Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
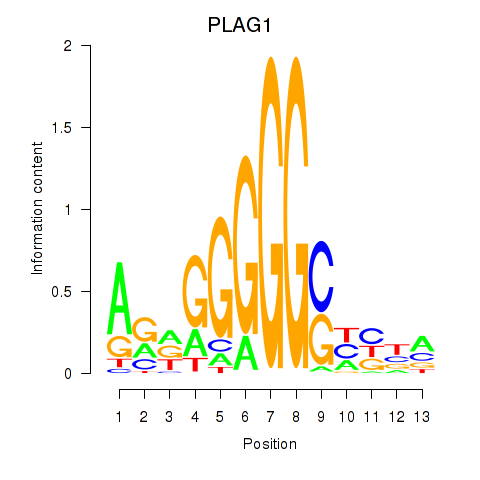
Results for PLAG1
Z-value: 0.30
Transcription factors associated with PLAG1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
PLAG1
|
ENSG00000181690.3 | PLAG1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| PLAG1 | hg19_v2_chr8_-_57123815_57123867 | 0.43 | 2.9e-01 | Click! |
Activity profile of PLAG1 motif
Sorted Z-values of PLAG1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of PLAG1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_-_31845914 | 0.46 |
ENST00000373713.2 |
FABP3 |
fatty acid binding protein 3, muscle and heart (mammary-derived growth inhibitor) |
| chr11_-_2160180 | 0.37 |
ENST00000381406.4 |
IGF2 |
insulin-like growth factor 2 (somatomedin A) |
| chr12_+_79258444 | 0.36 |
ENST00000261205.4 |
SYT1 |
synaptotagmin I |
| chr1_+_163038565 | 0.34 |
ENST00000421743.2 |
RGS4 |
regulator of G-protein signaling 4 |
| chr11_+_46299199 | 0.34 |
ENST00000529193.1 ENST00000288400.3 |
CREB3L1 |
cAMP responsive element binding protein 3-like 1 |
| chr4_+_55095264 | 0.33 |
ENST00000257290.5 |
PDGFRA |
platelet-derived growth factor receptor, alpha polypeptide |
| chr12_+_79258547 | 0.32 |
ENST00000457153.2 |
SYT1 |
synaptotagmin I |
| chr5_-_137368708 | 0.25 |
ENST00000033079.3 |
FAM13B |
family with sequence similarity 13, member B |
| chr12_-_108733078 | 0.23 |
ENST00000552995.1 ENST00000312143.7 ENST00000397688.2 ENST00000550402.1 |
CMKLR1 |
chemokine-like receptor 1 |
| chr12_-_58131931 | 0.23 |
ENST00000547588.1 |
AGAP2 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 2 |
| chr16_-_65155833 | 0.22 |
ENST00000566827.1 ENST00000394156.3 ENST00000562998.1 |
CDH11 |
cadherin 11, type 2, OB-cadherin (osteoblast) |
| chr19_+_55795493 | 0.20 |
ENST00000309383.1 |
BRSK1 |
BR serine/threonine kinase 1 |
| chr9_+_2621798 | 0.20 |
ENST00000382100.3 |
VLDLR |
very low density lipoprotein receptor |
| chr14_-_75079026 | 0.19 |
ENST00000261978.4 |
LTBP2 |
latent transforming growth factor beta binding protein 2 |
| chr9_+_139874683 | 0.18 |
ENST00000444903.1 |
PTGDS |
prostaglandin D2 synthase 21kDa (brain) |
| chr12_+_57522258 | 0.17 |
ENST00000553277.1 ENST00000243077.3 |
LRP1 |
low density lipoprotein receptor-related protein 1 |
| chr9_+_34652164 | 0.17 |
ENST00000441545.2 ENST00000553620.1 |
IL11RA |
interleukin 11 receptor, alpha |
| chr7_+_75024903 | 0.17 |
ENST00000323819.3 ENST00000430211.1 |
TRIM73 |
tripartite motif containing 73 |
| chr2_-_86850949 | 0.16 |
ENST00000237455.4 |
RNF103 |
ring finger protein 103 |
| chr5_+_75699149 | 0.16 |
ENST00000379730.3 |
IQGAP2 |
IQ motif containing GTPase activating protein 2 |
| chr7_+_75024337 | 0.16 |
ENST00000450434.1 |
TRIM73 |
tripartite motif containing 73 |
| chr12_+_57998595 | 0.16 |
ENST00000337737.3 ENST00000548198.1 ENST00000551632.1 |
DTX3 |
deltex homolog 3 (Drosophila) |
| chr13_-_49107303 | 0.16 |
ENST00000344532.3 |
RCBTB2 |
regulator of chromosome condensation (RCC1) and BTB (POZ) domain containing protein 2 |
| chr3_-_73673991 | 0.16 |
ENST00000308537.4 ENST00000263666.4 |
PDZRN3 |
PDZ domain containing ring finger 3 |
| chr2_-_30144432 | 0.15 |
ENST00000389048.3 |
ALK |
anaplastic lymphoma receptor tyrosine kinase |
| chr17_+_30593195 | 0.15 |
ENST00000431505.2 ENST00000269051.4 ENST00000538145.1 |
RHBDL3 |
rhomboid, veinlet-like 3 (Drosophila) |
| chrX_+_153146127 | 0.14 |
ENST00000452593.1 ENST00000357566.1 |
LCA10 |
Putative lung carcinoma-associated protein 10 |
| chr11_+_66610883 | 0.14 |
ENST00000309657.3 ENST00000524506.1 |
RCE1 |
Ras converting CAAX endopeptidase 1 |
| chr11_-_2323290 | 0.13 |
ENST00000381153.3 |
C11orf21 |
chromosome 11 open reading frame 21 |
| chr19_+_41725088 | 0.13 |
ENST00000301178.4 |
AXL |
AXL receptor tyrosine kinase |
| chr20_+_37353084 | 0.13 |
ENST00000217420.1 |
SLC32A1 |
solute carrier family 32 (GABA vesicular transporter), member 1 |
| chr20_+_35169885 | 0.13 |
ENST00000279022.2 ENST00000346786.2 |
MYL9 |
myosin, light chain 9, regulatory |
| chr6_+_38690622 | 0.12 |
ENST00000327475.6 |
DNAH8 |
dynein, axonemal, heavy chain 8 |
| chr1_-_68698222 | 0.12 |
ENST00000370976.3 ENST00000354777.2 ENST00000262348.4 ENST00000540432.1 |
WLS |
wntless Wnt ligand secretion mediator |
| chr12_+_57522439 | 0.12 |
ENST00000338962.4 |
LRP1 |
low density lipoprotein receptor-related protein 1 |
| chrX_+_23682379 | 0.12 |
ENST00000379349.1 |
PRDX4 |
peroxiredoxin 4 |
| chr19_+_40697514 | 0.12 |
ENST00000253055.3 |
MAP3K10 |
mitogen-activated protein kinase kinase kinase 10 |
| chr12_-_122018859 | 0.12 |
ENST00000536437.1 ENST00000377071.4 ENST00000538046.2 |
KDM2B |
lysine (K)-specific demethylase 2B |
| chr19_+_39904168 | 0.12 |
ENST00000438123.1 ENST00000409797.2 ENST00000451354.2 |
PLEKHG2 |
pleckstrin homology domain containing, family G (with RhoGef domain) member 2 |
| chr12_+_57998400 | 0.11 |
ENST00000548804.1 ENST00000550596.1 ENST00000551835.1 ENST00000549583.1 |
DTX3 |
deltex homolog 3 (Drosophila) |
| chr6_+_31730773 | 0.11 |
ENST00000415669.2 ENST00000425424.1 |
SAPCD1 |
suppressor APC domain containing 1 |
| chr14_-_51297197 | 0.11 |
ENST00000382043.4 |
NIN |
ninein (GSK3B interacting protein) |
| chr1_-_110613276 | 0.11 |
ENST00000369792.4 |
ALX3 |
ALX homeobox 3 |
| chr3_-_15374033 | 0.11 |
ENST00000253688.5 ENST00000383791.3 |
SH3BP5 |
SH3-domain binding protein 5 (BTK-associated) |
| chr12_-_112819896 | 0.11 |
ENST00000377560.5 ENST00000430131.2 ENST00000550722.1 ENST00000550724.1 |
HECTD4 |
HECT domain containing E3 ubiquitin protein ligase 4 |
| chr14_-_106518922 | 0.11 |
ENST00000390598.2 |
IGHV3-7 |
immunoglobulin heavy variable 3-7 |
| chr7_-_158380371 | 0.11 |
ENST00000389418.4 ENST00000389416.4 |
PTPRN2 |
protein tyrosine phosphatase, receptor type, N polypeptide 2 |
| chr15_+_92397051 | 0.11 |
ENST00000424469.2 |
SLCO3A1 |
solute carrier organic anion transporter family, member 3A1 |
| chr16_+_31483451 | 0.11 |
ENST00000565360.1 ENST00000361773.3 |
TGFB1I1 |
transforming growth factor beta 1 induced transcript 1 |
| chr16_+_31483374 | 0.10 |
ENST00000394863.3 |
TGFB1I1 |
transforming growth factor beta 1 induced transcript 1 |
| chr16_+_2521500 | 0.10 |
ENST00000293973.1 |
NTN3 |
netrin 3 |
| chr7_-_72439997 | 0.10 |
ENST00000285805.3 |
TRIM74 |
tripartite motif containing 74 |
| chr19_+_38924316 | 0.10 |
ENST00000355481.4 ENST00000360985.3 ENST00000359596.3 |
RYR1 |
ryanodine receptor 1 (skeletal) |
| chr2_-_233352531 | 0.10 |
ENST00000304546.1 |
ECEL1 |
endothelin converting enzyme-like 1 |
| chrX_-_16887963 | 0.10 |
ENST00000380084.4 |
RBBP7 |
retinoblastoma binding protein 7 |
| chr3_+_32726774 | 0.10 |
ENST00000538368.1 |
CNOT10 |
CCR4-NOT transcription complex, subunit 10 |
| chrX_+_51927919 | 0.10 |
ENST00000416960.1 |
MAGED4 |
melanoma antigen family D, 4 |
| chr19_+_41725140 | 0.10 |
ENST00000359092.3 |
AXL |
AXL receptor tyrosine kinase |
| chr4_-_5890145 | 0.09 |
ENST00000397890.2 |
CRMP1 |
collapsin response mediator protein 1 |
| chr14_-_75330537 | 0.09 |
ENST00000556084.2 ENST00000556489.2 ENST00000445876.1 |
PROX2 |
prospero homeobox 2 |
| chr7_-_158380465 | 0.09 |
ENST00000389413.3 ENST00000409483.1 |
PTPRN2 |
protein tyrosine phosphatase, receptor type, N polypeptide 2 |
| chrY_+_22737604 | 0.09 |
ENST00000361365.2 |
EIF1AY |
eukaryotic translation initiation factor 1A, Y-linked |
| chr22_-_31885514 | 0.09 |
ENST00000397525.1 |
EIF4ENIF1 |
eukaryotic translation initiation factor 4E nuclear import factor 1 |
| chr10_+_102891048 | 0.08 |
ENST00000467928.2 |
TLX1 |
T-cell leukemia homeobox 1 |
| chr18_-_51750948 | 0.08 |
ENST00000583046.1 ENST00000398398.2 |
MBD2 |
methyl-CpG binding domain protein 2 |
| chr1_-_153931052 | 0.08 |
ENST00000368630.3 ENST00000368633.1 |
CRTC2 |
CREB regulated transcription coactivator 2 |
| chr2_-_178937478 | 0.08 |
ENST00000286063.6 |
PDE11A |
phosphodiesterase 11A |
| chr6_-_150039249 | 0.08 |
ENST00000543571.1 |
LATS1 |
large tumor suppressor kinase 1 |
| chr16_+_29984962 | 0.08 |
ENST00000308893.4 |
TAOK2 |
TAO kinase 2 |
| chr6_+_31683117 | 0.08 |
ENST00000375825.3 ENST00000375824.1 |
LY6G6D |
lymphocyte antigen 6 complex, locus G6D |
| chr4_-_47840112 | 0.08 |
ENST00000504584.1 |
CORIN |
corin, serine peptidase |
| chr1_+_62902308 | 0.08 |
ENST00000339950.4 |
USP1 |
ubiquitin specific peptidase 1 |
| chr6_+_31543334 | 0.08 |
ENST00000449264.2 |
TNF |
tumor necrosis factor |
| chr3_-_116163830 | 0.08 |
ENST00000333617.4 |
LSAMP |
limbic system-associated membrane protein |
| chr16_+_33006369 | 0.07 |
ENST00000425181.3 |
IGHV3OR16-10 |
immunoglobulin heavy variable 3/OR16-10 (non-functional) |
| chr9_-_123476612 | 0.07 |
ENST00000426959.1 |
MEGF9 |
multiple EGF-like-domains 9 |
| chr16_+_2198604 | 0.07 |
ENST00000210187.6 |
RAB26 |
RAB26, member RAS oncogene family |
| chr16_-_18812746 | 0.07 |
ENST00000546206.2 ENST00000562819.1 ENST00000562234.2 ENST00000304414.7 ENST00000567078.2 |
ARL6IP1 RP11-1035H13.3 |
ADP-ribosylation factor-like 6 interacting protein 1 Uncharacterized protein |
| chr9_-_123476719 | 0.07 |
ENST00000373930.3 |
MEGF9 |
multiple EGF-like-domains 9 |
| chr17_-_50237374 | 0.07 |
ENST00000442502.2 |
CA10 |
carbonic anhydrase X |
| chr6_+_7727030 | 0.07 |
ENST00000283147.6 |
BMP6 |
bone morphogenetic protein 6 |
| chr1_-_20306909 | 0.07 |
ENST00000375111.3 ENST00000400520.3 |
PLA2G2A |
phospholipase A2, group IIA (platelets, synovial fluid) |
| chr9_-_135819987 | 0.07 |
ENST00000298552.3 ENST00000403810.1 |
TSC1 |
tuberous sclerosis 1 |
| chr7_-_72742085 | 0.07 |
ENST00000333149.2 |
TRIM50 |
tripartite motif containing 50 |
| chr5_-_59189545 | 0.06 |
ENST00000340635.6 |
PDE4D |
phosphodiesterase 4D, cAMP-specific |
| chr16_-_18573396 | 0.06 |
ENST00000543392.1 ENST00000381474.3 ENST00000330537.6 |
NOMO2 |
NODAL modulator 2 |
| chr6_-_150039170 | 0.06 |
ENST00000458696.2 ENST00000392273.3 |
LATS1 |
large tumor suppressor kinase 1 |
| chr15_-_43910438 | 0.06 |
ENST00000432436.1 |
STRC |
stereocilin |
| chr1_+_20617362 | 0.06 |
ENST00000375079.2 ENST00000289815.8 ENST00000375083.4 ENST00000289825.4 |
VWA5B1 |
von Willebrand factor A domain containing 5B1 |
| chr14_+_105047478 | 0.06 |
ENST00000410013.1 |
C14orf180 |
chromosome 14 open reading frame 180 |
| chr12_+_107168342 | 0.06 |
ENST00000392837.4 |
RIC8B |
RIC8 guanine nucleotide exchange factor B |
| chr5_+_10353780 | 0.06 |
ENST00000449913.2 ENST00000503788.1 ENST00000274140.5 |
MARCH6 |
membrane-associated ring finger (C3HC4) 6, E3 ubiquitin protein ligase |
| chr17_-_79805146 | 0.06 |
ENST00000415593.1 |
P4HB |
prolyl 4-hydroxylase, beta polypeptide |
| chr10_+_111767720 | 0.06 |
ENST00000356080.4 ENST00000277900.8 |
ADD3 |
adducin 3 (gamma) |
| chr1_-_68698197 | 0.06 |
ENST00000370973.2 ENST00000370971.1 |
WLS |
wntless Wnt ligand secretion mediator |
| chr2_-_74618964 | 0.06 |
ENST00000417090.1 ENST00000409868.1 |
DCTN1 |
dynactin 1 |
| chr1_+_151138500 | 0.06 |
ENST00000368905.4 |
SCNM1 |
sodium channel modifier 1 |
| chr1_+_22962948 | 0.06 |
ENST00000374642.3 |
C1QA |
complement component 1, q subcomponent, A chain |
| chr12_-_106641728 | 0.05 |
ENST00000378026.4 |
CKAP4 |
cytoskeleton-associated protein 4 |
| chr17_+_38278826 | 0.05 |
ENST00000577454.1 ENST00000578648.1 ENST00000579565.1 |
MSL1 |
male-specific lethal 1 homolog (Drosophila) |
| chr12_+_52282001 | 0.05 |
ENST00000340970.4 |
ANKRD33 |
ankyrin repeat domain 33 |
| chr1_-_92949505 | 0.05 |
ENST00000370332.1 |
GFI1 |
growth factor independent 1 transcription repressor |
| chr16_-_29910853 | 0.05 |
ENST00000308713.5 |
SEZ6L2 |
seizure related 6 homolog (mouse)-like 2 |
| chr15_+_41186609 | 0.05 |
ENST00000220509.5 |
VPS18 |
vacuolar protein sorting 18 homolog (S. cerevisiae) |
| chr16_-_29910365 | 0.05 |
ENST00000346932.5 ENST00000350527.3 ENST00000537485.1 ENST00000568380.1 |
SEZ6L2 |
seizure related 6 homolog (mouse)-like 2 |
| chr11_-_64511789 | 0.05 |
ENST00000419843.1 ENST00000394430.1 |
RASGRP2 |
RAS guanyl releasing protein 2 (calcium and DAG-regulated) |
| chr8_+_29952914 | 0.05 |
ENST00000321250.8 ENST00000518001.1 ENST00000520682.1 ENST00000442880.2 ENST00000523116.1 |
LEPROTL1 |
leptin receptor overlapping transcript-like 1 |
| chr2_-_128399706 | 0.05 |
ENST00000426981.1 |
LIMS2 |
LIM and senescent cell antigen-like domains 2 |
| chr15_-_41408339 | 0.05 |
ENST00000401393.3 |
INO80 |
INO80 complex subunit |
| chr21_-_15755446 | 0.05 |
ENST00000544452.1 ENST00000285667.3 |
HSPA13 |
heat shock protein 70kDa family, member 13 |
| chr1_+_151138526 | 0.05 |
ENST00000368902.1 |
SCNM1 |
sodium channel modifier 1 |
| chr3_+_32726620 | 0.05 |
ENST00000331889.6 ENST00000328834.5 |
CNOT10 |
CCR4-NOT transcription complex, subunit 10 |
| chr22_+_19744226 | 0.05 |
ENST00000332710.4 ENST00000329705.7 ENST00000359500.3 |
TBX1 |
T-box 1 |
| chr5_+_161275320 | 0.05 |
ENST00000437025.2 |
GABRA1 |
gamma-aminobutyric acid (GABA) A receptor, alpha 1 |
| chr22_-_30642782 | 0.05 |
ENST00000249075.3 |
LIF |
leukemia inhibitory factor |
| chr16_+_32063311 | 0.05 |
ENST00000426099.1 |
AC142381.1 |
AC142381.1 |
| chr17_+_20483037 | 0.05 |
ENST00000399044.1 |
CDRT15L2 |
CMT1A duplicated region transcript 15-like 2 |
| chr10_-_75571566 | 0.04 |
ENST00000299641.4 |
NDST2 |
N-deacetylase/N-sulfotransferase (heparan glucosaminyl) 2 |
| chr8_-_23261589 | 0.04 |
ENST00000524168.1 ENST00000523833.2 ENST00000519243.1 ENST00000389131.3 |
LOXL2 |
lysyl oxidase-like 2 |
| chr4_+_39046615 | 0.04 |
ENST00000261425.3 ENST00000508137.2 |
KLHL5 |
kelch-like family member 5 |
| chr17_+_73106035 | 0.04 |
ENST00000581078.1 ENST00000582136.1 ENST00000245543.1 |
ARMC7 |
armadillo repeat containing 7 |
| chr16_+_56485402 | 0.04 |
ENST00000566157.1 ENST00000562150.1 ENST00000561646.1 ENST00000568397.1 |
OGFOD1 |
2-oxoglutarate and iron-dependent oxygenase domain containing 1 |
| chr7_+_72742162 | 0.04 |
ENST00000431982.2 |
FKBP6 |
FK506 binding protein 6, 36kDa |
| chr21_+_17909594 | 0.04 |
ENST00000441820.1 ENST00000602280.1 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr2_-_74619152 | 0.04 |
ENST00000440727.1 ENST00000409240.1 |
DCTN1 |
dynactin 1 |
| chr2_-_85788652 | 0.04 |
ENST00000430215.3 |
GGCX |
gamma-glutamyl carboxylase |
| chr10_-_75571341 | 0.04 |
ENST00000309979.6 |
NDST2 |
N-deacetylase/N-sulfotransferase (heparan glucosaminyl) 2 |
| chr17_-_1588101 | 0.04 |
ENST00000577001.1 ENST00000572621.1 ENST00000304992.6 |
PRPF8 |
pre-mRNA processing factor 8 |
| chr4_+_55095428 | 0.04 |
ENST00000508170.1 ENST00000512143.1 |
PDGFRA |
platelet-derived growth factor receptor, alpha polypeptide |
| chr11_-_111170526 | 0.04 |
ENST00000355430.4 |
COLCA1 |
colorectal cancer associated 1 |
| chr14_+_79745746 | 0.04 |
ENST00000281127.7 |
NRXN3 |
neurexin 3 |
| chr19_+_45973120 | 0.04 |
ENST00000592811.1 ENST00000586615.1 |
FOSB |
FBJ murine osteosarcoma viral oncogene homolog B |
| chrX_+_110339439 | 0.03 |
ENST00000372010.1 ENST00000519681.1 ENST00000372007.5 |
PAK3 |
p21 protein (Cdc42/Rac)-activated kinase 3 |
| chr3_+_49711391 | 0.03 |
ENST00000296456.5 ENST00000449966.1 |
APEH |
acylaminoacyl-peptide hydrolase |
| chr14_-_25519317 | 0.03 |
ENST00000323944.5 |
STXBP6 |
syntaxin binding protein 6 (amisyn) |
| chr8_+_29953163 | 0.03 |
ENST00000518192.1 |
LEPROTL1 |
leptin receptor overlapping transcript-like 1 |
| chr6_-_43021612 | 0.03 |
ENST00000535468.1 |
CUL7 |
cullin 7 |
| chr12_+_58005204 | 0.03 |
ENST00000286494.4 |
ARHGEF25 |
Rho guanine nucleotide exchange factor (GEF) 25 |
| chr1_-_150947343 | 0.03 |
ENST00000271688.6 ENST00000368954.5 |
CERS2 |
ceramide synthase 2 |
| chr7_+_72742178 | 0.03 |
ENST00000442793.1 ENST00000413573.2 ENST00000252037.4 |
FKBP6 |
FK506 binding protein 6, 36kDa |
| chr11_-_57282349 | 0.03 |
ENST00000528450.1 |
SLC43A1 |
solute carrier family 43 (amino acid system L transporter), member 1 |
| chr4_-_47839966 | 0.03 |
ENST00000273857.4 ENST00000505909.1 ENST00000502252.1 |
CORIN |
corin, serine peptidase |
| chr4_+_36283213 | 0.03 |
ENST00000357504.3 |
DTHD1 |
death domain containing 1 |
| chr6_-_43021437 | 0.03 |
ENST00000265348.3 |
CUL7 |
cullin 7 |
| chr16_+_31191431 | 0.03 |
ENST00000254108.7 ENST00000380244.3 ENST00000568685.1 |
FUS |
fused in sarcoma |
| chr16_+_4896659 | 0.03 |
ENST00000592120.1 |
UBN1 |
ubinuclein 1 |
| chr12_-_12419703 | 0.02 |
ENST00000543091.1 ENST00000261349.4 |
LRP6 |
low density lipoprotein receptor-related protein 6 |
| chr6_+_43044003 | 0.02 |
ENST00000230419.4 ENST00000476760.1 ENST00000471863.1 ENST00000349241.2 ENST00000352931.2 ENST00000345201.2 |
PTK7 |
protein tyrosine kinase 7 |
| chrX_-_48776292 | 0.02 |
ENST00000376509.4 |
PIM2 |
pim-2 oncogene |
| chr14_+_79745682 | 0.02 |
ENST00000557594.1 |
NRXN3 |
neurexin 3 |
| chr12_+_121163538 | 0.02 |
ENST00000242592.4 |
ACADS |
acyl-CoA dehydrogenase, C-2 to C-3 short chain |
| chr3_-_19988462 | 0.02 |
ENST00000344838.4 |
EFHB |
EF-hand domain family, member B |
| chr9_+_131452239 | 0.02 |
ENST00000372688.4 ENST00000372686.5 |
SET |
SET nuclear oncogene |
| chr4_-_155533787 | 0.02 |
ENST00000407946.1 ENST00000405164.1 ENST00000336098.3 ENST00000393846.2 ENST00000404648.3 ENST00000443553.1 |
FGG |
fibrinogen gamma chain |
| chr5_+_7654057 | 0.02 |
ENST00000537121.1 |
ADCY2 |
adenylate cyclase 2 (brain) |
| chr19_-_35626104 | 0.02 |
ENST00000310123.3 ENST00000392225.3 |
LGI4 |
leucine-rich repeat LGI family, member 4 |
| chr17_-_56065484 | 0.02 |
ENST00000581208.1 |
VEZF1 |
vascular endothelial zinc finger 1 |
| chr14_-_25519095 | 0.02 |
ENST00000419632.2 ENST00000358326.2 ENST00000396700.1 ENST00000548724.1 |
STXBP6 |
syntaxin binding protein 6 (amisyn) |
| chr15_-_41408409 | 0.02 |
ENST00000361937.3 |
INO80 |
INO80 complex subunit |
| chr11_+_64053311 | 0.02 |
ENST00000540370.1 |
GPR137 |
G protein-coupled receptor 137 |
| chr19_+_40973049 | 0.02 |
ENST00000598249.1 ENST00000338932.3 ENST00000344104.3 |
SPTBN4 |
spectrin, beta, non-erythrocytic 4 |
| chr9_-_95527079 | 0.02 |
ENST00000356884.6 ENST00000375512.3 |
BICD2 |
bicaudal D homolog 2 (Drosophila) |
| chr17_-_1420006 | 0.02 |
ENST00000320345.6 ENST00000406424.4 |
INPP5K |
inositol polyphosphate-5-phosphatase K |
| chr1_-_212003556 | 0.01 |
ENST00000366996.1 |
LPGAT1 |
lysophosphatidylglycerol acyltransferase 1 |
| chr14_+_79746249 | 0.01 |
ENST00000428277.2 |
NRXN3 |
neurexin 3 |
| chr3_+_183853052 | 0.01 |
ENST00000273783.3 ENST00000432569.1 ENST00000444495.1 |
EIF2B5 |
eukaryotic translation initiation factor 2B, subunit 5 epsilon, 82kDa |
| chr2_-_152955213 | 0.01 |
ENST00000427385.1 |
CACNB4 |
calcium channel, voltage-dependent, beta 4 subunit |
| chr9_-_140115775 | 0.01 |
ENST00000391553.1 ENST00000392827.1 |
RNF208 |
ring finger protein 208 |
| chr16_+_46723552 | 0.01 |
ENST00000219097.2 ENST00000568364.2 |
ORC6 |
origin recognition complex, subunit 6 |
| chr3_-_197024394 | 0.01 |
ENST00000434148.1 ENST00000412364.2 ENST00000436682.1 ENST00000456699.2 ENST00000392380.2 |
DLG1 |
discs, large homolog 1 (Drosophila) |
| chr4_-_114682597 | 0.01 |
ENST00000394524.3 |
CAMK2D |
calcium/calmodulin-dependent protein kinase II delta |
| chr6_+_106534192 | 0.01 |
ENST00000369091.2 ENST00000369096.4 |
PRDM1 |
PR domain containing 1, with ZNF domain |
| chr19_-_51054299 | 0.01 |
ENST00000599957.1 |
LRRC4B |
leucine rich repeat containing 4B |
| chr5_+_134181755 | 0.01 |
ENST00000504727.1 ENST00000435259.2 ENST00000508791.1 |
C5orf24 |
chromosome 5 open reading frame 24 |
| chr13_+_53602894 | 0.01 |
ENST00000219022.2 |
OLFM4 |
olfactomedin 4 |
| chr4_-_114682936 | 0.01 |
ENST00000454265.2 ENST00000429180.1 ENST00000418639.2 ENST00000394526.2 ENST00000296402.5 |
CAMK2D |
calcium/calmodulin-dependent protein kinase II delta |
| chr20_+_18269121 | 0.01 |
ENST00000377671.3 ENST00000360010.5 ENST00000396026.3 ENST00000402618.2 ENST00000401790.1 ENST00000434018.1 ENST00000538547.1 ENST00000535822.1 |
ZNF133 |
zinc finger protein 133 |
| chrX_-_71526999 | 0.01 |
ENST00000453707.2 ENST00000373619.3 |
CITED1 |
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 1 |
| chr1_-_204654481 | 0.00 |
ENST00000367176.3 |
LRRN2 |
leucine rich repeat neuronal 2 |
| chr16_-_46723066 | 0.00 |
ENST00000299138.7 |
VPS35 |
vacuolar protein sorting 35 homolog (S. cerevisiae) |
| chr19_-_35264089 | 0.00 |
ENST00000588760.1 ENST00000329285.8 ENST00000587354.2 |
ZNF599 |
zinc finger protein 599 |
| chrX_-_137793826 | 0.00 |
ENST00000315930.6 |
FGF13 |
fibroblast growth factor 13 |
| chr15_+_29131103 | 0.00 |
ENST00000558402.1 ENST00000558330.1 |
APBA2 |
amyloid beta (A4) precursor protein-binding, family A, member 2 |
| chr17_-_4607335 | 0.00 |
ENST00000570571.1 ENST00000575101.1 ENST00000436683.2 ENST00000574876.1 |
PELP1 |
proline, glutamate and leucine rich protein 1 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0098746 | fast, calcium ion-dependent exocytosis of neurotransmitter(GO:0098746) |
| 0.2 | 0.5 | GO:2001245 | regulation of phosphatidylcholine biosynthetic process(GO:2001245) |
| 0.1 | 0.4 | GO:0072276 | metanephric glomerulus morphogenesis(GO:0072275) metanephric glomerulus vasculature morphogenesis(GO:0072276) metanephric glomerular capillary formation(GO:0072277) |
| 0.1 | 0.3 | GO:1905167 | positive regulation of lysosomal protein catabolic process(GO:1905167) |
| 0.0 | 0.2 | GO:0061357 | positive regulation of Wnt protein secretion(GO:0061357) |
| 0.0 | 0.4 | GO:0038028 | insulin receptor signaling pathway via phosphatidylinositol 3-kinase(GO:0038028) |
| 0.0 | 0.2 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.0 | 0.3 | GO:0070278 | extracellular matrix constituent secretion(GO:0070278) |
| 0.0 | 0.1 | GO:0003050 | regulation of systemic arterial blood pressure by atrial natriuretic peptide(GO:0003050) |
| 0.0 | 0.2 | GO:0034436 | glycoprotein transport(GO:0034436) |
| 0.0 | 0.2 | GO:2000669 | positive regulation of pinocytosis(GO:0048549) negative regulation of dendritic cell apoptotic process(GO:2000669) |
| 0.0 | 0.1 | GO:1904617 | negative regulation of actin filament binding(GO:1904530) negative regulation of actin binding(GO:1904617) |
| 0.0 | 0.1 | GO:0001828 | inner cell mass cell fate commitment(GO:0001827) inner cell mass cellular morphogenesis(GO:0001828) |
| 0.0 | 0.1 | GO:0010693 | positive regulation of chronic inflammatory response(GO:0002678) regulation of chronic inflammatory response to antigenic stimulus(GO:0002874) negative regulation of alkaline phosphatase activity(GO:0010693) positive regulation of translational initiation by iron(GO:0045994) positive regulation of vitamin metabolic process(GO:0046136) positive regulation of vitamin D biosynthetic process(GO:0060557) positive regulation of calcidiol 1-monooxygenase activity(GO:0060559) |
| 0.0 | 0.1 | GO:2000255 | negative regulation of male germ cell proliferation(GO:2000255) |
| 0.0 | 0.2 | GO:0030579 | ubiquitin-dependent SMAD protein catabolic process(GO:0030579) |
| 0.0 | 0.1 | GO:0021592 | midbrain-hindbrain boundary morphogenesis(GO:0021555) fourth ventricle development(GO:0021592) |
| 0.0 | 0.1 | GO:0010286 | heat acclimation(GO:0010286) cellular heat acclimation(GO:0070370) |
| 0.0 | 0.1 | GO:0090158 | endoplasmic reticulum membrane organization(GO:0090158) |
| 0.0 | 0.2 | GO:0021957 | corticospinal tract morphogenesis(GO:0021957) |
| 0.0 | 0.1 | GO:0070105 | positive regulation of interleukin-6-mediated signaling pathway(GO:0070105) |
| 0.0 | 0.1 | GO:0090063 | positive regulation of microtubule nucleation(GO:0090063) |
| 0.0 | 0.0 | GO:2001037 | tongue muscle cell differentiation(GO:0035981) positive regulation of skeletal muscle fiber differentiation(GO:1902811) regulation of tongue muscle cell differentiation(GO:2001035) positive regulation of tongue muscle cell differentiation(GO:2001037) |
| 0.0 | 0.1 | GO:0090031 | positive regulation of steroid hormone biosynthetic process(GO:0090031) |
| 0.0 | 0.1 | GO:0051029 | rRNA transport(GO:0051029) |
| 0.0 | 0.1 | GO:0007256 | activation of JNKK activity(GO:0007256) |
| 0.0 | 0.1 | GO:0098700 | neurotransmitter loading into synaptic vesicle(GO:0098700) |
| 0.0 | 0.1 | GO:0070309 | lens fiber cell morphogenesis(GO:0070309) |
| 0.0 | 0.0 | GO:0090118 | receptor-mediated endocytosis of low-density lipoprotein particle involved in cholesterol transport(GO:0090118) |
| 0.0 | 0.0 | GO:0060708 | spongiotrophoblast differentiation(GO:0060708) |
| 0.0 | 0.1 | GO:0034454 | microtubule anchoring at centrosome(GO:0034454) |
| 0.0 | 0.2 | GO:0032695 | negative regulation of interleukin-12 production(GO:0032695) |
| 0.0 | 0.2 | GO:0019371 | cyclooxygenase pathway(GO:0019371) |
| 0.0 | 0.1 | GO:0032511 | late endosome to vacuole transport via multivesicular body sorting pathway(GO:0032511) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | GO:0060200 | clathrin-sculpted acetylcholine transport vesicle(GO:0060200) clathrin-sculpted acetylcholine transport vesicle membrane(GO:0060201) |
| 0.0 | 0.1 | GO:1990425 | ryanodine receptor complex(GO:1990425) |
| 0.0 | 0.1 | GO:0033596 | TSC1-TSC2 complex(GO:0033596) |
| 0.0 | 0.1 | GO:0016222 | procollagen-proline 4-dioxygenase complex(GO:0016222) |
| 0.0 | 0.2 | GO:0033643 | host cell part(GO:0033643) |
| 0.0 | 0.1 | GO:0051286 | cell tip(GO:0051286) |
| 0.0 | 0.1 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.0 | 0.1 | GO:0005784 | Sec61 translocon complex(GO:0005784) translocon complex(GO:0071256) |
| 0.0 | 0.1 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.0 | 0.1 | GO:0005602 | complement component C1 complex(GO:0005602) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0070538 | oleic acid binding(GO:0070538) |
| 0.1 | 0.7 | GO:0030348 | syntaxin-3 binding(GO:0030348) |
| 0.1 | 0.4 | GO:0005017 | platelet-derived growth factor-activated receptor activity(GO:0005017) |
| 0.1 | 0.2 | GO:0034189 | very-low-density lipoprotein particle binding(GO:0034189) |
| 0.1 | 0.3 | GO:0042954 | apolipoprotein receptor activity(GO:0030226) lipoprotein transporter activity(GO:0042954) |
| 0.0 | 0.2 | GO:0004667 | prostaglandin-D synthase activity(GO:0004667) |
| 0.0 | 0.1 | GO:0005275 | amine transmembrane transporter activity(GO:0005275) |
| 0.0 | 0.1 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.0 | 0.1 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 0.0 | 0.1 | GO:0048763 | calcium-induced calcium release activity(GO:0048763) |
| 0.0 | 0.2 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.0 | 0.1 | GO:0050119 | N-acetylglucosamine deacetylase activity(GO:0050119) |
| 0.0 | 0.4 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.0 | 0.2 | GO:0005087 | Ran guanyl-nucleotide exchange factor activity(GO:0005087) |
| 0.0 | 0.2 | GO:0004704 | NF-kappaB-inducing kinase activity(GO:0004704) |
| 0.0 | 0.2 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.0 | 0.1 | GO:0004118 | cGMP-stimulated cyclic-nucleotide phosphodiesterase activity(GO:0004118) |
| 0.0 | 0.2 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) |
| 0.0 | 0.3 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.0 | 0.1 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.0 | 0.2 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.0 | 0.4 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 0.4 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | REACTOME DOPAMINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Dopamine Neurotransmitter Release Cycle |
| 0.0 | 0.4 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |