Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
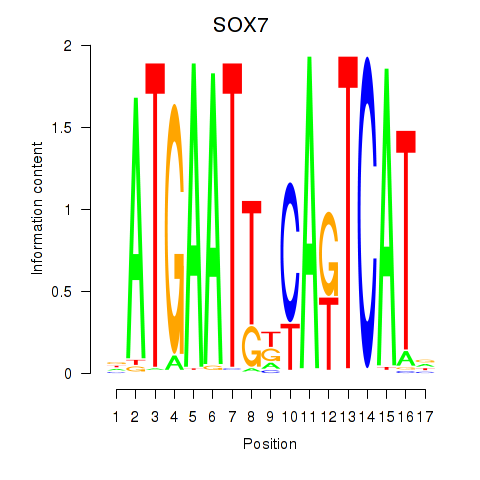
Results for SOX7
Z-value: 0.32
Transcription factors associated with SOX7
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SOX7
|
ENSG00000171056.6 | SOX7 |
|
SOX7
|
ENSG00000258724.1 | SOX7 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SOX7 | hg19_v2_chr8_-_10588010_10588030 | -0.61 | 1.1e-01 | Click! |
Activity profile of SOX7 motif
Sorted Z-values of SOX7 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of SOX7
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_223101757 | 0.22 |
ENST00000284476.6 |
DISP1 |
dispatched homolog 1 (Drosophila) |
| chr17_-_66951474 | 0.22 |
ENST00000269080.2 |
ABCA8 |
ATP-binding cassette, sub-family A (ABC1), member 8 |
| chr5_-_95158644 | 0.14 |
ENST00000237858.6 |
GLRX |
glutaredoxin (thioltransferase) |
| chr12_-_90049828 | 0.13 |
ENST00000261173.2 ENST00000348959.3 |
ATP2B1 |
ATPase, Ca++ transporting, plasma membrane 1 |
| chr12_-_90049878 | 0.13 |
ENST00000359142.3 |
ATP2B1 |
ATPase, Ca++ transporting, plasma membrane 1 |
| chr5_-_95158375 | 0.12 |
ENST00000512469.2 ENST00000379979.4 ENST00000505427.1 ENST00000508780.1 |
GLRX |
glutaredoxin (thioltransferase) |
| chr8_+_77593448 | 0.12 |
ENST00000521891.2 |
ZFHX4 |
zinc finger homeobox 4 |
| chr12_-_58027138 | 0.08 |
ENST00000341156.4 |
B4GALNT1 |
beta-1,4-N-acetyl-galactosaminyl transferase 1 |
| chr17_-_39623681 | 0.08 |
ENST00000225899.3 |
KRT32 |
keratin 32 |
| chr17_-_56082455 | 0.08 |
ENST00000578794.1 |
RP11-159D12.5 |
Uncharacterized protein |
| chr5_-_176778803 | 0.07 |
ENST00000303127.7 |
LMAN2 |
lectin, mannose-binding 2 |
| chr2_-_190445499 | 0.06 |
ENST00000261024.2 |
SLC40A1 |
solute carrier family 40 (iron-regulated transporter), member 1 |
| chr14_-_106573756 | 0.06 |
ENST00000390601.2 |
IGHV3-11 |
immunoglobulin heavy variable 3-11 (gene/pseudogene) |
| chr11_-_62474803 | 0.06 |
ENST00000533982.1 ENST00000360796.5 |
BSCL2 |
Berardinelli-Seip congenital lipodystrophy 2 (seipin) |
| chr6_+_80129989 | 0.05 |
ENST00000429444.1 |
RP1-232L24.3 |
RP1-232L24.3 |
| chr6_-_111804905 | 0.05 |
ENST00000358835.3 ENST00000435970.1 |
REV3L |
REV3-like, polymerase (DNA directed), zeta, catalytic subunit |
| chr10_-_12084770 | 0.05 |
ENST00000357604.5 |
UPF2 |
UPF2 regulator of nonsense transcripts homolog (yeast) |
| chr2_+_179317994 | 0.05 |
ENST00000375129.4 |
DFNB59 |
deafness, autosomal recessive 59 |
| chr6_+_131894284 | 0.04 |
ENST00000368087.3 ENST00000356962.2 |
ARG1 |
arginase 1 |
| chr6_+_147527103 | 0.04 |
ENST00000179882.6 |
STXBP5 |
syntaxin binding protein 5 (tomosyn) |
| chr19_-_38916839 | 0.04 |
ENST00000433821.2 ENST00000426920.2 ENST00000587753.1 ENST00000454404.2 ENST00000293062.9 |
RASGRP4 |
RAS guanyl releasing protein 4 |
| chr14_-_102771462 | 0.04 |
ENST00000522874.1 |
MOK |
MOK protein kinase |
| chr19_-_38916822 | 0.04 |
ENST00000586305.1 |
RASGRP4 |
RAS guanyl releasing protein 4 |
| chr19_+_37837185 | 0.04 |
ENST00000541583.2 |
HKR1 |
HKR1, GLI-Kruppel zinc finger family member |
| chr19_-_5784610 | 0.04 |
ENST00000390672.2 ENST00000419421.2 |
PRR22 |
proline rich 22 |
| chr2_-_88285309 | 0.04 |
ENST00000420840.2 |
RGPD2 |
RANBP2-like and GRIP domain containing 2 |
| chr10_-_17243579 | 0.04 |
ENST00000525762.1 ENST00000412821.3 ENST00000351358.4 ENST00000377766.5 ENST00000358282.7 ENST00000488990.1 ENST00000377799.3 |
TRDMT1 |
tRNA aspartic acid methyltransferase 1 |
| chrY_+_26997726 | 0.03 |
ENST00000382296.2 |
DAZ4 |
deleted in azoospermia 4 |
| chr1_+_199996733 | 0.03 |
ENST00000236914.3 |
NR5A2 |
nuclear receptor subfamily 5, group A, member 2 |
| chr4_-_39034542 | 0.03 |
ENST00000344606.6 |
TMEM156 |
transmembrane protein 156 |
| chr2_+_87144738 | 0.03 |
ENST00000559485.1 |
RGPD1 |
RANBP2-like and GRIP domain containing 1 |
| chr19_+_42349092 | 0.03 |
ENST00000269945.3 ENST00000596258.1 |
DMRTC2 |
DMRT-like family C2 |
| chr21_-_21630994 | 0.03 |
ENST00000436373.1 |
AP001171.1 |
AP001171.1 |
| chr19_-_22379753 | 0.03 |
ENST00000397121.2 |
ZNF676 |
zinc finger protein 676 |
| chr18_+_32820990 | 0.03 |
ENST00000601719.1 ENST00000591206.1 ENST00000330501.7 ENST00000261333.6 ENST00000355632.4 ENST00000585800.1 |
ZNF397 |
zinc finger protein 397 |
| chr16_-_28303360 | 0.03 |
ENST00000501520.1 |
RP11-57A19.2 |
RP11-57A19.2 |
| chr5_+_175511859 | 0.02 |
ENST00000503724.2 ENST00000253490.4 |
FAM153B |
family with sequence similarity 153, member B |
| chr12_-_57443886 | 0.02 |
ENST00000300119.3 |
MYO1A |
myosin IA |
| chr16_-_279405 | 0.02 |
ENST00000430864.1 ENST00000293872.8 ENST00000337351.4 ENST00000397783.1 |
LUC7L |
LUC7-like (S. cerevisiae) |
| chr16_+_28303804 | 0.02 |
ENST00000341901.4 |
SBK1 |
SH3 domain binding kinase 1 |
| chr19_-_38916802 | 0.02 |
ENST00000587738.1 |
RASGRP4 |
RAS guanyl releasing protein 4 |
| chr1_+_158901329 | 0.02 |
ENST00000368140.1 ENST00000368138.3 ENST00000392254.2 ENST00000392252.3 ENST00000368135.4 |
PYHIN1 |
pyrin and HIN domain family, member 1 |
| chr6_+_150690133 | 0.02 |
ENST00000392255.3 ENST00000500320.3 |
IYD |
iodotyrosine deiodinase |
| chr10_+_122610687 | 0.02 |
ENST00000263461.6 |
WDR11 |
WD repeat domain 11 |
| chr12_+_64173583 | 0.02 |
ENST00000261234.6 |
TMEM5 |
transmembrane protein 5 |
| chr13_-_28024681 | 0.01 |
ENST00000381116.1 ENST00000381120.3 ENST00000431572.2 |
MTIF3 |
mitochondrial translational initiation factor 3 |
| chr1_+_100598691 | 0.01 |
ENST00000370143.1 ENST00000370141.2 |
TRMT13 |
tRNA methyltransferase 13 homolog (S. cerevisiae) |
| chr15_+_63414760 | 0.01 |
ENST00000557972.1 |
LACTB |
lactamase, beta |
| chr9_-_70465758 | 0.01 |
ENST00000489273.1 |
CBWD5 |
COBW domain containing 5 |
| chr10_+_5005445 | 0.01 |
ENST00000380872.4 |
AKR1C1 |
aldo-keto reductase family 1, member C1 |
| chr6_+_140175987 | 0.01 |
ENST00000414038.1 ENST00000431609.1 |
RP5-899B16.1 |
RP5-899B16.1 |
| chr21_-_15918618 | 0.01 |
ENST00000400564.1 ENST00000400566.1 |
SAMSN1 |
SAM domain, SH3 domain and nuclear localization signals 1 |
| chr8_+_12803176 | 0.01 |
ENST00000524591.2 |
KIAA1456 |
KIAA1456 |
| chr8_-_19459993 | 0.01 |
ENST00000454498.2 ENST00000520003.1 |
CSGALNACT1 |
chondroitin sulfate N-acetylgalactosaminyltransferase 1 |
| chr1_+_10509971 | 0.01 |
ENST00000320498.4 |
CORT |
cortistatin |
| chr9_+_105757590 | 0.01 |
ENST00000374798.3 ENST00000487798.1 |
CYLC2 |
cylicin, basic protein of sperm head cytoskeleton 2 |
| chr6_-_86303833 | 0.01 |
ENST00000505648.1 |
SNX14 |
sorting nexin 14 |
| chr19_-_2783363 | 0.01 |
ENST00000221566.2 |
SGTA |
small glutamine-rich tetratricopeptide repeat (TPR)-containing, alpha |
| chr11_+_55944094 | 0.01 |
ENST00000312298.1 |
OR5J2 |
olfactory receptor, family 5, subfamily J, member 2 |
| chr13_-_37633567 | 0.01 |
ENST00000464744.1 |
SUPT20H |
suppressor of Ty 20 homolog (S. cerevisiae) |
| chr19_-_21950332 | 0.01 |
ENST00000598026.1 |
ZNF100 |
zinc finger protein 100 |
| chr6_-_46048116 | 0.00 |
ENST00000185206.6 |
CLIC5 |
chloride intracellular channel 5 |
| chr11_+_3011093 | 0.00 |
ENST00000332881.2 |
AC131971.1 |
HCG1782999; PRO0943; Uncharacterized protein |
| chr6_+_42531798 | 0.00 |
ENST00000372903.2 ENST00000372899.1 ENST00000372901.1 |
UBR2 |
ubiquitin protein ligase E3 component n-recognin 2 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | GO:1990034 | cellular response to corticosterone stimulus(GO:0071386) calcium ion export from cell(GO:1990034) |
| 0.0 | 0.1 | GO:0070839 | divalent metal ion export(GO:0070839) |
| 0.0 | 0.2 | GO:0042908 | xenobiotic transport(GO:0042908) |
| 0.0 | 0.0 | GO:0043091 | L-arginine import(GO:0043091) arginine import(GO:0090467) |
| 0.0 | 0.3 | GO:0045838 | positive regulation of membrane potential(GO:0045838) |
| 0.0 | 0.2 | GO:0060539 | diaphragm development(GO:0060539) |
| 0.0 | 0.1 | GO:0030259 | lipid glycosylation(GO:0030259) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | GO:0032591 | dendritic spine membrane(GO:0032591) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0003947 | (N-acetylneuraminyl)-galactosylglucosylceramide N-acetylgalactosaminyltransferase activity(GO:0003947) |
| 0.0 | 0.2 | GO:0008559 | xenobiotic-transporting ATPase activity(GO:0008559) |
| 0.0 | 0.3 | GO:0015038 | glutathione disulfide oxidoreductase activity(GO:0015038) |
| 0.0 | 0.2 | GO:0015197 | peptide transporter activity(GO:0015197) |