Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
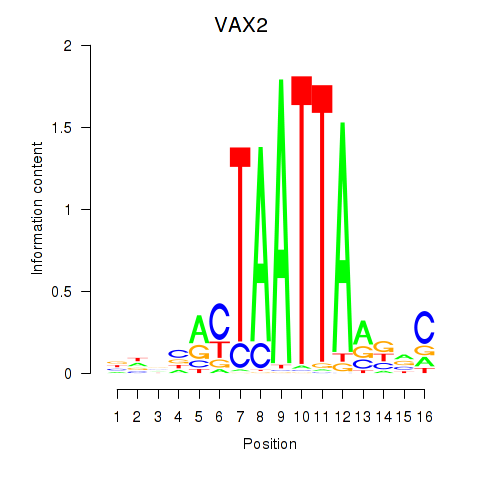
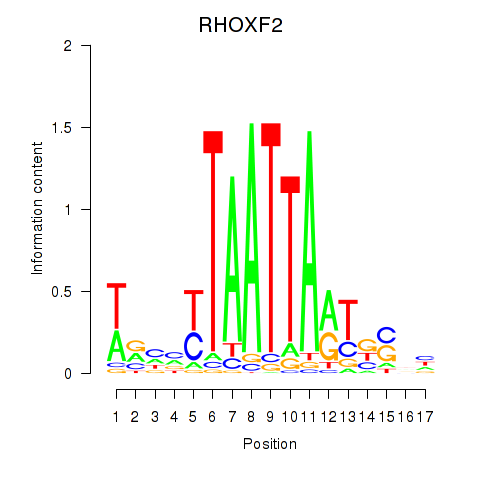
Results for VAX2_RHOXF2
Z-value: 0.25
Transcription factors associated with VAX2_RHOXF2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
VAX2
|
ENSG00000116035.2 | VAX2 |
|
RHOXF2
|
ENSG00000131721.4 | RHOXF2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| VAX2 | hg19_v2_chr2_+_71127699_71127744 | -0.41 | 3.1e-01 | Click! |
| RHOXF2 | hg19_v2_chrX_+_119292467_119292467 | 0.11 | 7.9e-01 | Click! |
Activity profile of VAX2_RHOXF2 motif
Sorted Z-values of VAX2_RHOXF2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of VAX2_RHOXF2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_-_57045228 | 0.54 |
ENST00000371250.3 |
PPAP2B |
phosphatidic acid phosphatase type 2B |
| chr5_-_20575959 | 0.46 |
ENST00000507958.1 |
CDH18 |
cadherin 18, type 2 |
| chr3_+_157154578 | 0.40 |
ENST00000295927.3 |
PTX3 |
pentraxin 3, long |
| chr4_-_186696425 | 0.32 |
ENST00000430503.1 ENST00000319454.6 ENST00000450341.1 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr16_+_53133070 | 0.27 |
ENST00000565832.1 |
CHD9 |
chromodomain helicase DNA binding protein 9 |
| chr9_-_95166841 | 0.23 |
ENST00000262551.4 |
OGN |
osteoglycin |
| chr4_-_138453606 | 0.19 |
ENST00000412923.2 ENST00000344876.4 ENST00000507846.1 ENST00000510305.1 |
PCDH18 |
protocadherin 18 |
| chr3_-_191000172 | 0.14 |
ENST00000427544.2 |
UTS2B |
urotensin 2B |
| chr4_-_66536057 | 0.14 |
ENST00000273854.3 |
EPHA5 |
EPH receptor A5 |
| chr4_-_138453559 | 0.14 |
ENST00000511115.1 |
PCDH18 |
protocadherin 18 |
| chr11_-_111794446 | 0.11 |
ENST00000527950.1 |
CRYAB |
crystallin, alpha B |
| chr19_+_50016610 | 0.11 |
ENST00000596975.1 |
FCGRT |
Fc fragment of IgG, receptor, transporter, alpha |
| chr7_-_92777606 | 0.10 |
ENST00000437805.1 ENST00000446959.1 ENST00000439952.1 ENST00000414791.1 ENST00000446033.1 ENST00000411955.1 ENST00000318238.4 |
SAMD9L |
sterile alpha motif domain containing 9-like |
| chr12_+_26348246 | 0.10 |
ENST00000422622.2 |
SSPN |
sarcospan |
| chr9_-_95166884 | 0.09 |
ENST00000375561.5 |
OGN |
osteoglycin |
| chr7_+_99717230 | 0.09 |
ENST00000262932.3 |
CNPY4 |
canopy FGF signaling regulator 4 |
| chr14_+_74034310 | 0.08 |
ENST00000538782.1 |
ACOT2 |
acyl-CoA thioesterase 2 |
| chr17_-_33390667 | 0.08 |
ENST00000378516.2 ENST00000268850.7 ENST00000394597.2 |
RFFL |
ring finger and FYVE-like domain containing E3 ubiquitin protein ligase |
| chr6_+_26440700 | 0.08 |
ENST00000494393.1 ENST00000482451.1 ENST00000244519.2 ENST00000339789.4 ENST00000471353.1 ENST00000361232.3 ENST00000487627.1 ENST00000496719.1 ENST00000490254.1 ENST00000487272.1 |
BTN3A3 |
butyrophilin, subfamily 3, member A3 |
| chrX_+_7137475 | 0.07 |
ENST00000217961.4 |
STS |
steroid sulfatase (microsomal), isozyme S |
| chr10_-_92681033 | 0.07 |
ENST00000371697.3 |
ANKRD1 |
ankyrin repeat domain 1 (cardiac muscle) |
| chr15_+_74218787 | 0.07 |
ENST00000261921.7 |
LOXL1 |
lysyl oxidase-like 1 |
| chr2_-_74618964 | 0.07 |
ENST00000417090.1 ENST00000409868.1 |
DCTN1 |
dynactin 1 |
| chr17_-_40337470 | 0.06 |
ENST00000293330.1 |
HCRT |
hypocretin (orexin) neuropeptide precursor |
| chr3_-_141747950 | 0.06 |
ENST00000497579.1 |
TFDP2 |
transcription factor Dp-2 (E2F dimerization partner 2) |
| chr20_+_15177480 | 0.06 |
ENST00000402914.1 |
MACROD2 |
MACRO domain containing 2 |
| chr20_-_33735070 | 0.06 |
ENST00000374491.3 ENST00000542871.1 ENST00000374492.3 |
EDEM2 |
ER degradation enhancer, mannosidase alpha-like 2 |
| chr8_-_42623747 | 0.05 |
ENST00000534622.1 |
CHRNA6 |
cholinergic receptor, nicotinic, alpha 6 (neuronal) |
| chr15_-_37393406 | 0.05 |
ENST00000338564.5 ENST00000558313.1 ENST00000340545.5 |
MEIS2 |
Meis homeobox 2 |
| chr2_-_74619152 | 0.05 |
ENST00000440727.1 ENST00000409240.1 |
DCTN1 |
dynactin 1 |
| chr21_+_17442799 | 0.05 |
ENST00000602580.1 ENST00000458468.1 ENST00000602935.1 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr7_+_138145076 | 0.05 |
ENST00000343526.4 |
TRIM24 |
tripartite motif containing 24 |
| chr10_-_27529486 | 0.05 |
ENST00000375888.1 |
ACBD5 |
acyl-CoA binding domain containing 5 |
| chr8_-_7243080 | 0.05 |
ENST00000400156.4 |
ZNF705G |
zinc finger protein 705G |
| chr12_-_78753496 | 0.05 |
ENST00000548512.1 |
RP11-38F22.1 |
RP11-38F22.1 |
| chr1_-_211307404 | 0.05 |
ENST00000367007.4 |
KCNH1 |
potassium voltage-gated channel, subfamily H (eag-related), member 1 |
| chr9_+_125133315 | 0.05 |
ENST00000223423.4 ENST00000362012.2 |
PTGS1 |
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) |
| chrX_+_43515467 | 0.05 |
ENST00000338702.3 ENST00000542639.1 |
MAOA |
monoamine oxidase A |
| chr12_+_26348429 | 0.05 |
ENST00000242729.2 |
SSPN |
sarcospan |
| chr2_+_169926047 | 0.04 |
ENST00000428522.1 ENST00000450153.1 ENST00000421653.1 |
DHRS9 |
dehydrogenase/reductase (SDR family) member 9 |
| chr11_+_33061543 | 0.04 |
ENST00000432887.1 ENST00000528898.1 ENST00000531632.2 |
TCP11L1 |
t-complex 11, testis-specific-like 1 |
| chr7_+_12727250 | 0.04 |
ENST00000404894.1 |
ARL4A |
ADP-ribosylation factor-like 4A |
| chr11_-_62521614 | 0.04 |
ENST00000527994.1 ENST00000394807.3 |
ZBTB3 |
zinc finger and BTB domain containing 3 |
| chr10_+_90660832 | 0.04 |
ENST00000371924.1 |
STAMBPL1 |
STAM binding protein-like 1 |
| chr12_+_131438443 | 0.04 |
ENST00000261654.5 |
GPR133 |
G protein-coupled receptor 133 |
| chr21_+_17443434 | 0.04 |
ENST00000400178.2 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr7_+_120628731 | 0.04 |
ENST00000310396.5 |
CPED1 |
cadherin-like and PC-esterase domain containing 1 |
| chr6_+_26402517 | 0.03 |
ENST00000414912.2 |
BTN3A1 |
butyrophilin, subfamily 3, member A1 |
| chr21_+_17443521 | 0.03 |
ENST00000456342.1 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr1_+_225600404 | 0.03 |
ENST00000366845.2 |
AC092811.1 |
AC092811.1 |
| chr1_+_35258592 | 0.03 |
ENST00000342280.4 ENST00000450137.1 |
GJA4 |
gap junction protein, alpha 4, 37kDa |
| chr14_-_53258314 | 0.03 |
ENST00000216410.3 ENST00000557604.1 |
GNPNAT1 |
glucosamine-phosphate N-acetyltransferase 1 |
| chr11_-_121986923 | 0.03 |
ENST00000560104.1 |
BLID |
BH3-like motif containing, cell death inducer |
| chr8_-_101661887 | 0.03 |
ENST00000311812.2 |
SNX31 |
sorting nexin 31 |
| chr17_+_66511540 | 0.03 |
ENST00000588188.2 |
PRKAR1A |
protein kinase, cAMP-dependent, regulatory, type I, alpha |
| chr1_+_205682497 | 0.03 |
ENST00000598338.1 |
AC119673.1 |
AC119673.1 |
| chr1_-_24151903 | 0.03 |
ENST00000436439.2 ENST00000374490.3 |
HMGCL |
3-hydroxymethyl-3-methylglutaryl-CoA lyase |
| chr8_+_7783859 | 0.03 |
ENST00000400120.3 |
ZNF705B |
zinc finger protein 705B |
| chr2_-_85555385 | 0.03 |
ENST00000377386.3 |
TGOLN2 |
trans-golgi network protein 2 |
| chr10_-_50396407 | 0.03 |
ENST00000374153.2 ENST00000374151.3 |
C10orf128 |
chromosome 10 open reading frame 128 |
| chr11_+_71900572 | 0.03 |
ENST00000312293.4 |
FOLR1 |
folate receptor 1 (adult) |
| chr15_-_98417780 | 0.03 |
ENST00000503874.3 |
LINC00923 |
long intergenic non-protein coding RNA 923 |
| chr11_+_60383204 | 0.03 |
ENST00000412599.1 ENST00000320202.4 |
LINC00301 |
long intergenic non-protein coding RNA 301 |
| chr12_+_31812121 | 0.03 |
ENST00000395763.3 |
METTL20 |
methyltransferase like 20 |
| chr7_-_99716952 | 0.02 |
ENST00000523306.1 ENST00000344095.4 ENST00000417349.1 ENST00000493322.1 ENST00000520135.1 ENST00000418432.2 ENST00000460673.2 ENST00000452041.1 ENST00000452438.2 ENST00000451699.1 ENST00000453269.2 |
TAF6 |
TAF6 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 80kDa |
| chr7_-_99717463 | 0.02 |
ENST00000437822.2 |
TAF6 |
TAF6 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 80kDa |
| chr5_+_140557371 | 0.02 |
ENST00000239444.2 |
PCDHB8 |
protocadherin beta 8 |
| chr7_+_100860949 | 0.02 |
ENST00000305105.2 |
ZNHIT1 |
zinc finger, HIT-type containing 1 |
| chrX_+_72783026 | 0.02 |
ENST00000373504.6 ENST00000373502.5 |
CHIC1 |
cysteine-rich hydrophobic domain 1 |
| chr11_+_71900703 | 0.02 |
ENST00000393681.2 |
FOLR1 |
folate receptor 1 (adult) |
| chr5_+_174151536 | 0.02 |
ENST00000239243.6 ENST00000507785.1 |
MSX2 |
msh homeobox 2 |
| chr4_+_169418255 | 0.02 |
ENST00000505667.1 ENST00000511948.1 |
PALLD |
palladin, cytoskeletal associated protein |
| chr20_-_29978286 | 0.02 |
ENST00000376315.2 |
DEFB119 |
defensin, beta 119 |
| chr2_-_40680578 | 0.02 |
ENST00000455476.1 |
SLC8A1 |
solute carrier family 8 (sodium/calcium exchanger), member 1 |
| chr21_-_37914898 | 0.02 |
ENST00000399136.1 |
CLDN14 |
claudin 14 |
| chr14_-_20923195 | 0.02 |
ENST00000206542.4 |
OSGEP |
O-sialoglycoprotein endopeptidase |
| chrX_+_36254051 | 0.02 |
ENST00000378657.4 |
CXorf30 |
chromosome X open reading frame 30 |
| chr1_+_54569968 | 0.02 |
ENST00000391366.1 |
AL161915.1 |
Uncharacterized protein |
| chrX_+_135730297 | 0.02 |
ENST00000370629.2 |
CD40LG |
CD40 ligand |
| chr6_+_26402465 | 0.02 |
ENST00000476549.2 ENST00000289361.6 ENST00000450085.2 ENST00000425234.2 ENST00000427334.1 ENST00000506698.1 |
BTN3A1 |
butyrophilin, subfamily 3, member A1 |
| chr14_+_20923350 | 0.02 |
ENST00000555414.1 ENST00000216714.3 ENST00000553681.1 ENST00000557344.1 ENST00000398030.4 ENST00000557181.1 ENST00000555839.1 ENST00000553368.1 ENST00000556054.1 ENST00000557054.1 ENST00000557592.1 ENST00000557150.1 |
APEX1 |
APEX nuclease (multifunctional DNA repair enzyme) 1 |
| chr5_-_149829294 | 0.02 |
ENST00000401695.3 |
RPS14 |
ribosomal protein S14 |
| chr1_+_212965170 | 0.02 |
ENST00000532324.1 ENST00000366974.4 ENST00000530441.1 ENST00000526641.1 ENST00000531963.1 ENST00000366973.4 ENST00000526997.1 ENST00000488246.2 |
TATDN3 |
TatD DNase domain containing 3 |
| chr10_-_17171817 | 0.02 |
ENST00000377833.4 |
CUBN |
cubilin (intrinsic factor-cobalamin receptor) |
| chrX_+_135730373 | 0.02 |
ENST00000370628.2 |
CD40LG |
CD40 ligand |
| chr5_-_149829244 | 0.02 |
ENST00000312037.5 |
RPS14 |
ribosomal protein S14 |
| chr7_+_6655225 | 0.01 |
ENST00000457543.3 |
ZNF853 |
zinc finger protein 853 |
| chr1_-_169429690 | 0.01 |
ENST00000445428.1 |
RP1-206D15.3 |
RP1-206D15.3 |
| chr19_+_12175504 | 0.01 |
ENST00000439326.3 |
ZNF844 |
zinc finger protein 844 |
| chr5_-_149829314 | 0.01 |
ENST00000407193.1 |
RPS14 |
ribosomal protein S14 |
| chr4_+_169418195 | 0.01 |
ENST00000261509.6 ENST00000335742.7 |
PALLD |
palladin, cytoskeletal associated protein |
| chr17_-_39191107 | 0.01 |
ENST00000344363.5 |
KRTAP1-3 |
keratin associated protein 1-3 |
| chr1_-_212965104 | 0.01 |
ENST00000422588.2 ENST00000366975.6 ENST00000366977.3 ENST00000366976.1 |
NSL1 |
NSL1, MIS12 kinetochore complex component |
| chr14_-_39639523 | 0.01 |
ENST00000330149.5 ENST00000554018.1 ENST00000347691.5 |
TRAPPC6B |
trafficking protein particle complex 6B |
| chr12_-_50643664 | 0.01 |
ENST00000550592.1 |
LIMA1 |
LIM domain and actin binding 1 |
| chr4_+_159727272 | 0.01 |
ENST00000379346.3 |
FNIP2 |
folliculin interacting protein 2 |
| chr19_-_46088068 | 0.01 |
ENST00000263275.4 ENST00000323060.3 |
OPA3 |
optic atrophy 3 (autosomal recessive, with chorea and spastic paraplegia) |
| chr16_+_12059050 | 0.01 |
ENST00000396495.3 |
TNFRSF17 |
tumor necrosis factor receptor superfamily, member 17 |
| chr15_+_58430368 | 0.01 |
ENST00000558772.1 ENST00000219919.4 |
AQP9 |
aquaporin 9 |
| chr15_+_58430567 | 0.01 |
ENST00000536493.1 |
AQP9 |
aquaporin 9 |
| chr16_+_69345243 | 0.01 |
ENST00000254950.11 |
VPS4A |
vacuolar protein sorting 4 homolog A (S. cerevisiae) |
| chr1_-_190446759 | 0.01 |
ENST00000367462.3 |
BRINP3 |
bone morphogenetic protein/retinoic acid inducible neural-specific 3 |
| chr13_+_36050881 | 0.01 |
ENST00000537702.1 |
NBEA |
neurobeachin |
| chr6_+_29079668 | 0.01 |
ENST00000377169.1 |
OR2J3 |
olfactory receptor, family 2, subfamily J, member 3 |
| chr12_-_118796910 | 0.01 |
ENST00000541186.1 ENST00000539872.1 |
TAOK3 |
TAO kinase 3 |
| chr17_-_39203519 | 0.01 |
ENST00000542137.1 ENST00000391419.3 |
KRTAP2-1 |
keratin associated protein 2-1 |
| chr15_+_80351977 | 0.01 |
ENST00000559157.1 ENST00000561012.1 ENST00000564367.1 ENST00000558494.1 |
ZFAND6 |
zinc finger, AN1-type domain 6 |
| chr9_+_131062367 | 0.01 |
ENST00000601297.1 |
AL359091.2 |
CDNA: FLJ21673 fis, clone COL09042; HCG2036511; Uncharacterized protein |
| chr3_+_152552685 | 0.01 |
ENST00000305097.3 |
P2RY1 |
purinergic receptor P2Y, G-protein coupled, 1 |
| chr17_+_43238438 | 0.01 |
ENST00000593138.1 ENST00000586681.1 |
HEXIM2 |
hexamethylene bis-acetamide inducible 2 |
| chr15_+_80351910 | 0.01 |
ENST00000261749.6 ENST00000561060.1 |
ZFAND6 |
zinc finger, AN1-type domain 6 |
| chr14_+_22465771 | 0.01 |
ENST00000390445.2 |
TRAV17 |
T cell receptor alpha variable 17 |
| chr12_+_81110684 | 0.01 |
ENST00000228644.3 |
MYF5 |
myogenic factor 5 |
| chrX_-_77225135 | 0.01 |
ENST00000458128.1 |
PGAM4 |
phosphoglycerate mutase family member 4 |
| chr4_-_69111401 | 0.01 |
ENST00000332644.5 |
TMPRSS11B |
transmembrane protease, serine 11B |
| chr20_-_29978383 | 0.01 |
ENST00000339144.3 ENST00000376321.3 |
DEFB119 |
defensin, beta 119 |
| chr5_+_176811431 | 0.01 |
ENST00000512593.1 ENST00000324417.5 |
SLC34A1 |
solute carrier family 34 (type II sodium/phosphate contransporter), member 1 |
| chr12_+_82347498 | 0.01 |
ENST00000550506.1 |
RP11-362A1.1 |
RP11-362A1.1 |
| chr16_+_72088376 | 0.01 |
ENST00000570083.1 ENST00000355906.5 ENST00000398131.2 ENST00000569639.1 ENST00000564499.1 ENST00000357763.4 ENST00000562526.1 ENST00000565574.1 ENST00000568417.2 ENST00000356967.5 |
HP HPR |
haptoglobin haptoglobin-related protein |
| chr10_-_99205607 | 0.01 |
ENST00000477692.2 ENST00000485122.2 ENST00000370886.5 ENST00000370885.4 ENST00000370902.3 ENST00000370884.5 |
EXOSC1 |
exosome component 1 |
| chr18_-_67624160 | 0.01 |
ENST00000581982.1 ENST00000280200.4 |
CD226 |
CD226 molecule |
| chr7_+_148936732 | 0.00 |
ENST00000335870.2 |
ZNF212 |
zinc finger protein 212 |
| chr9_-_21202204 | 0.00 |
ENST00000239347.3 |
IFNA7 |
interferon, alpha 7 |
| chr8_-_10512569 | 0.00 |
ENST00000382483.3 |
RP1L1 |
retinitis pigmentosa 1-like 1 |
| chr1_+_46972668 | 0.00 |
ENST00000371956.4 ENST00000360032.3 |
DMBX1 |
diencephalon/mesencephalon homeobox 1 |
| chr2_-_214016314 | 0.00 |
ENST00000434687.1 ENST00000374319.4 |
IKZF2 |
IKAROS family zinc finger 2 (Helios) |
| chr6_-_82957433 | 0.00 |
ENST00000306270.7 |
IBTK |
inhibitor of Bruton agammaglobulinemia tyrosine kinase |
| chr3_+_186353756 | 0.00 |
ENST00000431018.1 ENST00000450521.1 ENST00000539949.1 |
FETUB |
fetuin B |
| chr3_+_158519654 | 0.00 |
ENST00000415822.2 ENST00000392813.4 ENST00000264266.8 |
MFSD1 |
major facilitator superfamily domain containing 1 |
| chr16_-_28608424 | 0.00 |
ENST00000335715.4 |
SULT1A2 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 2 |
| chr5_+_129083772 | 0.00 |
ENST00000564719.1 |
KIAA1024L |
KIAA1024-like |
| chr7_-_44122063 | 0.00 |
ENST00000335195.6 ENST00000395831.3 ENST00000414235.1 ENST00000452049.1 ENST00000242248.5 |
POLM |
polymerase (DNA directed), mu |
| chr17_-_10372875 | 0.00 |
ENST00000255381.2 |
MYH4 |
myosin, heavy chain 4, skeletal muscle |
| chr17_+_48823975 | 0.00 |
ENST00000513969.1 ENST00000503728.1 |
LUC7L3 |
LUC7-like 3 (S. cerevisiae) |
| chr3_+_115342349 | 0.00 |
ENST00000393780.3 |
GAP43 |
growth associated protein 43 |
| chr17_+_4692230 | 0.00 |
ENST00000331264.7 |
GLTPD2 |
glycolipid transfer protein domain containing 2 |
| chr20_+_43990576 | 0.00 |
ENST00000372727.1 ENST00000414310.1 |
SYS1 |
SYS1 Golgi-localized integral membrane protein homolog (S. cerevisiae) |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0052199 | negative regulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052199) |
| 0.0 | 0.1 | GO:0036512 | trimming of terminal mannose on B branch(GO:0036509) trimming of first mannose on A branch(GO:0036511) trimming of second mannose on A branch(GO:0036512) |
| 0.0 | 0.3 | GO:0042340 | keratan sulfate catabolic process(GO:0042340) |
| 0.0 | 0.0 | GO:0003147 | neural crest cell migration involved in heart formation(GO:0003147) anterior neural tube closure(GO:0061713) |
| 0.0 | 0.1 | GO:2001271 | negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.0 | 0.1 | GO:0090063 | positive regulation of microtubule nucleation(GO:0090063) |
| 0.0 | 0.1 | GO:0002414 | immunoglobulin transcytosis in epithelial cells(GO:0002414) |
| 0.0 | 0.1 | GO:0051970 | negative regulation of transmission of nerve impulse(GO:0051970) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.0 | 0.1 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 0.0 | 0.1 | GO:0061629 | RNA polymerase II sequence-specific DNA binding transcription factor binding(GO:0061629) |
| 0.0 | 0.1 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.0 | 0.0 | GO:0061714 | folic acid receptor activity(GO:0061714) |
| 0.0 | 0.0 | GO:0005174 | CD40 receptor binding(GO:0005174) |
| 0.0 | 0.3 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.0 | 0.5 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 0.5 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |