Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
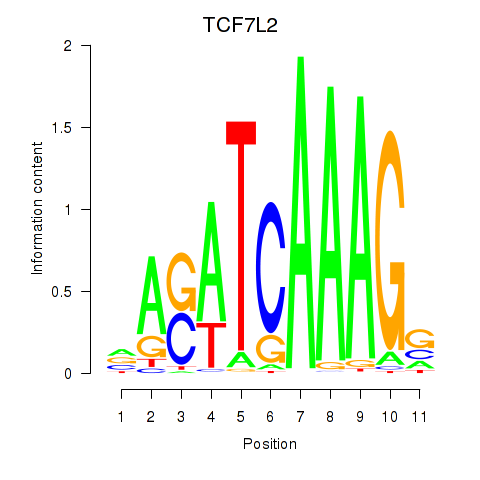
Results for TCF7L2
Z-value: 0.41
Transcription factors associated with TCF7L2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
TCF7L2
|
ENSG00000148737.11 | TCF7L2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| TCF7L2 | hg19_v2_chr10_+_114709999_114710031 | -0.83 | 1.1e-02 | Click! |
Activity profile of TCF7L2 motif
Sorted Z-values of TCF7L2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of TCF7L2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr13_+_102142296 | 0.74 |
ENST00000376162.3 |
ITGBL1 |
integrin, beta-like 1 (with EGF-like repeat domains) |
| chr13_+_76334498 | 0.54 |
ENST00000534657.1 |
LMO7 |
LIM domain 7 |
| chr13_+_24144509 | 0.44 |
ENST00000248484.4 |
TNFRSF19 |
tumor necrosis factor receptor superfamily, member 19 |
| chr13_+_24144796 | 0.39 |
ENST00000403372.2 |
TNFRSF19 |
tumor necrosis factor receptor superfamily, member 19 |
| chr12_+_1738363 | 0.37 |
ENST00000397196.2 |
WNT5B |
wingless-type MMTV integration site family, member 5B |
| chr13_+_76334795 | 0.37 |
ENST00000526202.1 ENST00000465261.2 |
LMO7 |
LIM domain 7 |
| chr13_+_76334567 | 0.32 |
ENST00000321797.8 |
LMO7 |
LIM domain 7 |
| chr5_-_111093167 | 0.32 |
ENST00000446294.2 ENST00000419114.2 |
NREP |
neuronal regeneration related protein |
| chr14_-_61124977 | 0.29 |
ENST00000554986.1 |
SIX1 |
SIX homeobox 1 |
| chr11_+_69455855 | 0.28 |
ENST00000227507.2 ENST00000536559.1 |
CCND1 |
cyclin D1 |
| chr1_+_223101757 | 0.28 |
ENST00000284476.6 |
DISP1 |
dispatched homolog 1 (Drosophila) |
| chr2_+_204801471 | 0.27 |
ENST00000316386.6 ENST00000435193.1 |
ICOS |
inducible T-cell co-stimulator |
| chrX_+_152760397 | 0.24 |
ENST00000331595.4 ENST00000431891.1 |
BGN |
biglycan |
| chr4_-_157892167 | 0.23 |
ENST00000541126.1 |
PDGFC |
platelet derived growth factor C |
| chr4_-_157892055 | 0.22 |
ENST00000422544.2 |
PDGFC |
platelet derived growth factor C |
| chr5_-_16742330 | 0.21 |
ENST00000505695.1 ENST00000427430.2 |
MYO10 |
myosin X |
| chr16_-_73082274 | 0.19 |
ENST00000268489.5 |
ZFHX3 |
zinc finger homeobox 3 |
| chr4_-_157892498 | 0.19 |
ENST00000502773.1 |
PDGFC |
platelet derived growth factor C |
| chr13_-_67802549 | 0.19 |
ENST00000328454.5 ENST00000377865.2 |
PCDH9 |
protocadherin 9 |
| chr17_-_67138015 | 0.18 |
ENST00000284425.2 ENST00000590645.1 |
ABCA6 |
ATP-binding cassette, sub-family A (ABC1), member 6 |
| chr6_-_34216766 | 0.18 |
ENST00000481533.1 ENST00000468145.1 ENST00000413013.2 ENST00000394990.4 ENST00000335352.3 |
C6orf1 |
chromosome 6 open reading frame 1 |
| chr10_-_25241499 | 0.17 |
ENST00000376378.1 ENST00000376376.3 ENST00000320152.6 |
PRTFDC1 |
phosphoribosyl transferase domain containing 1 |
| chr1_+_207039154 | 0.17 |
ENST00000367096.3 ENST00000391930.2 |
IL20 |
interleukin 20 |
| chr16_+_53242350 | 0.17 |
ENST00000565442.1 |
CHD9 |
chromodomain helicase DNA binding protein 9 |
| chr5_-_157002775 | 0.16 |
ENST00000257527.4 |
ADAM19 |
ADAM metallopeptidase domain 19 |
| chr1_+_61330931 | 0.15 |
ENST00000371191.1 |
NFIA |
nuclear factor I/A |
| chrX_-_39956656 | 0.15 |
ENST00000397354.3 ENST00000378444.4 |
BCOR |
BCL6 corepressor |
| chrX_-_63005405 | 0.15 |
ENST00000374878.1 ENST00000437457.2 |
ARHGEF9 |
Cdc42 guanine nucleotide exchange factor (GEF) 9 |
| chr5_+_60628074 | 0.15 |
ENST00000252744.5 |
ZSWIM6 |
zinc finger, SWIM-type containing 6 |
| chr2_+_69240511 | 0.15 |
ENST00000409349.3 |
ANTXR1 |
anthrax toxin receptor 1 |
| chr5_+_112074029 | 0.15 |
ENST00000512211.2 |
APC |
adenomatous polyposis coli |
| chr10_+_54074033 | 0.15 |
ENST00000373970.3 |
DKK1 |
dickkopf WNT signaling pathway inhibitor 1 |
| chr15_+_93447675 | 0.14 |
ENST00000536619.1 |
CHD2 |
chromodomain helicase DNA binding protein 2 |
| chr14_-_53258314 | 0.14 |
ENST00000216410.3 ENST00000557604.1 |
GNPNAT1 |
glucosamine-phosphate N-acetyltransferase 1 |
| chr5_-_157002749 | 0.14 |
ENST00000517905.1 ENST00000430702.2 ENST00000394020.1 |
ADAM19 |
ADAM metallopeptidase domain 19 |
| chr17_+_59529743 | 0.13 |
ENST00000589003.1 ENST00000393853.4 |
TBX4 |
T-box 4 |
| chr6_+_132873832 | 0.12 |
ENST00000275200.1 |
TAAR8 |
trace amine associated receptor 8 |
| chrX_+_54835493 | 0.12 |
ENST00000396224.1 |
MAGED2 |
melanoma antigen family D, 2 |
| chr15_-_90294523 | 0.12 |
ENST00000300057.4 |
MESP1 |
mesoderm posterior 1 homolog (mouse) |
| chr6_+_12717892 | 0.12 |
ENST00000379350.1 |
PHACTR1 |
phosphatase and actin regulator 1 |
| chr2_-_55647057 | 0.12 |
ENST00000436346.1 |
CCDC88A |
coiled-coil domain containing 88A |
| chr1_+_78383813 | 0.12 |
ENST00000342754.5 |
NEXN |
nexilin (F actin binding protein) |
| chr2_-_55646957 | 0.12 |
ENST00000263630.8 |
CCDC88A |
coiled-coil domain containing 88A |
| chr3_-_149375783 | 0.12 |
ENST00000467467.1 ENST00000460517.1 ENST00000360632.3 |
WWTR1 |
WW domain containing transcription regulator 1 |
| chr2_+_69240415 | 0.11 |
ENST00000409829.3 |
ANTXR1 |
anthrax toxin receptor 1 |
| chr2_+_69240302 | 0.11 |
ENST00000303714.4 |
ANTXR1 |
anthrax toxin receptor 1 |
| chr3_+_141106643 | 0.11 |
ENST00000514251.1 |
ZBTB38 |
zinc finger and BTB domain containing 38 |
| chr19_-_44008863 | 0.11 |
ENST00000601646.1 |
PHLDB3 |
pleckstrin homology-like domain, family B, member 3 |
| chr5_+_140734570 | 0.11 |
ENST00000571252.1 |
PCDHGA4 |
protocadherin gamma subfamily A, 4 |
| chr22_-_32860427 | 0.11 |
ENST00000534972.1 ENST00000397450.1 ENST00000397452.1 |
BPIFC |
BPI fold containing family C |
| chr12_-_49582978 | 0.11 |
ENST00000301071.7 |
TUBA1A |
tubulin, alpha 1a |
| chr5_+_102201509 | 0.11 |
ENST00000348126.2 ENST00000379787.4 |
PAM |
peptidylglycine alpha-amidating monooxygenase |
| chr5_+_102201430 | 0.11 |
ENST00000438793.3 ENST00000346918.2 |
PAM |
peptidylglycine alpha-amidating monooxygenase |
| chr17_+_42923686 | 0.10 |
ENST00000591513.1 |
HIGD1B |
HIG1 hypoxia inducible domain family, member 1B |
| chr4_-_69215699 | 0.10 |
ENST00000510746.1 ENST00000344157.4 ENST00000355665.3 |
YTHDC1 |
YTH domain containing 1 |
| chr5_-_135290705 | 0.10 |
ENST00000274507.1 |
LECT2 |
leukocyte cell-derived chemotaxin 2 |
| chr6_-_89927151 | 0.10 |
ENST00000454853.2 |
GABRR1 |
gamma-aminobutyric acid (GABA) A receptor, rho 1 |
| chr17_-_56065484 | 0.10 |
ENST00000581208.1 |
VEZF1 |
vascular endothelial zinc finger 1 |
| chr1_+_147013182 | 0.10 |
ENST00000234739.3 |
BCL9 |
B-cell CLL/lymphoma 9 |
| chr17_-_38020392 | 0.10 |
ENST00000346872.3 ENST00000439167.2 ENST00000377945.3 ENST00000394189.2 ENST00000377944.3 ENST00000377958.2 ENST00000535189.1 ENST00000377952.2 |
IKZF3 |
IKAROS family zinc finger 3 (Aiolos) |
| chr1_-_94079648 | 0.10 |
ENST00000370247.3 |
BCAR3 |
breast cancer anti-estrogen resistance 3 |
| chr20_-_656437 | 0.10 |
ENST00000488788.2 |
RP5-850E9.3 |
Uncharacterized protein |
| chr7_-_80141328 | 0.10 |
ENST00000398291.3 |
GNAT3 |
guanine nucleotide binding protein, alpha transducing 3 |
| chr7_-_148581251 | 0.10 |
ENST00000478654.1 ENST00000460911.1 ENST00000350995.2 |
EZH2 |
enhancer of zeste homolog 2 (Drosophila) |
| chr10_+_101542462 | 0.09 |
ENST00000370449.4 ENST00000370434.1 |
ABCC2 |
ATP-binding cassette, sub-family C (CFTR/MRP), member 2 |
| chr6_+_155537771 | 0.09 |
ENST00000275246.7 |
TIAM2 |
T-cell lymphoma invasion and metastasis 2 |
| chr4_+_86525299 | 0.09 |
ENST00000512201.1 |
ARHGAP24 |
Rho GTPase activating protein 24 |
| chr18_-_24445664 | 0.09 |
ENST00000578776.1 |
AQP4 |
aquaporin 4 |
| chr7_-_148581360 | 0.09 |
ENST00000320356.2 ENST00000541220.1 ENST00000483967.1 ENST00000536783.1 |
EZH2 |
enhancer of zeste homolog 2 (Drosophila) |
| chr17_+_42081914 | 0.09 |
ENST00000293404.3 ENST00000589767.1 |
NAGS |
N-acetylglutamate synthase |
| chr10_+_115312766 | 0.08 |
ENST00000351270.3 |
HABP2 |
hyaluronan binding protein 2 |
| chr15_-_82338460 | 0.08 |
ENST00000558133.1 ENST00000329713.4 |
MEX3B |
mex-3 RNA binding family member B |
| chr3_+_29323043 | 0.08 |
ENST00000452462.1 ENST00000456853.1 |
RBMS3 |
RNA binding motif, single stranded interacting protein 3 |
| chr3_-_134093275 | 0.07 |
ENST00000513145.1 ENST00000422605.2 |
AMOTL2 |
angiomotin like 2 |
| chr5_-_24645078 | 0.07 |
ENST00000264463.4 |
CDH10 |
cadherin 10, type 2 (T2-cadherin) |
| chr8_-_119964434 | 0.07 |
ENST00000297350.4 |
TNFRSF11B |
tumor necrosis factor receptor superfamily, member 11b |
| chr19_-_40791211 | 0.07 |
ENST00000579047.1 |
AKT2 |
v-akt murine thymoma viral oncogene homolog 2 |
| chr12_+_122326630 | 0.07 |
ENST00000541212.1 ENST00000340175.5 |
PSMD9 |
proteasome (prosome, macropain) 26S subunit, non-ATPase, 9 |
| chr22_+_19950060 | 0.07 |
ENST00000449653.1 |
COMT |
catechol-O-methyltransferase |
| chr2_+_228678550 | 0.07 |
ENST00000409189.3 ENST00000358813.4 |
CCL20 |
chemokine (C-C motif) ligand 20 |
| chr7_+_27282319 | 0.07 |
ENST00000222761.3 |
EVX1 |
even-skipped homeobox 1 |
| chr22_+_31742875 | 0.07 |
ENST00000504184.2 |
AC005003.1 |
CDNA FLJ20464 fis, clone KAT06158; HCG1777549; Uncharacterized protein |
| chr9_-_27005686 | 0.07 |
ENST00000380055.5 |
LRRC19 |
leucine rich repeat containing 19 |
| chr12_+_129178451 | 0.07 |
ENST00000537538.1 |
TMEM132C |
transmembrane protein 132C |
| chr11_-_62474803 | 0.07 |
ENST00000533982.1 ENST00000360796.5 |
BSCL2 |
Berardinelli-Seip congenital lipodystrophy 2 (seipin) |
| chr5_-_16936340 | 0.07 |
ENST00000507288.1 ENST00000513610.1 |
MYO10 |
myosin X |
| chr8_-_16859690 | 0.07 |
ENST00000180166.5 |
FGF20 |
fibroblast growth factor 20 |
| chr3_+_137483579 | 0.06 |
ENST00000306087.1 |
SOX14 |
SRY (sex determining region Y)-box 14 |
| chr17_-_63557759 | 0.06 |
ENST00000307078.5 |
AXIN2 |
axin 2 |
| chr9_-_15510287 | 0.06 |
ENST00000397519.2 |
PSIP1 |
PC4 and SFRS1 interacting protein 1 |
| chr18_+_29769978 | 0.06 |
ENST00000269202.6 ENST00000581447.1 |
MEP1B |
meprin A, beta |
| chrX_+_46937745 | 0.06 |
ENST00000397180.1 ENST00000457380.1 ENST00000352078.4 |
RGN |
regucalcin |
| chr12_+_122326662 | 0.06 |
ENST00000261817.2 ENST00000538613.1 ENST00000542602.1 |
PSMD9 |
proteasome (prosome, macropain) 26S subunit, non-ATPase, 9 |
| chr19_-_59030921 | 0.06 |
ENST00000354590.3 ENST00000596739.1 |
ZBTB45 |
zinc finger and BTB domain containing 45 |
| chr8_-_42234745 | 0.06 |
ENST00000220812.2 |
DKK4 |
dickkopf WNT signaling pathway inhibitor 4 |
| chr14_-_81425828 | 0.06 |
ENST00000555529.1 ENST00000556042.1 ENST00000556981.1 |
CEP128 |
centrosomal protein 128kDa |
| chr10_-_87551311 | 0.06 |
ENST00000536331.1 |
GRID1 |
glutamate receptor, ionotropic, delta 1 |
| chr11_-_6677018 | 0.06 |
ENST00000299441.3 |
DCHS1 |
dachsous cadherin-related 1 |
| chr12_+_52056548 | 0.06 |
ENST00000545061.1 ENST00000355133.3 |
SCN8A |
sodium channel, voltage gated, type VIII, alpha subunit |
| chr12_-_12491608 | 0.06 |
ENST00000545735.1 |
MANSC1 |
MANSC domain containing 1 |
| chr12_-_49582593 | 0.06 |
ENST00000295766.5 |
TUBA1A |
tubulin, alpha 1a |
| chr6_+_152130240 | 0.06 |
ENST00000427531.2 |
ESR1 |
estrogen receptor 1 |
| chr1_-_20250110 | 0.06 |
ENST00000375116.3 |
PLA2G2E |
phospholipase A2, group IIE |
| chr17_+_37618257 | 0.06 |
ENST00000447079.4 |
CDK12 |
cyclin-dependent kinase 12 |
| chr2_+_204732666 | 0.06 |
ENST00000295854.6 ENST00000472206.1 |
CTLA4 |
cytotoxic T-lymphocyte-associated protein 4 |
| chr5_+_140797296 | 0.06 |
ENST00000398594.2 |
PCDHGB7 |
protocadherin gamma subfamily B, 7 |
| chr3_-_134093395 | 0.05 |
ENST00000249883.5 |
AMOTL2 |
angiomotin like 2 |
| chr20_+_45947246 | 0.05 |
ENST00000599904.1 |
AL031666.2 |
HCG2018772; Uncharacterized protein; cDNA FLJ31609 fis, clone NT2RI2002852 |
| chr14_+_74815116 | 0.05 |
ENST00000256362.4 |
VRTN |
vertebrae development associated |
| chr14_+_75230011 | 0.05 |
ENST00000552421.1 ENST00000325680.7 ENST00000238571.3 |
YLPM1 |
YLP motif containing 1 |
| chr12_-_133464151 | 0.05 |
ENST00000315585.7 ENST00000266880.7 ENST00000443047.2 ENST00000432561.2 ENST00000450056.2 |
CHFR |
checkpoint with forkhead and ring finger domains, E3 ubiquitin protein ligase |
| chr5_-_58335281 | 0.05 |
ENST00000358923.6 |
PDE4D |
phosphodiesterase 4D, cAMP-specific |
| chr2_+_71357744 | 0.05 |
ENST00000498451.2 |
MPHOSPH10 |
M-phase phosphoprotein 10 (U3 small nucleolar ribonucleoprotein) |
| chr1_-_54304212 | 0.05 |
ENST00000540001.1 |
NDC1 |
NDC1 transmembrane nucleoporin |
| chr20_-_13971255 | 0.05 |
ENST00000284951.5 ENST00000378072.5 |
SEL1L2 |
sel-1 suppressor of lin-12-like 2 (C. elegans) |
| chr15_-_89764929 | 0.05 |
ENST00000268125.5 |
RLBP1 |
retinaldehyde binding protein 1 |
| chrX_-_153599578 | 0.04 |
ENST00000360319.4 ENST00000344736.4 |
FLNA |
filamin A, alpha |
| chr6_+_64281906 | 0.04 |
ENST00000370651.3 |
PTP4A1 |
protein tyrosine phosphatase type IVA, member 1 |
| chr20_+_18794370 | 0.04 |
ENST00000377428.2 |
SCP2D1 |
SCP2 sterol-binding domain containing 1 |
| chr3_-_33686925 | 0.04 |
ENST00000485378.2 ENST00000313350.6 ENST00000487200.1 |
CLASP2 |
cytoplasmic linker associated protein 2 |
| chr9_+_139221880 | 0.04 |
ENST00000392945.3 ENST00000440944.1 |
GPSM1 |
G-protein signaling modulator 1 |
| chr11_-_62572901 | 0.04 |
ENST00000439713.2 ENST00000531131.1 ENST00000530875.1 ENST00000531709.2 ENST00000294172.2 |
NXF1 |
nuclear RNA export factor 1 |
| chr5_+_54455946 | 0.04 |
ENST00000503787.1 ENST00000296734.6 ENST00000515370.1 |
GPX8 |
glutathione peroxidase 8 (putative) |
| chr2_-_233641265 | 0.04 |
ENST00000438786.1 ENST00000409779.1 ENST00000233826.3 |
KCNJ13 |
potassium inwardly-rectifying channel, subfamily J, member 13 |
| chr13_+_96743093 | 0.04 |
ENST00000376705.2 |
HS6ST3 |
heparan sulfate 6-O-sulfotransferase 3 |
| chr3_-_42452050 | 0.04 |
ENST00000441172.1 ENST00000287748.3 |
LYZL4 |
lysozyme-like 4 |
| chr21_+_18811205 | 0.04 |
ENST00000440664.1 |
C21orf37 |
chromosome 21 open reading frame 37 |
| chr20_+_59654146 | 0.04 |
ENST00000441660.1 |
RP5-827L5.1 |
RP5-827L5.1 |
| chr17_-_39316983 | 0.04 |
ENST00000390661.3 |
KRTAP4-4 |
keratin associated protein 4-4 |
| chr1_-_54303949 | 0.04 |
ENST00000234725.8 |
NDC1 |
NDC1 transmembrane nucleoporin |
| chr1_-_54303934 | 0.04 |
ENST00000537333.1 |
NDC1 |
NDC1 transmembrane nucleoporin |
| chr17_-_73389737 | 0.04 |
ENST00000392563.1 |
GRB2 |
growth factor receptor-bound protein 2 |
| chr1_-_166028709 | 0.04 |
ENST00000595430.1 |
AL626787.1 |
AL626787.1 |
| chr19_-_59031118 | 0.04 |
ENST00000600990.1 |
ZBTB45 |
zinc finger and BTB domain containing 45 |
| chr14_-_53258180 | 0.04 |
ENST00000554230.1 |
GNPNAT1 |
glucosamine-phosphate N-acetyltransferase 1 |
| chr15_-_31393910 | 0.03 |
ENST00000397795.2 ENST00000256552.6 ENST00000559179.1 |
TRPM1 |
transient receptor potential cation channel, subfamily M, member 1 |
| chr17_-_47308100 | 0.03 |
ENST00000503902.1 ENST00000512250.1 |
PHOSPHO1 |
phosphatase, orphan 1 |
| chr6_-_25930904 | 0.03 |
ENST00000377850.3 |
SLC17A2 |
solute carrier family 17, member 2 |
| chrX_-_43832711 | 0.03 |
ENST00000378062.5 |
NDP |
Norrie disease (pseudoglioma) |
| chr16_+_29819372 | 0.03 |
ENST00000568544.1 ENST00000569978.1 |
MAZ |
MYC-associated zinc finger protein (purine-binding transcription factor) |
| chr17_+_79679299 | 0.03 |
ENST00000331531.5 |
SLC25A10 |
solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 |
| chr5_-_175815565 | 0.03 |
ENST00000509257.1 ENST00000507413.1 ENST00000510123.1 |
NOP16 |
NOP16 nucleolar protein |
| chr4_+_170581213 | 0.03 |
ENST00000507875.1 |
CLCN3 |
chloride channel, voltage-sensitive 3 |
| chr12_+_6982725 | 0.03 |
ENST00000433346.1 |
LRRC23 |
leucine rich repeat containing 23 |
| chr6_-_25930819 | 0.03 |
ENST00000360488.3 |
SLC17A2 |
solute carrier family 17, member 2 |
| chr6_+_29427548 | 0.03 |
ENST00000377132.1 |
OR2H1 |
olfactory receptor, family 2, subfamily H, member 1 |
| chr2_-_88427568 | 0.03 |
ENST00000393750.3 ENST00000295834.3 |
FABP1 |
fatty acid binding protein 1, liver |
| chr19_-_3061397 | 0.03 |
ENST00000586839.1 |
AES |
amino-terminal enhancer of split |
| chrX_-_111923145 | 0.03 |
ENST00000371968.3 ENST00000536453.1 |
LHFPL1 |
lipoma HMGIC fusion partner-like 1 |
| chr17_-_47308128 | 0.03 |
ENST00000413580.1 ENST00000511066.1 |
PHOSPHO1 |
phosphatase, orphan 1 |
| chr16_+_29819446 | 0.03 |
ENST00000568282.1 |
MAZ |
MYC-associated zinc finger protein (purine-binding transcription factor) |
| chr17_-_73389854 | 0.03 |
ENST00000578961.1 ENST00000392564.1 ENST00000582582.1 |
GRB2 |
growth factor receptor-bound protein 2 |
| chr5_+_170846640 | 0.03 |
ENST00000274625.5 |
FGF18 |
fibroblast growth factor 18 |
| chr9_-_100707116 | 0.03 |
ENST00000259456.3 |
HEMGN |
hemogen |
| chr4_-_69215467 | 0.03 |
ENST00000579690.1 |
YTHDC1 |
YTH domain containing 1 |
| chr10_-_104179682 | 0.02 |
ENST00000406432.1 |
PSD |
pleckstrin and Sec7 domain containing |
| chrX_-_122866874 | 0.02 |
ENST00000245838.8 ENST00000355725.4 |
THOC2 |
THO complex 2 |
| chr17_-_40169659 | 0.02 |
ENST00000457167.4 |
DNAJC7 |
DnaJ (Hsp40) homolog, subfamily C, member 7 |
| chr3_+_155838337 | 0.02 |
ENST00000490337.1 ENST00000389636.5 |
KCNAB1 |
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chr19_+_51628165 | 0.02 |
ENST00000250360.3 ENST00000440804.3 |
SIGLEC9 |
sialic acid binding Ig-like lectin 9 |
| chr16_+_72088376 | 0.02 |
ENST00000570083.1 ENST00000355906.5 ENST00000398131.2 ENST00000569639.1 ENST00000564499.1 ENST00000357763.4 ENST00000562526.1 ENST00000565574.1 ENST00000568417.2 ENST00000356967.5 |
HP HPR |
haptoglobin haptoglobin-related protein |
| chr3_-_113464906 | 0.02 |
ENST00000477813.1 |
NAA50 |
N(alpha)-acetyltransferase 50, NatE catalytic subunit |
| chrX_+_2670066 | 0.02 |
ENST00000381174.5 ENST00000419513.2 ENST00000426774.1 |
XG |
Xg blood group |
| chr1_-_115053781 | 0.02 |
ENST00000358465.2 ENST00000369543.2 |
TRIM33 |
tripartite motif containing 33 |
| chr2_-_9770706 | 0.02 |
ENST00000381844.4 |
YWHAQ |
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, theta |
| chrX_-_100183894 | 0.02 |
ENST00000328526.5 ENST00000372956.2 |
XKRX |
XK, Kell blood group complex subunit-related, X-linked |
| chr13_-_72441315 | 0.02 |
ENST00000305425.4 ENST00000313174.7 ENST00000354591.4 |
DACH1 |
dachshund homolog 1 (Drosophila) |
| chr7_+_99613195 | 0.02 |
ENST00000324306.6 |
ZKSCAN1 |
zinc finger with KRAB and SCAN domains 1 |
| chr2_-_9771075 | 0.02 |
ENST00000446619.1 ENST00000238081.3 |
YWHAQ |
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, theta |
| chr8_-_6783588 | 0.02 |
ENST00000297436.2 |
DEFA6 |
defensin, alpha 6, Paneth cell-specific |
| chr2_+_204732487 | 0.02 |
ENST00000302823.3 |
CTLA4 |
cytotoxic T-lymphocyte-associated protein 4 |
| chr3_-_57199397 | 0.01 |
ENST00000296318.7 |
IL17RD |
interleukin 17 receptor D |
| chr12_+_85673868 | 0.01 |
ENST00000316824.3 |
ALX1 |
ALX homeobox 1 |
| chr3_-_33686743 | 0.01 |
ENST00000333778.6 ENST00000539981.1 |
CLASP2 |
cytoplasmic linker associated protein 2 |
| chr7_-_27219849 | 0.01 |
ENST00000396344.4 |
HOXA10 |
homeobox A10 |
| chrX_+_47004599 | 0.01 |
ENST00000329236.7 |
RBM10 |
RNA binding motif protein 10 |
| chr11_+_65408273 | 0.01 |
ENST00000394227.3 |
SIPA1 |
signal-induced proliferation-associated 1 |
| chr2_+_173686303 | 0.01 |
ENST00000397087.3 |
RAPGEF4 |
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr1_-_40367530 | 0.01 |
ENST00000372816.2 ENST00000372815.1 |
MYCL |
v-myc avian myelocytomatosis viral oncogene lung carcinoma derived homolog |
| chr2_+_71357434 | 0.01 |
ENST00000244230.2 |
MPHOSPH10 |
M-phase phosphoprotein 10 (U3 small nucleolar ribonucleoprotein) |
| chr10_+_7745232 | 0.01 |
ENST00000358415.4 |
ITIH2 |
inter-alpha-trypsin inhibitor heavy chain 2 |
| chr6_-_28220002 | 0.01 |
ENST00000377294.2 |
ZKSCAN4 |
zinc finger with KRAB and SCAN domains 4 |
| chr1_+_110091189 | 0.01 |
ENST00000369851.4 |
GNAI3 |
guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 3 |
| chr19_-_39924349 | 0.01 |
ENST00000602153.1 |
RPS16 |
ribosomal protein S16 |
| chr5_+_149546334 | 0.01 |
ENST00000231656.8 |
CDX1 |
caudal type homeobox 1 |
| chr10_+_114710516 | 0.01 |
ENST00000542695.1 ENST00000346198.4 |
TCF7L2 |
transcription factor 7-like 2 (T-cell specific, HMG-box) |
| chr7_-_148725733 | 0.01 |
ENST00000286091.4 |
PDIA4 |
protein disulfide isomerase family A, member 4 |
| chr7_+_99613212 | 0.01 |
ENST00000426572.1 ENST00000535170.1 |
ZKSCAN1 |
zinc finger with KRAB and SCAN domains 1 |
| chrX_+_47004639 | 0.01 |
ENST00000345781.6 |
RBM10 |
RNA binding motif protein 10 |
| chr15_+_57511609 | 0.01 |
ENST00000543579.1 ENST00000537840.1 ENST00000343827.3 |
TCF12 |
transcription factor 12 |
| chr22_+_29279552 | 0.01 |
ENST00000544604.2 |
ZNRF3 |
zinc and ring finger 3 |
| chr11_+_114128522 | 0.01 |
ENST00000535401.1 |
NNMT |
nicotinamide N-methyltransferase |
| chr17_-_39222131 | 0.01 |
ENST00000394015.2 |
KRTAP2-4 |
keratin associated protein 2-4 |
| chr12_-_6982442 | 0.01 |
ENST00000523102.1 ENST00000524270.1 ENST00000519357.1 |
SPSB2 |
splA/ryanodine receptor domain and SOCS box containing 2 |
| chr10_+_7745303 | 0.01 |
ENST00000429820.1 ENST00000379587.4 |
ITIH2 |
inter-alpha-trypsin inhibitor heavy chain 2 |
| chr13_-_72440901 | 0.01 |
ENST00000359684.2 |
DACH1 |
dachshund homolog 1 (Drosophila) |
| chr1_-_201096312 | 0.01 |
ENST00000449188.2 |
ASCL5 |
achaete-scute family bHLH transcription factor 5 |
| chr14_+_68189190 | 0.01 |
ENST00000539142.1 |
RDH12 |
retinol dehydrogenase 12 (all-trans/9-cis/11-cis) |
| chrX_-_110655306 | 0.01 |
ENST00000371993.2 |
DCX |
doublecortin |
| chr1_-_11907829 | 0.01 |
ENST00000376480.3 |
NPPA |
natriuretic peptide A |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:1901202 | negative regulation of extracellular matrix assembly(GO:1901202) |
| 0.1 | 0.3 | GO:0042663 | regulation of endodermal cell fate specification(GO:0042663) |
| 0.1 | 0.3 | GO:2000729 | positive regulation of mesenchymal cell proliferation involved in ureter development(GO:2000729) |
| 0.1 | 0.1 | GO:0021913 | regulation of transcription from RNA polymerase II promoter involved in ventral spinal cord interneuron specification(GO:0021913) |
| 0.1 | 0.2 | GO:0036333 | hepatocyte homeostasis(GO:0036333) response to tetrachloromethane(GO:1904772) |
| 0.1 | 0.2 | GO:1903566 | positive regulation of protein localization to cilium(GO:1903566) |
| 0.1 | 0.2 | GO:1901069 | guanosine-containing compound catabolic process(GO:1901069) |
| 0.1 | 0.2 | GO:0018032 | protein amidation(GO:0018032) |
| 0.0 | 0.1 | GO:0000415 | negative regulation of histone H3-K36 methylation(GO:0000415) |
| 0.0 | 0.2 | GO:1904059 | regulation of locomotor rhythm(GO:1904059) |
| 0.0 | 0.1 | GO:0051300 | spindle pole body duplication(GO:0030474) spindle pole body organization(GO:0051300) spindle pole body localization(GO:0070631) establishment of spindle pole body localization(GO:0070632) spindle pole body localization to nuclear envelope(GO:0071789) establishment of spindle pole body localization to nuclear envelope(GO:0071790) |
| 0.0 | 0.1 | GO:0050787 | antibiotic metabolic process(GO:0016999) detoxification of mercury ion(GO:0050787) |
| 0.0 | 0.2 | GO:0006041 | glucosamine metabolic process(GO:0006041) |
| 0.0 | 0.1 | GO:0070682 | proteasome regulatory particle assembly(GO:0070682) |
| 0.0 | 0.3 | GO:0070141 | response to UV-A(GO:0070141) |
| 0.0 | 0.1 | GO:0042489 | negative regulation of odontogenesis of dentin-containing tooth(GO:0042489) |
| 0.0 | 0.1 | GO:0045362 | regulation of interleukin-1 biosynthetic process(GO:0045360) positive regulation of interleukin-1 biosynthetic process(GO:0045362) |
| 0.0 | 0.1 | GO:1901671 | L-ascorbic acid biosynthetic process(GO:0019853) positive regulation of superoxide dismutase activity(GO:1901671) negative regulation of DNA catabolic process(GO:1903625) regulation of aminoacyl-tRNA ligase activity(GO:1903630) positive regulation of removal of superoxide radicals(GO:1904833) |
| 0.0 | 0.1 | GO:0003192 | mitral valve formation(GO:0003192) |
| 0.0 | 0.1 | GO:0050917 | sensory perception of sweet taste(GO:0050916) sensory perception of umami taste(GO:0050917) |
| 0.0 | 0.5 | GO:0007171 | activation of transmembrane receptor protein tyrosine kinase activity(GO:0007171) |
| 0.0 | 0.1 | GO:1904885 | beta-catenin destruction complex assembly(GO:1904885) |
| 0.0 | 0.1 | GO:1904338 | regulation of dopaminergic neuron differentiation(GO:1904338) |
| 0.0 | 0.3 | GO:0060539 | diaphragm development(GO:0060539) |
| 0.0 | 0.3 | GO:0070307 | lens fiber cell development(GO:0070307) |
| 0.0 | 0.0 | GO:0016479 | negative regulation of transcription from RNA polymerase I promoter(GO:0016479) |
| 0.0 | 0.1 | GO:0045590 | negative regulation of regulatory T cell differentiation(GO:0045590) |
| 0.0 | 0.1 | GO:0016036 | cellular response to phosphate starvation(GO:0016036) positive regulation of sulfur amino acid metabolic process(GO:0031337) positive regulation of homocysteine metabolic process(GO:0050668) |
| 0.0 | 0.3 | GO:0002517 | T cell tolerance induction(GO:0002517) |
| 0.0 | 0.1 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.0 | 0.0 | GO:0010746 | response to high light intensity(GO:0009644) regulation of plasma membrane long-chain fatty acid transport(GO:0010746) negative regulation of plasma membrane long-chain fatty acid transport(GO:0010748) cellular response to light intensity(GO:0071484) |
| 0.0 | 0.0 | GO:0071423 | thiosulfate transport(GO:0015709) oxaloacetate transport(GO:0015729) malate transport(GO:0015743) malate transmembrane transport(GO:0071423) oxaloacetate(2-) transmembrane transport(GO:1902356) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0070762 | nuclear pore transmembrane ring(GO:0070762) |
| 0.0 | 0.0 | GO:0031523 | Myb complex(GO:0031523) |
| 0.0 | 0.3 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.0 | 0.1 | GO:0016342 | catenin complex(GO:0016342) |
| 0.0 | 0.1 | GO:0034457 | Mpp10 complex(GO:0034457) |
| 0.0 | 0.0 | GO:0035841 | new growing cell tip(GO:0035841) |
| 0.0 | 0.1 | GO:0070436 | Grb2-EGFR complex(GO:0070436) |
| 0.0 | 0.2 | GO:0045120 | pronucleus(GO:0045120) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0045518 | interleukin-22 receptor binding(GO:0045518) |
| 0.1 | 0.2 | GO:0004504 | peptidylglycine monooxygenase activity(GO:0004504) peptidylamidoglycolate lyase activity(GO:0004598) |
| 0.0 | 0.3 | GO:0060002 | plus-end directed microfilament motor activity(GO:0060002) |
| 0.0 | 0.6 | GO:0005161 | platelet-derived growth factor receptor binding(GO:0005161) |
| 0.0 | 0.1 | GO:0031731 | CCR6 chemokine receptor binding(GO:0031731) |
| 0.0 | 0.9 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.0 | 0.1 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.0 | 0.2 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.0 | 0.3 | GO:0015197 | peptide transporter activity(GO:0015197) |
| 0.0 | 0.1 | GO:0016774 | phosphotransferase activity, carboxyl group as acceptor(GO:0016774) |
| 0.0 | 0.0 | GO:0017095 | heparan sulfate 6-O-sulfotransferase activity(GO:0017095) |
| 0.0 | 0.1 | GO:0004803 | transposase activity(GO:0004803) |
| 0.0 | 0.2 | GO:0043422 | protein kinase B binding(GO:0043422) |
| 0.0 | 0.1 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.0 | 0.0 | GO:0034988 | Fc-gamma receptor I complex binding(GO:0034988) |
| 0.0 | 0.0 | GO:0015131 | thiosulfate transmembrane transporter activity(GO:0015117) oxaloacetate transmembrane transporter activity(GO:0015131) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 0.4 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 0.2 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |