Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
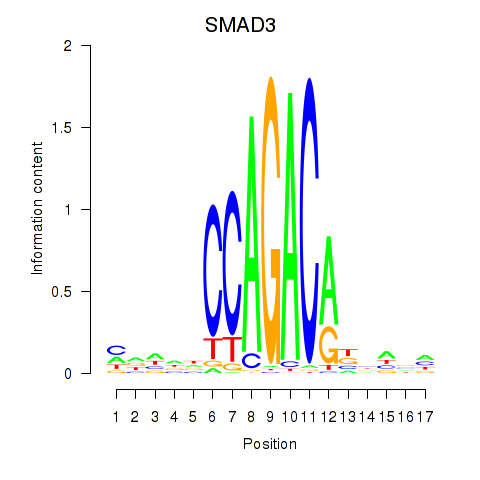
Results for SMAD3
Z-value: 0.74
Transcription factors associated with SMAD3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SMAD3
|
ENSG00000166949.11 | SMAD3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SMAD3 | hg19_v2_chr15_+_67358163_67358192 | 0.52 | 1.8e-01 | Click! |
Activity profile of SMAD3 motif
Sorted Z-values of SMAD3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of SMAD3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_209602771 | 0.93 |
ENST00000440276.1 |
MIR205HG |
MIR205 host gene (non-protein coding) |
| chr20_+_58251716 | 0.62 |
ENST00000355648.4 |
PHACTR3 |
phosphatase and actin regulator 3 |
| chr5_+_52776228 | 0.51 |
ENST00000256759.3 |
FST |
follistatin |
| chr4_-_90759440 | 0.43 |
ENST00000336904.3 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr19_-_11688447 | 0.36 |
ENST00000590420.1 |
ACP5 |
acid phosphatase 5, tartrate resistant |
| chr6_-_11807277 | 0.27 |
ENST00000379415.2 |
ADTRP |
androgen-dependent TFPI-regulating protein |
| chr6_-_27114577 | 0.25 |
ENST00000356950.1 ENST00000396891.4 |
HIST1H2BK |
histone cluster 1, H2bk |
| chr18_+_29598335 | 0.23 |
ENST00000217740.3 |
RNF125 |
ring finger protein 125, E3 ubiquitin protein ligase |
| chr1_+_153388993 | 0.23 |
ENST00000368729.4 |
S100A7A |
S100 calcium binding protein A7A |
| chr6_+_30850697 | 0.21 |
ENST00000509639.1 ENST00000412274.2 ENST00000507901.1 ENST00000507046.1 ENST00000437124.2 ENST00000454612.2 ENST00000396342.2 |
DDR1 |
discoidin domain receptor tyrosine kinase 1 |
| chr19_-_11688500 | 0.21 |
ENST00000433365.2 |
ACP5 |
acid phosphatase 5, tartrate resistant |
| chr6_-_27100529 | 0.20 |
ENST00000607124.1 ENST00000339812.2 ENST00000541790.1 |
HIST1H2BJ |
histone cluster 1, H2bj |
| chr6_+_26124373 | 0.20 |
ENST00000377791.2 ENST00000602637.1 |
HIST1H2AC |
histone cluster 1, H2ac |
| chr6_+_27791862 | 0.20 |
ENST00000355057.1 |
HIST1H4J |
histone cluster 1, H4j |
| chr6_-_133119668 | 0.19 |
ENST00000275227.4 ENST00000538764.1 |
SLC18B1 |
solute carrier family 18, subfamily B, member 1 |
| chr17_+_30813576 | 0.18 |
ENST00000313401.3 |
CDK5R1 |
cyclin-dependent kinase 5, regulatory subunit 1 (p35) |
| chr17_-_43487780 | 0.17 |
ENST00000532038.1 ENST00000528677.1 |
ARHGAP27 |
Rho GTPase activating protein 27 |
| chr20_-_23030296 | 0.16 |
ENST00000377103.2 |
THBD |
thrombomodulin |
| chr20_-_6103666 | 0.16 |
ENST00000536936.1 |
FERMT1 |
fermitin family member 1 |
| chr7_+_139529040 | 0.15 |
ENST00000455353.1 ENST00000458722.1 ENST00000411653.1 |
TBXAS1 |
thromboxane A synthase 1 (platelet) |
| chr6_+_47666275 | 0.15 |
ENST00000327753.3 ENST00000283303.2 |
GPR115 |
G protein-coupled receptor 115 |
| chr6_-_27775694 | 0.15 |
ENST00000377401.2 |
HIST1H2BL |
histone cluster 1, H2bl |
| chr16_+_618837 | 0.15 |
ENST00000409439.2 |
PIGQ |
phosphatidylinositol glycan anchor biosynthesis, class Q |
| chr1_-_9189229 | 0.15 |
ENST00000377411.4 |
GPR157 |
G protein-coupled receptor 157 |
| chr7_+_139528952 | 0.14 |
ENST00000416849.2 ENST00000436047.2 ENST00000414508.2 ENST00000448866.1 |
TBXAS1 |
thromboxane A synthase 1 (platelet) |
| chr20_-_43280361 | 0.14 |
ENST00000372874.4 |
ADA |
adenosine deaminase |
| chr17_-_73874654 | 0.14 |
ENST00000254816.2 |
TRIM47 |
tripartite motif containing 47 |
| chr15_+_81591757 | 0.14 |
ENST00000558332.1 |
IL16 |
interleukin 16 |
| chr17_-_43487741 | 0.13 |
ENST00000455881.1 |
ARHGAP27 |
Rho GTPase activating protein 27 |
| chr3_-_112565703 | 0.13 |
ENST00000488794.1 |
CD200R1L |
CD200 receptor 1-like |
| chr7_-_142247606 | 0.13 |
ENST00000390361.3 |
TRBV7-3 |
T cell receptor beta variable 7-3 |
| chr2_+_75061108 | 0.13 |
ENST00000290573.2 |
HK2 |
hexokinase 2 |
| chr6_-_66417107 | 0.13 |
ENST00000370621.3 ENST00000370618.3 ENST00000503581.1 ENST00000393380.2 |
EYS |
eyes shut homolog (Drosophila) |
| chr6_-_26216872 | 0.12 |
ENST00000244601.3 |
HIST1H2BG |
histone cluster 1, H2bg |
| chr19_-_36304201 | 0.12 |
ENST00000301175.3 |
PRODH2 |
proline dehydrogenase (oxidase) 2 |
| chr20_-_43280325 | 0.11 |
ENST00000537820.1 |
ADA |
adenosine deaminase |
| chr3_+_125687987 | 0.11 |
ENST00000514116.1 ENST00000251776.4 ENST00000504401.1 ENST00000513830.1 ENST00000508088.1 |
ROPN1B |
rhophilin associated tail protein 1B |
| chr7_-_19813192 | 0.11 |
ENST00000422233.1 ENST00000433641.1 |
TMEM196 |
transmembrane protein 196 |
| chr18_+_55862622 | 0.10 |
ENST00000456173.2 |
NEDD4L |
neural precursor cell expressed, developmentally down-regulated 4-like, E3 ubiquitin protein ligase |
| chr11_+_60869867 | 0.10 |
ENST00000347785.3 |
CD5 |
CD5 molecule |
| chr5_+_55147205 | 0.10 |
ENST00000396836.2 ENST00000396834.1 ENST00000447346.2 ENST00000359040.5 |
IL31RA |
interleukin 31 receptor A |
| chr14_+_91580777 | 0.10 |
ENST00000525393.2 ENST00000428926.2 ENST00000517362.1 |
C14orf159 |
chromosome 14 open reading frame 159 |
| chr17_-_39165366 | 0.10 |
ENST00000391588.1 |
KRTAP3-1 |
keratin associated protein 3-1 |
| chr14_+_91580708 | 0.10 |
ENST00000518868.1 |
C14orf159 |
chromosome 14 open reading frame 159 |
| chr20_+_44036900 | 0.10 |
ENST00000443296.1 |
DBNDD2 |
dysbindin (dystrobrevin binding protein 1) domain containing 2 |
| chr14_+_91581011 | 0.10 |
ENST00000523894.1 ENST00000522322.1 ENST00000523771.1 |
C14orf159 |
chromosome 14 open reading frame 159 |
| chr2_-_105030466 | 0.10 |
ENST00000449772.1 |
AC068535.3 |
AC068535.3 |
| chr19_-_42724261 | 0.10 |
ENST00000595337.1 |
DEDD2 |
death effector domain containing 2 |
| chr12_+_65563329 | 0.09 |
ENST00000308330.2 |
LEMD3 |
LEM domain containing 3 |
| chr6_+_26273144 | 0.09 |
ENST00000377733.2 |
HIST1H2BI |
histone cluster 1, H2bi |
| chr6_-_26124138 | 0.09 |
ENST00000314332.5 ENST00000396984.1 |
HIST1H2BC |
histone cluster 1, H2bc |
| chr4_-_157563459 | 0.08 |
ENST00000504237.1 ENST00000511399.1 |
RP11-171N4.2 |
Uncharacterized protein |
| chr1_+_149804218 | 0.08 |
ENST00000610125.1 |
HIST2H4A |
histone cluster 2, H4a |
| chr20_+_44036620 | 0.08 |
ENST00000372710.3 |
DBNDD2 |
dysbindin (dystrobrevin binding protein 1) domain containing 2 |
| chr17_-_39093672 | 0.08 |
ENST00000209718.3 ENST00000436344.3 ENST00000485751.1 |
KRT23 |
keratin 23 (histone deacetylase inducible) |
| chr7_+_39125365 | 0.08 |
ENST00000559001.1 ENST00000464276.2 |
POU6F2 |
POU class 6 homeobox 2 |
| chr4_-_38666430 | 0.07 |
ENST00000436901.1 |
AC021860.1 |
Uncharacterized protein |
| chr1_-_226129083 | 0.07 |
ENST00000420304.2 |
LEFTY2 |
left-right determination factor 2 |
| chr19_-_46148820 | 0.07 |
ENST00000587152.1 |
EML2 |
echinoderm microtubule associated protein like 2 |
| chr1_-_44482979 | 0.07 |
ENST00000360584.2 ENST00000357730.2 ENST00000528803.1 |
SLC6A9 |
solute carrier family 6 (neurotransmitter transporter, glycine), member 9 |
| chr2_-_208031943 | 0.07 |
ENST00000421199.1 ENST00000457962.1 |
KLF7 |
Kruppel-like factor 7 (ubiquitous) |
| chr11_-_133402410 | 0.07 |
ENST00000524381.1 |
OPCML |
opioid binding protein/cell adhesion molecule-like |
| chr6_+_27775899 | 0.07 |
ENST00000358739.3 |
HIST1H2AI |
histone cluster 1, H2ai |
| chr6_+_27806319 | 0.07 |
ENST00000606613.1 ENST00000396980.3 |
HIST1H2BN |
histone cluster 1, H2bn |
| chr3_+_51741072 | 0.07 |
ENST00000395052.3 |
GRM2 |
glutamate receptor, metabotropic 2 |
| chr15_+_22736484 | 0.07 |
ENST00000560659.2 |
GOLGA6L1 |
golgin A6 family-like 1 |
| chr18_+_44526786 | 0.07 |
ENST00000245121.5 ENST00000356157.7 |
KATNAL2 |
katanin p60 subunit A-like 2 |
| chr6_-_27782548 | 0.07 |
ENST00000333151.3 |
HIST1H2AJ |
histone cluster 1, H2aj |
| chr10_+_129796390 | 0.06 |
ENST00000455661.1 |
PTPRE |
protein tyrosine phosphatase, receptor type, E |
| chr15_+_22736246 | 0.06 |
ENST00000316397.3 |
GOLGA6L1 |
golgin A6 family-like 1 |
| chr14_-_65409502 | 0.06 |
ENST00000389614.5 |
GPX2 |
glutathione peroxidase 2 (gastrointestinal) |
| chr9_+_136399929 | 0.06 |
ENST00000393060.1 |
ADAMTSL2 |
ADAMTS-like 2 |
| chr6_+_139349817 | 0.06 |
ENST00000367660.3 |
ABRACL |
ABRA C-terminal like |
| chr6_+_27782788 | 0.06 |
ENST00000359465.4 |
HIST1H2BM |
histone cluster 1, H2bm |
| chr11_+_72983246 | 0.06 |
ENST00000393590.2 |
P2RY6 |
pyrimidinergic receptor P2Y, G-protein coupled, 6 |
| chr16_-_58328923 | 0.06 |
ENST00000567164.1 ENST00000219301.4 ENST00000569727.1 |
PRSS54 |
protease, serine, 54 |
| chr6_+_26217159 | 0.06 |
ENST00000303910.2 |
HIST1H2AE |
histone cluster 1, H2ae |
| chr11_-_107729887 | 0.06 |
ENST00000525815.1 |
SLC35F2 |
solute carrier family 35, member F2 |
| chr7_+_50348268 | 0.06 |
ENST00000438033.1 ENST00000439701.1 |
IKZF1 |
IKAROS family zinc finger 1 (Ikaros) |
| chr18_+_43753974 | 0.05 |
ENST00000282059.6 ENST00000321319.6 |
C18orf25 |
chromosome 18 open reading frame 25 |
| chr1_-_226129189 | 0.05 |
ENST00000366820.5 |
LEFTY2 |
left-right determination factor 2 |
| chr14_+_91580732 | 0.05 |
ENST00000519019.1 ENST00000523816.1 ENST00000517518.1 |
C14orf159 |
chromosome 14 open reading frame 159 |
| chr16_-_58328870 | 0.05 |
ENST00000543437.1 |
PRSS54 |
protease, serine, 54 |
| chr19_+_51728316 | 0.05 |
ENST00000436584.2 ENST00000421133.2 ENST00000391796.3 ENST00000262262.4 |
CD33 |
CD33 molecule |
| chr17_+_36452989 | 0.05 |
ENST00000312513.5 ENST00000582535.1 |
MRPL45 |
mitochondrial ribosomal protein L45 |
| chr20_-_60795296 | 0.05 |
ENST00000340177.5 |
HRH3 |
histamine receptor H3 |
| chr10_+_94451574 | 0.05 |
ENST00000492654.2 |
HHEX |
hematopoietically expressed homeobox |
| chr20_+_24449821 | 0.05 |
ENST00000376862.3 |
SYNDIG1 |
synapse differentiation inducing 1 |
| chr19_-_35166604 | 0.05 |
ENST00000601241.1 |
SCGB2B2 |
secretoglobin, family 2B, member 2 |
| chr12_-_323689 | 0.05 |
ENST00000428720.1 |
SLC6A12 |
solute carrier family 6 (neurotransmitter transporter), member 12 |
| chr6_-_27841289 | 0.05 |
ENST00000355981.2 |
HIST1H4L |
histone cluster 1, H4l |
| chr16_-_11036300 | 0.05 |
ENST00000331808.4 |
DEXI |
Dexi homolog (mouse) |
| chr15_+_63796779 | 0.05 |
ENST00000561442.1 ENST00000560070.1 ENST00000540797.1 ENST00000380324.3 ENST00000268049.7 ENST00000536001.1 ENST00000539772.1 |
USP3 |
ubiquitin specific peptidase 3 |
| chr2_+_28615669 | 0.04 |
ENST00000379619.1 ENST00000264716.4 |
FOSL2 |
FOS-like antigen 2 |
| chr2_+_220110177 | 0.04 |
ENST00000409638.3 ENST00000396738.2 ENST00000409516.3 |
STK16 |
serine/threonine kinase 16 |
| chrX_-_153523462 | 0.04 |
ENST00000361930.3 ENST00000369926.1 |
TEX28 |
testis expressed 28 |
| chr11_+_46383121 | 0.04 |
ENST00000454345.1 |
DGKZ |
diacylglycerol kinase, zeta |
| chr14_+_91580357 | 0.04 |
ENST00000298858.4 ENST00000521081.1 ENST00000520328.1 ENST00000256324.10 ENST00000524232.1 ENST00000522170.1 ENST00000519950.1 ENST00000523879.1 ENST00000521077.2 ENST00000518665.2 |
C14orf159 |
chromosome 14 open reading frame 159 |
| chr1_-_156399184 | 0.04 |
ENST00000368243.1 ENST00000357975.4 ENST00000310027.5 ENST00000400991.2 |
C1orf61 |
chromosome 1 open reading frame 61 |
| chr3_+_97868170 | 0.04 |
ENST00000437310.1 |
OR5H14 |
olfactory receptor, family 5, subfamily H, member 14 |
| chr9_+_37485932 | 0.04 |
ENST00000377798.4 ENST00000442009.2 |
POLR1E |
polymerase (RNA) I polypeptide E, 53kDa |
| chr1_-_89357179 | 0.04 |
ENST00000448623.1 ENST00000418217.1 ENST00000370500.5 |
GTF2B |
general transcription factor IIB |
| chr2_-_239140011 | 0.04 |
ENST00000409376.1 ENST00000409070.1 ENST00000409942.1 |
AC016757.3 |
Protein LOC151174 |
| chr14_-_65409438 | 0.04 |
ENST00000557049.1 |
GPX2 |
glutathione peroxidase 2 (gastrointestinal) |
| chr7_+_142364193 | 0.04 |
ENST00000390397.2 |
TRBV24-1 |
T cell receptor beta variable 24-1 |
| chr1_+_66797687 | 0.04 |
ENST00000371045.5 ENST00000531025.1 ENST00000526197.1 |
PDE4B |
phosphodiesterase 4B, cAMP-specific |
| chr2_+_96991033 | 0.03 |
ENST00000420176.1 ENST00000536814.1 ENST00000439118.2 |
ITPRIPL1 |
inositol 1,4,5-trisphosphate receptor interacting protein-like 1 |
| chr1_+_90287480 | 0.03 |
ENST00000394593.3 |
LRRC8D |
leucine rich repeat containing 8 family, member D |
| chr2_-_49381572 | 0.03 |
ENST00000454032.1 ENST00000304421.4 |
FSHR |
follicle stimulating hormone receptor |
| chr11_-_72145426 | 0.03 |
ENST00000535990.1 ENST00000437826.2 ENST00000340729.5 |
CLPB |
ClpB caseinolytic peptidase B homolog (E. coli) |
| chr6_-_99842041 | 0.03 |
ENST00000254759.3 ENST00000369242.1 |
COQ3 |
coenzyme Q3 methyltransferase |
| chr2_-_49381646 | 0.03 |
ENST00000346173.3 ENST00000406846.2 |
FSHR |
follicle stimulating hormone receptor |
| chr2_+_16080659 | 0.03 |
ENST00000281043.3 |
MYCN |
v-myc avian myelocytomatosis viral oncogene neuroblastoma derived homolog |
| chr9_+_37486005 | 0.03 |
ENST00000377792.3 |
POLR1E |
polymerase (RNA) I polypeptide E, 53kDa |
| chr6_+_41888926 | 0.03 |
ENST00000230340.4 |
BYSL |
bystin-like |
| chr12_+_67663056 | 0.03 |
ENST00000545606.1 |
CAND1 |
cullin-associated and neddylation-dissociated 1 |
| chr11_+_66406088 | 0.03 |
ENST00000310092.7 ENST00000396053.4 ENST00000408993.2 |
RBM4 |
RNA binding motif protein 4 |
| chr19_+_41594377 | 0.03 |
ENST00000330436.3 |
CYP2A13 |
cytochrome P450, family 2, subfamily A, polypeptide 13 |
| chr16_-_30773372 | 0.03 |
ENST00000545825.1 ENST00000541260.1 |
C16orf93 |
chromosome 16 open reading frame 93 |
| chr2_-_71062938 | 0.02 |
ENST00000410009.3 |
CD207 |
CD207 molecule, langerin |
| chr16_-_15180257 | 0.02 |
ENST00000540462.1 |
RRN3 |
RRN3 RNA polymerase I transcription factor homolog (S. cerevisiae) |
| chr1_-_11042094 | 0.02 |
ENST00000377004.4 ENST00000377008.4 |
C1orf127 |
chromosome 1 open reading frame 127 |
| chr11_+_35211511 | 0.02 |
ENST00000524922.1 |
CD44 |
CD44 molecule (Indian blood group) |
| chr8_-_80942061 | 0.02 |
ENST00000519386.1 |
MRPS28 |
mitochondrial ribosomal protein S28 |
| chr9_+_21967137 | 0.02 |
ENST00000441769.2 |
C9orf53 |
chromosome 9 open reading frame 53 |
| chr9_-_14910990 | 0.02 |
ENST00000380881.4 ENST00000422223.2 |
FREM1 |
FRAS1 related extracellular matrix 1 |
| chr7_-_22862448 | 0.02 |
ENST00000358435.4 |
TOMM7 |
translocase of outer mitochondrial membrane 7 homolog (yeast) |
| chr12_+_50135327 | 0.02 |
ENST00000549966.1 ENST00000547832.1 ENST00000547187.1 ENST00000548894.1 ENST00000546914.1 ENST00000552699.1 ENST00000267115.5 |
TMBIM6 |
transmembrane BAX inhibitor motif containing 6 |
| chrX_+_49832231 | 0.02 |
ENST00000376108.3 |
CLCN5 |
chloride channel, voltage-sensitive 5 |
| chr3_+_12838161 | 0.02 |
ENST00000456430.2 |
CAND2 |
cullin-associated and neddylation-dissociated 2 (putative) |
| chr10_-_102046417 | 0.02 |
ENST00000370372.2 |
BLOC1S2 |
biogenesis of lysosomal organelles complex-1, subunit 2 |
| chr2_-_239140276 | 0.02 |
ENST00000334973.4 |
AC016757.3 |
Protein LOC151174 |
| chr12_+_50135351 | 0.02 |
ENST00000549445.1 ENST00000550951.1 ENST00000549385.1 ENST00000548713.1 ENST00000548201.1 |
TMBIM6 |
transmembrane BAX inhibitor motif containing 6 |
| chr16_+_30483962 | 0.02 |
ENST00000356798.6 |
ITGAL |
integrin, alpha L (antigen CD11A (p180), lymphocyte function-associated antigen 1; alpha polypeptide) |
| chr16_-_3306587 | 0.02 |
ENST00000541159.1 ENST00000536379.1 ENST00000219596.1 ENST00000339854.4 |
MEFV |
Mediterranean fever |
| chr8_-_80942139 | 0.02 |
ENST00000521434.1 ENST00000519120.1 ENST00000520946.1 |
MRPS28 |
mitochondrial ribosomal protein S28 |
| chr7_-_22862406 | 0.01 |
ENST00000372879.4 |
TOMM7 |
translocase of outer mitochondrial membrane 7 homolog (yeast) |
| chr16_+_30484021 | 0.01 |
ENST00000358164.5 |
ITGAL |
integrin, alpha L (antigen CD11A (p180), lymphocyte function-associated antigen 1; alpha polypeptide) |
| chr1_-_9129631 | 0.01 |
ENST00000377414.3 |
SLC2A5 |
solute carrier family 2 (facilitated glucose/fructose transporter), member 5 |
| chr19_-_55691377 | 0.01 |
ENST00000589172.1 |
SYT5 |
synaptotagmin V |
| chr6_+_123038689 | 0.01 |
ENST00000354275.2 ENST00000368446.1 |
PKIB |
protein kinase (cAMP-dependent, catalytic) inhibitor beta |
| chr1_+_202385953 | 0.01 |
ENST00000466968.1 |
PPP1R12B |
protein phosphatase 1, regulatory subunit 12B |
| chr3_-_123710893 | 0.01 |
ENST00000467907.1 ENST00000459660.1 ENST00000495093.1 ENST00000460743.1 ENST00000405845.3 ENST00000484329.1 ENST00000479867.1 ENST00000496145.1 |
ROPN1 |
rhophilin associated tail protein 1 |
| chr6_+_146920097 | 0.01 |
ENST00000397944.3 ENST00000522242.1 |
ADGB |
androglobin |
| chr6_+_146920116 | 0.01 |
ENST00000367493.3 |
ADGB |
androglobin |
| chr6_-_27806117 | 0.01 |
ENST00000330180.2 |
HIST1H2AK |
histone cluster 1, H2ak |
| chr9_+_114287433 | 0.01 |
ENST00000358151.4 ENST00000355824.3 ENST00000374374.3 ENST00000309235.5 |
ZNF483 |
zinc finger protein 483 |
| chr6_+_27114861 | 0.01 |
ENST00000377459.1 |
HIST1H2AH |
histone cluster 1, H2ah |
| chr14_+_102786063 | 0.01 |
ENST00000442396.2 ENST00000262236.5 |
ZNF839 |
zinc finger protein 839 |
| chr16_-_84220604 | 0.01 |
ENST00000567759.1 |
TAF1C |
TATA box binding protein (TBP)-associated factor, RNA polymerase I, C, 110kDa |
| chr22_-_46644182 | 0.01 |
ENST00000404583.1 ENST00000404744.1 |
CDPF1 |
cysteine-rich, DPF motif domain containing 1 |
| chr6_-_46922659 | 0.01 |
ENST00000265417.7 |
GPR116 |
G protein-coupled receptor 116 |
| chr11_+_34938119 | 0.01 |
ENST00000227868.4 ENST00000430469.2 ENST00000533262.1 |
PDHX |
pyruvate dehydrogenase complex, component X |
| chr8_-_101117847 | 0.01 |
ENST00000523287.1 ENST00000519092.1 |
RGS22 |
regulator of G-protein signaling 22 |
| chr1_-_9129598 | 0.01 |
ENST00000535586.1 |
SLC2A5 |
solute carrier family 2 (facilitated glucose/fructose transporter), member 5 |
| chr3_+_14219858 | 0.01 |
ENST00000306024.3 |
LSM3 |
LSM3 homolog, U6 small nuclear RNA associated (S. cerevisiae) |
| chr8_-_101118210 | 0.01 |
ENST00000360863.6 |
RGS22 |
regulator of G-protein signaling 22 |
| chr8_-_101118185 | 0.01 |
ENST00000523437.1 |
RGS22 |
regulator of G-protein signaling 22 |
| chr7_+_141490017 | 0.01 |
ENST00000247883.4 |
TAS2R5 |
taste receptor, type 2, member 5 |
| chr6_+_27861190 | 0.01 |
ENST00000303806.4 |
HIST1H2BO |
histone cluster 1, H2bo |
| chr6_-_26285737 | 0.00 |
ENST00000377727.1 ENST00000289352.1 |
HIST1H4H |
histone cluster 1, H4h |
| chr15_-_26874230 | 0.00 |
ENST00000400188.3 |
GABRB3 |
gamma-aminobutyric acid (GABA) A receptor, beta 3 |
| chr16_+_30773636 | 0.00 |
ENST00000402121.3 ENST00000565995.1 ENST00000563683.1 ENST00000357890.5 ENST00000565931.1 |
RNF40 |
ring finger protein 40, E3 ubiquitin protein ligase |
| chr16_-_84220633 | 0.00 |
ENST00000566732.1 ENST00000561955.1 ENST00000564454.1 ENST00000341690.6 ENST00000541676.1 ENST00000570117.1 ENST00000564345.1 |
TAF1C |
TATA box binding protein (TBP)-associated factor, RNA polymerase I, C, 110kDa |
| chr7_+_133261209 | 0.00 |
ENST00000545148.1 |
EXOC4 |
exocyst complex component 4 |
| chr6_+_116937636 | 0.00 |
ENST00000368581.4 ENST00000229554.5 ENST00000368580.4 |
RSPH4A |
radial spoke head 4 homolog A (Chlamydomonas) |
| chr14_-_24551137 | 0.00 |
ENST00000396995.1 |
NRL |
neural retina leucine zipper |
| chr18_+_13612613 | 0.00 |
ENST00000586765.1 |
LDLRAD4 |
low density lipoprotein receptor class A domain containing 4 |
| chr15_-_83316254 | 0.00 |
ENST00000567678.1 ENST00000450751.2 |
CPEB1 |
cytoplasmic polyadenylation element binding protein 1 |
| chr16_-_20681177 | 0.00 |
ENST00000524149.1 |
ACSM1 |
acyl-CoA synthetase medium-chain family member 1 |
| chr6_+_27100811 | 0.00 |
ENST00000359193.2 |
HIST1H2AG |
histone cluster 1, H2ag |
| chr6_+_292051 | 0.00 |
ENST00000344450.5 |
DUSP22 |
dual specificity phosphatase 22 |
| chr2_+_204732487 | 0.00 |
ENST00000302823.3 |
CTLA4 |
cytotoxic T-lymphocyte-associated protein 4 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.1 | 0.4 | GO:0051620 | negative regulation of dopamine uptake involved in synaptic transmission(GO:0051585) norepinephrine uptake(GO:0051620) regulation of norepinephrine uptake(GO:0051621) negative regulation of norepinephrine uptake(GO:0051622) negative regulation of catecholamine uptake involved in synaptic transmission(GO:0051945) regulation of glutathione peroxidase activity(GO:1903282) positive regulation of glutathione peroxidase activity(GO:1903284) positive regulation of hydrogen peroxide catabolic process(GO:1903285) positive regulation of peroxidase activity(GO:2000470) |
| 0.1 | 0.3 | GO:0046103 | adenosine catabolic process(GO:0006154) inosine biosynthetic process(GO:0046103) |
| 0.1 | 0.6 | GO:0032926 | negative regulation of activin receptor signaling pathway(GO:0032926) |
| 0.1 | 0.2 | GO:0021718 | superior olivary nucleus development(GO:0021718) superior olivary nucleus maturation(GO:0021722) |
| 0.0 | 0.2 | GO:0048817 | negative regulation of hair follicle maturation(GO:0048817) |
| 0.0 | 0.5 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 0.0 | 0.1 | GO:0061536 | glycine secretion(GO:0061536) glycine secretion, neurotransmission(GO:0061537) |
| 0.0 | 0.2 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.0 | 0.1 | GO:2001288 | positive regulation of caveolin-mediated endocytosis(GO:2001288) |
| 0.0 | 0.2 | GO:0039536 | negative regulation of RIG-I signaling pathway(GO:0039536) |
| 0.0 | 0.1 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 0.0 | 0.1 | GO:0007402 | ganglion mother cell fate determination(GO:0007402) |
| 0.0 | 0.1 | GO:1904925 | negative regulation of mitochondrial membrane permeability(GO:0035795) positive regulation of macromitophagy(GO:1901526) positive regulation of mitophagy in response to mitochondrial depolarization(GO:1904925) |
| 0.0 | 0.1 | GO:0001188 | RNA polymerase I transcriptional preinitiation complex assembly(GO:0001188) RNA polymerase I transcriptional preinitiation complex assembly at the promoter for the nuclear large rRNA transcript(GO:0001189) |
| 0.0 | 0.3 | GO:0019371 | cyclooxygenase pathway(GO:0019371) |
| 0.0 | 0.0 | GO:0061010 | negative regulation of transcription by transcription factor localization(GO:0010621) gall bladder development(GO:0061010) |
| 0.0 | 0.2 | GO:0051918 | negative regulation of fibrinolysis(GO:0051918) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.0 | 0.2 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.0 | 1.0 | GO:0000786 | nucleosome(GO:0000786) |
| 0.0 | 0.4 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.0 | 0.2 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0036134 | thromboxane-A synthase activity(GO:0004796) 12-hydroxyheptadecatrienoic acid synthase activity(GO:0036134) |
| 0.1 | 0.4 | GO:0060961 | phospholipase D inhibitor activity(GO:0060961) |
| 0.1 | 0.4 | GO:0047820 | D-glutamate cyclase activity(GO:0047820) |
| 0.1 | 0.2 | GO:0016534 | cyclin-dependent protein kinase 5 activator activity(GO:0016534) |
| 0.0 | 0.2 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 0.0 | 0.6 | GO:0008199 | ferric iron binding(GO:0008199) |
| 0.0 | 0.1 | GO:0004657 | proline dehydrogenase activity(GO:0004657) |
| 0.0 | 0.5 | GO:0048185 | activin binding(GO:0048185) |
| 0.0 | 0.1 | GO:0015375 | glycine:sodium symporter activity(GO:0015375) |
| 0.0 | 0.6 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 0.1 | GO:0019158 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.0 | 0.3 | GO:0004000 | adenosine deaminase activity(GO:0004000) |
| 0.0 | 0.1 | GO:0001641 | group II metabotropic glutamate receptor activity(GO:0001641) |
| 0.0 | 0.2 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.0 | 0.1 | GO:0050436 | microfibril binding(GO:0050436) |
| 0.0 | 0.1 | GO:0045029 | UDP-activated nucleotide receptor activity(GO:0045029) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.3 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |