Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
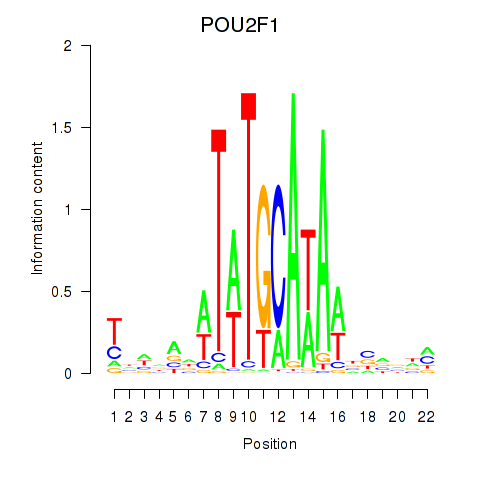
Results for POU2F1
Z-value: 0.43
Transcription factors associated with POU2F1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
POU2F1
|
ENSG00000143190.17 | POU2F1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| POU2F1 | hg19_v2_chr1_+_167190066_167190156 | 0.46 | 2.6e-01 | Click! |
Activity profile of POU2F1 motif
Sorted Z-values of POU2F1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of POU2F1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_62439037 | 0.47 |
ENST00000545929.1 |
INADL |
InaD-like (Drosophila) |
| chr10_+_118187379 | 0.38 |
ENST00000369230.3 |
PNLIPRP3 |
pancreatic lipase-related protein 3 |
| chr2_-_113594279 | 0.35 |
ENST00000416750.1 ENST00000418817.1 ENST00000263341.2 |
IL1B |
interleukin 1, beta |
| chr5_+_140227048 | 0.35 |
ENST00000532602.1 |
PCDHA9 |
protocadherin alpha 9 |
| chr3_-_111314230 | 0.28 |
ENST00000317012.4 |
ZBED2 |
zinc finger, BED-type containing 2 |
| chr1_+_44398943 | 0.26 |
ENST00000372359.5 ENST00000414809.3 |
ARTN |
artemin |
| chr6_-_11779403 | 0.23 |
ENST00000414691.3 |
ADTRP |
androgen-dependent TFPI-regulating protein |
| chr9_-_140196703 | 0.23 |
ENST00000356628.2 |
NRARP |
NOTCH-regulated ankyrin repeat protein |
| chr9_+_12693336 | 0.22 |
ENST00000381137.2 ENST00000388918.5 |
TYRP1 |
tyrosinase-related protein 1 |
| chr6_-_11779174 | 0.21 |
ENST00000379413.2 |
ADTRP |
androgen-dependent TFPI-regulating protein |
| chr6_-_11779014 | 0.21 |
ENST00000229583.5 |
ADTRP |
androgen-dependent TFPI-regulating protein |
| chr10_-_116286656 | 0.19 |
ENST00000428430.1 ENST00000369266.3 ENST00000392952.3 |
ABLIM1 |
actin binding LIM protein 1 |
| chr2_+_89890533 | 0.17 |
ENST00000429992.2 |
IGKV2D-40 |
immunoglobulin kappa variable 2D-40 |
| chr11_+_6897856 | 0.16 |
ENST00000379829.2 |
OR10A4 |
olfactory receptor, family 10, subfamily A, member 4 |
| chr18_-_52989217 | 0.15 |
ENST00000570287.2 |
TCF4 |
transcription factor 4 |
| chr2_+_102615416 | 0.15 |
ENST00000393414.2 |
IL1R2 |
interleukin 1 receptor, type II |
| chr8_+_86121448 | 0.14 |
ENST00000520225.1 |
E2F5 |
E2F transcription factor 5, p130-binding |
| chr9_+_116638562 | 0.14 |
ENST00000374126.5 ENST00000288466.7 |
ZNF618 |
zinc finger protein 618 |
| chr19_+_17858509 | 0.14 |
ENST00000594202.1 ENST00000252771.7 ENST00000389133.4 |
FCHO1 |
FCH domain only 1 |
| chr15_-_22448819 | 0.13 |
ENST00000604066.1 |
IGHV1OR15-1 |
immunoglobulin heavy variable 1/OR15-1 (non-functional) |
| chr3_-_155524049 | 0.13 |
ENST00000534941.1 ENST00000340171.2 |
C3orf33 |
chromosome 3 open reading frame 33 |
| chr2_+_191002486 | 0.13 |
ENST00000396974.2 |
C2orf88 |
chromosome 2 open reading frame 88 |
| chr19_+_17858547 | 0.12 |
ENST00000600676.1 ENST00000600209.1 ENST00000596309.1 ENST00000598539.1 ENST00000597474.1 ENST00000593385.1 ENST00000598067.1 ENST00000593833.1 |
FCHO1 |
FCH domain only 1 |
| chr11_-_58343319 | 0.12 |
ENST00000395074.2 |
LPXN |
leupaxin |
| chr2_+_90108504 | 0.12 |
ENST00000390271.2 |
IGKV6D-41 |
immunoglobulin kappa variable 6D-41 (non-functional) |
| chr10_-_116286563 | 0.12 |
ENST00000369253.2 |
ABLIM1 |
actin binding LIM protein 1 |
| chr2_+_90060377 | 0.11 |
ENST00000436451.2 |
IGKV6D-21 |
immunoglobulin kappa variable 6D-21 (non-functional) |
| chr11_+_5474638 | 0.11 |
ENST00000341449.2 |
OR51I2 |
olfactory receptor, family 51, subfamily I, member 2 |
| chr12_-_95510743 | 0.11 |
ENST00000551521.1 |
FGD6 |
FYVE, RhoGEF and PH domain containing 6 |
| chr1_-_109935819 | 0.10 |
ENST00000538502.1 |
SORT1 |
sortilin 1 |
| chrX_-_131262048 | 0.10 |
ENST00000298542.4 |
FRMD7 |
FERM domain containing 7 |
| chr17_-_41623716 | 0.10 |
ENST00000319349.5 |
ETV4 |
ets variant 4 |
| chr1_-_25291475 | 0.10 |
ENST00000338888.3 ENST00000399916.1 |
RUNX3 |
runt-related transcription factor 3 |
| chr1_-_226926864 | 0.10 |
ENST00000429204.1 ENST00000366784.1 |
ITPKB |
inositol-trisphosphate 3-kinase B |
| chr18_-_61311485 | 0.10 |
ENST00000436264.1 ENST00000356424.6 ENST00000341074.5 |
SERPINB4 |
serpin peptidase inhibitor, clade B (ovalbumin), member 4 |
| chr3_+_136676851 | 0.09 |
ENST00000309741.5 |
IL20RB |
interleukin 20 receptor beta |
| chr4_-_110723335 | 0.09 |
ENST00000394634.2 |
CFI |
complement factor I |
| chr4_-_110723134 | 0.08 |
ENST00000510800.1 ENST00000512148.1 |
CFI |
complement factor I |
| chr4_-_110723194 | 0.08 |
ENST00000394635.3 |
CFI |
complement factor I |
| chr2_-_89459813 | 0.08 |
ENST00000390256.2 |
IGKV6-21 |
immunoglobulin kappa variable 6-21 (non-functional) |
| chr4_+_88896819 | 0.08 |
ENST00000237623.7 ENST00000395080.3 ENST00000508233.1 ENST00000360804.4 |
SPP1 |
secreted phosphoprotein 1 |
| chr17_+_7387677 | 0.08 |
ENST00000322644.6 |
POLR2A |
polymerase (RNA) II (DNA directed) polypeptide A, 220kDa |
| chr3_+_136676707 | 0.08 |
ENST00000329582.4 |
IL20RB |
interleukin 20 receptor beta |
| chr6_+_135502466 | 0.08 |
ENST00000367814.4 |
MYB |
v-myb avian myeloblastosis viral oncogene homolog |
| chr11_-_107729887 | 0.08 |
ENST00000525815.1 |
SLC35F2 |
solute carrier family 35, member F2 |
| chr6_+_135502408 | 0.08 |
ENST00000341911.5 ENST00000442647.2 ENST00000316528.8 |
MYB |
v-myb avian myeloblastosis viral oncogene homolog |
| chrX_+_41548220 | 0.07 |
ENST00000378142.4 |
GPR34 |
G protein-coupled receptor 34 |
| chr15_+_80364901 | 0.07 |
ENST00000560228.1 ENST00000559835.1 ENST00000559775.1 ENST00000558688.1 ENST00000560392.1 ENST00000560976.1 ENST00000558272.1 ENST00000558390.1 |
ZFAND6 |
zinc finger, AN1-type domain 6 |
| chr14_-_106453155 | 0.07 |
ENST00000390594.2 |
IGHV1-2 |
immunoglobulin heavy variable 1-2 |
| chr13_-_20110902 | 0.07 |
ENST00000390680.2 ENST00000382977.4 ENST00000382975.4 ENST00000457266.2 |
TPTE2 |
transmembrane phosphoinositide 3-phosphatase and tensin homolog 2 |
| chr11_-_62609281 | 0.07 |
ENST00000525239.1 ENST00000538098.2 |
WDR74 |
WD repeat domain 74 |
| chr12_-_50616382 | 0.07 |
ENST00000552783.1 |
LIMA1 |
LIM domain and actin binding 1 |
| chr12_-_102874416 | 0.07 |
ENST00000392904.1 ENST00000337514.6 |
IGF1 |
insulin-like growth factor 1 (somatomedin C) |
| chr4_-_153274078 | 0.07 |
ENST00000263981.5 |
FBXW7 |
F-box and WD repeat domain containing 7, E3 ubiquitin protein ligase |
| chr15_-_45670924 | 0.07 |
ENST00000396659.3 |
GATM |
glycine amidinotransferase (L-arginine:glycine amidinotransferase) |
| chr17_+_38171681 | 0.07 |
ENST00000225474.2 ENST00000331769.2 ENST00000394148.3 ENST00000577675.1 |
CSF3 |
colony stimulating factor 3 (granulocyte) |
| chr7_+_143880559 | 0.06 |
ENST00000486333.1 |
CTAGE4 |
CTAGE family, member 4 |
| chr13_+_48611665 | 0.06 |
ENST00000258662.2 |
NUDT15 |
nudix (nucleoside diphosphate linked moiety X)-type motif 15 |
| chr12_-_11139511 | 0.06 |
ENST00000506868.1 |
TAS2R50 |
taste receptor, type 2, member 50 |
| chr8_-_86290333 | 0.06 |
ENST00000521846.1 ENST00000523022.1 ENST00000524324.1 ENST00000519991.1 ENST00000520663.1 ENST00000517590.1 ENST00000522579.1 ENST00000522814.1 ENST00000522662.1 ENST00000523858.1 ENST00000519129.1 |
CA1 |
carbonic anhydrase I |
| chr2_-_74007193 | 0.06 |
ENST00000377706.4 ENST00000443070.1 ENST00000272444.3 |
DUSP11 |
dual specificity phosphatase 11 (RNA/RNP complex 1-interacting) |
| chr6_-_132032206 | 0.06 |
ENST00000314099.8 |
CTAGE9 |
CTAGE family, member 9 |
| chr17_+_38171614 | 0.06 |
ENST00000583218.1 ENST00000394149.3 |
CSF3 |
colony stimulating factor 3 (granulocyte) |
| chr4_-_120550146 | 0.06 |
ENST00000354960.3 |
PDE5A |
phosphodiesterase 5A, cGMP-specific |
| chr1_-_106161540 | 0.06 |
ENST00000420901.1 ENST00000610126.1 ENST00000435253.2 |
RP11-251P6.1 |
RP11-251P6.1 |
| chr14_-_106967788 | 0.06 |
ENST00000390622.2 |
IGHV1-46 |
immunoglobulin heavy variable 1-46 |
| chr17_-_29624343 | 0.06 |
ENST00000247271.4 |
OMG |
oligodendrocyte myelin glycoprotein |
| chr5_-_78281775 | 0.06 |
ENST00000396151.3 ENST00000565165.1 |
ARSB |
arylsulfatase B |
| chr7_-_143966381 | 0.06 |
ENST00000487179.1 |
CTAGE8 |
CTAGE family, member 8 |
| chr1_+_152975488 | 0.06 |
ENST00000542696.1 |
SPRR3 |
small proline-rich protein 3 |
| chr4_+_100432161 | 0.06 |
ENST00000326581.4 ENST00000514652.1 |
C4orf17 |
chromosome 4 open reading frame 17 |
| chr3_-_74570291 | 0.06 |
ENST00000263665.6 |
CNTN3 |
contactin 3 (plasmacytoma associated) |
| chr6_+_135502501 | 0.05 |
ENST00000527615.1 ENST00000420123.2 ENST00000525369.1 ENST00000528774.1 ENST00000534121.1 ENST00000534044.1 ENST00000533624.1 |
MYB |
v-myb avian myeloblastosis viral oncogene homolog |
| chr3_-_121379739 | 0.05 |
ENST00000428394.2 ENST00000314583.3 |
HCLS1 |
hematopoietic cell-specific Lyn substrate 1 |
| chr11_-_101000445 | 0.05 |
ENST00000534013.1 |
PGR |
progesterone receptor |
| chr2_+_103035102 | 0.05 |
ENST00000264260.2 |
IL18RAP |
interleukin 18 receptor accessory protein |
| chr12_+_105380073 | 0.05 |
ENST00000552951.1 ENST00000280749.5 |
C12orf45 |
chromosome 12 open reading frame 45 |
| chr9_+_117904097 | 0.05 |
ENST00000374016.1 |
DEC1 |
deleted in esophageal cancer 1 |
| chr6_+_26217159 | 0.05 |
ENST00000303910.2 |
HIST1H2AE |
histone cluster 1, H2ae |
| chr12_-_102874378 | 0.05 |
ENST00000456098.1 |
IGF1 |
insulin-like growth factor 1 (somatomedin C) |
| chr2_-_152684977 | 0.05 |
ENST00000428992.2 ENST00000295087.8 |
ARL5A |
ADP-ribosylation factor-like 5A |
| chr11_-_58980342 | 0.05 |
ENST00000361050.3 |
MPEG1 |
macrophage expressed 1 |
| chr12_-_102874330 | 0.05 |
ENST00000307046.8 |
IGF1 |
insulin-like growth factor 1 (somatomedin C) |
| chr4_+_106631966 | 0.05 |
ENST00000360505.5 ENST00000510865.1 ENST00000509336.1 |
GSTCD |
glutathione S-transferase, C-terminal domain containing |
| chr3_-_148939835 | 0.05 |
ENST00000264613.6 |
CP |
ceruloplasmin (ferroxidase) |
| chr6_+_122720681 | 0.05 |
ENST00000368455.4 ENST00000452194.1 |
HSF2 |
heat shock transcription factor 2 |
| chr4_+_74301880 | 0.05 |
ENST00000395792.2 ENST00000226359.2 |
AFP |
alpha-fetoprotein |
| chr6_+_127898312 | 0.04 |
ENST00000329722.7 |
C6orf58 |
chromosome 6 open reading frame 58 |
| chr3_+_118892362 | 0.04 |
ENST00000497685.1 ENST00000264234.3 |
UPK1B |
uroplakin 1B |
| chr6_-_26124138 | 0.04 |
ENST00000314332.5 ENST00000396984.1 |
HIST1H2BC |
histone cluster 1, H2bc |
| chr16_-_20416053 | 0.04 |
ENST00000302451.4 |
PDILT |
protein disulfide isomerase-like, testis expressed |
| chr17_-_64225508 | 0.04 |
ENST00000205948.6 |
APOH |
apolipoprotein H (beta-2-glycoprotein I) |
| chr15_-_55489097 | 0.04 |
ENST00000260443.4 |
RSL24D1 |
ribosomal L24 domain containing 1 |
| chr11_+_74459876 | 0.04 |
ENST00000299563.4 |
RNF169 |
ring finger protein 169 |
| chr12_+_72056773 | 0.04 |
ENST00000308086.2 |
THAP2 |
THAP domain containing, apoptosis associated protein 2 |
| chr14_-_21270561 | 0.04 |
ENST00000412779.2 |
RNASE1 |
ribonuclease, RNase A family, 1 (pancreatic) |
| chr6_-_26216872 | 0.04 |
ENST00000244601.3 |
HIST1H2BG |
histone cluster 1, H2bg |
| chr11_-_128457446 | 0.04 |
ENST00000392668.4 |
ETS1 |
v-ets avian erythroblastosis virus E26 oncogene homolog 1 |
| chr3_+_63428752 | 0.04 |
ENST00000295894.5 |
SYNPR |
synaptoporin |
| chrX_+_135388147 | 0.04 |
ENST00000394141.1 |
GPR112 |
G protein-coupled receptor 112 |
| chr5_+_61874562 | 0.04 |
ENST00000334994.5 ENST00000409534.1 |
LRRC70 IPO11 |
leucine rich repeat containing 70 importin 11 |
| chr12_+_101988627 | 0.04 |
ENST00000547405.1 ENST00000452455.2 ENST00000441232.1 ENST00000360610.2 ENST00000392934.3 ENST00000547509.1 ENST00000361685.2 ENST00000549145.1 ENST00000553190.1 |
MYBPC1 |
myosin binding protein C, slow type |
| chr3_-_33686925 | 0.04 |
ENST00000485378.2 ENST00000313350.6 ENST00000487200.1 |
CLASP2 |
cytoplasmic linker associated protein 2 |
| chr1_-_113478603 | 0.04 |
ENST00000443580.1 |
SLC16A1 |
solute carrier family 16 (monocarboxylate transporter), member 1 |
| chrX_+_15518923 | 0.04 |
ENST00000348343.6 |
BMX |
BMX non-receptor tyrosine kinase |
| chr10_+_91589309 | 0.03 |
ENST00000448490.1 |
LINC00865 |
long intergenic non-protein coding RNA 865 |
| chr3_-_58200398 | 0.03 |
ENST00000318316.3 ENST00000460422.1 ENST00000483681.1 |
DNASE1L3 |
deoxyribonuclease I-like 3 |
| chr3_+_196466710 | 0.03 |
ENST00000327134.3 |
PAK2 |
p21 protein (Cdc42/Rac)-activated kinase 2 |
| chr3_-_33686743 | 0.03 |
ENST00000333778.6 ENST00000539981.1 |
CLASP2 |
cytoplasmic linker associated protein 2 |
| chr10_+_91589261 | 0.03 |
ENST00000448963.1 |
LINC00865 |
long intergenic non-protein coding RNA 865 |
| chr5_-_98262240 | 0.03 |
ENST00000284049.3 |
CHD1 |
chromodomain helicase DNA binding protein 1 |
| chr13_+_108870714 | 0.03 |
ENST00000375898.3 |
ABHD13 |
abhydrolase domain containing 13 |
| chr12_-_72057571 | 0.03 |
ENST00000548100.1 |
ZFC3H1 |
zinc finger, C3H1-type containing |
| chr1_-_223308098 | 0.03 |
ENST00000342210.6 |
TLR5 |
toll-like receptor 5 |
| chr14_-_21270995 | 0.03 |
ENST00000555698.1 ENST00000397970.4 ENST00000340900.3 |
RNASE1 |
ribonuclease, RNase A family, 1 (pancreatic) |
| chr2_-_89521942 | 0.03 |
ENST00000482769.1 |
IGKV2-28 |
immunoglobulin kappa variable 2-28 |
| chr9_+_135854091 | 0.03 |
ENST00000450530.1 ENST00000534944.1 |
GFI1B |
growth factor independent 1B transcription repressor |
| chrX_-_130423386 | 0.03 |
ENST00000370903.3 |
IGSF1 |
immunoglobulin superfamily, member 1 |
| chr3_+_151591422 | 0.03 |
ENST00000362032.5 |
SUCNR1 |
succinate receptor 1 |
| chr2_-_166930131 | 0.03 |
ENST00000303395.4 ENST00000409050.1 ENST00000423058.2 ENST00000375405.3 |
SCN1A |
sodium channel, voltage-gated, type I, alpha subunit |
| chr3_+_107318157 | 0.03 |
ENST00000406780.1 |
BBX |
bobby sox homolog (Drosophila) |
| chr3_+_63428982 | 0.03 |
ENST00000479198.1 ENST00000460711.1 ENST00000465156.1 |
SYNPR |
synaptoporin |
| chrX_-_130423200 | 0.03 |
ENST00000361420.3 |
IGSF1 |
immunoglobulin superfamily, member 1 |
| chr4_-_100356551 | 0.03 |
ENST00000209665.4 |
ADH7 |
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide |
| chr15_+_43425672 | 0.03 |
ENST00000260403.2 |
TMEM62 |
transmembrane protein 62 |
| chr9_-_116065551 | 0.03 |
ENST00000297894.5 |
RNF183 |
ring finger protein 183 |
| chr8_-_33370607 | 0.03 |
ENST00000360742.5 ENST00000523305.1 |
TTI2 |
TELO2 interacting protein 2 |
| chr1_-_201123586 | 0.03 |
ENST00000414605.2 ENST00000367334.5 ENST00000367332.1 |
TMEM9 |
transmembrane protein 9 |
| chr19_+_55315059 | 0.03 |
ENST00000359085.4 |
KIR2DL4 |
killer cell immunoglobulin-like receptor, two domains, long cytoplasmic tail, 4 |
| chr7_+_20686946 | 0.03 |
ENST00000443026.2 ENST00000406935.1 |
ABCB5 |
ATP-binding cassette, sub-family B (MDR/TAP), member 5 |
| chr14_+_89290965 | 0.03 |
ENST00000345383.5 ENST00000536576.1 ENST00000346301.4 ENST00000338104.6 ENST00000354441.6 ENST00000380656.2 ENST00000556651.1 ENST00000554686.1 |
TTC8 |
tetratricopeptide repeat domain 8 |
| chr1_-_201123546 | 0.03 |
ENST00000435310.1 ENST00000485839.2 ENST00000367330.1 |
TMEM9 |
transmembrane protein 9 |
| chr8_+_30244580 | 0.02 |
ENST00000523115.1 ENST00000519647.1 |
RBPMS |
RNA binding protein with multiple splicing |
| chr13_+_52436111 | 0.02 |
ENST00000242819.4 |
CCDC70 |
coiled-coil domain containing 70 |
| chrX_-_6146876 | 0.02 |
ENST00000381095.3 |
NLGN4X |
neuroligin 4, X-linked |
| chr16_+_19222479 | 0.02 |
ENST00000568433.1 |
SYT17 |
synaptotagmin XVII |
| chr1_-_160681593 | 0.02 |
ENST00000368045.3 ENST00000368046.3 |
CD48 |
CD48 molecule |
| chr10_-_52008313 | 0.02 |
ENST00000329428.6 ENST00000395526.4 ENST00000447815.1 |
ASAH2 |
N-acylsphingosine amidohydrolase (non-lysosomal ceramidase) 2 |
| chr5_-_176326333 | 0.02 |
ENST00000292432.5 |
HK3 |
hexokinase 3 (white cell) |
| chr12_-_111358372 | 0.02 |
ENST00000548438.1 ENST00000228841.8 |
MYL2 |
myosin, light chain 2, regulatory, cardiac, slow |
| chr17_-_39324424 | 0.02 |
ENST00000391356.2 |
KRTAP4-3 |
keratin associated protein 4-3 |
| chr4_+_70796784 | 0.02 |
ENST00000246891.4 ENST00000444405.3 |
CSN1S1 |
casein alpha s1 |
| chr18_+_616711 | 0.02 |
ENST00000579494.1 |
CLUL1 |
clusterin-like 1 (retinal) |
| chr18_+_3252206 | 0.02 |
ENST00000578562.2 |
MYL12A |
myosin, light chain 12A, regulatory, non-sarcomeric |
| chr12_-_11062161 | 0.02 |
ENST00000390677.2 |
TAS2R13 |
taste receptor, type 2, member 13 |
| chr15_+_48623208 | 0.02 |
ENST00000559935.1 ENST00000559416.1 |
DUT |
deoxyuridine triphosphatase |
| chr4_-_100356291 | 0.02 |
ENST00000476959.1 ENST00000482593.1 |
ADH7 |
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide |
| chr1_+_100436065 | 0.02 |
ENST00000370153.1 |
SLC35A3 |
solute carrier family 35 (UDP-N-acetylglucosamine (UDP-GlcNAc) transporter), member A3 |
| chr3_-_107777208 | 0.02 |
ENST00000398258.3 |
CD47 |
CD47 molecule |
| chr2_+_62900986 | 0.02 |
ENST00000405015.3 ENST00000413434.1 ENST00000426940.1 ENST00000449820.1 |
EHBP1 |
EH domain binding protein 1 |
| chr9_-_39239171 | 0.02 |
ENST00000358144.2 |
CNTNAP3 |
contactin associated protein-like 3 |
| chr22_+_24030321 | 0.02 |
ENST00000401461.1 |
RGL4 |
ral guanine nucleotide dissociation stimulator-like 4 |
| chr6_-_132834184 | 0.02 |
ENST00000367941.2 ENST00000367937.4 |
STX7 |
syntaxin 7 |
| chr19_+_55315087 | 0.02 |
ENST00000345540.5 ENST00000357494.4 ENST00000396293.1 ENST00000346587.4 ENST00000396289.5 |
KIR2DL4 |
killer cell immunoglobulin-like receptor, two domains, long cytoplasmic tail, 4 |
| chr20_+_61427797 | 0.02 |
ENST00000370487.3 |
MRGBP |
MRG/MORF4L binding protein |
| chr2_+_44001172 | 0.02 |
ENST00000260605.8 ENST00000406852.3 ENST00000443170.3 ENST00000398823.2 ENST00000605786.1 |
DYNC2LI1 |
dynein, cytoplasmic 2, light intermediate chain 1 |
| chr14_-_107283278 | 0.02 |
ENST00000390639.2 |
IGHV7-81 |
immunoglobulin heavy variable 7-81 (non-functional) |
| chr9_-_21166659 | 0.02 |
ENST00000380225.1 |
IFNA21 |
interferon, alpha 21 |
| chr4_-_177190364 | 0.02 |
ENST00000296525.3 |
ASB5 |
ankyrin repeat and SOCS box containing 5 |
| chr11_-_7818520 | 0.02 |
ENST00000329434.2 |
OR5P2 |
olfactory receptor, family 5, subfamily P, member 2 |
| chr4_+_70861647 | 0.02 |
ENST00000246895.4 ENST00000381060.2 |
STATH |
statherin |
| chr8_-_49833978 | 0.02 |
ENST00000020945.1 |
SNAI2 |
snail family zinc finger 2 |
| chr1_-_157567868 | 0.02 |
ENST00000271532.1 |
FCRL4 |
Fc receptor-like 4 |
| chr6_-_123958141 | 0.02 |
ENST00000334268.4 |
TRDN |
triadin |
| chr3_-_46249878 | 0.01 |
ENST00000296140.3 |
CCR1 |
chemokine (C-C motif) receptor 1 |
| chr2_-_89292422 | 0.01 |
ENST00000495489.1 |
IGKV1-8 |
immunoglobulin kappa variable 1-8 |
| chr12_-_118797475 | 0.01 |
ENST00000541786.1 ENST00000419821.2 ENST00000541878.1 |
TAOK3 |
TAO kinase 3 |
| chr12_-_110906027 | 0.01 |
ENST00000537466.2 ENST00000550974.1 ENST00000228827.3 |
GPN3 |
GPN-loop GTPase 3 |
| chr6_-_29055090 | 0.01 |
ENST00000377173.2 |
OR2B3 |
olfactory receptor, family 2, subfamily B, member 3 |
| chr1_-_153283194 | 0.01 |
ENST00000290722.1 |
PGLYRP3 |
peptidoglycan recognition protein 3 |
| chr1_+_25598872 | 0.01 |
ENST00000328664.4 |
RHD |
Rh blood group, D antigen |
| chr6_+_26124373 | 0.01 |
ENST00000377791.2 ENST00000602637.1 |
HIST1H2AC |
histone cluster 1, H2ac |
| chr5_+_125758813 | 0.01 |
ENST00000285689.3 ENST00000515200.1 |
GRAMD3 |
GRAM domain containing 3 |
| chr3_+_149530836 | 0.01 |
ENST00000466478.1 ENST00000491086.1 ENST00000467977.1 |
RNF13 |
ring finger protein 13 |
| chr14_-_106539557 | 0.01 |
ENST00000390599.2 |
IGHV1-8 |
immunoglobulin heavy variable 1-8 |
| chr2_+_89999259 | 0.01 |
ENST00000558026.1 |
IGKV2D-28 |
immunoglobulin kappa variable 2D-28 |
| chr6_-_123957942 | 0.01 |
ENST00000398178.3 |
TRDN |
triadin |
| chr11_-_110561721 | 0.01 |
ENST00000357139.3 |
ARHGAP20 |
Rho GTPase activating protein 20 |
| chr5_-_9630463 | 0.01 |
ENST00000382492.2 |
TAS2R1 |
taste receptor, type 2, member 1 |
| chr2_+_168043793 | 0.01 |
ENST00000409273.1 ENST00000409605.1 |
XIRP2 |
xin actin-binding repeat containing 2 |
| chr22_+_23089870 | 0.01 |
ENST00000390311.2 |
IGLV3-16 |
immunoglobulin lambda variable 3-16 |
| chr7_+_28452130 | 0.01 |
ENST00000357727.2 |
CREB5 |
cAMP responsive element binding protein 5 |
| chr11_+_3011093 | 0.01 |
ENST00000332881.2 |
AC131971.1 |
HCG1782999; PRO0943; Uncharacterized protein |
| chr9_-_35080013 | 0.01 |
ENST00000378643.3 |
FANCG |
Fanconi anemia, complementation group G |
| chr17_+_43239191 | 0.01 |
ENST00000589230.1 |
HEXIM2 |
hexamethylene bis-acetamide inducible 2 |
| chr16_-_18908196 | 0.01 |
ENST00000565324.1 ENST00000561947.1 |
SMG1 |
SMG1 phosphatidylinositol 3-kinase-related kinase |
| chr17_-_76573465 | 0.01 |
ENST00000585328.1 ENST00000389840.5 |
DNAH17 |
dynein, axonemal, heavy chain 17 |
| chr20_+_5731083 | 0.01 |
ENST00000445603.1 ENST00000442185.1 |
C20orf196 |
chromosome 20 open reading frame 196 |
| chr22_-_29949680 | 0.01 |
ENST00000397873.2 ENST00000490103.1 |
THOC5 |
THO complex 5 |
| chr7_-_103848405 | 0.01 |
ENST00000447452.2 ENST00000545943.1 ENST00000297431.4 |
ORC5 |
origin recognition complex, subunit 5 |
| chr17_-_7387524 | 0.01 |
ENST00000311403.4 |
ZBTB4 |
zinc finger and BTB domain containing 4 |
| chr3_+_133292574 | 0.01 |
ENST00000264993.3 |
CDV3 |
CDV3 homolog (mouse) |
| chr16_+_14294187 | 0.01 |
ENST00000573051.1 |
MKL2 |
MKL/myocardin-like 2 |
| chr14_-_68283291 | 0.01 |
ENST00000555452.1 ENST00000347230.4 |
ZFYVE26 |
zinc finger, FYVE domain containing 26 |
| chr2_+_36923933 | 0.01 |
ENST00000497382.1 ENST00000404084.1 ENST00000379241.3 ENST00000401530.1 |
VIT |
vitrin |
| chr10_+_95517566 | 0.01 |
ENST00000542308.1 |
LGI1 |
leucine-rich, glioma inactivated 1 |
| chr2_+_219524379 | 0.01 |
ENST00000443791.1 ENST00000359273.3 ENST00000392109.1 ENST00000392110.2 ENST00000423377.1 ENST00000392111.2 ENST00000412366.1 |
BCS1L |
BC1 (ubiquinol-cytochrome c reductase) synthesis-like |
| chr15_-_22473353 | 0.01 |
ENST00000557788.2 |
IGHV4OR15-8 |
immunoglobulin heavy variable 4/OR15-8 (non-functional) |
| chr11_-_14521379 | 0.01 |
ENST00000249923.3 ENST00000529866.1 ENST00000439561.2 ENST00000534771.1 |
COPB1 |
coatomer protein complex, subunit beta 1 |
| chr9_-_86593238 | 0.01 |
ENST00000351839.3 |
HNRNPK |
heterogeneous nuclear ribonucleoprotein K |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0060559 | positive regulation of vitamin metabolic process(GO:0046136) positive regulation of vitamin D biosynthetic process(GO:0060557) positive regulation of calcidiol 1-monooxygenase activity(GO:0060559) |
| 0.1 | 0.2 | GO:1904899 | regulation of hepatic stellate cell proliferation(GO:1904897) positive regulation of hepatic stellate cell proliferation(GO:1904899) hepatic stellate cell proliferation(GO:1990922) |
| 0.1 | 0.3 | GO:0061146 | Peyer's patch morphogenesis(GO:0061146) lymphocyte migration into lymphoid organs(GO:0097021) |
| 0.0 | 0.2 | GO:0001808 | negative regulation of type IV hypersensitivity(GO:0001808) |
| 0.0 | 0.1 | GO:0050712 | negative regulation of interleukin-1 alpha production(GO:0032690) negative regulation of interleukin-1 alpha secretion(GO:0050712) |
| 0.0 | 0.2 | GO:1904073 | trophectodermal cell proliferation(GO:0001834) regulation of trophectodermal cell proliferation(GO:1904073) positive regulation of trophectodermal cell proliferation(GO:1904075) |
| 0.0 | 0.1 | GO:0042270 | protection from natural killer cell mediated cytotoxicity(GO:0042270) |
| 0.0 | 0.1 | GO:2000866 | positive regulation of estrogen secretion(GO:2000863) positive regulation of estradiol secretion(GO:2000866) |
| 0.0 | 0.1 | GO:0001172 | transcription, RNA-templated(GO:0001172) |
| 0.0 | 0.2 | GO:0014807 | regulation of somitogenesis(GO:0014807) |
| 0.0 | 0.1 | GO:0061582 | intestinal epithelial cell migration(GO:0061582) |
| 0.0 | 0.2 | GO:0048023 | positive regulation of melanin biosynthetic process(GO:0048023) positive regulation of secondary metabolite biosynthetic process(GO:1900378) |
| 0.0 | 0.0 | GO:0010430 | fatty acid omega-oxidation(GO:0010430) |
| 0.0 | 0.1 | GO:0050859 | negative regulation of B cell receptor signaling pathway(GO:0050859) |
| 0.0 | 0.1 | GO:2000639 | regulation of SREBP signaling pathway(GO:2000638) negative regulation of SREBP signaling pathway(GO:2000639) |
| 0.0 | 0.1 | GO:0060748 | tertiary branching involved in mammary gland duct morphogenesis(GO:0060748) |
| 0.0 | 0.1 | GO:0006203 | dGTP catabolic process(GO:0006203) |
| 0.0 | 0.1 | GO:0033029 | regulation of neutrophil apoptotic process(GO:0033029) common myeloid progenitor cell proliferation(GO:0035726) |
| 0.0 | 0.0 | GO:0051780 | mevalonate transport(GO:0015728) behavioral response to nutrient(GO:0051780) |
| 0.0 | 0.1 | GO:0030223 | neutrophil differentiation(GO:0030223) |
| 0.0 | 0.1 | GO:0051005 | negative regulation of lipoprotein lipase activity(GO:0051005) |
| 0.0 | 0.0 | GO:0034146 | toll-like receptor 5 signaling pathway(GO:0034146) |
| 0.0 | 0.0 | GO:1902626 | assembly of large subunit precursor of preribosome(GO:1902626) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | GO:0035867 | alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0004910 | interleukin-1, Type II, blocking receptor activity(GO:0004910) |
| 0.0 | 0.2 | GO:0042015 | interleukin-20 binding(GO:0042015) |
| 0.0 | 0.1 | GO:0003968 | RNA-directed RNA polymerase activity(GO:0003968) |
| 0.0 | 0.1 | GO:0008413 | 8-oxo-7,8-dihydroguanosine triphosphate pyrophosphatase activity(GO:0008413) 8-oxo-7,8-dihydrodeoxyguanosine triphosphate pyrophosphatase activity(GO:0035539) |
| 0.0 | 0.1 | GO:0010465 | nerve growth factor receptor activity(GO:0010465) |
| 0.0 | 0.1 | GO:0003943 | N-acetylgalactosamine-4-sulfatase activity(GO:0003943) |
| 0.0 | 0.4 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 0.1 | GO:0051800 | phosphatidylinositol-3,4-bisphosphate 3-phosphatase activity(GO:0051800) |
| 0.0 | 0.2 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.0 | 0.3 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.0 | 0.1 | GO:0098518 | polynucleotide phosphatase activity(GO:0098518) |
| 0.0 | 0.4 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.0 | GO:0004031 | aldehyde oxidase activity(GO:0004031) |
| 0.0 | 0.0 | GO:0015130 | mevalonate transmembrane transporter activity(GO:0015130) |
| 0.0 | 0.1 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.0 | 0.1 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.0 | 0.1 | GO:0004522 | ribonuclease A activity(GO:0004522) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |