Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
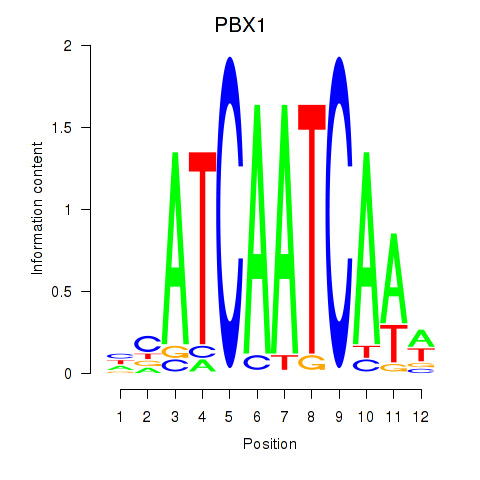
Results for PBX1
Z-value: 1.12
Transcription factors associated with PBX1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
PBX1
|
ENSG00000185630.14 | PBX1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| PBX1 | hg19_v2_chr1_+_164528866_164528877 | 0.87 | 4.9e-03 | Click! |
Activity profile of PBX1 motif
Sorted Z-values of PBX1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of PBX1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr14_-_60097297 | 1.11 |
ENST00000395090.1 |
RTN1 |
reticulon 1 |
| chr8_-_108510224 | 1.10 |
ENST00000517746.1 ENST00000297450.3 |
ANGPT1 |
angiopoietin 1 |
| chr2_-_190044480 | 1.07 |
ENST00000374866.3 |
COL5A2 |
collagen, type V, alpha 2 |
| chr10_+_124320156 | 1.04 |
ENST00000338354.3 ENST00000344338.3 ENST00000330163.4 ENST00000368909.3 ENST00000368955.3 ENST00000368956.2 |
DMBT1 |
deleted in malignant brain tumors 1 |
| chr10_+_69865866 | 1.02 |
ENST00000354393.2 |
MYPN |
myopalladin |
| chr10_+_124320195 | 1.01 |
ENST00000359586.6 |
DMBT1 |
deleted in malignant brain tumors 1 |
| chr16_+_15596123 | 0.94 |
ENST00000452191.2 |
C16orf45 |
chromosome 16 open reading frame 45 |
| chr17_+_1665345 | 0.84 |
ENST00000576406.1 ENST00000571149.1 |
SERPINF1 |
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 |
| chr6_-_167040731 | 0.75 |
ENST00000265678.4 |
RPS6KA2 |
ribosomal protein S6 kinase, 90kDa, polypeptide 2 |
| chr12_+_21679220 | 0.67 |
ENST00000256969.2 |
C12orf39 |
chromosome 12 open reading frame 39 |
| chr16_+_55512742 | 0.64 |
ENST00000568715.1 ENST00000219070.4 |
MMP2 |
matrix metallopeptidase 2 (gelatinase A, 72kDa gelatinase, 72kDa type IV collagenase) |
| chr2_-_200323414 | 0.63 |
ENST00000443023.1 |
SATB2 |
SATB homeobox 2 |
| chr7_-_27142290 | 0.62 |
ENST00000222718.5 |
HOXA2 |
homeobox A2 |
| chr13_-_33760216 | 0.57 |
ENST00000255486.4 |
STARD13 |
StAR-related lipid transfer (START) domain containing 13 |
| chr7_+_120629653 | 0.53 |
ENST00000450913.2 ENST00000340646.5 |
CPED1 |
cadherin-like and PC-esterase domain containing 1 |
| chrX_-_92928557 | 0.52 |
ENST00000373079.3 ENST00000475430.2 |
NAP1L3 |
nucleosome assembly protein 1-like 3 |
| chr5_-_10761206 | 0.51 |
ENST00000432074.2 ENST00000230895.6 |
DAP |
death-associated protein |
| chr2_-_200322723 | 0.50 |
ENST00000417098.1 |
SATB2 |
SATB homeobox 2 |
| chr2_+_20646824 | 0.48 |
ENST00000272233.4 |
RHOB |
ras homolog family member B |
| chr4_-_70626430 | 0.47 |
ENST00000310613.3 |
SULT1B1 |
sulfotransferase family, cytosolic, 1B, member 1 |
| chr16_-_88729473 | 0.46 |
ENST00000301012.3 ENST00000569177.1 |
MVD |
mevalonate (diphospho) decarboxylase |
| chr7_-_7782204 | 0.45 |
ENST00000418534.2 |
AC007161.5 |
AC007161.5 |
| chr14_-_74551096 | 0.41 |
ENST00000350259.4 |
ALDH6A1 |
aldehyde dehydrogenase 6 family, member A1 |
| chr17_+_58755184 | 0.39 |
ENST00000589222.1 ENST00000407086.3 ENST00000390652.5 |
BCAS3 |
breast carcinoma amplified sequence 3 |
| chr2_+_12857015 | 0.37 |
ENST00000155926.4 |
TRIB2 |
tribbles pseudokinase 2 |
| chr7_-_102184083 | 0.36 |
ENST00000379357.5 |
POLR2J3 |
polymerase (RNA) II (DNA directed) polypeptide J3 |
| chr2_+_12857043 | 0.36 |
ENST00000381465.2 |
TRIB2 |
tribbles pseudokinase 2 |
| chr3_-_169587621 | 0.36 |
ENST00000523069.1 ENST00000316428.5 ENST00000264676.5 |
LRRC31 |
leucine rich repeat containing 31 |
| chrX_-_70474377 | 0.35 |
ENST00000373978.1 ENST00000373981.1 |
ZMYM3 |
zinc finger, MYM-type 3 |
| chr2_+_62132800 | 0.32 |
ENST00000538736.1 |
COMMD1 |
copper metabolism (Murr1) domain containing 1 |
| chr15_-_37393406 | 0.32 |
ENST00000338564.5 ENST00000558313.1 ENST00000340545.5 |
MEIS2 |
Meis homeobox 2 |
| chr14_+_88851874 | 0.32 |
ENST00000393545.4 ENST00000356583.5 ENST00000555401.1 ENST00000553885.1 |
SPATA7 |
spermatogenesis associated 7 |
| chr2_+_201997492 | 0.31 |
ENST00000494258.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr22_-_39190116 | 0.31 |
ENST00000406622.1 ENST00000216068.4 ENST00000406199.3 |
SUN2 DNAL4 |
Sad1 and UNC84 domain containing 2 dynein, axonemal, light chain 4 |
| chr12_+_79258444 | 0.31 |
ENST00000261205.4 |
SYT1 |
synaptotagmin I |
| chr6_+_151561085 | 0.30 |
ENST00000402676.2 |
AKAP12 |
A kinase (PRKA) anchor protein 12 |
| chr14_-_74551172 | 0.30 |
ENST00000553458.1 |
ALDH6A1 |
aldehyde dehydrogenase 6 family, member A1 |
| chr1_-_146068184 | 0.30 |
ENST00000604894.1 ENST00000369323.3 ENST00000479926.2 |
NBPF11 |
neuroblastoma breakpoint family, member 11 |
| chrX_-_70473957 | 0.30 |
ENST00000373984.3 ENST00000314425.5 ENST00000373982.1 |
ZMYM3 |
zinc finger, MYM-type 3 |
| chrX_-_63005405 | 0.29 |
ENST00000374878.1 ENST00000437457.2 |
ARHGEF9 |
Cdc42 guanine nucleotide exchange factor (GEF) 9 |
| chr11_+_114166536 | 0.29 |
ENST00000299964.3 |
NNMT |
nicotinamide N-methyltransferase |
| chr2_+_62132781 | 0.29 |
ENST00000311832.5 |
COMMD1 |
copper metabolism (Murr1) domain containing 1 |
| chr6_-_15586238 | 0.28 |
ENST00000462989.2 |
DTNBP1 |
dystrobrevin binding protein 1 |
| chr12_-_56615693 | 0.28 |
ENST00000394013.2 ENST00000345093.4 ENST00000551711.1 ENST00000552656.1 |
RNF41 |
ring finger protein 41 |
| chrX_-_102757802 | 0.27 |
ENST00000372633.1 |
RAB40A |
RAB40A, member RAS oncogene family |
| chr11_-_34937858 | 0.26 |
ENST00000278359.5 |
APIP |
APAF1 interacting protein |
| chr21_+_27011899 | 0.26 |
ENST00000425221.2 |
JAM2 |
junctional adhesion molecule 2 |
| chr7_-_120498357 | 0.26 |
ENST00000415871.1 ENST00000222747.3 ENST00000430985.1 |
TSPAN12 |
tetraspanin 12 |
| chr14_+_29236269 | 0.26 |
ENST00000313071.4 |
FOXG1 |
forkhead box G1 |
| chr14_+_24583836 | 0.26 |
ENST00000559115.1 ENST00000558215.1 ENST00000557810.1 ENST00000561375.1 ENST00000446197.3 ENST00000559796.1 ENST00000560713.1 ENST00000560901.1 ENST00000559382.1 |
DCAF11 |
DDB1 and CUL4 associated factor 11 |
| chr12_+_56401268 | 0.26 |
ENST00000262032.5 |
IKZF4 |
IKAROS family zinc finger 4 (Eos) |
| chr9_-_14308004 | 0.25 |
ENST00000493697.1 |
NFIB |
nuclear factor I/B |
| chr5_-_171881362 | 0.24 |
ENST00000519643.1 |
SH3PXD2B |
SH3 and PX domains 2B |
| chr7_+_134331550 | 0.24 |
ENST00000344924.3 ENST00000418040.1 ENST00000393132.2 |
BPGM |
2,3-bisphosphoglycerate mutase |
| chr22_-_30866564 | 0.24 |
ENST00000435069.1 ENST00000415957.2 ENST00000540910.1 |
SEC14L3 |
SEC14-like 3 (S. cerevisiae) |
| chr17_+_4853442 | 0.24 |
ENST00000522301.1 |
ENO3 |
enolase 3 (beta, muscle) |
| chr2_+_201997676 | 0.24 |
ENST00000462763.1 ENST00000479953.2 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr15_+_44069276 | 0.23 |
ENST00000381359.1 |
SERF2 |
small EDRK-rich factor 2 |
| chr1_-_147610081 | 0.23 |
ENST00000369226.3 |
NBPF24 |
neuroblastoma breakpoint family, member 24 |
| chr5_+_43603229 | 0.23 |
ENST00000344920.4 ENST00000512996.2 |
NNT |
nicotinamide nucleotide transhydrogenase |
| chr12_+_123944070 | 0.23 |
ENST00000412157.2 |
SNRNP35 |
small nuclear ribonucleoprotein 35kDa (U11/U12) |
| chr4_+_83956237 | 0.22 |
ENST00000264389.2 |
COPS4 |
COP9 signalosome subunit 4 |
| chr6_+_27114861 | 0.22 |
ENST00000377459.1 |
HIST1H2AH |
histone cluster 1, H2ah |
| chr1_+_104159999 | 0.22 |
ENST00000414303.2 ENST00000423678.1 |
AMY2A |
amylase, alpha 2A (pancreatic) |
| chr12_-_91573249 | 0.22 |
ENST00000550099.1 ENST00000546391.1 ENST00000551354.1 |
DCN |
decorin |
| chr2_+_201997595 | 0.22 |
ENST00000470178.2 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr12_+_79258547 | 0.22 |
ENST00000457153.2 |
SYT1 |
synaptotagmin I |
| chr1_-_54405773 | 0.21 |
ENST00000371376.1 |
HSPB11 |
heat shock protein family B (small), member 11 |
| chr8_-_116681221 | 0.21 |
ENST00000395715.3 |
TRPS1 |
trichorhinophalangeal syndrome I |
| chr12_-_102455846 | 0.21 |
ENST00000545679.1 |
CCDC53 |
coiled-coil domain containing 53 |
| chr22_+_35695793 | 0.21 |
ENST00000456128.1 ENST00000449058.2 ENST00000411850.1 ENST00000425375.1 ENST00000436462.2 ENST00000382034.5 |
TOM1 |
target of myb1 (chicken) |
| chr1_+_104293028 | 0.21 |
ENST00000370079.3 |
AMY1C |
amylase, alpha 1C (salivary) |
| chr10_+_50887683 | 0.20 |
ENST00000374113.3 ENST00000374111.3 ENST00000374112.3 ENST00000535836.1 |
C10orf53 |
chromosome 10 open reading frame 53 |
| chr2_+_44396000 | 0.20 |
ENST00000409895.4 ENST00000409432.3 ENST00000282412.4 ENST00000378551.2 ENST00000345249.4 |
PPM1B |
protein phosphatase, Mg2+/Mn2+ dependent, 1B |
| chr12_-_102455902 | 0.20 |
ENST00000240079.6 |
CCDC53 |
coiled-coil domain containing 53 |
| chrX_+_120181457 | 0.20 |
ENST00000328078.1 |
GLUD2 |
glutamate dehydrogenase 2 |
| chr21_+_27011584 | 0.20 |
ENST00000400532.1 ENST00000480456.1 ENST00000312957.5 |
JAM2 |
junctional adhesion molecule 2 |
| chr4_+_83956312 | 0.19 |
ENST00000509317.1 ENST00000503682.1 ENST00000511653.1 |
COPS4 |
COP9 signalosome subunit 4 |
| chrX_-_70474499 | 0.19 |
ENST00000353904.2 |
ZMYM3 |
zinc finger, MYM-type 3 |
| chr5_-_125930929 | 0.19 |
ENST00000553117.1 ENST00000447989.2 ENST00000409134.3 |
ALDH7A1 |
aldehyde dehydrogenase 7 family, member A1 |
| chr6_+_36839616 | 0.18 |
ENST00000359359.2 ENST00000510325.2 |
C6orf89 |
chromosome 6 open reading frame 89 |
| chr10_-_31288398 | 0.18 |
ENST00000538351.2 |
ZNF438 |
zinc finger protein 438 |
| chr9_-_73736511 | 0.18 |
ENST00000377110.3 ENST00000377111.2 |
TRPM3 |
transient receptor potential cation channel, subfamily M, member 3 |
| chrX_-_65253506 | 0.18 |
ENST00000427538.1 |
VSIG4 |
V-set and immunoglobulin domain containing 4 |
| chr3_+_148508845 | 0.18 |
ENST00000491148.1 |
CPB1 |
carboxypeptidase B1 (tissue) |
| chr12_-_54653313 | 0.18 |
ENST00000550411.1 ENST00000439541.2 |
CBX5 |
chromobox homolog 5 |
| chr21_-_35014027 | 0.17 |
ENST00000399442.1 ENST00000413017.2 ENST00000445393.1 ENST00000417979.1 ENST00000426935.1 ENST00000381540.3 ENST00000361534.2 ENST00000381554.3 |
CRYZL1 |
crystallin, zeta (quinone reductase)-like 1 |
| chr6_+_27775899 | 0.17 |
ENST00000358739.3 |
HIST1H2AI |
histone cluster 1, H2ai |
| chr3_+_129159039 | 0.17 |
ENST00000507564.1 ENST00000431818.2 ENST00000504021.1 ENST00000349441.2 ENST00000348417.2 ENST00000440957.2 |
IFT122 |
intraflagellar transport 122 homolog (Chlamydomonas) |
| chr5_-_133702761 | 0.17 |
ENST00000521118.1 ENST00000265334.4 ENST00000435211.1 |
CDKL3 |
cyclin-dependent kinase-like 3 |
| chr19_-_58400148 | 0.17 |
ENST00000595048.1 ENST00000600634.1 ENST00000595295.1 ENST00000596604.1 ENST00000597342.1 ENST00000597807.1 |
ZNF814 |
zinc finger protein 814 |
| chr5_+_162864575 | 0.16 |
ENST00000512163.1 ENST00000393929.1 ENST00000340828.2 ENST00000511683.2 ENST00000510097.1 ENST00000511490.2 ENST00000510664.1 |
CCNG1 |
cyclin G1 |
| chr3_+_129158926 | 0.16 |
ENST00000347300.2 ENST00000296266.3 |
IFT122 |
intraflagellar transport 122 homolog (Chlamydomonas) |
| chr4_-_152147579 | 0.16 |
ENST00000304527.4 ENST00000455740.1 ENST00000424281.1 ENST00000409598.4 |
SH3D19 |
SH3 domain containing 19 |
| chr16_-_30546141 | 0.16 |
ENST00000535210.1 ENST00000395094.3 |
ZNF747 |
zinc finger protein 747 |
| chr5_+_43602750 | 0.16 |
ENST00000505678.2 ENST00000512422.1 ENST00000264663.5 |
NNT |
nicotinamide nucleotide transhydrogenase |
| chr4_-_185776789 | 0.15 |
ENST00000510284.1 |
RP11-701P16.5 |
RP11-701P16.5 |
| chr4_-_185776854 | 0.15 |
ENST00000511703.1 |
RP11-701P16.5 |
RP11-701P16.5 |
| chr12_-_76478686 | 0.15 |
ENST00000261182.8 |
NAP1L1 |
nucleosome assembly protein 1-like 1 |
| chr19_+_44645700 | 0.15 |
ENST00000592437.1 |
ZNF234 |
zinc finger protein 234 |
| chr11_+_71903169 | 0.15 |
ENST00000393676.3 |
FOLR1 |
folate receptor 1 (adult) |
| chr11_-_34938039 | 0.15 |
ENST00000395787.3 |
APIP |
APAF1 interacting protein |
| chr7_-_72971934 | 0.15 |
ENST00000411832.1 |
BCL7B |
B-cell CLL/lymphoma 7B |
| chr19_-_51141196 | 0.15 |
ENST00000338916.4 |
SYT3 |
synaptotagmin III |
| chr1_-_1690014 | 0.15 |
ENST00000400922.2 ENST00000342348.5 |
NADK |
NAD kinase |
| chr14_+_50234309 | 0.14 |
ENST00000298307.5 |
KLHDC2 |
kelch domain containing 2 |
| chr11_+_95523823 | 0.14 |
ENST00000538658.1 |
CEP57 |
centrosomal protein 57kDa |
| chr2_-_242626127 | 0.14 |
ENST00000445261.1 |
DTYMK |
deoxythymidylate kinase (thymidylate kinase) |
| chr4_+_120056939 | 0.14 |
ENST00000307128.5 |
MYOZ2 |
myozenin 2 |
| chr10_-_31146615 | 0.14 |
ENST00000444692.2 |
ZNF438 |
zinc finger protein 438 |
| chr3_+_159557637 | 0.14 |
ENST00000445224.2 |
SCHIP1 |
schwannomin interacting protein 1 |
| chr6_+_151561506 | 0.14 |
ENST00000253332.1 |
AKAP12 |
A kinase (PRKA) anchor protein 12 |
| chr16_+_16484691 | 0.13 |
ENST00000344087.4 |
NPIPA7 |
nuclear pore complex interacting protein family, member A7 |
| chr5_-_179045199 | 0.13 |
ENST00000523921.1 |
HNRNPH1 |
heterogeneous nuclear ribonucleoprotein H1 (H) |
| chr14_-_69619823 | 0.13 |
ENST00000341516.5 |
DCAF5 |
DDB1 and CUL4 associated factor 5 |
| chr11_-_2162162 | 0.13 |
ENST00000381389.1 |
IGF2 |
insulin-like growth factor 2 (somatomedin A) |
| chr15_+_43477455 | 0.13 |
ENST00000300213.4 |
CCNDBP1 |
cyclin D-type binding-protein 1 |
| chr3_-_187388173 | 0.13 |
ENST00000287641.3 |
SST |
somatostatin |
| chr6_+_167704838 | 0.13 |
ENST00000366829.2 |
UNC93A |
unc-93 homolog A (C. elegans) |
| chr10_+_63422695 | 0.12 |
ENST00000330194.2 ENST00000389639.3 |
C10orf107 |
chromosome 10 open reading frame 107 |
| chr1_-_115323245 | 0.12 |
ENST00000060969.5 ENST00000369528.5 |
SIKE1 |
suppressor of IKBKE 1 |
| chr7_-_7680601 | 0.12 |
ENST00000396682.2 |
RPA3 |
replication protein A3, 14kDa |
| chr4_-_140005443 | 0.12 |
ENST00000510408.1 ENST00000420916.2 ENST00000358635.3 |
ELF2 |
E74-like factor 2 (ets domain transcription factor) |
| chr1_-_146054494 | 0.12 |
ENST00000401009.2 |
NBPF11 |
neuroblastoma breakpoint family, member 11 |
| chr16_+_19079215 | 0.12 |
ENST00000544894.2 ENST00000561858.1 |
COQ7 |
coenzyme Q7 homolog, ubiquinone (yeast) |
| chr2_-_154335300 | 0.12 |
ENST00000325926.3 |
RPRM |
reprimo, TP53 dependent G2 arrest mediator candidate |
| chr17_+_43239231 | 0.11 |
ENST00000591576.1 ENST00000591070.1 ENST00000592695.1 |
HEXIM2 |
hexamethylene bis-acetamide inducible 2 |
| chr13_-_24895566 | 0.11 |
ENST00000422229.2 |
AL359736.1 |
protein PCOTH isoform 1 |
| chr13_-_46679144 | 0.11 |
ENST00000181383.4 |
CPB2 |
carboxypeptidase B2 (plasma) |
| chr11_-_126870655 | 0.11 |
ENST00000525144.2 |
KIRREL3 |
kin of IRRE like 3 (Drosophila) |
| chr17_-_40828969 | 0.11 |
ENST00000591022.1 ENST00000587627.1 ENST00000293349.6 |
PLEKHH3 |
pleckstrin homology domain containing, family H (with MyTH4 domain) member 3 |
| chr16_+_69221028 | 0.11 |
ENST00000336278.4 |
SNTB2 |
syntrophin, beta 2 (dystrophin-associated protein A1, 59kDa, basic component 2) |
| chr3_+_52570610 | 0.11 |
ENST00000307106.3 ENST00000477703.1 ENST00000476842.1 |
SMIM4 |
small integral membrane protein 4 |
| chr11_-_31531121 | 0.10 |
ENST00000532287.1 ENST00000526776.1 ENST00000534812.1 ENST00000529749.1 ENST00000278200.1 ENST00000530023.1 ENST00000533642.1 |
IMMP1L |
IMP1 inner mitochondrial membrane peptidase-like (S. cerevisiae) |
| chr6_-_110736742 | 0.10 |
ENST00000368924.3 ENST00000368923.3 |
DDO |
D-aspartate oxidase |
| chrX_-_50386648 | 0.10 |
ENST00000460112.3 |
SHROOM4 |
shroom family member 4 |
| chr16_+_69458537 | 0.10 |
ENST00000515314.1 ENST00000561792.1 ENST00000568237.1 |
CYB5B |
cytochrome b5 type B (outer mitochondrial membrane) |
| chr2_-_88427568 | 0.10 |
ENST00000393750.3 ENST00000295834.3 |
FABP1 |
fatty acid binding protein 1, liver |
| chr13_-_46679185 | 0.10 |
ENST00000439329.3 |
CPB2 |
carboxypeptidase B2 (plasma) |
| chr5_+_98109322 | 0.10 |
ENST00000513185.1 |
RGMB |
repulsive guidance molecule family member b |
| chr19_-_5903714 | 0.09 |
ENST00000586349.1 ENST00000585661.1 ENST00000308961.4 ENST00000592634.1 ENST00000418389.2 ENST00000252675.5 |
AC024592.12 NDUFA11 FUT5 |
Uncharacterized protein NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 11, 14.7kDa fucosyltransferase 5 (alpha (1,3) fucosyltransferase) |
| chr4_-_52883786 | 0.09 |
ENST00000343457.3 |
LRRC66 |
leucine rich repeat containing 66 |
| chr9_-_32573130 | 0.09 |
ENST00000350021.2 ENST00000379847.3 |
NDUFB6 |
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 6, 17kDa |
| chr14_+_50065459 | 0.09 |
ENST00000318317.4 |
LRR1 |
leucine rich repeat protein 1 |
| chr21_+_44313375 | 0.09 |
ENST00000354250.2 ENST00000340344.4 |
NDUFV3 |
NADH dehydrogenase (ubiquinone) flavoprotein 3, 10kDa |
| chrM_-_14670 | 0.09 |
ENST00000361681.2 |
MT-ND6 |
mitochondrially encoded NADH dehydrogenase 6 |
| chr2_-_27886676 | 0.09 |
ENST00000337768.5 |
SUPT7L |
suppressor of Ty 7 (S. cerevisiae)-like |
| chr13_+_98086445 | 0.09 |
ENST00000245304.4 |
RAP2A |
RAP2A, member of RAS oncogene family |
| chr3_-_71353892 | 0.09 |
ENST00000484350.1 |
FOXP1 |
forkhead box P1 |
| chr2_-_27886460 | 0.09 |
ENST00000404798.2 ENST00000405491.1 ENST00000464789.2 ENST00000406540.1 |
SUPT7L |
suppressor of Ty 7 (S. cerevisiae)-like |
| chr20_+_44441271 | 0.09 |
ENST00000335046.3 ENST00000243893.6 |
UBE2C |
ubiquitin-conjugating enzyme E2C |
| chr14_+_50065376 | 0.09 |
ENST00000298288.6 |
LRR1 |
leucine rich repeat protein 1 |
| chr2_+_79412357 | 0.08 |
ENST00000466387.1 |
CTNNA2 |
catenin (cadherin-associated protein), alpha 2 |
| chr4_-_113437328 | 0.08 |
ENST00000313341.3 |
NEUROG2 |
neurogenin 2 |
| chr5_+_68513622 | 0.08 |
ENST00000512880.1 ENST00000602380.1 |
MRPS36 |
mitochondrial ribosomal protein S36 |
| chr18_-_48346415 | 0.08 |
ENST00000431965.2 ENST00000436348.2 |
MRO |
maestro |
| chr4_+_175839551 | 0.08 |
ENST00000404450.4 ENST00000514159.1 |
ADAM29 |
ADAM metallopeptidase domain 29 |
| chr4_-_140005341 | 0.08 |
ENST00000379549.2 ENST00000512627.1 |
ELF2 |
E74-like factor 2 (ets domain transcription factor) |
| chr16_-_20364030 | 0.08 |
ENST00000396134.2 ENST00000573567.1 ENST00000570757.1 ENST00000424589.1 ENST00000302509.4 ENST00000571174.1 ENST00000576688.1 |
UMOD |
uromodulin |
| chr17_-_40829026 | 0.08 |
ENST00000412503.1 |
PLEKHH3 |
pleckstrin homology domain containing, family H (with MyTH4 domain) member 3 |
| chr2_+_211342432 | 0.08 |
ENST00000430249.2 |
CPS1 |
carbamoyl-phosphate synthase 1, mitochondrial |
| chr7_+_80275752 | 0.08 |
ENST00000419819.2 |
CD36 |
CD36 molecule (thrombospondin receptor) |
| chr12_+_133657461 | 0.08 |
ENST00000412146.2 ENST00000544426.1 ENST00000440984.2 ENST00000319849.3 ENST00000440550.2 |
ZNF140 |
zinc finger protein 140 |
| chrX_+_53123314 | 0.08 |
ENST00000605526.1 ENST00000604062.1 ENST00000604369.1 ENST00000366185.2 ENST00000604849.1 |
RP11-258C19.5 |
long intergenic non-protein coding RNA 1155 |
| chr2_-_10587897 | 0.08 |
ENST00000405333.1 ENST00000443218.1 |
ODC1 |
ornithine decarboxylase 1 |
| chr2_+_169926047 | 0.08 |
ENST00000428522.1 ENST00000450153.1 ENST00000421653.1 |
DHRS9 |
dehydrogenase/reductase (SDR family) member 9 |
| chr11_+_95523621 | 0.07 |
ENST00000325542.5 ENST00000325486.5 ENST00000544522.1 ENST00000541365.1 |
CEP57 |
centrosomal protein 57kDa |
| chr12_+_21525818 | 0.07 |
ENST00000240652.3 ENST00000542023.1 ENST00000537593.1 |
IAPP |
islet amyloid polypeptide |
| chr1_+_226250379 | 0.07 |
ENST00000366815.3 ENST00000366814.3 |
H3F3A |
H3 histone, family 3A |
| chr7_-_77325545 | 0.07 |
ENST00000447009.1 ENST00000416650.1 ENST00000440088.1 ENST00000430801.1 ENST00000398043.2 |
RSBN1L-AS1 |
RSBN1L antisense RNA 1 |
| chr16_-_20364122 | 0.07 |
ENST00000396138.4 ENST00000577168.1 |
UMOD |
uromodulin |
| chr3_+_155860751 | 0.07 |
ENST00000471742.1 |
KCNAB1 |
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chr19_+_49999631 | 0.07 |
ENST00000270625.2 ENST00000596873.1 ENST00000594493.1 ENST00000599561.1 |
RPS11 |
ribosomal protein S11 |
| chr17_-_26694979 | 0.07 |
ENST00000438614.1 |
VTN |
vitronectin |
| chr5_-_42825983 | 0.07 |
ENST00000506577.1 |
SEPP1 |
selenoprotein P, plasma, 1 |
| chr15_+_34638066 | 0.06 |
ENST00000333756.4 |
NUTM1 |
NUT midline carcinoma, family member 1 |
| chr3_+_159570722 | 0.06 |
ENST00000482804.1 |
SCHIP1 |
schwannomin interacting protein 1 |
| chr16_+_19078960 | 0.06 |
ENST00000568985.1 ENST00000566110.1 |
COQ7 |
coenzyme Q7 homolog, ubiquinone (yeast) |
| chr17_+_8191815 | 0.06 |
ENST00000226105.6 ENST00000407006.4 ENST00000580434.1 ENST00000439238.3 |
RANGRF |
RAN guanine nucleotide release factor |
| chr9_-_123676827 | 0.06 |
ENST00000546084.1 |
TRAF1 |
TNF receptor-associated factor 1 |
| chr2_+_264869 | 0.06 |
ENST00000272067.6 ENST00000272065.5 ENST00000407983.3 |
ACP1 |
acid phosphatase 1, soluble |
| chr17_-_59668550 | 0.06 |
ENST00000521764.1 |
NACA2 |
nascent polypeptide-associated complex alpha subunit 2 |
| chr6_-_36725157 | 0.06 |
ENST00000393189.2 |
CPNE5 |
copine V |
| chr14_-_36789783 | 0.06 |
ENST00000605579.1 ENST00000604336.1 ENST00000359527.7 ENST00000603139.1 ENST00000318473.7 |
MBIP |
MAP3K12 binding inhibitory protein 1 |
| chr11_-_104035088 | 0.06 |
ENST00000302251.5 |
PDGFD |
platelet derived growth factor D |
| chr12_+_48722763 | 0.06 |
ENST00000335017.1 |
H1FNT |
H1 histone family, member N, testis-specific |
| chr19_-_39881777 | 0.06 |
ENST00000595564.1 ENST00000221265.3 |
PAF1 |
Paf1, RNA polymerase II associated factor, homolog (S. cerevisiae) |
| chr17_+_7328925 | 0.06 |
ENST00000333870.3 ENST00000574034.1 |
C17orf74 |
chromosome 17 open reading frame 74 |
| chr10_+_60028818 | 0.06 |
ENST00000333926.5 |
CISD1 |
CDGSH iron sulfur domain 1 |
| chr19_-_49121054 | 0.06 |
ENST00000546623.1 ENST00000084795.5 |
RPL18 |
ribosomal protein L18 |
| chrX_+_56590002 | 0.06 |
ENST00000338222.5 |
UBQLN2 |
ubiquilin 2 |
| chr12_+_54402790 | 0.06 |
ENST00000040584.4 |
HOXC8 |
homeobox C8 |
| chr17_-_26695013 | 0.05 |
ENST00000555059.2 |
CTB-96E2.2 |
Homeobox protein SEBOX |
| chr16_-_52119019 | 0.05 |
ENST00000561513.1 ENST00000565742.1 |
LINC00919 |
long intergenic non-protein coding RNA 919 |
| chr10_+_48189612 | 0.05 |
ENST00000453919.1 |
AGAP9 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 9 |
| chr12_-_21654479 | 0.05 |
ENST00000421138.2 ENST00000444129.2 ENST00000539672.1 ENST00000542432.1 ENST00000536964.1 ENST00000536240.1 ENST00000396093.3 ENST00000314748.6 |
RECQL |
RecQ protein-like (DNA helicase Q1-like) |
| chr4_+_37962018 | 0.05 |
ENST00000504686.1 |
PTTG2 |
pituitary tumor-transforming 2 |
| chr1_+_203463245 | 0.05 |
ENST00000367222.2 |
OPTC |
opticin |
| chr12_+_12870055 | 0.05 |
ENST00000228872.4 |
CDKN1B |
cyclin-dependent kinase inhibitor 1B (p27, Kip1) |
| chrX_+_95939711 | 0.05 |
ENST00000373049.4 ENST00000324765.8 |
DIAPH2 |
diaphanous-related formin 2 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.1 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.2 | 0.6 | GO:0021569 | rhombomere 3 development(GO:0021569) central nervous system segmentation(GO:0035283) brain segmentation(GO:0035284) |
| 0.2 | 2.0 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.2 | 0.9 | GO:1903976 | negative regulation of glial cell migration(GO:1903976) |
| 0.2 | 0.7 | GO:0006210 | pyrimidine nucleobase catabolic process(GO:0006208) thymine catabolic process(GO:0006210) thymine metabolic process(GO:0019859) |
| 0.2 | 0.5 | GO:0098746 | fast, calcium ion-dependent exocytosis of neurotransmitter(GO:0098746) |
| 0.2 | 0.8 | GO:0071279 | cellular response to cobalt ion(GO:0071279) |
| 0.2 | 1.1 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.2 | 0.5 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 0.1 | 0.7 | GO:0042091 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 0.1 | 1.1 | GO:0021902 | commitment of neuronal cell to specific neuron type in forebrain(GO:0021902) |
| 0.1 | 0.6 | GO:0097498 | endothelial tube lumen extension(GO:0097498) |
| 0.1 | 0.3 | GO:0035720 | intraciliary anterograde transport(GO:0035720) ciliary receptor clustering involved in smoothened signaling pathway(GO:0060830) |
| 0.1 | 0.6 | GO:0001957 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.1 | 0.8 | GO:1903944 | regulation of hepatocyte apoptotic process(GO:1903943) negative regulation of hepatocyte apoptotic process(GO:1903944) |
| 0.1 | 0.4 | GO:0006740 | NADPH regeneration(GO:0006740) |
| 0.1 | 0.2 | GO:0019285 | glycine betaine biosynthetic process from choline(GO:0019285) glycine betaine metabolic process(GO:0031455) glycine betaine biosynthetic process(GO:0031456) |
| 0.1 | 0.2 | GO:0043456 | regulation of pentose-phosphate shunt(GO:0043456) |
| 0.1 | 1.0 | GO:0048739 | cardiac muscle fiber development(GO:0048739) |
| 0.1 | 0.2 | GO:0003330 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) |
| 0.1 | 0.3 | GO:0031022 | nuclear migration along microfilament(GO:0031022) |
| 0.0 | 0.1 | GO:0061713 | neural crest cell migration involved in heart formation(GO:0003147) cell migration involved in heart formation(GO:0060974) anterior neural tube closure(GO:0061713) |
| 0.0 | 0.1 | GO:0009138 | pyrimidine nucleoside diphosphate metabolic process(GO:0009138) |
| 0.0 | 0.4 | GO:0021730 | trigeminal sensory nucleus development(GO:0021730) principal sensory nucleus of trigeminal nerve development(GO:0021740) |
| 0.0 | 0.4 | GO:0070269 | L-methionine biosynthetic process from methylthioadenosine(GO:0019509) pyroptosis(GO:0070269) |
| 0.0 | 0.6 | GO:0048227 | plasma membrane to endosome transport(GO:0048227) |
| 0.0 | 0.1 | GO:0046416 | D-amino acid metabolic process(GO:0046416) |
| 0.0 | 0.5 | GO:0010739 | positive regulation of protein kinase A signaling(GO:0010739) |
| 0.0 | 0.2 | GO:0006499 | N-terminal protein myristoylation(GO:0006499) |
| 0.0 | 0.2 | GO:0044245 | polysaccharide digestion(GO:0044245) |
| 0.0 | 0.3 | GO:1901525 | negative regulation of macromitophagy(GO:1901525) |
| 0.0 | 0.2 | GO:1901727 | positive regulation of histone deacetylase activity(GO:1901727) |
| 0.0 | 0.1 | GO:0072023 | thick ascending limb development(GO:0072023) metanephric thick ascending limb development(GO:0072233) |
| 0.0 | 0.2 | GO:0006537 | glutamate biosynthetic process(GO:0006537) |
| 0.0 | 0.3 | GO:1903546 | protein localization to photoreceptor outer segment(GO:1903546) |
| 0.0 | 0.1 | GO:0008628 | hormone-mediated apoptotic signaling pathway(GO:0008628) |
| 0.0 | 0.2 | GO:0002051 | osteoblast fate commitment(GO:0002051) |
| 0.0 | 0.4 | GO:0000338 | protein deneddylation(GO:0000338) |
| 0.0 | 0.6 | GO:0061154 | endothelial tube morphogenesis(GO:0061154) |
| 0.0 | 0.1 | GO:0031508 | pericentric heterochromatin assembly(GO:0031508) |
| 0.0 | 0.1 | GO:0006741 | NADP biosynthetic process(GO:0006741) |
| 0.0 | 0.1 | GO:0098905 | regulation of bundle of His cell action potential(GO:0098905) |
| 0.0 | 0.4 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.0 | 0.1 | GO:0070543 | response to linoleic acid(GO:0070543) |
| 0.0 | 0.1 | GO:2000439 | positive regulation of monocyte extravasation(GO:2000439) |
| 0.0 | 0.1 | GO:0071400 | cellular response to oleic acid(GO:0071400) |
| 0.0 | 0.1 | GO:0033387 | putrescine biosynthetic process from ornithine(GO:0033387) |
| 0.0 | 0.3 | GO:0060155 | platelet dense granule organization(GO:0060155) |
| 0.0 | 0.1 | GO:1904020 | regulation of G-protein coupled receptor internalization(GO:1904020) |
| 0.0 | 0.5 | GO:0006068 | ethanol catabolic process(GO:0006068) |
| 0.0 | 0.1 | GO:2000467 | positive regulation of glycogen (starch) synthase activity(GO:2000467) |
| 0.0 | 0.3 | GO:0034356 | NAD biosynthesis via nicotinamide riboside salvage pathway(GO:0034356) |
| 0.0 | 0.1 | GO:0007231 | osmosensory signaling pathway(GO:0007231) |
| 0.0 | 0.2 | GO:1902548 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) negative regulation of cellular response to vascular endothelial growth factor stimulus(GO:1902548) |
| 0.0 | 0.1 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.0 | 0.0 | GO:0002314 | germinal center B cell differentiation(GO:0002314) |
| 0.0 | 0.2 | GO:0070986 | left/right axis specification(GO:0070986) |
| 0.0 | 0.1 | GO:0032000 | positive regulation of fatty acid beta-oxidation(GO:0032000) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.1 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.1 | 2.0 | GO:0042589 | zymogen granule membrane(GO:0042589) |
| 0.1 | 0.5 | GO:0060201 | clathrin-sculpted acetylcholine transport vesicle(GO:0060200) clathrin-sculpted acetylcholine transport vesicle membrane(GO:0060201) |
| 0.1 | 0.8 | GO:0043203 | axon hillock(GO:0043203) |
| 0.0 | 0.6 | GO:0031462 | Cul2-RING ubiquitin ligase complex(GO:0031462) |
| 0.0 | 0.8 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) ripoptosome(GO:0097342) |
| 0.0 | 0.1 | GO:0042720 | mitochondrial inner membrane peptidase complex(GO:0042720) |
| 0.0 | 0.2 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.0 | 0.3 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.0 | 0.3 | GO:0034992 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.0 | 0.1 | GO:0071062 | rough endoplasmic reticulum lumen(GO:0048237) alphav-beta3 integrin-vitronectin complex(GO:0071062) |
| 0.0 | 0.2 | GO:0000243 | commitment complex(GO:0000243) |
| 0.0 | 0.2 | GO:0005589 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.0 | 0.2 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 0.0 | 0.3 | GO:0045009 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.0 | 0.4 | GO:0035327 | transcriptionally active chromatin(GO:0035327) |
| 0.0 | 0.1 | GO:0005947 | mitochondrial alpha-ketoglutarate dehydrogenase complex(GO:0005947) |
| 0.0 | 0.1 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.0 | 0.1 | GO:0045179 | apical cortex(GO:0045179) |
| 0.0 | 0.4 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.0 | 0.2 | GO:0030914 | STAGA complex(GO:0030914) |
| 0.0 | 1.3 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 0.4 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0008746 | NAD(P)+ transhydrogenase activity(GO:0008746) oxidoreductase activity, acting on NAD(P)H, NAD(P) as acceptor(GO:0016652) |
| 0.1 | 0.4 | GO:0010698 | acetyltransferase activator activity(GO:0010698) |
| 0.1 | 0.5 | GO:0030348 | syntaxin-3 binding(GO:0030348) |
| 0.1 | 0.3 | GO:0008112 | nicotinamide N-methyltransferase activity(GO:0008112) pyridine N-methyltransferase activity(GO:0030760) |
| 0.1 | 2.1 | GO:0038187 | signaling pattern recognition receptor activity(GO:0008329) pattern recognition receptor activity(GO:0038187) |
| 0.1 | 0.2 | GO:0046538 | bisphosphoglycerate mutase activity(GO:0004082) phosphoglycerate mutase activity(GO:0004619) 2,3-bisphosphoglycerate-dependent phosphoglycerate mutase activity(GO:0046538) |
| 0.1 | 0.2 | GO:0016160 | amylase activity(GO:0016160) |
| 0.0 | 0.2 | GO:0016639 | oxidoreductase activity, acting on the CH-NH2 group of donors, NAD or NADP as acceptor(GO:0016639) |
| 0.0 | 0.7 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.0 | 0.4 | GO:0019784 | NEDD8-specific protease activity(GO:0019784) |
| 0.0 | 0.1 | GO:0061714 | folic acid receptor activity(GO:0061714) |
| 0.0 | 0.1 | GO:0009041 | uridylate kinase activity(GO:0009041) |
| 0.0 | 0.2 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.0 | 0.5 | GO:0070513 | death domain binding(GO:0070513) |
| 0.0 | 0.1 | GO:0004087 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 0.0 | 0.4 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.0 | 0.6 | GO:0070300 | phosphatidic acid binding(GO:0070300) |
| 0.0 | 0.3 | GO:0031386 | protein tag(GO:0031386) |
| 0.0 | 0.8 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.0 | 0.1 | GO:0051373 | FATZ binding(GO:0051373) |
| 0.0 | 0.6 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.0 | 0.1 | GO:0030366 | molybdopterin synthase activity(GO:0030366) |
| 0.0 | 0.5 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.0 | 0.6 | GO:0000062 | fatty-acyl-CoA binding(GO:0000062) |
| 0.0 | 0.2 | GO:0003960 | NADPH:quinone reductase activity(GO:0003960) |
| 0.0 | 0.1 | GO:0017060 | 3-galactosyl-N-acetylglucosaminide 4-alpha-L-fucosyltransferase activity(GO:0017060) |
| 0.0 | 0.2 | GO:0010314 | phosphatidylinositol-5-phosphate binding(GO:0010314) |
| 0.0 | 0.4 | GO:0001055 | RNA polymerase II activity(GO:0001055) |
| 0.0 | 0.1 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.0 | 0.3 | GO:0043495 | protein anchor(GO:0043495) |
| 0.0 | 0.1 | GO:0000285 | 1-phosphatidylinositol-3-phosphate 5-kinase activity(GO:0000285) |
| 0.0 | 1.1 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.0 | 0.4 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.0 | 0.3 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.8 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 1.9 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 0.8 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.0 | 1.6 | PID INTEGRIN1 PATHWAY | Beta1 integrin cell surface interactions |
| 0.0 | 0.5 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.0 | 1.1 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.0 | 0.8 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.0 | 0.5 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 0.8 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 0.7 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 0.8 | REACTOME ERK MAPK TARGETS | Genes involved in ERK/MAPK targets |
| 0.0 | 0.5 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.0 | 1.0 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 0.4 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 0.4 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.0 | 0.4 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.3 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.0 | 0.5 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.0 | 0.1 | REACTOME REMOVAL OF THE FLAP INTERMEDIATE FROM THE C STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 0.0 | 0.3 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |