1_+_9648921 | 0.11 |
ENST00000377376.4 ENST00000340305.5 ENST00000340381.6 |
TMEM201 |
transmembrane protein 201 |
| chr7_-_140624499 | 0.11 |
ENST00000288602.6 |
BRAF |
v-raf murine sarcoma viral oncogene homolog B |
| chr7_-_81399411 | 0.10 |
ENST00000423064.2 |
HGF |
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr18_-_12377283 | 0.10 |
ENST00000269143.3 |
AFG3L2 |
AFG3-like AAA ATPase 2 |
| chr12_-_8088871 | 0.09 |
ENST00000075120.7 |
SLC2A3 |
solute carrier family 2 (facilitated glucose transporter), member 3 |
| chr1_+_212738676 | 0.09 |
ENST00000366981.4 ENST00000366987.2 |
ATF3 |
activating transcription factor 3 |
| chr8_+_145582633 | 0.09 |
ENST00000540505.1 |
SLC52A2 |
solute carrier family 52 (riboflavin transporter), member 2 |
| chr22_+_51176624 | 0.09 |
ENST00000216139.5 ENST00000529621.1 |
ACR |
acrosin |
| chr20_-_62130474 | 0.09 |
ENST00000217182.3 |
EEF1A2 |
eukaryotic translation elongation factor 1 alpha 2 |
| chr17_+_79670386 | 0.09 |
ENST00000333676.3 ENST00000571730.1 ENST00000541223.1 |
MRPL12 SLC25A10 SLC25A10 |
mitochondrial ribosomal protein L12 Mitochondrial dicarboxylate carrier; Uncharacterized protein; cDNA FLJ60124, highly similar to Mitochondrial dicarboxylate carrier solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 |
| chr6_-_107077347 | 0.09 |
ENST00000369063.3 ENST00000539449.1 |
RTN4IP1 |
reticulon 4 interacting protein 1 |
| chr7_+_150782945 | 0.09 |
ENST00000463381.1 |
AGAP3 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 3 |
| chr19_-_36054555 | 0.09 |
ENST00000262623.3 |
ATP4A |
ATPase, H+/K+ exchanging, alpha polypeptide |
| chr9_-_39239171 | 0.09 |
ENST00000358144.2 |
CNTNAP3 |
contactin associated protein-like 3 |
| chr4_+_24797085 | 0.09 |
ENST00000382120.3 |
SOD3 |
superoxide dismutase 3, extracellular |
| chr10_-_101841588 | 0.09 |
ENST00000370418.3 |
CPN1 |
carboxypeptidase N, polypeptide 1 |
| chr8_+_145149930 | 0.08 |
ENST00000318911.4 |
CYC1 |
cytochrome c-1 |
| chr17_-_56358287 | 0.08 |
ENST00000225275.3 ENST00000340482.3 |
MPO |
myeloperoxidase |
| chr17_-_29624343 | 0.08 |
ENST00000247271.4 |
OMG |
oligodendrocyte myelin glycoprotein |
| chr6_-_39290316 | 0.08 |
ENST00000425054.2 ENST00000373227.4 ENST00000373229.5 ENST00000437525.2 |
KCNK16 |
potassium channel, subfamily K, member 16 |
| chr6_+_149068464 | 0.08 |
ENST00000367463.4 |
UST |
uronyl-2-sulfotransferase |
| chr11_+_66624527 | 0.08 |
ENST00000393952.3 |
LRFN4 |
leucine rich repeat and fibronectin type III domain containing 4 |
| chr22_-_29784519 | 0.07 |
ENST00000357586.2 ENST00000356015.2 ENST00000432560.2 ENST00000317368.7 |
AP1B1 |
adaptor-related protein complex 1, beta 1 subunit |
| chr1_-_157108266 | 0.07 |
ENST00000326786.4 |
ETV3 |
ets variant 3 |
| chr16_+_691792 | 0.07 |
ENST00000307650.4 |
FAM195A |
family with sequence similarity 195, member A |
| chr1_-_205904950 | 0.07 |
ENST00000340781.4 |
SLC26A9 |
solute carrier family 26 (anion exchanger), member 9 |
| chr22_+_30821732 | 0.07 |
ENST00000355143.4 |
MTFP1 |
mitochondrial fission process 1 |
| chr11_+_3876859 | 0.07 |
ENST00000300737.4 |
STIM1 |
stromal interaction molecule 1 |
| chr1_+_43803475 | 0.07 |
ENST00000372470.3 ENST00000413998.2 |
MPL |
myeloproliferative leukemia virus oncogene |
| chr2_-_207024134 | 0.07 |
ENST00000457011.1 ENST00000440274.1 ENST00000432169.1 |
NDUFS1 |
NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) |
| chr1_-_226129083 | 0.07 |
ENST00000420304.2 |
LEFTY2 |
left-right determination factor 2 |
| chr22_+_50354104 | 0.07 |
ENST00000360612.4 |
PIM3 |
pim-3 oncogene |
| chr3_-_167452262 | 0.07 |
ENST00000487947.2 |
PDCD10 |
programmed cell death 10 |
| chr12_-_16759711 | 0.07 |
ENST00000447609.1 |
LMO3 |
LIM domain only 3 (rhombotin-like 2) |
| chr3_+_9745487 | 0.07 |
ENST00000383832.3 |
CPNE9 |
copine family member IX |
| chr15_+_78441663 | 0.07 |
ENST00000299518.2 ENST00000558554.1 ENST00000557826.1 ENST00000561279.1 ENST00000559186.1 ENST00000560770.1 ENST00000559881.1 ENST00000559205.1 ENST00000441490.2 |
IDH3A |
isocitrate dehydrogenase 3 (NAD+) alpha |
| chr2_+_234627424 | 0.07 |
ENST00000373409.3 |
UGT1A4 |
UDP glucuronosyltransferase 1 family, polypeptide A4 |
| chr4_+_81118647 | 0.07 |
ENST00000415738.2 |
PRDM8 |
PR domain containing 8 |
| chr2_+_234621551 | 0.07 |
ENST00000608381.1 ENST00000373414.3 |
UGT1A1 UGT1A5 |
UDP glucuronosyltransferase 1 family, polypeptide A8 UDP glucuronosyltransferase 1 family, polypeptide A5 |
| chr3_-_113465065 | 0.07 |
ENST00000497255.1 ENST00000478020.1 ENST00000240922.3 ENST00000493900.1 |
NAA50 |
N(alpha)-acetyltransferase 50, NatE catalytic subunit |
| chr3_+_10857885 | 0.07 |
ENST00000254488.2 ENST00000454147.1 |
SLC6A11 |
solute carrier family 6 (neurotransmitter transporter), member 11 |
| chr1_-_157108130 | 0.07 |
ENST00000368192.4 |
ETV3 |
ets variant 3 |
| chr10_-_50970382 | 0.07 |
ENST00000419399.1 ENST00000432695.1 |
OGDHL |
oxoglutarate dehydrogenase-like |
| chr4_-_151936416 | 0.07 |
ENST00000510413.1 ENST00000507224.1 |
LRBA |
LPS-responsive vesicle trafficking, beach and anchor containing |
| chr4_-_151936865 | 0.06 |
ENST00000535741.1 |
LRBA |
LPS-responsive vesicle trafficking, beach and anchor containing |
| chr12_+_93963590 | 0.06 |
ENST00000340600.2 |
SOCS2 |
suppressor of cytokine signaling 2 |
| chr3_+_9745510 | 0.06 |
ENST00000383831.3 |
CPNE9 |
copine family member IX |
| chr10_-_50970322 | 0.06 |
ENST00000374103.4 |
OGDHL |
oxoglutarate dehydrogenase-like |
| chr1_+_206858328 | 0.06 |
ENST00000367103.3 |
MAPKAPK2 |
mitogen-activated protein kinase-activated protein kinase 2 |
| chr4_+_146560245 | 0.06 |
ENST00000541599.1 |
MMAA |
methylmalonic aciduria (cobalamin deficiency) cblA type |
| chr19_-_46476791 | 0.06 |
ENST00000263257.5 |
NOVA2 |
neuro-oncological ventral antigen 2 |
| chr1_-_226129189 | 0.06 |
ENST00000366820.5 |
LEFTY2 |
left-right determination factor 2 |
| chrY_-_15591485 | 0.06 |
ENST00000382896.4 ENST00000537580.1 ENST00000540140.1 ENST00000545955.1 ENST00000538878.1 |
UTY |
ubiquitously transcribed tetratricopeptide repeat containing, Y-linked |
| chr2_-_207023918 | 0.06 |
ENST00000455934.2 ENST00000449699.1 ENST00000454195.1 |
NDUFS1 |
NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) |
| chr2_+_103378472 | 0.06 |
ENST00000412401.2 |
TMEM182 |
transmembrane protein 182 |
| chr1_-_234667504 | 0.06 |
ENST00000421207.1 ENST00000435574.1 |
RP5-855F14.1 |
RP5-855F14.1 |
| chr7_+_117864708 | 0.06 |
ENST00000357099.4 ENST00000265224.4 ENST00000486422.1 ENST00000417525.1 |
ANKRD7 |
ankyrin repeat domain 7 |
| chr3_-_197025447 | 0.06 |
ENST00000346964.2 ENST00000357674.4 ENST00000314062.3 ENST00000448528.2 ENST00000419553.1 |
DLG1 |
discs, large homolog 1 (Drosophila) |
| chr16_+_11038403 | 0.06 |
ENST00000409552.3 |
CLEC16A |
C-type lectin domain family 16, member A |
| chr18_-_19284724 | 0.06 |
ENST00000580981.1 ENST00000289119.2 |
ABHD3 |
abhydrolase domain containing 3 |
| chr15_-_41408339 | 0.06 |
ENST00000401393.3 |
INO80 |
INO80 complex subunit |
| chr3_+_57541975 | 0.05 |
ENST00000487257.1 ENST00000311180.8 |
PDE12 |
phosphodiesterase 12 |
| chr7_+_112090483 | 0.05 |
ENST00000403825.3 ENST00000429071.1 |
IFRD1 |
interferon-related developmental regulator 1 |
| chr17_+_48423453 | 0.05 |
ENST00000017003.2 ENST00000509778.1 ENST00000507602.1 |
XYLT2 |
xylosyltransferase II |
| chr4_-_122872909 | 0.05 |
ENST00000379645.3 |
TRPC3 |
transient receptor potential cation channel, subfamily C, member 3 |
| chr13_+_97928395 | 0.05 |
ENST00000445661.2 |
MBNL2 |
muscleblind-like splicing regulator 2 |
| chr15_-_41408409 | 0.05 |
ENST00000361937.3 |
INO80 |
INO80 complex subunit |
| chr3_+_52813932 | 0.05 |
ENST00000537050.1 |
ITIH1 |
inter-alpha-trypsin inhibitor heavy chain 1 |
| chr2_+_133874577 | 0.05 |
ENST00000596384.1 |
AC011755.1 |
HCG2006742; Protein LOC100996685 |
| chr4_+_41614720 | 0.05 |
ENST00000509277.1 |
LIMCH1 |
LIM and calponin homology domains 1 |
| chr2_+_191334212 | 0.05 |
ENST00000444317.1 ENST00000535751.1 |
MFSD6 |
major facilitator superfamily domain containing 6 |
| chr3_-_197024965 | 0.05 |
ENST00000392382.2 |
DLG1 |
discs, large homolog 1 (Drosophila) |
| chr1_+_169075554 | 0.05 |
ENST00000367815.4 |
ATP1B1 |
ATPase, Na+/K+ transporting, beta 1 polypeptide |
| chr16_-_58328923 | 0.05 |
ENST00000567164.1 ENST00000219301.4 ENST00000569727.1 |
PRSS54 |
protease, serine, 54 |
| chr19_+_589893 | 0.05 |
ENST00000251287.2 |
HCN2 |
hyperpolarization activated cyclic nucleotide-gated potassium channel 2 |
| chr7_+_39663485 | 0.05 |
ENST00000436179.1 |
RALA |
v-ral simian leukemia viral oncogene homolog A (ras related) |
| chr2_+_219745020 | 0.05 |
ENST00000258411.3 |
WNT10A |
wingless-type MMTV integration site family, member 10A |
| chr21_+_35014706 | 0.05 |
ENST00000399353.1 ENST00000444491.1 ENST00000381318.3 |
ITSN1 |
intersectin 1 (SH3 domain protein) |
| chr15_-_41120896 | 0.05 |
ENST00000299174.5 ENST00000427255.2 |
PPP1R14D |
protein phosphatase 1, regulatory (inhibitor) subunit 14D |
| chr16_+_6069586 | 0.05 |
ENST00000547372.1 |
RBFOX1 |
RNA binding protein, fox-1 homolog (C. elegans) 1 |
| chr10_-_128359074 | 0.05 |
ENST00000544758.1 |
C10orf90 |
chromosome 10 open reading frame 90 |
| chr16_-_70729496 | 0.04 |
ENST00000567648.1 |
VAC14 |
Vac14 homolog (S. cerevisiae) |
| chr3_-_167452298 | 0.04 |
ENST00000475915.2 ENST00000462725.2 ENST00000461494.1 |
PDCD10 |
programmed cell death 10 |
| chr12_-_16761007 | 0.04 |
ENST00000354662.1 ENST00000441439.2 |
LMO3 |
LIM domain only 3 (rhombotin-like 2) |
| chr6_-_31628512 | 0.04 |
ENST00000375911.1 |
C6orf47 |
chromosome 6 open reading frame 47 |
| chr9_+_113431059 | 0.04 |
ENST00000416899.2 |
MUSK |
muscle, skeletal, receptor tyrosine kinase |
| chr19_+_1407517 | 0.04 |
ENST00000336761.6 ENST00000233078.4 |
DAZAP1 |
DAZ associated protein 1 |
| chr20_-_13765526 | 0.04 |
ENST00000202816.1 |
ESF1 |
ESF1, nucleolar pre-rRNA processing protein, homolog (S. cerevisiae) |
| chr12_-_108955070 | 0.04 |
ENST00000228284.3 ENST00000546611.1 |
SART3 |
squamous cell carcinoma antigen recognized by T cells 3 |
| chr5_-_149792295 | 0.04 |
ENST00000518797.1 ENST00000524315.1 ENST00000009530.7 ENST00000377795.3 |
CD74 |
CD74 molecule, major histocompatibility complex, class II invariant chain |
| chr12_+_120875887 | 0.04 |
ENST00000229379.2 |
COX6A1 |
cytochrome c oxidase subunit VIa polypeptide 1 |
| chr10_-_51623203 | 0.04 |
ENST00000444743.1 ENST00000374065.3 ENST00000374064.3 ENST00000260867.4 |
TIMM23 |
translocase of inner mitochondrial membrane 23 homolog (yeast) |
| chr1_-_17380630 | 0.04 |
ENST00000375499.3 |
SDHB |
succinate dehydrogenase complex, subunit B, iron sulfur (Ip) |
| chr8_-_145018905 | 0.04 |
ENST00000398774.2 |
PLEC |
plectin |
| chr2_+_119981384 | 0.04 |
ENST00000393108.2 ENST00000354888.5 ENST00000450943.2 ENST00000393110.2 ENST00000393106.2 ENST00000409811.1 ENST00000393107.2 |
STEAP3 |
STEAP family member 3, metalloreductase |
| chr1_+_206858232 | 0.04 |
ENST00000294981.4 |
MAPKAPK2 |
mitogen-activated protein kinase-activated protein kinase 2 |
| chr17_+_79679369 | 0.04 |
ENST00000350690.5 |
SLC25A10 |
solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 |
| chr20_-_1373606 | 0.04 |
ENST00000381715.1 ENST00000439640.2 ENST00000381719.3 |
FKBP1A |
FK506 binding protein 1A, 12kDa |
| chr7_-_81399329 | 0.04 |
ENST00000453411.1 ENST00000444829.2 |
HGF |
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr6_-_34664612 | 0.04 |
ENST00000374023.3 ENST00000374026.3 |
C6orf106 |
chromosome 6 open reading frame 106 |
| chrX_+_44732757 | 0.04 |
ENST00000377967.4 ENST00000536777.1 ENST00000382899.4 ENST00000543216.1 |
KDM6A |
lysine (K)-specific demethylase 6A |
| chr1_+_155179012 | 0.04 |
ENST00000609421.1 |
MTX1 |
metaxin 1 |
| chr6_-_9933500 | 0.04 |
ENST00000492169.1 |
OFCC1 |
orofacial cleft 1 candidate 1 |
| chr10_+_51371390 | 0.04 |
ENST00000478381.1 ENST00000451577.2 ENST00000374098.2 ENST00000374097.2 |
TIMM23B |
translocase of inner mitochondrial membrane 23 homolog B (yeast) |
| chr1_+_26348259 | 0.04 |
ENST00000374280.3 |
EXTL1 |
exostosin-like glycosyltransferase 1 |
| chr5_+_218356 | 0.04 |
ENST00000264932.6 ENST00000504309.1 ENST00000510361.1 |
SDHA |
succinate dehydrogenase complex, subunit A, flavoprotein (Fp) |
| chr19_-_46272106 | 0.04 |
ENST00000560168.1 |
SIX5 |
SIX homeobox 5 |
| chr20_+_47538357 | 0.04 |
ENST00000371917.4 |
ARFGEF2 |
ADP-ribosylation factor guanine nucleotide-exchange factor 2 (brefeldin A-inhibited) |
| chr12_+_120875910 | 0.04 |
ENST00000551806.1 |
AL021546.6 |
Glutamyl-tRNA(Gln) amidotransferase subunit C, mitochondrial |
| chr10_+_80828774 | 0.04 |
ENST00000334512.5 |
ZMIZ1 |
zinc finger, MIZ-type containing 1 |
| chr3_-_42743006 | 0.04 |
ENST00000310417.5 |
HHATL |
hedgehog acyltransferase-like |
| chr19_-_2328572 | 0.04 |
ENST00000252622.10 |
LSM7 |
LSM7 homolog, U6 small nuclear RNA associated (S. cerevisiae) |
| chr1_+_32084641 | 0.04 |
ENST00000373706.5 |
HCRTR1 |
hypocretin (orexin) receptor 1 |
| chr5_-_169626104 | 0.03 |
ENST00000520275.1 ENST00000506431.2 |
CTB-27N1.1 |
CTB-27N1.1 |
| chr1_-_8877692 | 0.03 |
ENST00000400908.2 |
RERE |
arginine-glutamic acid dipeptide (RE) repeats |
| chr9_-_34048873 | 0.03 |
ENST00000449054.1 ENST00000379239.4 ENST00000539807.1 ENST00000379238.1 ENST00000418786.2 ENST00000360802.1 ENST00000412543.1 |
UBAP2 |
ubiquitin associated protein 2 |
| chr10_-_120938303 | 0.03 |
ENST00000356951.3 ENST00000298510.2 |
PRDX3 |
peroxiredoxin 3 |
| chr9_-_138853156 | 0.03 |
ENST00000371756.3 |
UBAC1 |
UBA domain containing 1 |
| chr2_+_220299547 | 0.03 |
ENST00000312358.7 |
SPEG |
SPEG complex locus |
| chr19_-_8008533 | 0.03 |
ENST00000597926.1 |
TIMM44 |
translocase of inner mitochondrial membrane 44 homolog (yeast) |
| chr1_-_229569834 | 0.03 |
ENST00000366684.3 ENST00000366683.2 |
ACTA1 |
actin, alpha 1, skeletal muscle |
| chr16_+_30937213 | 0.03 |
ENST00000427128.1 |
FBXL19 |
F-box and leucine-rich repeat protein 19 |
| chr1_-_20987851 | 0.03 |
ENST00000464364.1 ENST00000602624.2 |
DDOST |
dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit (non-catalytic) |
| chr17_-_10276319 | 0.03 |
ENST00000252172.4 ENST00000418404.3 |
MYH13 |
myosin, heavy chain 13, skeletal muscle |
| chr17_+_79679299 | 0.03 |
ENST00000331531.5 |
SLC25A10 |
solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 |
| chr9_+_113431029 | 0.03 |
ENST00000189978.5 ENST00000374448.4 ENST00000374440.3 |
MUSK |
muscle, skeletal, receptor tyrosine kinase |
| chr14_+_42076765 | 0.03 |
ENST00000298119.4 |
LRFN5 |
leucine rich repeat and fibronectin type III domain containing 5 |
| chr1_+_155583012 | 0.03 |
ENST00000462250.2 |
MSTO1 |
misato 1, mitochondrial distribution and morphology regulator |
| chr1_-_201140673 | 0.03 |
ENST00000367333.2 |
TMEM9 |
transmembrane protein 9 |
| chr6_+_107077435 | 0.03 |
ENST00000369046.4 |
QRSL1 |
glutaminyl-tRNA synthase (glutamine-hydrolyzing)-like 1 |
| chr2_+_234686976 | 0.03 |
ENST00000389758.3 |
MROH2A |
maestro heat-like repeat family member 2A |
| chr6_+_107077471 | 0.03 |
ENST00000369044.1 |
QRSL1 |
glutaminyl-tRNA synthase (glutamine-hydrolyzing)-like 1 |
| chr7_-_81399355 | 0.03 |
ENST00000457544.2 |
HGF |
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr9_+_129567282 | 0.03 |
ENST00000449886.1 ENST00000373464.4 ENST00000450858.1 |
ZBTB43 |
zinc finger and BTB domain containing 43 |
| chr16_-_58328870 | 0.03 |
ENST00000543437.1 |
PRSS54 |
protease, serine, 54 |
| chr19_+_8509842 | 0.03 |
ENST00000325495.4 ENST00000600092.1 ENST00000594907.1 ENST00000596984.1 ENST00000601645.1 |
HNRNPM |
heterogeneous nuclear ribonucleoprotein M |
| chr1_+_45140360 | 0.03 |
ENST00000418644.1 ENST00000458657.2 ENST00000441519.1 ENST00000535358.1 ENST00000445071.1 |
C1orf228 |
chromosome 1 open reading frame 228 |
| chr3_-_46000064 | 0.03 |
ENST00000433878.1 |
FYCO1 |
FYVE and coiled-coil domain containing 1 |
| chr22_-_38506619 | 0.03 |
ENST00000332536.5 ENST00000381669.3 |
BAIAP2L2 |
BAI1-associated protein 2-like 2 |
| chr11_+_86748998 | 0.03 |
ENST00000525018.1 ENST00000355734.4 |
TMEM135 |
transmembrane protein 135 |
| chr1_-_20987889 | 0.03 |
ENST00000415136.2 |
DDOST |
dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit (non-catalytic) |
| chr10_+_14880157 | 0.03 |
ENST00000378372.3 |
HSPA14 |
heat shock 70kDa protein 14 |
| chr2_+_219135115 | 0.03 |
ENST00000248451.3 ENST00000273077.4 |
PNKD |
paroxysmal nonkinesigenic dyskinesia |
| chr1_+_201924619 | 0.02 |
ENST00000367287.4 |
TIMM17A |
translocase of inner mitochondrial membrane 17 homolog A (yeast) |
| chr5_-_137911049 | 0.02 |
ENST00000297185.3 |
HSPA9 |
heat shock 70kDa protein 9 (mortalin) |
| chr11_+_34938119 | 0.02 |
ENST00000227868.4 ENST00000430469.2 ENST00000533262.1 |
PDHX |
pyruvate dehydrogenase complex, component X |
| chr4_-_71705060 | 0.02 |
ENST00000514161.1 |
GRSF1 |
G-rich RNA sequence binding factor 1 |
| chr14_+_27342334 | 0.02 |
ENST00000548170.1 ENST00000552926.1 |
RP11-384J4.1 |
RP11-384J4.1 |
| chr16_-_67427389 | 0.02 |
ENST00000562206.1 ENST00000290942.5 ENST00000393957.2 |
TPPP3 |
tubulin polymerization-promoting protein family member 3 |
| chr22_-_27620603 | 0.02 |
ENST00000418271.1 ENST00000444114.1 |
RP5-1172A22.1 |
RP5-1172A22.1 |
| chr8_+_99439214 | 0.02 |
ENST00000287042.4 |
KCNS2 |
potassium voltage-gated channel, delayed-rectifier, subfamily S, member 2 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:2000276 | negative regulation of oxidative phosphorylation uncoupler activity(GO:2000276) |
| 0.1 | 1.3 | GO:0006600 | creatine metabolic process(GO:0006600) |
| 0.1 | 0.3 | GO:0014846 | esophagus smooth muscle contraction(GO:0014846) |
| 0.1 | 0.2 | GO:1903028 | positive regulation of opsonization(GO:1903028) |
| 0.1 | 0.2 | GO:0021966 | cerebral cortex tangential migration using cell-cell interactions(GO:0021823) postnatal olfactory bulb interneuron migration(GO:0021827) chemorepulsion involved in postnatal olfactory bulb interneuron migration(GO:0021836) corticospinal neuron axon guidance(GO:0021966) negative regulation of granulocyte chemotaxis(GO:0071623) negative regulation of mononuclear cell migration(GO:0071676) negative regulation of neutrophil chemotaxis(GO:0090024) negative regulation of neutrophil migration(GO:1902623) |
| 0.1 | 0.2 | GO:0071423 | thiosulfate transport(GO:0015709) oxaloacetate transport(GO:0015729) malate transport(GO:0015743) malate transmembrane transport(GO:0071423) oxaloacetate(2-) transmembrane transport(GO:1902356) |
| 0.1 | 0.2 | GO:0018894 | dibenzo-p-dioxin metabolic process(GO:0018894) cellular alkene metabolic process(GO:0043449) |
| 0.0 | 0.3 | GO:0035709 | memory T cell activation(GO:0035709) regulation of memory T cell activation(GO:2000567) positive regulation of memory T cell activation(GO:2000568) |
| 0.0 | 0.1 | GO:0051066 | dihydrobiopterin metabolic process(GO:0051066) |
| 0.0 | 0.1 | GO:0071557 | histone H3-K27 demethylation(GO:0071557) |
| 0.0 | 0.1 | GO:0070837 | dehydroascorbic acid transport(GO:0070837) |
| 0.0 | 0.1 | GO:0061394 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) |
| 0.0 | 0.2 | GO:0070560 | protein secretion by platelet(GO:0070560) |
| 0.0 | 0.2 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.0 | 0.1 | GO:0038163 | thrombopoietin-mediated signaling pathway(GO:0038163) |
| 0.0 | 0.2 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.0 | 0.2 | GO:0001878 | response to yeast(GO:0001878) |
| 0.0 | 0.1 | GO:0006789 | bilirubin conjugation(GO:0006789) |
| 0.0 | 0.1 | GO:2000301 | negative regulation of synaptic vesicle exocytosis(GO:2000301) |
| 0.0 | 0.1 | GO:0030070 | insulin processing(GO:0030070) |
| 0.0 | 0.2 | GO:0086068 | Purkinje myocyte to ventricular cardiac muscle cell signaling(GO:0086029) Purkinje myocyte to ventricular cardiac muscle cell communication(GO:0086068) |
| 0.0 | 0.1 | GO:0070681 | glutaminyl-tRNAGln biosynthesis via transamidation(GO:0070681) |
| 0.0 | 0.1 | GO:1903588 | negative regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903588) |
| 0.0 | 0.2 | GO:0060665 | regulation of branching involved in salivary gland morphogenesis by mesenchymal-epithelial signaling(GO:0060665) |
| 0.0 | 0.1 | GO:0032218 | riboflavin transport(GO:0032218) |
| 0.0 | 0.1 | GO:0007518 | myoblast fate determination(GO:0007518) |
| 0.0 | 0.3 | GO:0090361 | platelet-derived growth factor production(GO:0090360) regulation of platelet-derived growth factor production(GO:0090361) |
| 0.0 | 0.3 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.0 | 0.4 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.0 | 0.3 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.0 | 0.1 | GO:0009236 | cobalamin biosynthetic process(GO:0009236) |
| 0.0 | 0.1 | GO:1901097 | negative regulation of autophagosome maturation(GO:1901097) |
| 0.0 | 0.2 | GO:1903286 | regulation of potassium ion import(GO:1903286) |
| 0.0 | 0.1 | GO:2000324 | positive regulation of glucocorticoid receptor signaling pathway(GO:2000324) |
| 0.0 | 0.2 | GO:0046512 | diol biosynthetic process(GO:0034312) sphingosine biosynthetic process(GO:0046512) |
| 0.0 | 0.1 | GO:0007341 | penetration of zona pellucida(GO:0007341) |
| 0.0 | 0.4 | GO:0060445 | branching involved in salivary gland morphogenesis(GO:0060445) |
| 0.0 | 0.1 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 0.0 | 0.0 | GO:0006500 | N-terminal protein palmitoylation(GO:0006500) |
| 0.0 | 0.1 | GO:0032049 | cardiolipin biosynthetic process(GO:0032049) |
| 0.0 | 0.1 | GO:0010571 | positive regulation of nuclear cell cycle DNA replication(GO:0010571) |
| 0.0 | 0.1 | GO:0010615 | positive regulation of cardiac muscle adaptation(GO:0010615) positive regulation of cardiac muscle hypertrophy in response to stress(GO:1903244) |
| 0.0 | 0.2 | GO:0006853 | carnitine shuttle(GO:0006853) |
| 0.0 | 0.1 | GO:0002767 | immune response-inhibiting cell surface receptor signaling pathway(GO:0002767) |
| 0.0 | 0.1 | GO:0032237 | activation of store-operated calcium channel activity(GO:0032237) positive regulation of store-operated calcium channel activity(GO:1901341) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0090571 | RNA polymerase II transcription repressor complex(GO:0090571) |
| 0.0 | 0.1 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 0.0 | 0.1 | GO:0043159 | acrosomal matrix(GO:0043159) |
| 0.0 | 0.1 | GO:0070470 | plasma membrane respiratory chain(GO:0070470) |
| 0.0 | 0.2 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 0.1 | GO:0030956 | glutamyl-tRNA(Gln) amidotransferase complex(GO:0030956) |
| 0.0 | 0.0 | GO:0098855 | HCN channel complex(GO:0098855) |
| 0.0 | 0.1 | GO:0071001 | U4/U6 snRNP(GO:0071001) |
| 0.0 | 0.0 | GO:0005749 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 0.0 | 0.4 | GO:0005753 | mitochondrial proton-transporting ATP synthase complex(GO:0005753) |
| 0.0 | 0.1 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 0.0 | 0.1 | GO:0030314 | junctional membrane complex(GO:0030314) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.3 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.1 | 0.2 | GO:0015131 | thiosulfate transmembrane transporter activity(GO:0015117) oxaloacetate transmembrane transporter activity(GO:0015131) |
| 0.0 | 0.3 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.0 | 0.3 | GO:0023025 | MHC class Ib protein complex binding(GO:0023025) MHC class Ib protein binding, via antigen binding groove(GO:0023030) |
| 0.0 | 0.1 | GO:0004040 | amidase activity(GO:0004040) |
| 0.0 | 0.2 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.0 | 0.5 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.0 | 0.1 | GO:0071558 | histone demethylase activity (H3-K27 specific)(GO:0071558) |
| 0.0 | 0.1 | GO:0001588 | dopamine neurotransmitter receptor activity, coupled via Gs(GO:0001588) |
| 0.0 | 0.2 | GO:0070576 | vitamin D 24-hydroxylase activity(GO:0070576) |
| 0.0 | 0.1 | GO:0033300 | dehydroascorbic acid transporter activity(GO:0033300) |
| 0.0 | 0.1 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.0 | 0.1 | GO:0005294 | neutral L-amino acid secondary active transmembrane transporter activity(GO:0005294) |
| 0.0 | 0.1 | GO:0045155 | electron transporter, transferring electrons from CoQH2-cytochrome c reductase complex and cytochrome c oxidase complex activity(GO:0045155) |
| 0.0 | 0.1 | GO:0008177 | succinate dehydrogenase (ubiquinone) activity(GO:0008177) |
| 0.0 | 0.3 | GO:0015266 | protein channel activity(GO:0015266) |
| 0.0 | 0.1 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 0.0 | 0.2 | GO:0008607 | phosphorylase kinase regulator activity(GO:0008607) |
| 0.0 | 0.1 | GO:0008900 | hydrogen:potassium-exchanging ATPase activity(GO:0008900) |
| 0.0 | 0.1 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 0.0 | 0.1 | GO:0032217 | riboflavin transporter activity(GO:0032217) |
| 0.0 | 0.2 | GO:0043237 | laminin-1 binding(GO:0043237) Roundabout binding(GO:0048495) |
| 0.0 | 0.1 | GO:0030158 | protein xylosyltransferase activity(GO:0030158) |
| 0.0 | 0.6 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 0.1 | GO:0004449 | isocitrate dehydrogenase (NAD+) activity(GO:0004449) |
| 0.0 | 0.4 | GO:0001608 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.0 | 0.5 | GO:0008171 | O-methyltransferase activity(GO:0008171) |
| 0.0 | 0.1 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 0.0 | 0.0 | GO:0005222 | intracellular cAMP activated cation channel activity(GO:0005222) |
| 0.0 | 0.1 | GO:0052740 | 1-acyl-2-lysophosphatidylserine acylhydrolase activity(GO:0052740) |
| 0.0 | 0.0 | GO:0008267 | poly-glutamine tract binding(GO:0008267) |
| 0.0 | 0.1 | GO:0005332 | gamma-aminobutyric acid:sodium symporter activity(GO:0005332) |
| 0.0 | 0.1 | GO:0008269 | JAK pathway signal transduction adaptor activity(GO:0008269) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.0 | 0.2 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |