Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
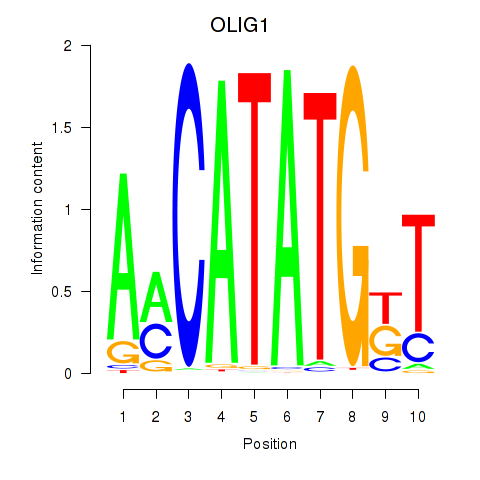
Results for OLIG1
Z-value: 0.86
Transcription factors associated with OLIG1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
OLIG1
|
ENSG00000184221.8 | OLIG1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| OLIG1 | hg19_v2_chr21_+_34442439_34442455 | -0.73 | 4.0e-02 | Click! |
Activity profile of OLIG1 motif
Sorted Z-values of OLIG1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of OLIG1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_-_91573132 | 1.69 |
ENST00000550563.1 ENST00000546370.1 |
DCN |
decorin |
| chr5_-_111312622 | 1.35 |
ENST00000395634.3 |
NREP |
neuronal regeneration related protein |
| chr2_-_188378368 | 1.19 |
ENST00000392365.1 ENST00000435414.1 |
TFPI |
tissue factor pathway inhibitor (lipoprotein-associated coagulation inhibitor) |
| chr5_-_75919253 | 0.97 |
ENST00000296641.4 |
F2RL2 |
coagulation factor II (thrombin) receptor-like 2 |
| chr3_-_195310802 | 0.77 |
ENST00000421243.1 ENST00000453131.1 |
APOD |
apolipoprotein D |
| chr3_-_18480260 | 0.71 |
ENST00000454909.2 |
SATB1 |
SATB homeobox 1 |
| chr8_+_70378852 | 0.65 |
ENST00000525061.1 ENST00000458141.2 ENST00000260128.4 |
SULF1 |
sulfatase 1 |
| chr4_-_159094194 | 0.64 |
ENST00000592057.1 ENST00000585682.1 ENST00000393807.5 |
FAM198B |
family with sequence similarity 198, member B |
| chr2_+_210444748 | 0.61 |
ENST00000392194.1 |
MAP2 |
microtubule-associated protein 2 |
| chr5_-_75919217 | 0.60 |
ENST00000504899.1 |
F2RL2 |
coagulation factor II (thrombin) receptor-like 2 |
| chr9_-_95244781 | 0.55 |
ENST00000375544.3 ENST00000375543.1 ENST00000395538.3 ENST00000450139.2 |
ASPN |
asporin |
| chr2_+_152214098 | 0.55 |
ENST00000243347.3 |
TNFAIP6 |
tumor necrosis factor, alpha-induced protein 6 |
| chr9_+_18474098 | 0.51 |
ENST00000327883.7 ENST00000431052.2 ENST00000380570.4 |
ADAMTSL1 |
ADAMTS-like 1 |
| chr6_+_73076432 | 0.51 |
ENST00000414192.2 |
RIMS1 |
regulating synaptic membrane exocytosis 1 |
| chr12_-_108714412 | 0.47 |
ENST00000412676.1 ENST00000550573.1 |
CMKLR1 |
chemokine-like receptor 1 |
| chr6_-_52859046 | 0.47 |
ENST00000457564.1 ENST00000541324.1 ENST00000370960.1 |
GSTA4 |
glutathione S-transferase alpha 4 |
| chr17_-_7145475 | 0.46 |
ENST00000571129.1 ENST00000571253.1 ENST00000573928.1 |
GABARAP |
GABA(A) receptor-associated protein |
| chr15_-_63448973 | 0.46 |
ENST00000462430.1 |
RPS27L |
ribosomal protein S27-like |
| chr2_-_79313973 | 0.43 |
ENST00000454188.1 |
REG1B |
regenerating islet-derived 1 beta |
| chr12_+_26126681 | 0.41 |
ENST00000542865.1 |
RASSF8 |
Ras association (RalGDS/AF-6) domain family (N-terminal) member 8 |
| chr3_-_114477962 | 0.37 |
ENST00000471418.1 |
ZBTB20 |
zinc finger and BTB domain containing 20 |
| chr3_-_114477787 | 0.36 |
ENST00000464560.1 |
ZBTB20 |
zinc finger and BTB domain containing 20 |
| chr2_-_175462934 | 0.36 |
ENST00000392546.2 ENST00000436221.1 |
WIPF1 |
WAS/WASL interacting protein family, member 1 |
| chr3_-_121379739 | 0.36 |
ENST00000428394.2 ENST00000314583.3 |
HCLS1 |
hematopoietic cell-specific Lyn substrate 1 |
| chr19_+_18208603 | 0.36 |
ENST00000262811.6 |
MAST3 |
microtubule associated serine/threonine kinase 3 |
| chr13_+_49551020 | 0.35 |
ENST00000541916.1 |
FNDC3A |
fibronectin type III domain containing 3A |
| chr16_+_21716284 | 0.34 |
ENST00000388957.3 |
OTOA |
otoancorin |
| chr12_+_7052974 | 0.34 |
ENST00000544681.1 ENST00000537087.1 |
C12orf57 |
chromosome 12 open reading frame 57 |
| chr12_+_7053172 | 0.33 |
ENST00000229281.5 |
C12orf57 |
chromosome 12 open reading frame 57 |
| chr1_+_104293028 | 0.33 |
ENST00000370079.3 |
AMY1C |
amylase, alpha 1C (salivary) |
| chr6_-_31648150 | 0.31 |
ENST00000375858.3 ENST00000383237.4 |
LY6G5C |
lymphocyte antigen 6 complex, locus G5C |
| chr2_+_54342574 | 0.31 |
ENST00000303536.4 ENST00000394666.3 |
ACYP2 |
acylphosphatase 2, muscle type |
| chr1_-_198509804 | 0.31 |
ENST00000489986.1 ENST00000367382.1 |
ATP6V1G3 |
ATPase, H+ transporting, lysosomal 13kDa, V1 subunit G3 |
| chr5_+_135496675 | 0.30 |
ENST00000507637.1 |
SMAD5 |
SMAD family member 5 |
| chr12_+_7053228 | 0.30 |
ENST00000540506.2 |
C12orf57 |
chromosome 12 open reading frame 57 |
| chr11_+_75428857 | 0.29 |
ENST00000198801.5 |
MOGAT2 |
monoacylglycerol O-acyltransferase 2 |
| chr4_+_70894130 | 0.29 |
ENST00000526767.1 ENST00000530128.1 ENST00000381057.3 |
HTN3 |
histatin 3 |
| chr7_-_38403077 | 0.29 |
ENST00000426402.2 |
TRGV2 |
T cell receptor gamma variable 2 |
| chr10_+_85954377 | 0.29 |
ENST00000332904.3 ENST00000372117.3 |
CDHR1 |
cadherin-related family member 1 |
| chr4_+_160188889 | 0.29 |
ENST00000264431.4 |
RAPGEF2 |
Rap guanine nucleotide exchange factor (GEF) 2 |
| chr14_-_23822080 | 0.29 |
ENST00000397267.1 ENST00000354772.3 |
SLC22A17 |
solute carrier family 22, member 17 |
| chr19_+_44764031 | 0.28 |
ENST00000592581.1 ENST00000590668.1 ENST00000588489.1 ENST00000391958.2 |
ZNF233 |
zinc finger protein 233 |
| chr6_+_69942298 | 0.28 |
ENST00000238918.8 |
BAI3 |
brain-specific angiogenesis inhibitor 3 |
| chrX_+_70798261 | 0.28 |
ENST00000373696.3 |
ACRC |
acidic repeat containing |
| chr2_+_201994042 | 0.28 |
ENST00000417748.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr2_-_163099885 | 0.27 |
ENST00000443424.1 |
FAP |
fibroblast activation protein, alpha |
| chr3_-_49466686 | 0.26 |
ENST00000273598.3 ENST00000436744.2 |
NICN1 |
nicolin 1 |
| chr9_-_130635741 | 0.26 |
ENST00000223836.10 |
AK1 |
adenylate kinase 1 |
| chr5_+_140739537 | 0.26 |
ENST00000522605.1 |
PCDHGB2 |
protocadherin gamma subfamily B, 2 |
| chr11_-_134123142 | 0.26 |
ENST00000392595.2 ENST00000341541.3 ENST00000352327.5 ENST00000392594.3 |
THYN1 |
thymocyte nuclear protein 1 |
| chr12_+_4699244 | 0.26 |
ENST00000540757.2 |
DYRK4 |
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 4 |
| chr3_-_178984759 | 0.25 |
ENST00000349697.2 ENST00000497599.1 |
KCNMB3 |
potassium large conductance calcium-activated channel, subfamily M beta member 3 |
| chr2_-_163100045 | 0.25 |
ENST00000188790.4 |
FAP |
fibroblast activation protein, alpha |
| chr14_+_95047725 | 0.25 |
ENST00000554760.1 ENST00000554866.1 ENST00000329597.7 ENST00000556775.1 |
SERPINA5 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 5 |
| chr14_+_95047744 | 0.25 |
ENST00000553511.1 ENST00000554633.1 ENST00000555681.1 ENST00000554276.1 |
SERPINA5 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 5 |
| chr22_-_19435755 | 0.24 |
ENST00000542103.1 ENST00000399562.4 |
C22orf39 |
chromosome 22 open reading frame 39 |
| chrX_+_120181457 | 0.24 |
ENST00000328078.1 |
GLUD2 |
glutamate dehydrogenase 2 |
| chr12_+_55248289 | 0.23 |
ENST00000308796.6 |
MUCL1 |
mucin-like 1 |
| chr14_+_88471468 | 0.23 |
ENST00000267549.3 |
GPR65 |
G protein-coupled receptor 65 |
| chr10_+_5135981 | 0.22 |
ENST00000380554.3 |
AKR1C3 |
aldo-keto reductase family 1, member C3 |
| chr2_-_152382500 | 0.22 |
ENST00000434685.1 |
NEB |
nebulin |
| chr10_+_81462983 | 0.22 |
ENST00000448135.1 ENST00000429828.1 ENST00000372321.1 |
NUTM2B |
NUT family member 2B |
| chr16_-_30905584 | 0.21 |
ENST00000380317.4 |
BCL7C |
B-cell CLL/lymphoma 7C |
| chr2_+_201994569 | 0.21 |
ENST00000457277.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr10_-_31288398 | 0.21 |
ENST00000538351.2 |
ZNF438 |
zinc finger protein 438 |
| chr2_-_48468122 | 0.21 |
ENST00000447571.1 |
AC079807.4 |
AC079807.4 |
| chr13_+_28519343 | 0.21 |
ENST00000381026.3 |
ATP5EP2 |
ATP synthase, H+ transporting, mitochondrial F1 complex, epsilon subunit pseudogene 2 |
| chr12_-_91573249 | 0.20 |
ENST00000550099.1 ENST00000546391.1 ENST00000551354.1 |
DCN |
decorin |
| chrX_-_69509738 | 0.20 |
ENST00000374454.1 ENST00000239666.4 |
PDZD11 |
PDZ domain containing 11 |
| chr9_-_96215822 | 0.19 |
ENST00000375412.5 |
FAM120AOS |
family with sequence similarity 120A opposite strand |
| chr18_-_56296182 | 0.19 |
ENST00000361673.3 |
ALPK2 |
alpha-kinase 2 |
| chr3_-_53916202 | 0.19 |
ENST00000335754.3 |
ACTR8 |
ARP8 actin-related protein 8 homolog (yeast) |
| chrX_-_102757802 | 0.19 |
ENST00000372633.1 |
RAB40A |
RAB40A, member RAS oncogene family |
| chr7_+_140396465 | 0.19 |
ENST00000476279.1 ENST00000247866.4 ENST00000461457.1 ENST00000465506.1 ENST00000204307.5 ENST00000464566.1 |
NDUFB2 |
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 2, 8kDa |
| chr1_+_104198377 | 0.19 |
ENST00000370083.4 |
AMY1A |
amylase, alpha 1A (salivary) |
| chr11_+_134123389 | 0.19 |
ENST00000281182.4 ENST00000537423.1 ENST00000543332.1 ENST00000374752.4 |
ACAD8 |
acyl-CoA dehydrogenase family, member 8 |
| chr4_-_89744314 | 0.19 |
ENST00000508369.1 |
FAM13A |
family with sequence similarity 13, member A |
| chr5_-_133510456 | 0.18 |
ENST00000520417.1 |
SKP1 |
S-phase kinase-associated protein 1 |
| chr1_-_186344802 | 0.18 |
ENST00000451586.1 |
TPR |
translocated promoter region, nuclear basket protein |
| chr12_-_11002063 | 0.18 |
ENST00000544994.1 ENST00000228811.4 ENST00000540107.1 |
PRR4 |
proline rich 4 (lacrimal) |
| chrX_+_118108571 | 0.18 |
ENST00000304778.7 |
LONRF3 |
LON peptidase N-terminal domain and ring finger 3 |
| chr10_+_180987 | 0.17 |
ENST00000381591.1 |
ZMYND11 |
zinc finger, MYND-type containing 11 |
| chr17_-_47022140 | 0.17 |
ENST00000290330.3 |
SNF8 |
SNF8, ESCRT-II complex subunit |
| chrX_+_118108601 | 0.17 |
ENST00000371628.3 |
LONRF3 |
LON peptidase N-terminal domain and ring finger 3 |
| chr15_+_26360970 | 0.17 |
ENST00000556159.1 ENST00000557523.1 |
LINC00929 |
long intergenic non-protein coding RNA 929 |
| chr6_+_3000057 | 0.17 |
ENST00000397717.2 |
NQO2 |
NAD(P)H dehydrogenase, quinone 2 |
| chr8_-_135522425 | 0.17 |
ENST00000521673.1 |
ZFAT |
zinc finger and AT hook domain containing |
| chr5_+_118812237 | 0.17 |
ENST00000513628.1 |
HSD17B4 |
hydroxysteroid (17-beta) dehydrogenase 4 |
| chr2_-_183387430 | 0.17 |
ENST00000410103.1 |
PDE1A |
phosphodiesterase 1A, calmodulin-dependent |
| chr6_+_97010424 | 0.16 |
ENST00000541107.1 ENST00000450218.1 ENST00000326771.2 |
FHL5 |
four and a half LIM domains 5 |
| chr6_+_3000195 | 0.16 |
ENST00000338130.2 |
NQO2 |
NAD(P)H dehydrogenase, quinone 2 |
| chr6_+_292051 | 0.16 |
ENST00000344450.5 |
DUSP22 |
dual specificity phosphatase 22 |
| chr19_+_18496957 | 0.16 |
ENST00000252809.3 |
GDF15 |
growth differentiation factor 15 |
| chr12_-_11175219 | 0.16 |
ENST00000390673.2 |
TAS2R19 |
taste receptor, type 2, member 19 |
| chr15_-_50557863 | 0.15 |
ENST00000543581.1 |
HDC |
histidine decarboxylase |
| chr16_-_4896205 | 0.15 |
ENST00000589389.1 |
GLYR1 |
glyoxylate reductase 1 homolog (Arabidopsis) |
| chr5_-_171433579 | 0.15 |
ENST00000265094.5 ENST00000393802.2 |
FBXW11 |
F-box and WD repeat domain containing 11 |
| chr4_-_89744365 | 0.15 |
ENST00000513837.1 ENST00000503556.1 |
FAM13A |
family with sequence similarity 13, member A |
| chr10_+_180405 | 0.15 |
ENST00000439456.1 ENST00000397962.3 ENST00000309776.4 ENST00000381602.4 |
ZMYND11 |
zinc finger, MYND-type containing 11 |
| chr1_+_36335351 | 0.15 |
ENST00000373206.1 |
AGO1 |
argonaute RISC catalytic component 1 |
| chr3_+_185300391 | 0.15 |
ENST00000545472.1 |
SENP2 |
SUMO1/sentrin/SMT3 specific peptidase 2 |
| chr6_+_3000218 | 0.15 |
ENST00000380441.1 ENST00000380455.4 ENST00000380454.4 |
NQO2 |
NAD(P)H dehydrogenase, quinone 2 |
| chr6_+_57182400 | 0.15 |
ENST00000607273.1 |
PRIM2 |
primase, DNA, polypeptide 2 (58kDa) |
| chr10_+_133918175 | 0.14 |
ENST00000298622.4 |
JAKMIP3 |
Janus kinase and microtubule interacting protein 3 |
| chr19_-_37019562 | 0.14 |
ENST00000523638.1 |
ZNF260 |
zinc finger protein 260 |
| chr11_+_65405556 | 0.14 |
ENST00000534313.1 ENST00000533361.1 ENST00000526137.1 |
SIPA1 |
signal-induced proliferation-associated 1 |
| chr14_-_106494587 | 0.14 |
ENST00000390597.2 |
IGHV2-5 |
immunoglobulin heavy variable 2-5 |
| chr2_-_97760576 | 0.14 |
ENST00000414820.1 ENST00000272610.3 |
FAHD2B |
fumarylacetoacetate hydrolase domain containing 2B |
| chr9_-_21217310 | 0.14 |
ENST00000380216.1 |
IFNA16 |
interferon, alpha 16 |
| chr5_+_118812294 | 0.14 |
ENST00000509514.1 |
HSD17B4 |
hydroxysteroid (17-beta) dehydrogenase 4 |
| chr12_-_95941987 | 0.14 |
ENST00000537435.2 |
USP44 |
ubiquitin specific peptidase 44 |
| chr6_-_33714752 | 0.14 |
ENST00000451316.1 |
IP6K3 |
inositol hexakisphosphate kinase 3 |
| chr1_-_8000872 | 0.14 |
ENST00000377507.3 |
TNFRSF9 |
tumor necrosis factor receptor superfamily, member 9 |
| chr2_-_70418032 | 0.13 |
ENST00000425268.1 ENST00000428751.1 ENST00000417203.1 ENST00000417865.1 ENST00000428010.1 ENST00000447804.1 ENST00000264434.2 |
C2orf42 |
chromosome 2 open reading frame 42 |
| chr2_-_158345462 | 0.13 |
ENST00000439355.1 ENST00000540637.1 |
CYTIP |
cytohesin 1 interacting protein |
| chr4_-_99064387 | 0.13 |
ENST00000295268.3 |
STPG2 |
sperm-tail PG-rich repeat containing 2 |
| chr15_-_55700216 | 0.13 |
ENST00000569205.1 |
CCPG1 |
cell cycle progression 1 |
| chr5_+_157158205 | 0.13 |
ENST00000231198.7 |
THG1L |
tRNA-histidine guanylyltransferase 1-like (S. cerevisiae) |
| chr7_+_140396756 | 0.13 |
ENST00000460088.1 ENST00000472695.1 |
NDUFB2 |
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 2, 8kDa |
| chr15_+_74466012 | 0.13 |
ENST00000249842.3 |
ISLR |
immunoglobulin superfamily containing leucine-rich repeat |
| chr16_+_2933229 | 0.13 |
ENST00000573965.1 ENST00000572006.1 |
FLYWCH2 |
FLYWCH family member 2 |
| chr4_-_89744457 | 0.13 |
ENST00000395002.2 |
FAM13A |
family with sequence similarity 13, member A |
| chr19_+_49990811 | 0.13 |
ENST00000391857.4 ENST00000467825.2 |
RPL13A |
ribosomal protein L13a |
| chr19_-_42806919 | 0.13 |
ENST00000595530.1 ENST00000538771.1 ENST00000601865.1 |
PAFAH1B3 |
platelet-activating factor acetylhydrolase 1b, catalytic subunit 3 (29kDa) |
| chr17_+_42923686 | 0.13 |
ENST00000591513.1 |
HIGD1B |
HIG1 hypoxia inducible domain family, member 1B |
| chr20_-_32031680 | 0.13 |
ENST00000217381.2 |
SNTA1 |
syntrophin, alpha 1 |
| chr13_-_46679185 | 0.13 |
ENST00000439329.3 |
CPB2 |
carboxypeptidase B2 (plasma) |
| chr1_-_111970353 | 0.13 |
ENST00000369732.3 |
OVGP1 |
oviductal glycoprotein 1, 120kDa |
| chr19_-_39390350 | 0.13 |
ENST00000447739.1 ENST00000358931.5 ENST00000407552.1 |
SIRT2 |
sirtuin 2 |
| chr3_+_9839335 | 0.12 |
ENST00000453882.1 |
ARPC4-TTLL3 |
ARPC4-TTLL3 readthrough |
| chr10_-_1071796 | 0.12 |
ENST00000277517.1 |
IDI2 |
isopentenyl-diphosphate delta isomerase 2 |
| chr5_-_171433819 | 0.12 |
ENST00000296933.6 |
FBXW11 |
F-box and WD repeat domain containing 11 |
| chr3_-_128690173 | 0.12 |
ENST00000508239.1 |
RP11-723O4.6 |
Uncharacterized protein FLJ43738 |
| chr10_-_69597915 | 0.12 |
ENST00000225171.2 |
DNAJC12 |
DnaJ (Hsp40) homolog, subfamily C, member 12 |
| chr2_+_241807870 | 0.12 |
ENST00000307503.3 |
AGXT |
alanine-glyoxylate aminotransferase |
| chr12_-_95397442 | 0.11 |
ENST00000547157.1 ENST00000547986.1 ENST00000327772.2 |
NDUFA12 |
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 12 |
| chr17_+_60758814 | 0.11 |
ENST00000579432.1 ENST00000446119.2 |
MRC2 |
mannose receptor, C type 2 |
| chr18_-_59274139 | 0.11 |
ENST00000586949.1 |
RP11-879F14.2 |
RP11-879F14.2 |
| chr3_-_48936272 | 0.11 |
ENST00000544097.1 ENST00000430379.1 ENST00000319017.4 |
SLC25A20 |
solute carrier family 25 (carnitine/acylcarnitine translocase), member 20 |
| chr2_+_201994208 | 0.11 |
ENST00000440180.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr20_+_48429233 | 0.11 |
ENST00000417961.1 |
SLC9A8 |
solute carrier family 9, subfamily A (NHE8, cation proton antiporter 8), member 8 |
| chr6_-_33714667 | 0.11 |
ENST00000293756.4 |
IP6K3 |
inositol hexakisphosphate kinase 3 |
| chr12_+_133657461 | 0.11 |
ENST00000412146.2 ENST00000544426.1 ENST00000440984.2 ENST00000319849.3 ENST00000440550.2 |
ZNF140 |
zinc finger protein 140 |
| chr4_+_2470664 | 0.11 |
ENST00000314289.8 ENST00000541204.1 ENST00000502316.1 ENST00000507247.1 ENST00000509258.1 ENST00000511859.1 |
RNF4 |
ring finger protein 4 |
| chr1_+_201159914 | 0.11 |
ENST00000335211.4 ENST00000451870.2 ENST00000295591.8 |
IGFN1 |
immunoglobulin-like and fibronectin type III domain containing 1 |
| chrX_+_101975643 | 0.11 |
ENST00000361229.4 |
BHLHB9 |
basic helix-loop-helix domain containing, class B, 9 |
| chr14_+_23344345 | 0.11 |
ENST00000551466.1 |
LRP10 |
low density lipoprotein receptor-related protein 10 |
| chr15_+_63354769 | 0.10 |
ENST00000558910.1 |
TPM1 |
tropomyosin 1 (alpha) |
| chr14_-_69864993 | 0.10 |
ENST00000555373.1 |
ERH |
enhancer of rudimentary homolog (Drosophila) |
| chr13_+_38923959 | 0.10 |
ENST00000379649.1 ENST00000239878.4 ENST00000437952.1 ENST00000379641.1 |
UFM1 |
ubiquitin-fold modifier 1 |
| chr8_-_102181718 | 0.10 |
ENST00000565617.1 |
KB-1460A1.5 |
KB-1460A1.5 |
| chr3_+_62304648 | 0.10 |
ENST00000462069.1 ENST00000232519.5 ENST00000465142.1 |
C3orf14 |
chromosome 3 open reading frame 14 |
| chr11_-_6633799 | 0.10 |
ENST00000299424.4 |
TAF10 |
TAF10 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 30kDa |
| chr1_-_53387386 | 0.10 |
ENST00000467988.1 ENST00000358358.5 ENST00000371522.4 |
ECHDC2 |
enoyl CoA hydratase domain containing 2 |
| chrX_+_101975619 | 0.09 |
ENST00000457056.1 |
BHLHB9 |
basic helix-loop-helix domain containing, class B, 9 |
| chr7_+_148287657 | 0.09 |
ENST00000307003.2 |
C7orf33 |
chromosome 7 open reading frame 33 |
| chr7_-_72972319 | 0.09 |
ENST00000223368.2 |
BCL7B |
B-cell CLL/lymphoma 7B |
| chr5_+_140560980 | 0.09 |
ENST00000361016.2 |
PCDHB16 |
protocadherin beta 16 |
| chr1_+_203274639 | 0.09 |
ENST00000290551.4 |
BTG2 |
BTG family, member 2 |
| chr15_-_89755034 | 0.09 |
ENST00000563254.1 |
RLBP1 |
retinaldehyde binding protein 1 |
| chr17_-_34344991 | 0.09 |
ENST00000591423.1 |
CCL23 |
chemokine (C-C motif) ligand 23 |
| chr2_+_223162866 | 0.09 |
ENST00000295226.1 |
CCDC140 |
coiled-coil domain containing 140 |
| chr18_-_10748498 | 0.09 |
ENST00000579949.1 |
PIEZO2 |
piezo-type mechanosensitive ion channel component 2 |
| chr8_+_48173514 | 0.09 |
ENST00000518074.1 ENST00000541342.1 |
SPIDR |
scaffolding protein involved in DNA repair |
| chr16_+_31119615 | 0.09 |
ENST00000394950.3 ENST00000287507.3 ENST00000219794.6 ENST00000561755.1 |
BCKDK |
branched chain ketoacid dehydrogenase kinase |
| chr8_-_145701718 | 0.09 |
ENST00000377317.4 |
FOXH1 |
forkhead box H1 |
| chr12_+_21284118 | 0.08 |
ENST00000256958.2 |
SLCO1B1 |
solute carrier organic anion transporter family, member 1B1 |
| chr5_-_137674000 | 0.08 |
ENST00000510119.1 ENST00000513970.1 |
CDC25C |
cell division cycle 25C |
| chr4_+_25915896 | 0.08 |
ENST00000514384.1 |
SMIM20 |
small integral membrane protein 20 |
| chr1_-_24306835 | 0.08 |
ENST00000484146.2 |
SRSF10 |
serine/arginine-rich splicing factor 10 |
| chr2_-_32490859 | 0.08 |
ENST00000404025.2 |
NLRC4 |
NLR family, CARD domain containing 4 |
| chr17_+_49243639 | 0.08 |
ENST00000512737.1 ENST00000503064.1 |
NME1-NME2 |
NME1-NME2 readthrough |
| chr1_+_159272111 | 0.08 |
ENST00000368114.1 |
FCER1A |
Fc fragment of IgE, high affinity I, receptor for; alpha polypeptide |
| chr12_+_10124001 | 0.08 |
ENST00000396507.3 ENST00000304361.4 ENST00000434319.2 |
CLEC12A |
C-type lectin domain family 12, member A |
| chr3_+_62936098 | 0.08 |
ENST00000475886.1 ENST00000465684.1 ENST00000465262.1 ENST00000468072.1 |
LINC00698 |
long intergenic non-protein coding RNA 698 |
| chr2_-_183387283 | 0.08 |
ENST00000435564.1 |
PDE1A |
phosphodiesterase 1A, calmodulin-dependent |
| chr12_+_117348742 | 0.08 |
ENST00000309909.5 ENST00000455858.2 |
FBXW8 |
F-box and WD repeat domain containing 8 |
| chr15_-_98417780 | 0.08 |
ENST00000503874.3 |
LINC00923 |
long intergenic non-protein coding RNA 923 |
| chr22_+_22749343 | 0.08 |
ENST00000390298.2 |
IGLV7-43 |
immunoglobulin lambda variable 7-43 |
| chr11_-_26743546 | 0.08 |
ENST00000280467.6 ENST00000396005.3 |
SLC5A12 |
solute carrier family 5 (sodium/monocarboxylate cotransporter), member 12 |
| chr2_-_32490801 | 0.08 |
ENST00000360906.5 ENST00000342905.6 |
NLRC4 |
NLR family, CARD domain containing 4 |
| chr19_-_54974894 | 0.07 |
ENST00000333834.4 |
LENG9 |
leukocyte receptor cluster (LRC) member 9 |
| chr4_-_171011084 | 0.07 |
ENST00000337664.4 |
AADAT |
aminoadipate aminotransferase |
| chr1_-_146054494 | 0.07 |
ENST00000401009.2 |
NBPF11 |
neuroblastoma breakpoint family, member 11 |
| chr1_+_50569575 | 0.07 |
ENST00000371827.1 |
ELAVL4 |
ELAV like neuron-specific RNA binding protein 4 |
| chr1_-_44818599 | 0.07 |
ENST00000537474.1 |
ERI3 |
ERI1 exoribonuclease family member 3 |
| chr1_-_24306798 | 0.07 |
ENST00000374452.5 ENST00000492112.2 ENST00000343255.5 ENST00000344989.6 |
SRSF10 |
serine/arginine-rich splicing factor 10 |
| chr19_-_46999755 | 0.07 |
ENST00000599531.1 |
PNMAL2 |
paraneoplastic Ma antigen family-like 2 |
| chr9_+_18474163 | 0.07 |
ENST00000380566.4 |
ADAMTSL1 |
ADAMTS-like 1 |
| chr11_-_88796803 | 0.07 |
ENST00000418177.2 ENST00000455756.2 |
GRM5 |
glutamate receptor, metabotropic 5 |
| chr19_+_57901208 | 0.07 |
ENST00000366197.5 ENST00000596282.1 ENST00000597400.1 ENST00000598895.1 ENST00000336128.7 ENST00000596617.1 |
ZNF548 AC003002.6 |
zinc finger protein 548 Uncharacterized protein |
| chr9_-_125675576 | 0.07 |
ENST00000373659.3 |
ZBTB6 |
zinc finger and BTB domain containing 6 |
| chr2_-_182545603 | 0.07 |
ENST00000295108.3 |
NEUROD1 |
neuronal differentiation 1 |
| chr10_-_7513904 | 0.07 |
ENST00000420395.1 |
RP5-1031D4.2 |
RP5-1031D4.2 |
| chr14_+_77292715 | 0.07 |
ENST00000393774.3 ENST00000555189.1 ENST00000450042.2 |
C14orf166B |
chromosome 14 open reading frame 166B |
| chr20_-_30060816 | 0.07 |
ENST00000317676.2 |
DEFB124 |
defensin, beta 124 |
| chrX_-_13835147 | 0.07 |
ENST00000493677.1 ENST00000355135.2 |
GPM6B |
glycoprotein M6B |
| chrX_-_13752675 | 0.07 |
ENST00000380579.1 ENST00000458511.2 ENST00000519885.1 ENST00000358231.5 ENST00000518847.1 ENST00000453655.2 ENST00000359680.5 |
TRAPPC2 |
trafficking protein particle complex 2 |
| chr17_+_32907768 | 0.06 |
ENST00000321639.5 |
TMEM132E |
transmembrane protein 132E |
| chr2_+_186603355 | 0.06 |
ENST00000343098.5 |
FSIP2 |
fibrous sheath interacting protein 2 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.3 | GO:0048022 | negative regulation of melanin biosynthetic process(GO:0048022) negative regulation of secondary metabolite biosynthetic process(GO:1900377) |
| 0.3 | 0.8 | GO:2000097 | regulation of smooth muscle cell-matrix adhesion(GO:2000097) |
| 0.2 | 0.7 | GO:0014846 | esophagus smooth muscle contraction(GO:0014846) |
| 0.1 | 1.2 | GO:0007598 | blood coagulation, extrinsic pathway(GO:0007598) |
| 0.1 | 1.0 | GO:0036343 | psychomotor behavior(GO:0036343) |
| 0.1 | 0.6 | GO:0070171 | negative regulation of tooth mineralization(GO:0070171) |
| 0.1 | 0.5 | GO:0010716 | negative regulation of extracellular matrix disassembly(GO:0010716) melanocyte proliferation(GO:0097325) |
| 0.1 | 1.9 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.1 | 0.5 | GO:0061107 | seminal vesicle development(GO:0061107) |
| 0.1 | 0.4 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.1 | 1.6 | GO:0070493 | thrombin receptor signaling pathway(GO:0070493) |
| 0.1 | 0.6 | GO:1903944 | regulation of hepatocyte apoptotic process(GO:1903943) negative regulation of hepatocyte apoptotic process(GO:1903944) |
| 0.1 | 0.2 | GO:0045799 | mRNA export from nucleus in response to heat stress(GO:0031990) negative regulation of nucleobase-containing compound transport(GO:0032240) positive regulation of chromatin assembly or disassembly(GO:0045799) negative regulation of RNA export from nucleus(GO:0046832) |
| 0.1 | 0.3 | GO:0015688 | iron chelate transport(GO:0015688) siderophore transport(GO:0015891) |
| 0.1 | 0.2 | GO:0016095 | polyprenol catabolic process(GO:0016095) |
| 0.1 | 0.2 | GO:0001694 | histamine biosynthetic process(GO:0001694) |
| 0.1 | 0.3 | GO:0001880 | Mullerian duct regression(GO:0001880) |
| 0.1 | 0.3 | GO:0036111 | very long-chain fatty-acyl-CoA metabolic process(GO:0036111) |
| 0.0 | 0.3 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 0.0 | 0.2 | GO:0007525 | somatic muscle development(GO:0007525) |
| 0.0 | 0.1 | GO:2000777 | positive regulation of oocyte maturation(GO:1900195) positive regulation of proteasomal ubiquitin-dependent protein catabolic process involved in cellular response to hypoxia(GO:2000777) |
| 0.0 | 0.6 | GO:0036066 | protein O-linked fucosylation(GO:0036066) |
| 0.0 | 0.1 | GO:0009436 | glyoxylate catabolic process(GO:0009436) |
| 0.0 | 0.1 | GO:1902616 | acyl carnitine transport(GO:0006844) acyl carnitine transmembrane transport(GO:1902616) |
| 0.0 | 0.7 | GO:0043374 | CD8-positive, alpha-beta T cell differentiation(GO:0043374) |
| 0.0 | 0.2 | GO:0043328 | protein targeting to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043328) |
| 0.0 | 0.2 | GO:0006537 | glutamate biosynthetic process(GO:0006537) |
| 0.0 | 0.1 | GO:0003331 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) negative regulation of plasminogen activation(GO:0010757) |
| 0.0 | 0.1 | GO:2000360 | negative regulation of binding of sperm to zona pellucida(GO:2000360) |
| 0.0 | 0.1 | GO:0033512 | L-lysine catabolic process to acetyl-CoA via saccharopine(GO:0033512) |
| 0.0 | 0.2 | GO:0006574 | valine catabolic process(GO:0006574) |
| 0.0 | 0.1 | GO:0042631 | cellular response to water deprivation(GO:0042631) |
| 0.0 | 0.1 | GO:2000824 | negative regulation of androgen receptor activity(GO:2000824) |
| 0.0 | 0.5 | GO:0006995 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.0 | 0.5 | GO:0098828 | positive regulation of inhibitory postsynaptic potential(GO:0097151) modulation of inhibitory postsynaptic potential(GO:0098828) |
| 0.0 | 0.1 | GO:0090234 | regulation of kinetochore assembly(GO:0090234) |
| 0.0 | 0.1 | GO:0001812 | positive regulation of type I hypersensitivity(GO:0001812) |
| 0.0 | 0.1 | GO:2001106 | regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001106) |
| 0.0 | 0.1 | GO:0009240 | isopentenyl diphosphate biosynthetic process(GO:0009240) isopentenyl diphosphate metabolic process(GO:0046490) |
| 0.0 | 0.1 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 0.0 | 0.6 | GO:0032695 | negative regulation of interleukin-12 production(GO:0032695) |
| 0.0 | 0.1 | GO:0007206 | phospholipase C-activating G-protein coupled glutamate receptor signaling pathway(GO:0007206) |
| 0.0 | 0.4 | GO:0030854 | positive regulation of granulocyte differentiation(GO:0030854) |
| 0.0 | 0.2 | GO:1903361 | protein localization to basolateral plasma membrane(GO:1903361) |
| 0.0 | 0.4 | GO:0060009 | Sertoli cell development(GO:0060009) |
| 0.0 | 0.1 | GO:0090625 | mRNA cleavage involved in gene silencing by siRNA(GO:0090625) |
| 0.0 | 0.5 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.0 | 0.1 | GO:0018094 | protein polyglycylation(GO:0018094) |
| 0.0 | 0.1 | GO:1902630 | regulation of membrane hyperpolarization(GO:1902630) |
| 0.0 | 0.1 | GO:0051612 | negative regulation of neurotransmitter uptake(GO:0051581) serotonin uptake(GO:0051610) regulation of serotonin uptake(GO:0051611) negative regulation of serotonin uptake(GO:0051612) |
| 0.0 | 0.2 | GO:0010265 | SCF complex assembly(GO:0010265) |
| 0.0 | 0.5 | GO:1904707 | positive regulation of vascular smooth muscle cell proliferation(GO:1904707) |
| 0.0 | 0.4 | GO:0060712 | spongiotrophoblast layer development(GO:0060712) |
| 0.0 | 0.5 | GO:0030728 | ovulation(GO:0030728) |
| 0.0 | 0.2 | GO:0070269 | pyroptosis(GO:0070269) |
| 0.0 | 0.1 | GO:0043323 | regulation of natural killer cell degranulation(GO:0043321) positive regulation of natural killer cell degranulation(GO:0043323) |
| 0.0 | 0.3 | GO:0046069 | cGMP catabolic process(GO:0046069) |
| 0.0 | 0.1 | GO:0072757 | cellular response to camptothecin(GO:0072757) |
| 0.0 | 0.1 | GO:1990592 | protein polyufmylation(GO:1990564) protein K69-linked ufmylation(GO:1990592) |
| 0.0 | 0.1 | GO:0060729 | intestinal epithelial structure maintenance(GO:0060729) |
| 0.0 | 0.1 | GO:0003065 | positive regulation of heart rate by epinephrine(GO:0003065) |
| 0.0 | 0.3 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.0 | 0.2 | GO:0000244 | spliceosomal tri-snRNP complex assembly(GO:0000244) |
| 0.0 | 0.1 | GO:0003070 | age-dependent response to oxidative stress(GO:0001306) age-dependent response to reactive oxygen species(GO:0001315) regulation of systemic arterial blood pressure by acetylcholine(GO:0003068) vasodilation by acetylcholine involved in regulation of systemic arterial blood pressure(GO:0003069) regulation of systemic arterial blood pressure by neurotransmitter(GO:0003070) age-dependent general metabolic decline(GO:0007571) |
| 0.0 | 0.0 | GO:0009138 | pyrimidine nucleoside diphosphate metabolic process(GO:0009138) |
| 0.0 | 0.5 | GO:1901685 | glutathione derivative metabolic process(GO:1901685) glutathione derivative biosynthetic process(GO:1901687) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0036026 | protein C inhibitor-TMPRSS7 complex(GO:0036024) protein C inhibitor-TMPRSS11E complex(GO:0036025) protein C inhibitor-PLAT complex(GO:0036026) protein C inhibitor-PLAU complex(GO:0036027) protein C inhibitor-thrombin complex(GO:0036028) protein C inhibitor-KLK3 complex(GO:0036029) protein C inhibitor-plasma kallikrein complex(GO:0036030) serine protease inhibitor complex(GO:0097180) protein C inhibitor-coagulation factor V complex(GO:0097181) protein C inhibitor-coagulation factor Xa complex(GO:0097182) protein C inhibitor-coagulation factor XI complex(GO:0097183) |
| 0.2 | 0.6 | GO:0097442 | CA3 pyramidal cell dendrite(GO:0097442) |
| 0.1 | 1.9 | GO:0098647 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.1 | 0.2 | GO:0000814 | ESCRT II complex(GO:0000814) |
| 0.0 | 0.5 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.0 | 0.2 | GO:0000275 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) |
| 0.0 | 0.6 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) ripoptosome(GO:0097342) |
| 0.0 | 0.3 | GO:0031467 | Cul7-RING ubiquitin ligase complex(GO:0031467) |
| 0.0 | 0.1 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.0 | 0.4 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.0 | 0.1 | GO:0097452 | GAIT complex(GO:0097452) |
| 0.0 | 0.3 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.0 | 0.8 | GO:0005720 | nuclear heterochromatin(GO:0005720) |
| 0.0 | 0.2 | GO:0072557 | IPAF inflammasome complex(GO:0072557) |
| 0.0 | 0.1 | GO:0032444 | activin responsive factor complex(GO:0032444) |
| 0.0 | 0.1 | GO:0016013 | syntrophin complex(GO:0016013) |
| 0.0 | 0.1 | GO:0005947 | mitochondrial alpha-ketoglutarate dehydrogenase complex(GO:0005947) |
| 0.0 | 0.1 | GO:0044305 | calyx of Held(GO:0044305) |
| 0.0 | 0.3 | GO:0016471 | vacuolar proton-transporting V-type ATPase complex(GO:0016471) |
| 0.0 | 0.1 | GO:0000125 | PCAF complex(GO:0000125) |
| 0.0 | 0.1 | GO:0035068 | micro-ribonucleoprotein complex(GO:0035068) |
| 0.0 | 0.5 | GO:0048786 | presynaptic active zone(GO:0048786) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.6 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.1 | 0.5 | GO:0001512 | dihydronicotinamide riboside quinone reductase activity(GO:0001512) melatonin binding(GO:1904408) |
| 0.1 | 0.7 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.1 | 0.2 | GO:0052836 | inositol 5-diphosphate pentakisphosphate 5-kinase activity(GO:0052836) inositol diphosphate tetrakisphosphate kinase activity(GO:0052839) |
| 0.1 | 0.2 | GO:0016639 | oxidoreductase activity, acting on the CH-NH2 group of donors, NAD or NADP as acceptor(GO:0016639) |
| 0.1 | 0.3 | GO:0031697 | beta-1 adrenergic receptor binding(GO:0031697) |
| 0.1 | 0.2 | GO:0047718 | indanol dehydrogenase activity(GO:0047718) |
| 0.1 | 0.3 | GO:0003998 | acylphosphatase activity(GO:0003998) |
| 0.1 | 0.3 | GO:0033989 | 3alpha,7alpha,12alpha-trihydroxy-5beta-cholest-24-enoyl-CoA hydratase activity(GO:0033989) 17-beta-hydroxysteroid dehydrogenase (NAD+) activity(GO:0044594) |
| 0.0 | 0.5 | GO:0032190 | acrosin binding(GO:0032190) |
| 0.0 | 0.3 | GO:0003846 | 2-acylglycerol O-acyltransferase activity(GO:0003846) |
| 0.0 | 0.2 | GO:0016160 | amylase activity(GO:0016160) |
| 0.0 | 0.1 | GO:0008192 | RNA guanylyltransferase activity(GO:0008192) |
| 0.0 | 0.1 | GO:0034739 | histone deacetylase activity (H4-K16 specific)(GO:0034739) |
| 0.0 | 0.2 | GO:0004616 | phosphogluconate dehydrogenase (decarboxylating) activity(GO:0004616) |
| 0.0 | 0.1 | GO:0015227 | acyl carnitine transmembrane transporter activity(GO:0015227) |
| 0.0 | 0.3 | GO:0048101 | calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 0.0 | 0.1 | GO:0003896 | DNA primase activity(GO:0003896) |
| 0.0 | 0.3 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.2 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.0 | 0.1 | GO:0032184 | SUMO polymer binding(GO:0032184) |
| 0.0 | 0.1 | GO:0047179 | platelet-activating factor acetyltransferase activity(GO:0047179) |
| 0.0 | 0.1 | GO:0070137 | ubiquitin-like protein-specific endopeptidase activity(GO:0070137) SUMO-specific endopeptidase activity(GO:0070139) |
| 0.0 | 0.1 | GO:0016212 | kynurenine-oxoglutarate transaminase activity(GO:0016212) kynurenine aminotransferase activity(GO:0036137) |
| 0.0 | 0.1 | GO:0008453 | alanine-glyoxylate transaminase activity(GO:0008453) |
| 0.0 | 0.5 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.0 | 1.9 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.1 | GO:0070735 | protein-glycine ligase activity(GO:0070735) |
| 0.0 | 0.5 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.0 | 0.4 | GO:0005522 | profilin binding(GO:0005522) |
| 0.0 | 0.5 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.0 | 0.6 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.0 | 0.5 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 0.1 | GO:0019863 | IgE binding(GO:0019863) |
| 0.0 | 0.3 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.0 | 0.1 | GO:0005502 | 11-cis retinal binding(GO:0005502) |
| 0.0 | 0.1 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) |
| 0.0 | 0.4 | GO:0001637 | G-protein coupled chemoattractant receptor activity(GO:0001637) chemokine receptor activity(GO:0004950) |
| 0.0 | 0.2 | GO:0030618 | transforming growth factor beta receptor, pathway-specific cytoplasmic mediator activity(GO:0030618) |
| 0.0 | 0.8 | GO:0015485 | cholesterol binding(GO:0015485) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.6 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.1 | 1.2 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 2.4 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.6 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.9 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.0 | 0.5 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.0 | 1.2 | REACTOME FORMATION OF FIBRIN CLOT CLOTTING CASCADE | Genes involved in Formation of Fibrin Clot (Clotting Cascade) |
| 0.0 | 1.5 | REACTOME THROMBIN SIGNALLING THROUGH PROTEINASE ACTIVATED RECEPTORS PARS | Genes involved in Thrombin signalling through proteinase activated receptors (PARs) |
| 0.0 | 0.5 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 0.6 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 0.1 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 0.0 | 0.3 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.0 | 0.5 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |