Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
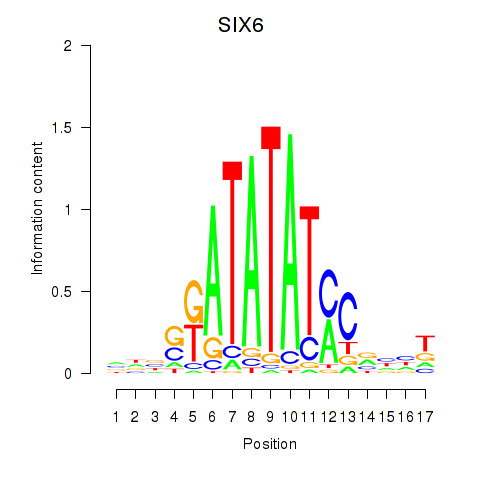
Results for SIX6
Z-value: 0.70
Transcription factors associated with SIX6
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SIX6
|
ENSG00000184302.6 | SIX6 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SIX6 | hg19_v2_chr14_+_60975644_60975673 | -0.23 | 5.8e-01 | Click! |
Activity profile of SIX6 motif
Sorted Z-values of SIX6 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of SIX6
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_-_28123206 | 0.54 |
ENST00000542963.1 ENST00000535992.1 |
PTHLH |
parathyroid hormone-like hormone |
| chr12_-_28122980 | 0.53 |
ENST00000395868.3 ENST00000534890.1 |
PTHLH |
parathyroid hormone-like hormone |
| chr9_-_23821842 | 0.43 |
ENST00000544538.1 |
ELAVL2 |
ELAV like neuron-specific RNA binding protein 2 |
| chr3_-_111314230 | 0.38 |
ENST00000317012.4 |
ZBED2 |
zinc finger, BED-type containing 2 |
| chr14_+_21525981 | 0.33 |
ENST00000308227.2 |
RNASE8 |
ribonuclease, RNase A family, 8 |
| chr8_+_101170257 | 0.29 |
ENST00000251809.3 |
SPAG1 |
sperm associated antigen 1 |
| chr6_+_37012607 | 0.24 |
ENST00000423336.1 |
COX6A1P2 |
cytochrome c oxidase subunit VIa polypeptide 1 pseudogene 2 |
| chr4_-_155511887 | 0.24 |
ENST00000302053.3 ENST00000403106.3 |
FGA |
fibrinogen alpha chain |
| chr5_+_55147205 | 0.23 |
ENST00000396836.2 ENST00000396834.1 ENST00000447346.2 ENST00000359040.5 |
IL31RA |
interleukin 31 receptor A |
| chr14_+_21510385 | 0.21 |
ENST00000298690.4 |
RNASE7 |
ribonuclease, RNase A family, 7 |
| chr13_+_73629107 | 0.20 |
ENST00000539231.1 |
KLF5 |
Kruppel-like factor 5 (intestinal) |
| chr11_-_104916034 | 0.20 |
ENST00000528513.1 ENST00000375706.2 ENST00000375704.3 |
CARD16 |
caspase recruitment domain family, member 16 |
| chr1_+_86934526 | 0.20 |
ENST00000394711.1 |
CLCA1 |
chloride channel accessory 1 |
| chr1_+_212738676 | 0.19 |
ENST00000366981.4 ENST00000366987.2 |
ATF3 |
activating transcription factor 3 |
| chr1_+_156119466 | 0.19 |
ENST00000414683.1 |
SEMA4A |
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A |
| chr1_+_156119798 | 0.19 |
ENST00000355014.2 |
SEMA4A |
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A |
| chr19_-_16008880 | 0.18 |
ENST00000011989.7 ENST00000221700.6 |
CYP4F2 |
cytochrome P450, family 4, subfamily F, polypeptide 2 |
| chr2_+_177134134 | 0.17 |
ENST00000249442.6 ENST00000392529.2 ENST00000443241.1 |
MTX2 |
metaxin 2 |
| chr11_-_104972158 | 0.16 |
ENST00000598974.1 ENST00000593315.1 ENST00000594519.1 ENST00000415981.2 ENST00000525374.1 ENST00000375707.1 |
CASP1 CARD16 CARD17 |
caspase 1, apoptosis-related cysteine peptidase caspase recruitment domain family, member 16 caspase recruitment domain family, member 17 |
| chr13_-_20080080 | 0.16 |
ENST00000400103.2 |
TPTE2 |
transmembrane phosphoinositide 3-phosphatase and tensin homolog 2 |
| chr1_+_160709076 | 0.15 |
ENST00000359331.4 ENST00000495334.1 |
SLAMF7 |
SLAM family member 7 |
| chr8_+_32579341 | 0.15 |
ENST00000519240.1 ENST00000539990.1 |
NRG1 |
neuregulin 1 |
| chr8_+_1993173 | 0.15 |
ENST00000523438.1 |
MYOM2 |
myomesin 2 |
| chr12_-_87232644 | 0.15 |
ENST00000549405.2 |
MGAT4C |
mannosyl (alpha-1,3-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase, isozyme C (putative) |
| chr8_+_1993152 | 0.14 |
ENST00000262113.4 |
MYOM2 |
myomesin 2 |
| chr19_+_3136115 | 0.13 |
ENST00000262958.3 |
GNA15 |
guanine nucleotide binding protein (G protein), alpha 15 (Gq class) |
| chr1_+_160709055 | 0.13 |
ENST00000368043.3 ENST00000368042.3 ENST00000458602.2 ENST00000458104.2 |
SLAMF7 |
SLAM family member 7 |
| chr2_+_177134201 | 0.13 |
ENST00000452865.1 |
MTX2 |
metaxin 2 |
| chr22_+_18721427 | 0.12 |
ENST00000342888.3 |
AC008132.1 |
Uncharacterized protein |
| chr20_+_54987305 | 0.12 |
ENST00000371336.3 ENST00000434344.1 |
CASS4 |
Cas scaffolding protein family member 4 |
| chr1_-_161015752 | 0.11 |
ENST00000435396.1 ENST00000368021.3 |
USF1 |
upstream transcription factor 1 |
| chr1_+_145413268 | 0.11 |
ENST00000421822.2 ENST00000336751.5 ENST00000497365.1 ENST00000475797.1 |
HFE2 |
hemochromatosis type 2 (juvenile) |
| chr1_+_31885963 | 0.11 |
ENST00000373709.3 |
SERINC2 |
serine incorporator 2 |
| chr11_+_33902189 | 0.10 |
ENST00000330381.2 |
AC132216.1 |
HCG1785179; PRO1787; Uncharacterized protein |
| chrX_+_8433376 | 0.10 |
ENST00000440654.2 ENST00000381029.4 |
VCX3B |
variable charge, X-linked 3B |
| chr1_+_234509186 | 0.10 |
ENST00000366615.4 |
COA6 |
cytochrome c oxidase assembly factor 6 homolog (S. cerevisiae) |
| chrX_-_20236970 | 0.10 |
ENST00000379548.4 |
RPS6KA3 |
ribosomal protein S6 kinase, 90kDa, polypeptide 3 |
| chr20_+_54987168 | 0.09 |
ENST00000360314.3 |
CASS4 |
Cas scaffolding protein family member 4 |
| chr5_+_140201183 | 0.09 |
ENST00000529619.1 ENST00000529859.1 ENST00000378126.3 |
PCDHA5 |
protocadherin alpha 5 |
| chr8_-_141774467 | 0.08 |
ENST00000520151.1 ENST00000519024.1 ENST00000519465.1 |
PTK2 |
protein tyrosine kinase 2 |
| chr16_+_19467772 | 0.08 |
ENST00000219821.5 ENST00000561503.1 ENST00000564959.1 |
TMC5 |
transmembrane channel-like 5 |
| chr5_-_78281775 | 0.07 |
ENST00000396151.3 ENST00000565165.1 |
ARSB |
arylsulfatase B |
| chr7_+_117251671 | 0.07 |
ENST00000468795.1 |
CFTR |
cystic fibrosis transmembrane conductance regulator (ATP-binding cassette sub-family C, member 7) |
| chr17_+_7788104 | 0.07 |
ENST00000380358.4 |
CHD3 |
chromodomain helicase DNA binding protein 3 |
| chr19_-_1650666 | 0.07 |
ENST00000588136.1 |
TCF3 |
transcription factor 3 |
| chr19_-_14628645 | 0.07 |
ENST00000598235.1 |
DNAJB1 |
DnaJ (Hsp40) homolog, subfamily B, member 1 |
| chr17_-_42327236 | 0.07 |
ENST00000399246.2 |
AC003102.1 |
AC003102.1 |
| chr1_+_234509413 | 0.07 |
ENST00000366613.1 ENST00000366612.1 |
COA6 |
cytochrome c oxidase assembly factor 6 homolog (S. cerevisiae) |
| chr4_+_74301880 | 0.07 |
ENST00000395792.2 ENST00000226359.2 |
AFP |
alpha-fetoprotein |
| chr14_+_20937538 | 0.06 |
ENST00000361505.5 ENST00000553591.1 |
PNP |
purine nucleoside phosphorylase |
| chr3_+_125694347 | 0.06 |
ENST00000505382.1 ENST00000511082.1 |
ROPN1B |
rhophilin associated tail protein 1B |
| chr12_-_14849470 | 0.06 |
ENST00000261170.3 |
GUCY2C |
guanylate cyclase 2C (heat stable enterotoxin receptor) |
| chr16_-_20364122 | 0.06 |
ENST00000396138.4 ENST00000577168.1 |
UMOD |
uromodulin |
| chr1_+_160709029 | 0.06 |
ENST00000444090.2 ENST00000441662.2 |
SLAMF7 |
SLAM family member 7 |
| chr2_-_134326009 | 0.06 |
ENST00000409261.1 ENST00000409213.1 |
NCKAP5 |
NCK-associated protein 5 |
| chr2_-_77749474 | 0.06 |
ENST00000409093.1 ENST00000409088.3 |
LRRTM4 |
leucine rich repeat transmembrane neuronal 4 |
| chr9_-_14910990 | 0.06 |
ENST00000380881.4 ENST00000422223.2 |
FREM1 |
FRAS1 related extracellular matrix 1 |
| chr4_-_123377880 | 0.05 |
ENST00000226730.4 |
IL2 |
interleukin 2 |
| chr1_-_67142710 | 0.05 |
ENST00000502413.2 |
AL139147.1 |
Uncharacterized protein |
| chr16_-_20364030 | 0.05 |
ENST00000396134.2 ENST00000573567.1 ENST00000570757.1 ENST00000424589.1 ENST00000302509.4 ENST00000571174.1 ENST00000576688.1 |
UMOD |
uromodulin |
| chrX_-_118699325 | 0.05 |
ENST00000320339.4 ENST00000371594.4 ENST00000536133.1 |
CXorf56 |
chromosome X open reading frame 56 |
| chr13_+_34392185 | 0.05 |
ENST00000380071.3 |
RFC3 |
replication factor C (activator 1) 3, 38kDa |
| chr15_+_58430567 | 0.05 |
ENST00000536493.1 |
AQP9 |
aquaporin 9 |
| chr17_-_5342380 | 0.05 |
ENST00000225698.4 |
C1QBP |
complement component 1, q subcomponent binding protein |
| chr15_-_80263506 | 0.05 |
ENST00000335661.6 |
BCL2A1 |
BCL2-related protein A1 |
| chrY_-_16098393 | 0.05 |
ENST00000250825.4 |
VCY |
variable charge, Y-linked |
| chr15_+_58430368 | 0.05 |
ENST00000558772.1 ENST00000219919.4 |
AQP9 |
aquaporin 9 |
| chr5_+_40909354 | 0.05 |
ENST00000313164.9 |
C7 |
complement component 7 |
| chrX_+_38660704 | 0.04 |
ENST00000378474.3 |
MID1IP1 |
MID1 interacting protein 1 |
| chr1_+_63063152 | 0.04 |
ENST00000371129.3 |
ANGPTL3 |
angiopoietin-like 3 |
| chr22_-_19137796 | 0.04 |
ENST00000086933.2 |
GSC2 |
goosecoid homeobox 2 |
| chrY_+_16168097 | 0.04 |
ENST00000250823.4 |
VCY1B |
variable charge, Y-linked 1B |
| chr10_+_91061712 | 0.04 |
ENST00000371826.3 |
IFIT2 |
interferon-induced protein with tetratricopeptide repeats 2 |
| chr12_-_121477039 | 0.04 |
ENST00000257570.5 |
OASL |
2'-5'-oligoadenylate synthetase-like |
| chr6_-_108145499 | 0.04 |
ENST00000369020.3 ENST00000369022.2 |
SCML4 |
sex comb on midleg-like 4 (Drosophila) |
| chr12_+_19282643 | 0.04 |
ENST00000317589.4 ENST00000355397.3 ENST00000359180.3 ENST00000309364.4 ENST00000540972.1 ENST00000429027.2 |
PLEKHA5 |
pleckstrin homology domain containing, family A member 5 |
| chr2_-_239140011 | 0.04 |
ENST00000409376.1 ENST00000409070.1 ENST00000409942.1 |
AC016757.3 |
Protein LOC151174 |
| chr4_-_71532339 | 0.03 |
ENST00000254801.4 |
IGJ |
immunoglobulin J polypeptide, linker protein for immunoglobulin alpha and mu polypeptides |
| chr4_+_88532028 | 0.03 |
ENST00000282478.7 |
DSPP |
dentin sialophosphoprotein |
| chr17_-_67057114 | 0.03 |
ENST00000370732.2 |
ABCA9 |
ATP-binding cassette, sub-family A (ABC1), member 9 |
| chr4_+_113739244 | 0.03 |
ENST00000503271.1 ENST00000503423.1 ENST00000506722.1 |
ANK2 |
ankyrin 2, neuronal |
| chr1_+_53308398 | 0.03 |
ENST00000371528.1 |
ZYG11A |
zyg-11 family member A, cell cycle regulator |
| chrY_-_24038660 | 0.03 |
ENST00000382677.3 |
RBMY1D |
RNA binding motif protein, Y-linked, family 1, member D |
| chrY_+_23698778 | 0.03 |
ENST00000303902.5 |
RBMY1A1 |
RNA binding motif protein, Y-linked, family 1, member A1 |
| chr6_+_127898312 | 0.03 |
ENST00000329722.7 |
C6orf58 |
chromosome 6 open reading frame 58 |
| chr12_-_121476959 | 0.03 |
ENST00000339275.5 |
OASL |
2'-5'-oligoadenylate synthetase-like |
| chr15_+_69857515 | 0.03 |
ENST00000559477.1 |
RP11-279F6.1 |
RP11-279F6.1 |
| chr6_+_52051171 | 0.03 |
ENST00000340057.1 |
IL17A |
interleukin 17A |
| chrX_+_64708615 | 0.03 |
ENST00000338957.4 ENST00000423889.3 |
ZC3H12B |
zinc finger CCCH-type containing 12B |
| chr12_-_122018346 | 0.03 |
ENST00000377069.4 |
KDM2B |
lysine (K)-specific demethylase 2B |
| chr8_+_84824920 | 0.03 |
ENST00000523678.1 |
RP11-120I21.2 |
RP11-120I21.2 |
| chr10_-_61720640 | 0.02 |
ENST00000521074.1 ENST00000444900.1 |
C10orf40 |
chromosome 10 open reading frame 40 |
| chr7_-_44530479 | 0.02 |
ENST00000355451.7 |
NUDCD3 |
NudC domain containing 3 |
| chr1_-_13673511 | 0.02 |
ENST00000344998.3 ENST00000334600.6 |
PRAMEF14 |
PRAME family member 14 |
| chr4_+_74347400 | 0.02 |
ENST00000226355.3 |
AFM |
afamin |
| chr12_-_121476750 | 0.02 |
ENST00000543677.1 |
OASL |
2'-5'-oligoadenylate synthetase-like |
| chr9_-_21351377 | 0.02 |
ENST00000380210.1 |
IFNA6 |
interferon, alpha 6 |
| chr9_-_7800067 | 0.02 |
ENST00000358227.4 |
TMEM261 |
transmembrane protein 261 |
| chr17_-_67057047 | 0.02 |
ENST00000495634.1 ENST00000453985.2 ENST00000585714.1 |
ABCA9 |
ATP-binding cassette, sub-family A (ABC1), member 9 |
| chr1_+_159557607 | 0.02 |
ENST00000255040.2 |
APCS |
amyloid P component, serum |
| chr19_-_44031375 | 0.02 |
ENST00000292147.2 |
ETHE1 |
ethylmalonic encephalopathy 1 |
| chr9_-_99382065 | 0.02 |
ENST00000265659.2 ENST00000375241.1 ENST00000375236.1 |
CDC14B |
cell division cycle 14B |
| chr4_-_42154895 | 0.02 |
ENST00000502486.1 ENST00000504360.1 |
BEND4 |
BEN domain containing 4 |
| chrX_+_128674213 | 0.02 |
ENST00000371113.4 ENST00000357121.5 |
OCRL |
oculocerebrorenal syndrome of Lowe |
| chr10_+_52094298 | 0.02 |
ENST00000595931.1 |
AC069547.1 |
HCG1745369; PRO3073; Uncharacterized protein |
| chr11_+_74699942 | 0.02 |
ENST00000526068.1 ENST00000532963.1 ENST00000531619.1 ENST00000534628.1 ENST00000545272.1 |
NEU3 |
sialidase 3 (membrane sialidase) |
| chr1_+_112016414 | 0.01 |
ENST00000343534.5 ENST00000369718.3 |
C1orf162 |
chromosome 1 open reading frame 162 |
| chr11_+_74699703 | 0.01 |
ENST00000529024.1 ENST00000544263.1 |
NEU3 |
sialidase 3 (membrane sialidase) |
| chr19_+_46000479 | 0.01 |
ENST00000456399.2 |
PPM1N |
protein phosphatase, Mg2+/Mn2+ dependent, 1N (putative) |
| chrX_-_55020511 | 0.01 |
ENST00000375006.3 ENST00000374992.2 |
PFKFB1 |
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 |
| chr2_+_210636697 | 0.01 |
ENST00000439458.1 ENST00000272845.6 |
UNC80 |
unc-80 homolog (C. elegans) |
| chr18_+_55018044 | 0.01 |
ENST00000324000.3 |
ST8SIA3 |
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 3 |
| chr9_-_100684845 | 0.01 |
ENST00000375119.3 |
C9orf156 |
chromosome 9 open reading frame 156 |
| chr4_-_139051839 | 0.01 |
ENST00000514600.1 ENST00000513895.1 ENST00000512536.1 |
LINC00616 |
long intergenic non-protein coding RNA 616 |
| chr16_+_72090053 | 0.01 |
ENST00000576168.2 ENST00000567185.3 ENST00000567612.2 |
HP |
haptoglobin |
| chr6_-_8102279 | 0.00 |
ENST00000488226.2 |
EEF1E1 |
eukaryotic translation elongation factor 1 epsilon 1 |
| chr16_-_52119019 | 0.00 |
ENST00000561513.1 ENST00000565742.1 |
LINC00919 |
long intergenic non-protein coding RNA 919 |
| chr7_-_92855762 | 0.00 |
ENST00000453812.2 ENST00000394468.2 |
HEPACAM2 |
HEPACAM family member 2 |
| chr2_-_197457335 | 0.00 |
ENST00000260983.3 |
HECW2 |
HECT, C2 and WW domain containing E3 ubiquitin protein ligase 2 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0061760 | antifungal innate immune response(GO:0061760) |
| 0.1 | 0.2 | GO:0061394 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) |
| 0.1 | 0.2 | GO:0032304 | negative regulation of icosanoid secretion(GO:0032304) |
| 0.0 | 0.4 | GO:0071494 | cellular response to UV-C(GO:0071494) |
| 0.0 | 0.3 | GO:0002074 | extraocular skeletal muscle development(GO:0002074) |
| 0.0 | 1.0 | GO:0032331 | negative regulation of chondrocyte differentiation(GO:0032331) |
| 0.0 | 0.2 | GO:0051673 | membrane disruption in other organism(GO:0051673) |
| 0.0 | 0.1 | GO:0061582 | intestinal epithelial cell migration(GO:0061582) |
| 0.0 | 0.1 | GO:1902161 | positive regulation of cyclic nucleotide-gated ion channel activity(GO:1902161) |
| 0.0 | 0.2 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.0 | 0.1 | GO:0072023 | thick ascending limb development(GO:0072023) metanephric thick ascending limb development(GO:0072233) |
| 0.0 | 0.2 | GO:0032534 | regulation of microvillus assembly(GO:0032534) |
| 0.0 | 0.2 | GO:0021842 | directional guidance of interneurons involved in migration from the subpallium to the cortex(GO:0021840) chemorepulsion involved in interneuron migration from the subpallium to the cortex(GO:0021842) |
| 0.0 | 0.1 | GO:0006738 | nicotinamide riboside catabolic process(GO:0006738) nicotinamide riboside metabolic process(GO:0046495) pyridine nucleoside metabolic process(GO:0070637) pyridine nucleoside catabolic process(GO:0070638) |
| 0.0 | 0.1 | GO:1904219 | regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904217) positive regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904219) positive regulation of serine C-palmitoyltransferase activity(GO:1904222) |
| 0.0 | 0.1 | GO:0015722 | canalicular bile acid transport(GO:0015722) |
| 0.0 | 0.1 | GO:0000432 | regulation of transcription from RNA polymerase II promoter by glucose(GO:0000430) positive regulation of transcription from RNA polymerase II promoter by glucose(GO:0000432) |
| 0.0 | 0.1 | GO:0007207 | phospholipase C-activating G-protein coupled acetylcholine receptor signaling pathway(GO:0007207) |
| 0.0 | 0.1 | GO:0046013 | regulation of T cell homeostatic proliferation(GO:0046013) |
| 0.0 | 0.2 | GO:0043031 | negative regulation of macrophage activation(GO:0043031) |
| 0.0 | 0.0 | GO:1901165 | negative regulation of MDA-5 signaling pathway(GO:0039534) positive regulation of trophoblast cell migration(GO:1901165) |
| 0.0 | 0.0 | GO:0036371 | protein localization to T-tubule(GO:0036371) |
| 0.0 | 0.4 | GO:0045063 | T-helper 1 cell differentiation(GO:0045063) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 0.1 | 0.4 | GO:0097179 | protease inhibitor complex(GO:0097179) |
| 0.0 | 0.2 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.0 | 0.1 | GO:0031205 | endoplasmic reticulum Sec complex(GO:0031205) |
| 0.0 | 0.2 | GO:0042589 | zymogen granule membrane(GO:0042589) |
| 0.0 | 0.1 | GO:0016589 | NURF complex(GO:0016589) |
| 0.0 | 0.3 | GO:0032982 | myosin filament(GO:0032982) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0052870 | tocopherol omega-hydroxylase activity(GO:0052870) alpha-tocopherol omega-hydroxylase activity(GO:0052871) 20-hydroxy-leukotriene B4 omega oxidase activity(GO:0097258) 20-aldehyde-leukotriene B4 20-monooxygenase activity(GO:0097259) |
| 0.0 | 0.2 | GO:0051800 | phosphatidylinositol-3,4-bisphosphate 3-phosphatase activity(GO:0051800) |
| 0.0 | 1.1 | GO:0051428 | peptide hormone receptor binding(GO:0051428) |
| 0.0 | 0.2 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.0 | 0.4 | GO:0089720 | caspase binding(GO:0089720) |
| 0.0 | 0.2 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.0 | 0.1 | GO:0003943 | N-acetylgalactosamine-4-sulfatase activity(GO:0003943) |
| 0.0 | 0.1 | GO:0005275 | amine transmembrane transporter activity(GO:0005275) |
| 0.0 | 0.1 | GO:0005260 | channel-conductance-controlling ATPase activity(GO:0005260) |
| 0.0 | 0.1 | GO:0004731 | purine-nucleoside phosphorylase activity(GO:0004731) |
| 0.0 | 0.4 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.0 | 0.0 | GO:0030984 | kininogen binding(GO:0030984) |
| 0.0 | 0.2 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.0 | 0.1 | GO:0098821 | BMP receptor activity(GO:0098821) |
| 0.0 | 0.1 | GO:0070644 | vitamin D response element binding(GO:0070644) |
| 0.0 | 0.2 | GO:0042834 | peptidoglycan binding(GO:0042834) |
| 0.0 | 0.3 | GO:0097493 | structural molecule activity conferring elasticity(GO:0097493) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.1 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 0.2 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.0 | 0.1 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |