Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011): : averaged replicates
Navigation
Downloads
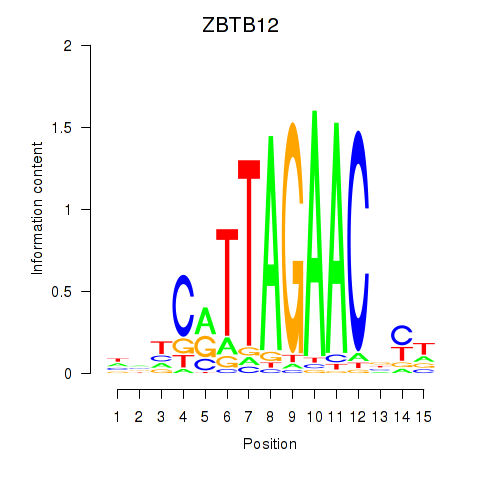
Results for ZBTB12
Z-value: 1.27
Transcription factors associated with ZBTB12
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ZBTB12
|
ENSG00000204366.3 | ZBTB12 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ZBTB12 | hg19_v2_chr6_-_31869769_31869880 | 0.12 | 7.8e-01 | Click! |
Activity profile of ZBTB12 motif
Sorted Z-values of ZBTB12 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ZBTB12
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_-_39677971 | 1.27 |
ENST00000393976.2 |
KRT15 |
keratin 15 |
| chr10_-_54215029 | 1.20 |
ENST00000435813.2 |
RP11-346D6.6 |
RP11-346D6.6 |
| chr1_+_35220613 | 1.02 |
ENST00000338513.1 |
GJB5 |
gap junction protein, beta 5, 31.1kDa |
| chr6_+_36097992 | 0.88 |
ENST00000211287.4 |
MAPK13 |
mitogen-activated protein kinase 13 |
| chr1_-_117210290 | 0.71 |
ENST00000369483.1 ENST00000369486.3 |
IGSF3 |
immunoglobulin superfamily, member 3 |
| chr19_-_6767431 | 0.71 |
ENST00000437152.3 ENST00000597687.1 |
SH2D3A |
SH2 domain containing 3A |
| chr19_-_6767516 | 0.69 |
ENST00000245908.6 |
SH2D3A |
SH2 domain containing 3A |
| chr4_-_36246060 | 0.50 |
ENST00000303965.4 |
ARAP2 |
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 2 |
| chr3_-_190167571 | 0.50 |
ENST00000354905.2 |
TMEM207 |
transmembrane protein 207 |
| chr7_-_142139783 | 0.49 |
ENST00000390374.3 |
TRBV7-6 |
T cell receptor beta variable 7-6 |
| chr8_+_118533049 | 0.47 |
ENST00000522839.1 |
MED30 |
mediator complex subunit 30 |
| chrX_-_24690771 | 0.47 |
ENST00000379145.1 |
PCYT1B |
phosphate cytidylyltransferase 1, choline, beta |
| chr22_+_38035459 | 0.46 |
ENST00000357436.4 |
SH3BP1 |
SH3-domain binding protein 1 |
| chr8_+_118532937 | 0.44 |
ENST00000297347.3 |
MED30 |
mediator complex subunit 30 |
| chr1_-_185597619 | 0.36 |
ENST00000608417.1 ENST00000436955.1 |
GS1-204I12.1 |
GS1-204I12.1 |
| chr6_-_170599561 | 0.34 |
ENST00000366756.3 |
DLL1 |
delta-like 1 (Drosophila) |
| chr19_-_11688500 | 0.34 |
ENST00000433365.2 |
ACP5 |
acid phosphatase 5, tartrate resistant |
| chr13_+_97928395 | 0.33 |
ENST00000445661.2 |
MBNL2 |
muscleblind-like splicing regulator 2 |
| chr8_+_104892639 | 0.32 |
ENST00000436393.2 |
RIMS2 |
regulating synaptic membrane exocytosis 2 |
| chr10_+_5454505 | 0.32 |
ENST00000355029.4 |
NET1 |
neuroepithelial cell transforming 1 |
| chr14_-_91526462 | 0.31 |
ENST00000536315.2 |
RPS6KA5 |
ribosomal protein S6 kinase, 90kDa, polypeptide 5 |
| chr13_-_46716969 | 0.29 |
ENST00000435666.2 |
LCP1 |
lymphocyte cytosolic protein 1 (L-plastin) |
| chr19_-_51531272 | 0.29 |
ENST00000319720.7 |
KLK11 |
kallikrein-related peptidase 11 |
| chr6_+_18387570 | 0.28 |
ENST00000259939.3 |
RNF144B |
ring finger protein 144B |
| chr7_-_142247606 | 0.28 |
ENST00000390361.3 |
TRBV7-3 |
T cell receptor beta variable 7-3 |
| chr19_-_51531210 | 0.27 |
ENST00000391804.3 |
KLK11 |
kallikrein-related peptidase 11 |
| chr19_-_13227534 | 0.26 |
ENST00000588229.1 ENST00000357720.4 |
TRMT1 |
tRNA methyltransferase 1 homolog (S. cerevisiae) |
| chr19_-_51530916 | 0.25 |
ENST00000594768.1 |
KLK11 |
kallikrein-related peptidase 11 |
| chr17_-_7518145 | 0.25 |
ENST00000250113.7 ENST00000571597.1 |
FXR2 |
fragile X mental retardation, autosomal homolog 2 |
| chr8_+_1993152 | 0.24 |
ENST00000262113.4 |
MYOM2 |
myomesin 2 |
| chr1_-_109203997 | 0.24 |
ENST00000370032.5 |
HENMT1 |
HEN1 methyltransferase homolog 1 (Arabidopsis) |
| chr11_+_112047087 | 0.24 |
ENST00000526088.1 ENST00000532593.1 ENST00000531169.1 |
BCO2 |
beta-carotene oxygenase 2 |
| chr5_-_156772729 | 0.24 |
ENST00000312349.4 |
FNDC9 |
fibronectin type III domain containing 9 |
| chr13_+_45694583 | 0.23 |
ENST00000340473.6 |
GTF2F2 |
general transcription factor IIF, polypeptide 2, 30kDa |
| chr1_-_32264356 | 0.22 |
ENST00000452755.2 |
SPOCD1 |
SPOC domain containing 1 |
| chr8_+_1993173 | 0.22 |
ENST00000523438.1 |
MYOM2 |
myomesin 2 |
| chrX_+_41548259 | 0.22 |
ENST00000378138.5 |
GPR34 |
G protein-coupled receptor 34 |
| chr19_+_11485333 | 0.22 |
ENST00000312423.2 |
SWSAP1 |
SWIM-type zinc finger 7 associated protein 1 |
| chr2_+_11674213 | 0.21 |
ENST00000381486.2 |
GREB1 |
growth regulation by estrogen in breast cancer 1 |
| chr11_-_59952106 | 0.21 |
ENST00000529054.1 ENST00000530839.1 |
MS4A6A |
membrane-spanning 4-domains, subfamily A, member 6A |
| chr4_-_87028478 | 0.21 |
ENST00000515400.1 ENST00000395157.3 |
MAPK10 |
mitogen-activated protein kinase 10 |
| chr1_+_84873913 | 0.20 |
ENST00000370662.3 |
DNASE2B |
deoxyribonuclease II beta |
| chr14_-_21270995 | 0.20 |
ENST00000555698.1 ENST00000397970.4 ENST00000340900.3 |
RNASE1 |
ribonuclease, RNase A family, 1 (pancreatic) |
| chr3_-_72149553 | 0.20 |
ENST00000468646.2 ENST00000464271.1 |
LINC00877 |
long intergenic non-protein coding RNA 877 |
| chr11_+_60145997 | 0.19 |
ENST00000530614.1 ENST00000530027.1 ENST00000530234.2 ENST00000528215.1 ENST00000531787.1 |
MS4A7 MS4A14 |
membrane-spanning 4-domains, subfamily A, member 7 membrane-spanning 4-domains, subfamily A, member 14 |
| chr16_+_66914264 | 0.19 |
ENST00000311765.2 ENST00000568869.1 ENST00000561704.1 ENST00000568398.1 ENST00000566776.1 |
PDP2 |
pyruvate dehyrogenase phosphatase catalytic subunit 2 |
| chr5_-_38557561 | 0.19 |
ENST00000511561.1 |
LIFR |
leukemia inhibitory factor receptor alpha |
| chr1_+_151009054 | 0.18 |
ENST00000295294.7 |
BNIPL |
BCL2/adenovirus E1B 19kD interacting protein like |
| chr1_+_151009035 | 0.18 |
ENST00000368931.3 |
BNIPL |
BCL2/adenovirus E1B 19kD interacting protein like |
| chr6_-_114292449 | 0.18 |
ENST00000519065.1 |
HDAC2 |
histone deacetylase 2 |
| chr4_-_99851766 | 0.18 |
ENST00000450253.2 |
EIF4E |
eukaryotic translation initiation factor 4E |
| chr4_-_85419603 | 0.18 |
ENST00000295886.4 |
NKX6-1 |
NK6 homeobox 1 |
| chr1_+_100503643 | 0.17 |
ENST00000370152.3 |
HIAT1 |
hippocampus abundant transcript 1 |
| chr5_-_88179302 | 0.16 |
ENST00000504921.2 |
MEF2C |
myocyte enhancer factor 2C |
| chr4_+_184826418 | 0.16 |
ENST00000308497.4 ENST00000438269.1 |
STOX2 |
storkhead box 2 |
| chr7_+_129847688 | 0.16 |
ENST00000297819.3 |
SSMEM1 |
serine-rich single-pass membrane protein 1 |
| chr7_-_93520191 | 0.15 |
ENST00000545378.1 |
TFPI2 |
tissue factor pathway inhibitor 2 |
| chr15_+_40453204 | 0.15 |
ENST00000287598.6 ENST00000412359.3 |
BUB1B |
BUB1 mitotic checkpoint serine/threonine kinase B |
| chr19_-_36019123 | 0.15 |
ENST00000588674.1 ENST00000452271.2 ENST00000518157.1 |
SBSN |
suprabasin |
| chr7_+_23637763 | 0.15 |
ENST00000410069.1 |
CCDC126 |
coiled-coil domain containing 126 |
| chr19_-_46148820 | 0.14 |
ENST00000587152.1 |
EML2 |
echinoderm microtubule associated protein like 2 |
| chr18_-_53177984 | 0.14 |
ENST00000543082.1 |
TCF4 |
transcription factor 4 |
| chr4_-_96470108 | 0.13 |
ENST00000513796.1 |
UNC5C |
unc-5 homolog C (C. elegans) |
| chr20_-_36793663 | 0.13 |
ENST00000536701.1 ENST00000536724.1 |
TGM2 |
transglutaminase 2 |
| chr11_+_64323098 | 0.12 |
ENST00000301891.4 |
SLC22A11 |
solute carrier family 22 (organic anion/urate transporter), member 11 |
| chr11_-_101000445 | 0.12 |
ENST00000534013.1 |
PGR |
progesterone receptor |
| chr11_+_64323156 | 0.12 |
ENST00000377585.3 |
SLC22A11 |
solute carrier family 22 (organic anion/urate transporter), member 11 |
| chr14_+_22966642 | 0.11 |
ENST00000390496.1 |
TRAJ41 |
T cell receptor alpha joining 41 |
| chrX_+_140096761 | 0.11 |
ENST00000370530.1 |
SPANXB1 |
SPANX family, member B1 |
| chrX_+_149861836 | 0.10 |
ENST00000542156.1 ENST00000370390.3 ENST00000490316.2 ENST00000445323.2 ENST00000544228.1 ENST00000451863.2 |
MTMR1 |
myotubularin related protein 1 |
| chr1_+_22778337 | 0.10 |
ENST00000404138.1 ENST00000400239.2 ENST00000375647.4 ENST00000374651.4 |
ZBTB40 |
zinc finger and BTB domain containing 40 |
| chr11_-_65769594 | 0.10 |
ENST00000532707.1 ENST00000533544.1 ENST00000526451.1 ENST00000312234.2 ENST00000530462.1 ENST00000525767.1 ENST00000529964.1 ENST00000527249.1 |
EIF1AD |
eukaryotic translation initiation factor 1A domain containing |
| chr7_+_129984630 | 0.10 |
ENST00000355388.3 ENST00000497503.1 ENST00000463587.1 ENST00000461828.1 ENST00000494311.1 ENST00000466363.2 ENST00000485477.1 ENST00000431780.2 ENST00000474905.1 |
CPA5 |
carboxypeptidase A5 |
| chr7_-_142176790 | 0.10 |
ENST00000390369.2 |
TRBV7-4 |
T cell receptor beta variable 7-4 (gene/pseudogene) |
| chr3_-_149051194 | 0.10 |
ENST00000470080.1 |
TM4SF18 |
transmembrane 4 L six family member 18 |
| chrX_+_133930798 | 0.09 |
ENST00000414371.2 |
FAM122C |
family with sequence similarity 122C |
| chr14_-_54908043 | 0.09 |
ENST00000556113.1 ENST00000553660.1 ENST00000395573.4 ENST00000557690.1 ENST00000216416.4 |
CNIH1 |
cornichon family AMPA receptor auxiliary protein 1 |
| chr2_+_89999259 | 0.09 |
ENST00000558026.1 |
IGKV2D-28 |
immunoglobulin kappa variable 2D-28 |
| chr7_-_93520259 | 0.08 |
ENST00000222543.5 |
TFPI2 |
tissue factor pathway inhibitor 2 |
| chr15_+_66797455 | 0.08 |
ENST00000446801.2 |
ZWILCH |
zwilch kinetochore protein |
| chr1_+_158979792 | 0.08 |
ENST00000359709.3 ENST00000430894.2 |
IFI16 |
interferon, gamma-inducible protein 16 |
| chr11_+_111126707 | 0.08 |
ENST00000280325.4 |
C11orf53 |
chromosome 11 open reading frame 53 |
| chr14_-_96743060 | 0.08 |
ENST00000554299.1 |
DKFZP434O1614 |
DKFZP434O1614 |
| chr8_-_141728760 | 0.08 |
ENST00000430260.2 |
PTK2 |
protein tyrosine kinase 2 |
| chr17_-_5487768 | 0.08 |
ENST00000269280.4 ENST00000345221.3 ENST00000262467.5 |
NLRP1 |
NLR family, pyrin domain containing 1 |
| chr2_+_234545092 | 0.08 |
ENST00000344644.5 |
UGT1A10 |
UDP glucuronosyltransferase 1 family, polypeptide A10 |
| chr13_+_46039037 | 0.07 |
ENST00000349995.5 |
COG3 |
component of oligomeric golgi complex 3 |
| chr17_-_7142725 | 0.07 |
ENST00000571362.1 ENST00000576955.1 ENST00000320316.3 |
PHF23 |
PHD finger protein 23 |
| chr17_+_73257945 | 0.07 |
ENST00000579002.1 |
MRPS7 |
mitochondrial ribosomal protein S7 |
| chr2_+_190722119 | 0.07 |
ENST00000452382.1 |
PMS1 |
PMS1 postmeiotic segregation increased 1 (S. cerevisiae) |
| chr14_-_22938665 | 0.07 |
ENST00000535880.2 |
TRDV3 |
T cell receptor delta variable 3 |
| chr3_+_35682913 | 0.07 |
ENST00000449196.1 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chr1_+_32757668 | 0.07 |
ENST00000373548.3 |
HDAC1 |
histone deacetylase 1 |
| chr17_+_7517264 | 0.07 |
ENST00000593717.1 ENST00000572182.1 ENST00000574539.1 ENST00000576728.1 ENST00000575314.1 ENST00000570547.1 ENST00000572262.1 ENST00000576478.1 |
AC007421.1 SHBG |
AC007421.1 sex hormone-binding globulin |
| chr14_+_96949319 | 0.07 |
ENST00000554706.1 |
AK7 |
adenylate kinase 7 |
| chr14_-_23876801 | 0.07 |
ENST00000356287.3 |
MYH6 |
myosin, heavy chain 6, cardiac muscle, alpha |
| chr1_+_45212051 | 0.07 |
ENST00000372222.3 |
KIF2C |
kinesin family member 2C |
| chr1_-_198990166 | 0.07 |
ENST00000427439.1 |
RP11-16L9.3 |
RP11-16L9.3 |
| chr14_-_23877474 | 0.06 |
ENST00000405093.3 |
MYH6 |
myosin, heavy chain 6, cardiac muscle, alpha |
| chr1_-_94344754 | 0.06 |
ENST00000436063.2 |
DNTTIP2 |
deoxynucleotidyltransferase, terminal, interacting protein 2 |
| chr3_-_149051444 | 0.06 |
ENST00000296059.2 |
TM4SF18 |
transmembrane 4 L six family member 18 |
| chr2_+_197577841 | 0.06 |
ENST00000409270.1 |
CCDC150 |
coiled-coil domain containing 150 |
| chr22_-_42084863 | 0.06 |
ENST00000401959.1 ENST00000355257.3 |
NHP2L1 |
NHP2 non-histone chromosome protein 2-like 1 (S. cerevisiae) |
| chr17_-_27405875 | 0.06 |
ENST00000359450.6 |
TIAF1 |
TGFB1-induced anti-apoptotic factor 1 |
| chr2_-_214013353 | 0.06 |
ENST00000451136.2 ENST00000421754.2 ENST00000374327.4 ENST00000413091.3 |
IKZF2 |
IKAROS family zinc finger 2 (Helios) |
| chr12_-_7077125 | 0.06 |
ENST00000545555.2 |
PHB2 |
prohibitin 2 |
| chr3_-_50336826 | 0.06 |
ENST00000443842.1 ENST00000354862.4 ENST00000443094.2 ENST00000415204.1 ENST00000336307.1 |
NAT6 HYAL3 |
N-acetyltransferase 6 (GCN5-related) hyaluronoglucosaminidase 3 |
| chr1_+_45212074 | 0.06 |
ENST00000372217.1 |
KIF2C |
kinesin family member 2C |
| chr11_-_119252359 | 0.06 |
ENST00000455332.2 |
USP2 |
ubiquitin specific peptidase 2 |
| chr22_+_18121356 | 0.05 |
ENST00000317582.5 ENST00000543133.1 ENST00000538149.1 ENST00000337612.5 ENST00000493680.1 |
BCL2L13 |
BCL2-like 13 (apoptosis facilitator) |
| chr22_+_51039098 | 0.05 |
ENST00000399912.1 ENST00000329492.3 ENST00000442429.2 ENST00000341339.4 |
MAPK8IP2 |
mitogen-activated protein kinase 8 interacting protein 2 |
| chr1_+_100598691 | 0.05 |
ENST00000370143.1 ENST00000370141.2 |
TRMT13 |
tRNA methyltransferase 13 homolog (S. cerevisiae) |
| chr1_+_171107241 | 0.05 |
ENST00000236166.3 |
FMO6P |
flavin containing monooxygenase 6 pseudogene |
| chr19_+_46195895 | 0.05 |
ENST00000366382.4 |
QPCTL |
glutaminyl-peptide cyclotransferase-like |
| chr17_+_4643337 | 0.05 |
ENST00000592813.1 |
ZMYND15 |
zinc finger, MYND-type containing 15 |
| chr5_+_29143603 | 0.05 |
ENST00000602470.1 ENST00000602801.1 |
CTD-2118P12.1 |
CTD-2118P12.1 |
| chr15_+_66797627 | 0.05 |
ENST00000565627.1 ENST00000564179.1 |
ZWILCH |
zwilch kinetochore protein |
| chr15_-_66797172 | 0.05 |
ENST00000569438.1 ENST00000569696.1 ENST00000307961.6 |
RPL4 |
ribosomal protein L4 |
| chr11_+_60145967 | 0.05 |
ENST00000534016.1 |
MS4A7 |
membrane-spanning 4-domains, subfamily A, member 7 |
| chr11_-_104972158 | 0.05 |
ENST00000598974.1 ENST00000593315.1 ENST00000594519.1 ENST00000415981.2 ENST00000525374.1 ENST00000375707.1 |
CASP1 CARD16 CARD17 |
caspase 1, apoptosis-related cysteine peptidase caspase recruitment domain family, member 16 caspase recruitment domain family, member 17 |
| chrX_+_50653784 | 0.05 |
ENST00000252677.3 |
BMP15 |
bone morphogenetic protein 15 |
| chr12_+_123459127 | 0.05 |
ENST00000397389.2 ENST00000538755.1 ENST00000536150.1 ENST00000545056.1 ENST00000545612.1 ENST00000538628.1 ENST00000545317.1 |
OGFOD2 |
2-oxoglutarate and iron-dependent oxygenase domain containing 2 |
| chr5_-_35089722 | 0.05 |
ENST00000511486.1 ENST00000310101.5 ENST00000231423.3 ENST00000513753.1 ENST00000348262.3 ENST00000397391.3 ENST00000542609.1 |
PRLR |
prolactin receptor |
| chr6_+_26158343 | 0.05 |
ENST00000377777.4 ENST00000289316.2 |
HIST1H2BD |
histone cluster 1, H2bd |
| chr15_-_35280426 | 0.05 |
ENST00000559564.1 ENST00000356321.4 |
ZNF770 |
zinc finger protein 770 |
| chr3_-_52804872 | 0.05 |
ENST00000535191.1 ENST00000461689.1 ENST00000383721.4 ENST00000233027.5 |
NEK4 |
NIMA-related kinase 4 |
| chr22_-_50524298 | 0.05 |
ENST00000311597.5 |
MLC1 |
megalencephalic leukoencephalopathy with subcortical cysts 1 |
| chr10_+_76969909 | 0.04 |
ENST00000298468.5 ENST00000543351.1 |
VDAC2 |
voltage-dependent anion channel 2 |
| chrX_+_23352133 | 0.04 |
ENST00000379361.4 |
PTCHD1 |
patched domain containing 1 |
| chr18_+_39766626 | 0.04 |
ENST00000593234.1 ENST00000585627.1 ENST00000591199.1 ENST00000586990.1 ENST00000593051.1 ENST00000593316.1 ENST00000591381.1 ENST00000585639.1 ENST00000589068.1 |
LINC00907 |
long intergenic non-protein coding RNA 907 |
| chr12_+_14572070 | 0.04 |
ENST00000545769.1 ENST00000428217.2 ENST00000396279.2 ENST00000542514.1 ENST00000536279.1 |
ATF7IP |
activating transcription factor 7 interacting protein |
| chr1_+_158979680 | 0.04 |
ENST00000368131.4 ENST00000340979.6 |
IFI16 |
interferon, gamma-inducible protein 16 |
| chr10_-_12084770 | 0.04 |
ENST00000357604.5 |
UPF2 |
UPF2 regulator of nonsense transcripts homolog (yeast) |
| chr2_+_166152283 | 0.04 |
ENST00000375427.2 |
SCN2A |
sodium channel, voltage-gated, type II, alpha subunit |
| chrX_+_70338525 | 0.04 |
ENST00000374102.1 |
MED12 |
mediator complex subunit 12 |
| chrX_+_101478829 | 0.04 |
ENST00000372763.1 ENST00000372758.1 |
NXF2 |
nuclear RNA export factor 2 |
| chr3_+_49236910 | 0.04 |
ENST00000452691.2 ENST00000366429.2 |
CCDC36 |
coiled-coil domain containing 36 |
| chr1_+_158979686 | 0.04 |
ENST00000368132.3 ENST00000295809.7 |
IFI16 |
interferon, gamma-inducible protein 16 |
| chr11_-_104905840 | 0.03 |
ENST00000526568.1 ENST00000393136.4 ENST00000531166.1 ENST00000534497.1 ENST00000527979.1 ENST00000446369.1 ENST00000353247.5 ENST00000528974.1 ENST00000533400.1 ENST00000525825.1 ENST00000436863.3 |
CASP1 |
caspase 1, apoptosis-related cysteine peptidase |
| chr18_-_47018769 | 0.03 |
ENST00000583637.1 ENST00000578528.1 ENST00000578532.1 ENST00000580387.1 ENST00000579248.1 ENST00000581373.1 |
RPL17 |
ribosomal protein L17 |
| chr1_-_157811588 | 0.03 |
ENST00000368174.4 |
CD5L |
CD5 molecule-like |
| chr6_-_31782813 | 0.03 |
ENST00000375654.4 |
HSPA1L |
heat shock 70kDa protein 1-like |
| chr15_-_55562479 | 0.03 |
ENST00000564609.1 |
RAB27A |
RAB27A, member RAS oncogene family |
| chr1_+_26644441 | 0.03 |
ENST00000374213.2 |
CD52 |
CD52 molecule |
| chr21_-_32253874 | 0.02 |
ENST00000332378.4 |
KRTAP11-1 |
keratin associated protein 11-1 |
| chr1_+_155583012 | 0.02 |
ENST00000462250.2 |
MSTO1 |
misato 1, mitochondrial distribution and morphology regulator |
| chr12_-_62586543 | 0.02 |
ENST00000416284.3 |
FAM19A2 |
family with sequence similarity 19 (chemokine (C-C motif)-like), member A2 |
| chr3_+_26735991 | 0.02 |
ENST00000456208.2 |
LRRC3B |
leucine rich repeat containing 3B |
| chr8_-_52721975 | 0.02 |
ENST00000356297.4 ENST00000543296.1 |
PXDNL |
peroxidasin homolog (Drosophila)-like |
| chr4_-_168155417 | 0.02 |
ENST00000511269.1 ENST00000506697.1 ENST00000512042.1 |
SPOCK3 |
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3 |
| chr3_-_50336548 | 0.01 |
ENST00000450489.1 ENST00000513170.1 ENST00000450982.1 |
NAT6 HYAL3 |
N-acetyltransferase 6 (GCN5-related) hyaluronoglucosaminidase 3 |
| chr19_+_52264449 | 0.01 |
ENST00000599326.1 ENST00000598953.1 |
FPR2 |
formyl peptide receptor 2 |
| chr3_-_151047327 | 0.01 |
ENST00000325602.5 |
P2RY13 |
purinergic receptor P2Y, G-protein coupled, 13 |
| chr14_+_67707826 | 0.01 |
ENST00000261681.4 |
MPP5 |
membrane protein, palmitoylated 5 (MAGUK p55 subfamily member 5) |
| chr8_-_30670384 | 0.01 |
ENST00000221138.4 ENST00000518243.1 |
PPP2CB |
protein phosphatase 2, catalytic subunit, beta isozyme |
| chrX_-_122866874 | 0.01 |
ENST00000245838.8 ENST00000355725.4 |
THOC2 |
THO complex 2 |
| chr1_+_46049706 | 0.00 |
ENST00000527470.1 ENST00000525515.1 ENST00000537798.1 ENST00000402363.3 ENST00000528238.1 ENST00000350030.3 ENST00000470768.1 ENST00000372052.4 ENST00000351223.3 |
NASP |
nuclear autoantigenic sperm protein (histone-binding) |
| chr11_-_59951889 | 0.00 |
ENST00000532169.1 ENST00000534596.1 |
MS4A6A |
membrane-spanning 4-domains, subfamily A, member 6A |
| chr9_-_28026318 | 0.00 |
ENST00000308675.3 |
LINGO2 |
leucine rich repeat and Ig domain containing 2 |
| chr11_+_60145948 | 0.00 |
ENST00000300184.3 ENST00000358246.1 |
MS4A7 |
membrane-spanning 4-domains, subfamily A, member 7 |
| chr3_-_196242233 | 0.00 |
ENST00000397537.2 |
SMCO1 |
single-pass membrane protein with coiled-coil domains 1 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.0 | GO:0060708 | spongiotrophoblast differentiation(GO:0060708) |
| 0.1 | 0.3 | GO:0048633 | negative regulation of auditory receptor cell differentiation(GO:0045608) positive regulation of skeletal muscle tissue growth(GO:0048633) |
| 0.1 | 0.2 | GO:0042214 | terpene metabolic process(GO:0042214) |
| 0.1 | 0.3 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.1 | 0.3 | GO:0043988 | histone H3-S28 phosphorylation(GO:0043988) |
| 0.1 | 0.4 | GO:0040009 | regulation of growth rate(GO:0040009) |
| 0.1 | 0.7 | GO:0032808 | lacrimal gland development(GO:0032808) |
| 0.1 | 0.2 | GO:0072560 | glandular epithelial cell maturation(GO:0002071) regulation of transcription from RNA polymerase II promoter involved in ventral spinal cord interneuron specification(GO:0021913) type B pancreatic cell maturation(GO:0072560) |
| 0.1 | 0.5 | GO:0002074 | extraocular skeletal muscle development(GO:0002074) |
| 0.0 | 0.2 | GO:0061198 | fungiform papilla formation(GO:0061198) |
| 0.0 | 0.2 | GO:0048861 | leukemia inhibitory factor signaling pathway(GO:0048861) |
| 0.0 | 0.1 | GO:0018153 | isopeptide cross-linking via N6-(L-isoglutamyl)-L-lysine(GO:0018153) isopeptide cross-linking(GO:0018262) |
| 0.0 | 0.3 | GO:0061669 | spontaneous neurotransmitter secretion(GO:0061669) spontaneous synaptic transmission(GO:0098814) |
| 0.0 | 0.5 | GO:0006657 | CDP-choline pathway(GO:0006657) |
| 0.0 | 0.2 | GO:0051754 | meiotic sister chromatid cohesion, centromeric(GO:0051754) |
| 0.0 | 0.1 | GO:0060748 | tertiary branching involved in mammary gland duct morphogenesis(GO:0060748) |
| 0.0 | 0.2 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 0.0 | 0.1 | GO:0050717 | positive regulation of interleukin-1 alpha secretion(GO:0050717) |
| 0.0 | 0.3 | GO:0051451 | myoblast migration(GO:0051451) |
| 0.0 | 0.2 | GO:0015747 | urate transport(GO:0015747) |
| 0.0 | 0.3 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.0 | 0.1 | GO:0017186 | peptidyl-pyroglutamic acid biosynthetic process, using glutaminyl-peptide cyclotransferase(GO:0017186) |
| 0.0 | 0.2 | GO:0032968 | positive regulation of transcription elongation from RNA polymerase II promoter(GO:0032968) |
| 0.0 | 0.1 | GO:1901097 | negative regulation of autophagosome maturation(GO:1901097) |
| 0.0 | 0.1 | GO:0055009 | atrial cardiac muscle tissue development(GO:0003228) atrial cardiac muscle tissue morphogenesis(GO:0055009) |
| 0.0 | 0.1 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.0 | 0.1 | GO:0010968 | regulation of microtubule nucleation(GO:0010968) |
| 0.0 | 0.0 | GO:0042138 | meiotic DNA double-strand break formation(GO:0042138) |
| 0.0 | 0.2 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.0 | 0.2 | GO:0010510 | regulation of acetyl-CoA biosynthetic process from pyruvate(GO:0010510) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0005674 | transcription factor TFIIF complex(GO:0005674) |
| 0.1 | 0.2 | GO:0097196 | Shu complex(GO:0097196) |
| 0.0 | 1.0 | GO:0005922 | connexon complex(GO:0005922) |
| 0.0 | 0.1 | GO:1990423 | RZZ complex(GO:1990423) |
| 0.0 | 0.2 | GO:0072558 | NLRP1 inflammasome complex(GO:0072558) |
| 0.0 | 0.9 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.6 | GO:0032982 | myosin filament(GO:0032982) |
| 0.0 | 0.2 | GO:0060293 | P granule(GO:0043186) pole plasm(GO:0045495) germ plasm(GO:0060293) |
| 0.0 | 0.3 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.0 | 0.2 | GO:0000778 | condensed nuclear chromosome kinetochore(GO:0000778) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 0.1 | 0.2 | GO:0004531 | deoxyribonuclease II activity(GO:0004531) |
| 0.1 | 0.3 | GO:0004809 | tRNA (guanine-N2-)-methyltransferase activity(GO:0004809) |
| 0.1 | 0.2 | GO:0004923 | leukemia inhibitory factor receptor activity(GO:0004923) |
| 0.1 | 0.9 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.0 | 1.0 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.0 | 0.2 | GO:0004741 | [pyruvate dehydrogenase (lipoamide)] phosphatase activity(GO:0004741) |
| 0.0 | 0.2 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.0 | 0.1 | GO:0030144 | alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase activity(GO:0030144) |
| 0.0 | 0.3 | GO:0008199 | ferric iron binding(GO:0008199) |
| 0.0 | 0.1 | GO:0017018 | myosin phosphatase activity(GO:0017018) |
| 0.0 | 0.2 | GO:0000990 | transcription factor activity, core RNA polymerase binding(GO:0000990) |
| 0.0 | 0.3 | GO:0030957 | Tat protein binding(GO:0030957) |
| 0.0 | 0.2 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.0 | 0.2 | GO:1901702 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.0 | 0.1 | GO:0033906 | hyaluronoglucuronidase activity(GO:0033906) |
| 0.0 | 0.1 | GO:0016603 | glutaminyl-peptide cyclotransferase activity(GO:0016603) |
| 0.0 | 0.5 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.0 | 1.3 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.0 | 0.3 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 0.1 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.0 | 0.2 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 0.1 | GO:0019237 | centromeric DNA binding(GO:0019237) |
| 0.0 | 1.4 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.0 | 0.1 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.9 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 0.3 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.0 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.0 | 0.9 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.0 | 0.3 | REACTOME RECEPTOR LIGAND BINDING INITIATES THE SECOND PROTEOLYTIC CLEAVAGE OF NOTCH RECEPTOR | Genes involved in Receptor-ligand binding initiates the second proteolytic cleavage of Notch receptor |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.0 | 0.2 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |