Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
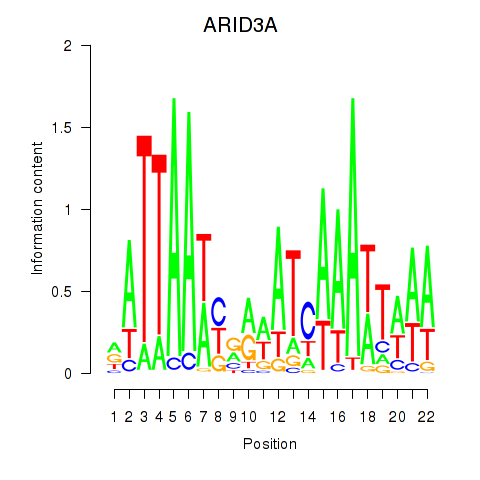
Results for ARID3A
Z-value: 1.02
Transcription factors associated with ARID3A
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ARID3A
|
ENSG00000116017.6 | ARID3A |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ARID3A | hg19_v2_chr19_+_926000_926046 | -0.84 | 9.1e-03 | Click! |
Activity profile of ARID3A motif
Sorted Z-values of ARID3A motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ARID3A
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr18_+_61554932 | 3.19 |
ENST00000299502.4 ENST00000457692.1 ENST00000413956.1 |
SERPINB2 |
serpin peptidase inhibitor, clade B (ovalbumin), member 2 |
| chr1_+_77333117 | 1.96 |
ENST00000477717.1 |
ST6GALNAC5 |
ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 5 |
| chr2_-_165424973 | 1.82 |
ENST00000543549.1 |
GRB14 |
growth factor receptor-bound protein 14 |
| chr17_-_3599696 | 1.62 |
ENST00000225328.5 |
P2RX5 |
purinergic receptor P2X, ligand-gated ion channel, 5 |
| chr17_-_3599327 | 1.60 |
ENST00000551178.1 ENST00000552276.1 ENST00000547178.1 |
P2RX5 |
purinergic receptor P2X, ligand-gated ion channel, 5 |
| chr17_-_3599492 | 1.56 |
ENST00000435558.1 ENST00000345901.3 |
P2RX5 |
purinergic receptor P2X, ligand-gated ion channel, 5 |
| chr5_+_140220769 | 1.35 |
ENST00000531613.1 ENST00000378123.3 |
PCDHA8 |
protocadherin alpha 8 |
| chr6_-_136847099 | 1.27 |
ENST00000438100.2 |
MAP7 |
microtubule-associated protein 7 |
| chr12_-_28125638 | 1.19 |
ENST00000545234.1 |
PTHLH |
parathyroid hormone-like hormone |
| chr16_+_76587314 | 1.17 |
ENST00000563764.1 |
RP11-58C22.1 |
Uncharacterized protein |
| chr8_+_18248786 | 1.07 |
ENST00000520116.1 |
NAT2 |
N-acetyltransferase 2 (arylamine N-acetyltransferase) |
| chr8_+_18248755 | 1.06 |
ENST00000286479.3 |
NAT2 |
N-acetyltransferase 2 (arylamine N-acetyltransferase) |
| chr20_+_58179582 | 1.06 |
ENST00000371015.1 ENST00000395639.4 |
PHACTR3 |
phosphatase and actin regulator 3 |
| chr1_-_207206092 | 0.96 |
ENST00000359470.5 ENST00000461135.2 |
C1orf116 |
chromosome 1 open reading frame 116 |
| chr11_+_67351213 | 0.85 |
ENST00000398603.1 |
GSTP1 |
glutathione S-transferase pi 1 |
| chrX_-_114252193 | 0.85 |
ENST00000243213.1 |
IL13RA2 |
interleukin 13 receptor, alpha 2 |
| chr6_+_47624172 | 0.83 |
ENST00000507065.1 ENST00000296862.1 |
GPR111 |
G protein-coupled receptor 111 |
| chr11_+_67351019 | 0.78 |
ENST00000398606.3 |
GSTP1 |
glutathione S-transferase pi 1 |
| chr7_-_139763521 | 0.76 |
ENST00000263549.3 |
PARP12 |
poly (ADP-ribose) polymerase family, member 12 |
| chr7_+_107224364 | 0.73 |
ENST00000491150.1 |
BCAP29 |
B-cell receptor-associated protein 29 |
| chr17_-_56494713 | 0.71 |
ENST00000407977.2 |
RNF43 |
ring finger protein 43 |
| chr17_-_56494882 | 0.71 |
ENST00000584437.1 |
RNF43 |
ring finger protein 43 |
| chr17_-_56494908 | 0.71 |
ENST00000577716.1 |
RNF43 |
ring finger protein 43 |
| chr12_-_95044309 | 0.68 |
ENST00000261226.4 |
TMCC3 |
transmembrane and coiled-coil domain family 3 |
| chr3_+_36421826 | 0.68 |
ENST00000273183.3 |
STAC |
SH3 and cysteine rich domain |
| chr4_+_100737954 | 0.67 |
ENST00000296414.7 ENST00000512369.1 |
DAPP1 |
dual adaptor of phosphotyrosine and 3-phosphoinositides |
| chr19_-_55652290 | 0.66 |
ENST00000589745.1 |
TNNT1 |
troponin T type 1 (skeletal, slow) |
| chr11_+_18154059 | 0.64 |
ENST00000531264.1 |
MRGPRX3 |
MAS-related GPR, member X3 |
| chr2_+_106468204 | 0.61 |
ENST00000425756.1 ENST00000393349.2 |
NCK2 |
NCK adaptor protein 2 |
| chr7_-_22539771 | 0.60 |
ENST00000406890.2 ENST00000424363.1 |
STEAP1B |
STEAP family member 1B |
| chr8_+_32579341 | 0.59 |
ENST00000519240.1 ENST00000539990.1 |
NRG1 |
neuregulin 1 |
| chrX_-_117119243 | 0.58 |
ENST00000539496.1 ENST00000469946.1 |
KLHL13 |
kelch-like family member 13 |
| chr5_+_140180635 | 0.57 |
ENST00000522353.2 ENST00000532566.2 |
PCDHA3 |
protocadherin alpha 3 |
| chr4_-_76957214 | 0.55 |
ENST00000306621.3 |
CXCL11 |
chemokine (C-X-C motif) ligand 11 |
| chr1_+_207943667 | 0.53 |
ENST00000462968.2 |
CD46 |
CD46 molecule, complement regulatory protein |
| chr19_-_48614063 | 0.52 |
ENST00000599921.1 ENST00000599111.1 |
PLA2G4C |
phospholipase A2, group IVC (cytosolic, calcium-independent) |
| chr19_-_48614033 | 0.51 |
ENST00000354276.3 |
PLA2G4C |
phospholipase A2, group IVC (cytosolic, calcium-independent) |
| chr7_-_38305279 | 0.51 |
ENST00000443402.2 |
TRGC1 |
T cell receptor gamma constant 1 |
| chr16_+_19421803 | 0.47 |
ENST00000541464.1 |
TMC5 |
transmembrane channel-like 5 |
| chr6_+_121756809 | 0.46 |
ENST00000282561.3 |
GJA1 |
gap junction protein, alpha 1, 43kDa |
| chr21_+_34442439 | 0.46 |
ENST00000382348.1 ENST00000333063.5 |
OLIG1 |
oligodendrocyte transcription factor 1 |
| chr12_+_113344582 | 0.45 |
ENST00000202917.5 ENST00000445409.2 ENST00000452357.2 |
OAS1 |
2'-5'-oligoadenylate synthetase 1, 40/46kDa |
| chr4_-_80994619 | 0.44 |
ENST00000404191.1 |
ANTXR2 |
anthrax toxin receptor 2 |
| chr6_-_116381918 | 0.43 |
ENST00000606080.1 |
FRK |
fyn-related kinase |
| chr8_+_104892639 | 0.43 |
ENST00000436393.2 |
RIMS2 |
regulating synaptic membrane exocytosis 2 |
| chr4_-_80994210 | 0.43 |
ENST00000403729.2 |
ANTXR2 |
anthrax toxin receptor 2 |
| chr7_+_89783689 | 0.41 |
ENST00000297205.2 |
STEAP1 |
six transmembrane epithelial antigen of the prostate 1 |
| chr2_+_191002486 | 0.40 |
ENST00000396974.2 |
C2orf88 |
chromosome 2 open reading frame 88 |
| chr4_+_74301880 | 0.40 |
ENST00000395792.2 ENST00000226359.2 |
AFP |
alpha-fetoprotein |
| chr1_-_100231349 | 0.39 |
ENST00000287474.5 ENST00000414213.1 |
FRRS1 |
ferric-chelate reductase 1 |
| chr3_+_118892362 | 0.39 |
ENST00000497685.1 ENST00000264234.3 |
UPK1B |
uroplakin 1B |
| chr17_-_15466742 | 0.38 |
ENST00000584811.1 ENST00000419890.2 |
TVP23C |
trans-golgi network vesicle protein 23 homolog C (S. cerevisiae) |
| chr12_-_39734783 | 0.38 |
ENST00000552961.1 |
KIF21A |
kinesin family member 21A |
| chr4_+_71200681 | 0.38 |
ENST00000273936.5 |
CABS1 |
calcium-binding protein, spermatid-specific 1 |
| chr2_-_130031335 | 0.38 |
ENST00000375987.3 |
AC079586.1 |
AC079586.1 |
| chr14_+_39734482 | 0.37 |
ENST00000554392.1 ENST00000555716.1 ENST00000341749.3 ENST00000557038.1 |
CTAGE5 |
CTAGE family, member 5 |
| chr17_+_7531281 | 0.36 |
ENST00000575729.1 ENST00000340624.5 |
SHBG |
sex hormone-binding globulin |
| chr4_-_80994471 | 0.36 |
ENST00000295465.4 |
ANTXR2 |
anthrax toxin receptor 2 |
| chr1_-_235098935 | 0.35 |
ENST00000423175.1 |
RP11-443B7.1 |
RP11-443B7.1 |
| chr2_-_160473114 | 0.35 |
ENST00000392783.2 |
BAZ2B |
bromodomain adjacent to zinc finger domain, 2B |
| chr19_+_9296279 | 0.35 |
ENST00000344248.2 |
OR7D2 |
olfactory receptor, family 7, subfamily D, member 2 |
| chr4_-_170948361 | 0.34 |
ENST00000393702.3 |
MFAP3L |
microfibrillar-associated protein 3-like |
| chr11_+_112046190 | 0.34 |
ENST00000357685.5 ENST00000393032.2 ENST00000361053.4 |
BCO2 |
beta-carotene oxygenase 2 |
| chr4_+_74606223 | 0.34 |
ENST00000307407.3 ENST00000401931.1 |
IL8 |
interleukin 8 |
| chr15_+_75491213 | 0.32 |
ENST00000360639.2 |
C15orf39 |
chromosome 15 open reading frame 39 |
| chr11_+_118175132 | 0.31 |
ENST00000361763.4 |
CD3E |
CD3e molecule, epsilon (CD3-TCR complex) |
| chr17_+_18684563 | 0.30 |
ENST00000476139.1 |
TVP23B |
trans-golgi network vesicle protein 23 homolog B (S. cerevisiae) |
| chr16_+_19422035 | 0.30 |
ENST00000381414.4 ENST00000396229.2 |
TMC5 |
transmembrane channel-like 5 |
| chr9_-_21351377 | 0.29 |
ENST00000380210.1 |
IFNA6 |
interferon, alpha 6 |
| chr11_+_45825616 | 0.29 |
ENST00000442528.2 ENST00000456334.1 ENST00000526817.1 |
SLC35C1 |
solute carrier family 35 (GDP-fucose transporter), member C1 |
| chr6_-_46703430 | 0.28 |
ENST00000537365.1 |
PLA2G7 |
phospholipase A2, group VII (platelet-activating factor acetylhydrolase, plasma) |
| chr17_-_29641104 | 0.28 |
ENST00000577894.1 ENST00000330927.4 |
EVI2B |
ecotropic viral integration site 2B |
| chr17_-_18430160 | 0.27 |
ENST00000392176.3 |
FAM106A |
family with sequence similarity 106, member A |
| chr16_+_81812863 | 0.27 |
ENST00000359376.3 |
PLCG2 |
phospholipase C, gamma 2 (phosphatidylinositol-specific) |
| chr10_+_5238793 | 0.27 |
ENST00000263126.1 |
AKR1C4 |
aldo-keto reductase family 1, member C4 |
| chr7_-_100253993 | 0.27 |
ENST00000461605.1 ENST00000160382.5 |
ACTL6B |
actin-like 6B |
| chr11_+_35201826 | 0.26 |
ENST00000531873.1 |
CD44 |
CD44 molecule (Indian blood group) |
| chr18_+_61144160 | 0.26 |
ENST00000489441.1 ENST00000424602.1 |
SERPINB5 |
serpin peptidase inhibitor, clade B (ovalbumin), member 5 |
| chr3_-_139396853 | 0.26 |
ENST00000406164.1 ENST00000406824.1 |
NMNAT3 |
nicotinamide nucleotide adenylyltransferase 3 |
| chr7_-_38289173 | 0.26 |
ENST00000436911.2 |
TRGC2 |
T cell receptor gamma constant 2 |
| chr10_+_26727125 | 0.26 |
ENST00000376236.4 |
APBB1IP |
amyloid beta (A4) precursor protein-binding, family B, member 1 interacting protein |
| chr7_-_92855762 | 0.26 |
ENST00000453812.2 ENST00000394468.2 |
HEPACAM2 |
HEPACAM family member 2 |
| chr6_+_111303218 | 0.25 |
ENST00000441448.2 |
RPF2 |
ribosome production factor 2 homolog (S. cerevisiae) |
| chr3_+_43328004 | 0.25 |
ENST00000454177.1 ENST00000429705.2 ENST00000296088.7 ENST00000437827.1 |
SNRK |
SNF related kinase |
| chr22_+_18721427 | 0.25 |
ENST00000342888.3 |
AC008132.1 |
Uncharacterized protein |
| chr12_+_113682066 | 0.24 |
ENST00000392569.4 ENST00000552542.1 |
TPCN1 |
two pore segment channel 1 |
| chr17_-_41050716 | 0.24 |
ENST00000417193.1 ENST00000301683.3 ENST00000436546.1 ENST00000431109.2 |
LINC00671 |
long intergenic non-protein coding RNA 671 |
| chr7_+_76090993 | 0.23 |
ENST00000425780.1 ENST00000456590.1 ENST00000451769.1 ENST00000324432.5 ENST00000307569.8 ENST00000457529.1 ENST00000446600.1 ENST00000413936.2 ENST00000423646.1 ENST00000438930.1 ENST00000430490.2 |
DTX2 |
deltex homolog 2 (Drosophila) |
| chr8_+_86099884 | 0.23 |
ENST00000517476.1 ENST00000521429.1 |
E2F5 |
E2F transcription factor 5, p130-binding |
| chr9_-_21385396 | 0.23 |
ENST00000380206.2 |
IFNA2 |
interferon, alpha 2 |
| chr12_+_122688090 | 0.23 |
ENST00000324189.4 ENST00000546192.1 |
B3GNT4 |
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 4 |
| chr3_-_139396801 | 0.23 |
ENST00000413939.2 ENST00000339837.5 ENST00000512391.1 |
NMNAT3 |
nicotinamide nucleotide adenylyltransferase 3 |
| chr8_-_62559366 | 0.23 |
ENST00000522919.1 |
ASPH |
aspartate beta-hydroxylase |
| chr1_-_39395165 | 0.23 |
ENST00000372985.3 |
RHBDL2 |
rhomboid, veinlet-like 2 (Drosophila) |
| chr19_+_11200038 | 0.23 |
ENST00000558518.1 ENST00000557933.1 ENST00000455727.2 ENST00000535915.1 ENST00000545707.1 ENST00000558013.1 |
LDLR |
low density lipoprotein receptor |
| chr11_-_5255861 | 0.22 |
ENST00000380299.3 |
HBD |
hemoglobin, delta |
| chr1_+_196743912 | 0.22 |
ENST00000367425.4 |
CFHR3 |
complement factor H-related 3 |
| chr11_+_112047087 | 0.22 |
ENST00000526088.1 ENST00000532593.1 ENST00000531169.1 |
BCO2 |
beta-carotene oxygenase 2 |
| chr6_-_29324054 | 0.22 |
ENST00000543825.1 |
OR5V1 |
olfactory receptor, family 5, subfamily V, member 1 |
| chr5_+_126988732 | 0.21 |
ENST00000395322.3 |
CTXN3 |
cortexin 3 |
| chr8_-_87242589 | 0.21 |
ENST00000419776.2 ENST00000297524.3 |
SLC7A13 |
solute carrier family 7 (anionic amino acid transporter), member 13 |
| chr1_-_235098861 | 0.21 |
ENST00000458044.1 |
RP11-443B7.1 |
RP11-443B7.1 |
| chr14_+_97263641 | 0.21 |
ENST00000216639.3 |
VRK1 |
vaccinia related kinase 1 |
| chr12_+_69186125 | 0.21 |
ENST00000399333.3 |
AC124890.1 |
HCG1774533, isoform CRA_a; PRO2268; Uncharacterized protein |
| chr12_+_119772502 | 0.21 |
ENST00000536742.1 ENST00000327554.2 |
CCDC60 |
coiled-coil domain containing 60 |
| chr18_-_13915530 | 0.21 |
ENST00000327606.3 |
MC2R |
melanocortin 2 receptor (adrenocorticotropic hormone) |
| chr22_-_40289759 | 0.21 |
ENST00000325157.6 |
ENTHD1 |
ENTH domain containing 1 |
| chr1_+_196743943 | 0.20 |
ENST00000471440.2 ENST00000391985.3 |
CFHR3 |
complement factor H-related 3 |
| chr3_-_191000172 | 0.20 |
ENST00000427544.2 |
UTS2B |
urotensin 2B |
| chr6_+_24403144 | 0.20 |
ENST00000274747.7 ENST00000543597.1 ENST00000535061.1 ENST00000378353.1 ENST00000378386.3 ENST00000443868.2 |
MRS2 |
MRS2 magnesium transporter |
| chr6_+_31105426 | 0.19 |
ENST00000547221.1 |
PSORS1C1 |
psoriasis susceptibility 1 candidate 1 |
| chr2_+_109335929 | 0.19 |
ENST00000283195.6 |
RANBP2 |
RAN binding protein 2 |
| chr4_-_57844989 | 0.19 |
ENST00000264230.4 |
NOA1 |
nitric oxide associated 1 |
| chr1_+_73771844 | 0.19 |
ENST00000440762.1 ENST00000444827.1 ENST00000415686.1 ENST00000411903.1 |
RP4-598G3.1 |
RP4-598G3.1 |
| chr14_-_53162361 | 0.19 |
ENST00000395686.3 |
ERO1L |
ERO1-like (S. cerevisiae) |
| chr3_+_142442841 | 0.18 |
ENST00000476941.1 ENST00000273482.6 |
TRPC1 |
transient receptor potential cation channel, subfamily C, member 1 |
| chr4_-_169239921 | 0.18 |
ENST00000514995.1 ENST00000393743.3 |
DDX60 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 60 |
| chr4_-_68749745 | 0.18 |
ENST00000283916.6 |
TMPRSS11D |
transmembrane protease, serine 11D |
| chr15_-_49103235 | 0.18 |
ENST00000380950.2 |
CEP152 |
centrosomal protein 152kDa |
| chr19_+_45542295 | 0.18 |
ENST00000221455.3 ENST00000391953.4 ENST00000588936.1 |
CLASRP |
CLK4-associating serine/arginine rich protein |
| chr6_+_26087509 | 0.18 |
ENST00000397022.3 ENST00000353147.5 ENST00000352392.4 ENST00000349999.4 ENST00000317896.7 ENST00000357618.5 ENST00000470149.1 ENST00000336625.8 ENST00000461397.1 ENST00000488199.1 |
HFE |
hemochromatosis |
| chr12_-_11287243 | 0.17 |
ENST00000539585.1 |
TAS2R30 |
taste receptor, type 2, member 30 |
| chr12_-_120765565 | 0.17 |
ENST00000423423.3 ENST00000308366.4 |
PLA2G1B |
phospholipase A2, group IB (pancreas) |
| chrX_+_139791917 | 0.17 |
ENST00000607004.1 ENST00000370535.3 |
LINC00632 |
long intergenic non-protein coding RNA 632 |
| chr12_-_4488872 | 0.17 |
ENST00000237837.1 |
FGF23 |
fibroblast growth factor 23 |
| chr19_+_11670245 | 0.17 |
ENST00000585493.1 |
ZNF627 |
zinc finger protein 627 |
| chr8_-_102218292 | 0.17 |
ENST00000518336.1 ENST00000520454.1 |
ZNF706 |
zinc finger protein 706 |
| chr12_-_113772835 | 0.17 |
ENST00000552014.1 ENST00000548186.1 ENST00000202831.3 ENST00000549181.1 |
SLC8B1 |
solute carrier family 8 (sodium/lithium/calcium exchanger), member B1 |
| chr16_+_30935418 | 0.17 |
ENST00000338343.4 |
FBXL19 |
F-box and leucine-rich repeat protein 19 |
| chr14_+_70918874 | 0.17 |
ENST00000603540.1 |
ADAM21 |
ADAM metallopeptidase domain 21 |
| chr2_-_219031709 | 0.17 |
ENST00000295683.2 |
CXCR1 |
chemokine (C-X-C motif) receptor 1 |
| chr4_+_15341442 | 0.16 |
ENST00000397700.2 ENST00000295297.4 |
C1QTNF7 |
C1q and tumor necrosis factor related protein 7 |
| chr16_+_56970567 | 0.16 |
ENST00000563911.1 |
HERPUD1 |
homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 |
| chr12_+_32832134 | 0.16 |
ENST00000452533.2 |
DNM1L |
dynamin 1-like |
| chr9_-_19786926 | 0.16 |
ENST00000341998.2 ENST00000286344.3 |
SLC24A2 |
solute carrier family 24 (sodium/potassium/calcium exchanger), member 2 |
| chrX_+_83116142 | 0.15 |
ENST00000329312.4 |
CYLC1 |
cylicin, basic protein of sperm head cytoskeleton 1 |
| chrX_-_2882296 | 0.15 |
ENST00000438544.1 ENST00000381134.3 ENST00000545496.1 |
ARSE |
arylsulfatase E (chondrodysplasia punctata 1) |
| chr4_-_96470108 | 0.15 |
ENST00000513796.1 |
UNC5C |
unc-5 homolog C (C. elegans) |
| chr12_-_71182695 | 0.15 |
ENST00000342084.4 |
PTPRR |
protein tyrosine phosphatase, receptor type, R |
| chr4_+_159131630 | 0.15 |
ENST00000504569.1 ENST00000509278.1 ENST00000514558.1 ENST00000503200.1 |
TMEM144 |
transmembrane protein 144 |
| chr10_-_11574274 | 0.15 |
ENST00000277575.5 |
USP6NL |
USP6 N-terminal like |
| chr4_-_100356291 | 0.14 |
ENST00000476959.1 ENST00000482593.1 |
ADH7 |
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide |
| chr1_-_55230165 | 0.14 |
ENST00000371279.3 |
PARS2 |
prolyl-tRNA synthetase 2, mitochondrial (putative) |
| chr16_-_84220604 | 0.14 |
ENST00000567759.1 |
TAF1C |
TATA box binding protein (TBP)-associated factor, RNA polymerase I, C, 110kDa |
| chr14_+_22694606 | 0.14 |
ENST00000390463.3 |
TRAV36DV7 |
T cell receptor alpha variable 36/delta variable 7 |
| chr12_+_32832203 | 0.14 |
ENST00000553257.1 ENST00000549701.1 ENST00000358214.5 ENST00000266481.6 ENST00000551476.1 ENST00000550154.1 ENST00000547312.1 ENST00000414834.2 ENST00000381000.4 ENST00000548750.1 |
DNM1L |
dynamin 1-like |
| chr5_-_75013193 | 0.14 |
ENST00000514838.2 ENST00000506164.1 ENST00000502826.1 ENST00000503835.1 ENST00000428202.2 ENST00000380475.2 |
POC5 |
POC5 centriolar protein |
| chr4_-_69434245 | 0.14 |
ENST00000317746.2 |
UGT2B17 |
UDP glucuronosyltransferase 2 family, polypeptide B17 |
| chr7_+_115862858 | 0.14 |
ENST00000393481.2 |
TES |
testis derived transcript (3 LIM domains) |
| chr4_-_110723194 | 0.14 |
ENST00000394635.3 |
CFI |
complement factor I |
| chr3_-_139396787 | 0.14 |
ENST00000296202.7 ENST00000509291.1 |
NMNAT3 |
nicotinamide nucleotide adenylyltransferase 3 |
| chrX_+_21857717 | 0.14 |
ENST00000379484.5 |
MBTPS2 |
membrane-bound transcription factor peptidase, site 2 |
| chr11_-_5207612 | 0.13 |
ENST00000380367.1 |
OR52A1 |
olfactory receptor, family 52, subfamily A, member 1 |
| chr16_-_11375179 | 0.13 |
ENST00000312511.3 |
PRM1 |
protamine 1 |
| chr6_+_26087646 | 0.13 |
ENST00000309234.6 |
HFE |
hemochromatosis |
| chr1_-_205391178 | 0.13 |
ENST00000367153.4 ENST00000367151.2 ENST00000391936.2 ENST00000367149.3 |
LEMD1 |
LEM domain containing 1 |
| chr6_-_100016678 | 0.13 |
ENST00000523799.1 ENST00000520429.1 |
CCNC |
cyclin C |
| chr11_-_5345582 | 0.13 |
ENST00000328813.2 |
OR51B2 |
olfactory receptor, family 51, subfamily B, member 2 |
| chrX_+_134975858 | 0.13 |
ENST00000537770.1 |
SAGE1 |
sarcoma antigen 1 |
| chrX_+_134975753 | 0.13 |
ENST00000535938.1 |
SAGE1 |
sarcoma antigen 1 |
| chr4_+_57845043 | 0.13 |
ENST00000433463.1 ENST00000314595.5 |
POLR2B |
polymerase (RNA) II (DNA directed) polypeptide B, 140kDa |
| chr18_-_64271363 | 0.12 |
ENST00000262150.2 |
CDH19 |
cadherin 19, type 2 |
| chr6_-_100016527 | 0.12 |
ENST00000523985.1 ENST00000518714.1 ENST00000520371.1 |
CCNC |
cyclin C |
| chr19_-_55458860 | 0.12 |
ENST00000592784.1 ENST00000448121.2 ENST00000340844.2 |
NLRP7 |
NLR family, pyrin domain containing 7 |
| chr8_-_59412717 | 0.12 |
ENST00000301645.3 |
CYP7A1 |
cytochrome P450, family 7, subfamily A, polypeptide 1 |
| chr3_+_57875738 | 0.12 |
ENST00000417128.1 ENST00000438794.1 |
SLMAP |
sarcolemma associated protein |
| chr18_-_19994830 | 0.12 |
ENST00000525417.1 |
CTAGE1 |
cutaneous T-cell lymphoma-associated antigen 1 |
| chr13_-_102068769 | 0.12 |
ENST00000376196.3 |
NALCN |
sodium leak channel, non-selective |
| chr16_+_84801852 | 0.12 |
ENST00000569925.1 ENST00000567526.1 |
USP10 |
ubiquitin specific peptidase 10 |
| chr10_+_62538089 | 0.12 |
ENST00000519078.2 ENST00000395284.3 ENST00000316629.4 |
CDK1 |
cyclin-dependent kinase 1 |
| chr15_-_45670924 | 0.12 |
ENST00000396659.3 |
GATM |
glycine amidinotransferase (L-arginine:glycine amidinotransferase) |
| chr4_-_110723335 | 0.12 |
ENST00000394634.2 |
CFI |
complement factor I |
| chr9_-_86571628 | 0.11 |
ENST00000376344.3 |
C9orf64 |
chromosome 9 open reading frame 64 |
| chr10_+_7745232 | 0.11 |
ENST00000358415.4 |
ITIH2 |
inter-alpha-trypsin inhibitor heavy chain 2 |
| chr10_+_118305435 | 0.11 |
ENST00000369221.2 |
PNLIP |
pancreatic lipase |
| chr1_+_53308398 | 0.11 |
ENST00000371528.1 |
ZYG11A |
zyg-11 family member A, cell cycle regulator |
| chr10_+_7745303 | 0.11 |
ENST00000429820.1 ENST00000379587.4 |
ITIH2 |
inter-alpha-trypsin inhibitor heavy chain 2 |
| chr2_+_102413726 | 0.11 |
ENST00000350878.4 |
MAP4K4 |
mitogen-activated protein kinase kinase kinase kinase 4 |
| chr16_+_30034655 | 0.11 |
ENST00000300575.2 |
C16orf92 |
chromosome 16 open reading frame 92 |
| chr4_+_5527117 | 0.11 |
ENST00000505296.1 |
C4orf6 |
chromosome 4 open reading frame 6 |
| chr17_-_27333311 | 0.11 |
ENST00000317338.12 ENST00000585644.1 |
SEZ6 |
seizure related 6 homolog (mouse) |
| chr9_-_27005686 | 0.11 |
ENST00000380055.5 |
LRRC19 |
leucine rich repeat containing 19 |
| chr16_-_84220633 | 0.11 |
ENST00000566732.1 ENST00000561955.1 ENST00000564454.1 ENST00000341690.6 ENST00000541676.1 ENST00000570117.1 ENST00000564345.1 |
TAF1C |
TATA box binding protein (TBP)-associated factor, RNA polymerase I, C, 110kDa |
| chr1_-_31661000 | 0.11 |
ENST00000263693.1 ENST00000398657.2 ENST00000526106.1 |
NKAIN1 |
Na+/K+ transporting ATPase interacting 1 |
| chr17_-_15466850 | 0.11 |
ENST00000438826.3 ENST00000225576.3 ENST00000519970.1 ENST00000518321.1 ENST00000428082.2 ENST00000522212.2 |
TVP23C TVP23C-CDRT4 |
trans-golgi network vesicle protein 23 homolog C (S. cerevisiae) TVP23C-CDRT4 readthrough |
| chr3_+_169629354 | 0.10 |
ENST00000428432.2 ENST00000335556.3 |
SAMD7 |
sterile alpha motif domain containing 7 |
| chr11_+_22688150 | 0.10 |
ENST00000454584.2 |
GAS2 |
growth arrest-specific 2 |
| chr4_-_185275104 | 0.10 |
ENST00000317596.3 |
RP11-290F5.2 |
RP11-290F5.2 |
| chrY_-_20935572 | 0.10 |
ENST00000382852.1 ENST00000344884.4 ENST00000304790.3 |
HSFY2 |
heat shock transcription factor, Y linked 2 |
| chr4_+_5526883 | 0.10 |
ENST00000195455.2 |
C4orf6 |
chromosome 4 open reading frame 6 |
| chr7_-_33080506 | 0.10 |
ENST00000381626.2 ENST00000409467.1 ENST00000449201.1 |
NT5C3A |
5'-nucleotidase, cytosolic IIIA |
| chr4_+_70796784 | 0.10 |
ENST00000246891.4 ENST00000444405.3 |
CSN1S1 |
casein alpha s1 |
| chr2_-_208994548 | 0.10 |
ENST00000282141.3 |
CRYGC |
crystallin, gamma C |
| chrY_+_20708557 | 0.10 |
ENST00000307393.2 ENST00000309834.4 ENST00000382856.2 |
HSFY1 |
heat shock transcription factor, Y-linked 1 |
| chr15_-_34880646 | 0.10 |
ENST00000543376.1 |
GOLGA8A |
golgin A8 family, member A |
| chr2_+_90121477 | 0.09 |
ENST00000483379.1 |
IGKV1D-17 |
immunoglobulin kappa variable 1D-17 |
| chr12_+_96883347 | 0.09 |
ENST00000524981.4 ENST00000298953.3 |
C12orf55 |
chromosome 12 open reading frame 55 |
| chrX_+_48242863 | 0.09 |
ENST00000376886.2 ENST00000375517.3 |
SSX4 |
synovial sarcoma, X breakpoint 4 |
| chr9_-_139343294 | 0.09 |
ENST00000313084.5 |
SEC16A |
SEC16 homolog A (S. cerevisiae) |
| chr5_+_140227357 | 0.09 |
ENST00000378122.3 |
PCDHA9 |
protocadherin alpha 9 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 2.1 | GO:0038018 | Wnt receptor catabolic process(GO:0038018) |
| 0.4 | 1.6 | GO:0051771 | negative regulation of nitric-oxide synthase biosynthetic process(GO:0051771) |
| 0.2 | 0.6 | GO:0042214 | terpene metabolic process(GO:0042214) |
| 0.2 | 0.5 | GO:0010645 | regulation of cell communication by chemical coupling(GO:0010645) positive regulation of cell communication by chemical coupling(GO:0010652) |
| 0.1 | 1.0 | GO:0098706 | ferric iron import into cell(GO:0097461) ferric iron import across plasma membrane(GO:0098706) |
| 0.1 | 4.4 | GO:0035590 | purinergic nucleotide receptor signaling pathway(GO:0035590) |
| 0.1 | 1.1 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.1 | 0.3 | GO:0002589 | regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002589) negative regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002590) regulation of T cell antigen processing and presentation(GO:0002625) positive regulation of iron ion transport(GO:0034758) positive regulation of iron ion transmembrane transport(GO:0034761) regulation of iron ion import(GO:1900390) regulation of ferrous iron import into cell(GO:1903989) positive regulation of ferrous iron import into cell(GO:1903991) regulation of ferrous iron binding(GO:1904432) positive regulation of ferrous iron binding(GO:1904434) regulation of transferrin receptor binding(GO:1904435) positive regulation of transferrin receptor binding(GO:1904437) regulation of ferrous iron import across plasma membrane(GO:1904438) positive regulation of ferrous iron import across plasma membrane(GO:1904440) response to iron ion starvation(GO:1990641) |
| 0.1 | 0.6 | GO:0036493 | positive regulation of translation in response to endoplasmic reticulum stress(GO:0036493) |
| 0.1 | 0.3 | GO:0090149 | synaptic vesicle recycling via endosome(GO:0036466) mitochondrial membrane fission(GO:0090149) |
| 0.1 | 0.8 | GO:0043305 | negative regulation of mast cell degranulation(GO:0043305) |
| 0.1 | 0.6 | GO:0021840 | directional guidance of interneurons involved in migration from the subpallium to the cortex(GO:0021840) chemorepulsion involved in interneuron migration from the subpallium to the cortex(GO:0021842) |
| 0.1 | 3.2 | GO:0042730 | fibrinolysis(GO:0042730) |
| 0.1 | 0.5 | GO:0034628 | nicotinamide nucleotide biosynthetic process from aspartate(GO:0019355) 'de novo' NAD biosynthetic process from aspartate(GO:0034628) |
| 0.1 | 2.0 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.1 | 0.7 | GO:0031444 | slow-twitch skeletal muscle fiber contraction(GO:0031444) |
| 0.1 | 0.3 | GO:0036079 | GDP-fucose transport(GO:0015783) purine nucleotide-sugar transport(GO:0036079) |
| 0.1 | 0.5 | GO:0043950 | positive regulation of cAMP-mediated signaling(GO:0043950) |
| 0.1 | 0.3 | GO:0002316 | follicular B cell differentiation(GO:0002316) |
| 0.1 | 0.3 | GO:0002913 | positive regulation of T cell anergy(GO:0002669) positive regulation of lymphocyte anergy(GO:0002913) |
| 0.1 | 1.2 | GO:0036149 | phosphatidylinositol acyl-chain remodeling(GO:0036149) |
| 0.1 | 0.2 | GO:0002725 | negative regulation of T cell cytokine production(GO:0002725) |
| 0.1 | 0.2 | GO:0042369 | vitamin D catabolic process(GO:0042369) |
| 0.1 | 0.2 | GO:0030311 | poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 0.1 | 0.2 | GO:0042264 | peptidyl-aspartic acid hydroxylation(GO:0042264) |
| 0.1 | 0.2 | GO:0090118 | receptor-mediated endocytosis of low-density lipoprotein particle involved in cholesterol transport(GO:0090118) positive regulation of lysosomal protein catabolic process(GO:1905167) |
| 0.1 | 0.3 | GO:0046469 | platelet activating factor metabolic process(GO:0046469) |
| 0.1 | 0.2 | GO:0098759 | response to interleukin-8(GO:0098758) cellular response to interleukin-8(GO:0098759) |
| 0.1 | 0.4 | GO:0098814 | spontaneous neurotransmitter secretion(GO:0061669) spontaneous synaptic transmission(GO:0098814) |
| 0.0 | 0.1 | GO:0035526 | retrograde transport, plasma membrane to Golgi(GO:0035526) |
| 0.0 | 0.1 | GO:0010430 | fatty acid omega-oxidation(GO:0010430) |
| 0.0 | 0.5 | GO:0043382 | positive regulation of memory T cell differentiation(GO:0043382) |
| 0.0 | 0.2 | GO:0090526 | regulation of gluconeogenesis involved in cellular glucose homeostasis(GO:0090526) |
| 0.0 | 0.3 | GO:0090166 | Golgi disassembly(GO:0090166) |
| 0.0 | 0.1 | GO:0006433 | prolyl-tRNA aminoacylation(GO:0006433) |
| 0.0 | 0.2 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 0.0 | 2.3 | GO:1902475 | L-alpha-amino acid transmembrane transport(GO:1902475) |
| 0.0 | 0.3 | GO:0009753 | response to jasmonic acid(GO:0009753) cellular response to jasmonic acid stimulus(GO:0071395) |
| 0.0 | 1.2 | GO:0032331 | negative regulation of chondrocyte differentiation(GO:0032331) |
| 0.0 | 1.9 | GO:0046627 | negative regulation of insulin receptor signaling pathway(GO:0046627) |
| 0.0 | 0.3 | GO:1900625 | regulation of monocyte aggregation(GO:1900623) positive regulation of monocyte aggregation(GO:1900625) |
| 0.0 | 0.2 | GO:1905150 | regulation of voltage-gated sodium channel activity(GO:1905150) |
| 0.0 | 0.4 | GO:0001542 | ovulation from ovarian follicle(GO:0001542) |
| 0.0 | 0.2 | GO:0098535 | de novo centriole assembly(GO:0098535) |
| 0.0 | 0.3 | GO:1901409 | positive regulation of phosphorylation of RNA polymerase II C-terminal domain(GO:1901409) |
| 0.0 | 0.1 | GO:2000863 | positive regulation of estrogen secretion(GO:2000863) positive regulation of estradiol secretion(GO:2000866) |
| 0.0 | 0.2 | GO:1902570 | protein localization to nucleolus(GO:1902570) |
| 0.0 | 0.1 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.0 | 0.1 | GO:1903385 | regulation of homophilic cell adhesion(GO:1903385) |
| 0.0 | 0.2 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.0 | 1.2 | GO:1901998 | toxin transport(GO:1901998) |
| 0.0 | 0.1 | GO:1904204 | regulation of skeletal muscle hypertrophy(GO:1904204) |
| 0.0 | 0.2 | GO:1902455 | negative regulation of stem cell population maintenance(GO:1902455) |
| 0.0 | 0.2 | GO:0036500 | ATF6-mediated unfolded protein response(GO:0036500) |
| 0.0 | 0.2 | GO:0030099 | myeloid cell differentiation(GO:0030099) |
| 0.0 | 0.2 | GO:0002291 | T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:0002291) |
| 0.0 | 0.7 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
| 0.0 | 0.1 | GO:0000023 | maltose metabolic process(GO:0000023) |
| 0.0 | 0.0 | GO:0045065 | cytotoxic T cell differentiation(GO:0045065) |
| 0.0 | 0.1 | GO:0006600 | creatine metabolic process(GO:0006600) |
| 0.0 | 0.1 | GO:0035063 | nuclear speck organization(GO:0035063) |
| 0.0 | 0.0 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.0 | 0.2 | GO:0008217 | regulation of blood pressure(GO:0008217) |
| 0.0 | 0.2 | GO:0006089 | lactate metabolic process(GO:0006089) |
| 0.0 | 0.1 | GO:2001206 | positive regulation of osteoclast development(GO:2001206) |
| 0.0 | 0.2 | GO:1903071 | positive regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903071) |
| 0.0 | 0.1 | GO:0016127 | cholesterol catabolic process(GO:0006707) sterol catabolic process(GO:0016127) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.6 | GO:0097057 | TRAF2-GSTP1 complex(GO:0097057) |
| 0.2 | 4.8 | GO:0005639 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) |
| 0.1 | 0.5 | GO:0002079 | inner acrosomal membrane(GO:0002079) |
| 0.1 | 0.2 | GO:1990666 | PCSK9-LDLR complex(GO:1990666) |
| 0.1 | 0.3 | GO:1990357 | terminal web(GO:1990357) |
| 0.1 | 0.2 | GO:0043159 | cytoskeletal calyx(GO:0033150) acrosomal matrix(GO:0043159) |
| 0.0 | 0.3 | GO:0035692 | macrophage migration inhibitory factor receptor complex(GO:0035692) |
| 0.0 | 0.2 | GO:0098536 | deuterosome(GO:0098536) |
| 0.0 | 0.2 | GO:0044614 | nuclear pore cytoplasmic filaments(GO:0044614) |
| 0.0 | 0.2 | GO:0000120 | RNA polymerase I transcription factor complex(GO:0000120) |
| 0.0 | 0.5 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.0 | 0.2 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.0 | 0.3 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.0 | 0.7 | GO:0005861 | troponin complex(GO:0005861) |
| 0.0 | 0.2 | GO:0031314 | extrinsic component of mitochondrial inner membrane(GO:0031314) |
| 0.0 | 0.2 | GO:1990037 | Lewy body core(GO:1990037) |
| 0.0 | 0.1 | GO:0097125 | cyclin B1-CDK1 complex(GO:0097125) |
| 0.0 | 0.2 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.0 | 0.2 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 0.1 | GO:0005712 | chiasma(GO:0005712) |
| 0.0 | 0.6 | GO:0030673 | axolemma(GO:0030673) |
| 0.0 | 0.4 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.0 | 0.8 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.0 | 0.3 | GO:0034362 | low-density lipoprotein particle(GO:0034362) |
| 0.0 | 0.3 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.0 | 0.2 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 0.0 | 0.1 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.0 | 0.3 | GO:0071565 | nBAF complex(GO:0071565) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.1 | GO:0004060 | arylamine N-acetyltransferase activity(GO:0004060) |
| 0.4 | 4.8 | GO:0035381 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.3 | 1.6 | GO:0070026 | nitric oxide binding(GO:0070026) |
| 0.3 | 2.0 | GO:0001665 | alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase activity(GO:0001665) |
| 0.1 | 1.0 | GO:0008823 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 0.1 | 1.3 | GO:0047499 | calcium-independent phospholipase A2 activity(GO:0047499) |
| 0.1 | 0.5 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 0.1 | 0.5 | GO:0000309 | nicotinamide-nucleotide adenylyltransferase activity(GO:0000309) |
| 0.1 | 0.2 | GO:0004597 | peptide-aspartate beta-dioxygenase activity(GO:0004597) |
| 0.1 | 0.3 | GO:0036080 | GDP-fucose transmembrane transporter activity(GO:0005457) purine nucleotide-sugar transmembrane transporter activity(GO:0036080) |
| 0.1 | 0.3 | GO:0047023 | androsterone dehydrogenase activity(GO:0047023) |
| 0.1 | 0.5 | GO:0086075 | gap junction channel activity involved in cardiac conduction electrical coupling(GO:0086075) |
| 0.1 | 0.4 | GO:0000293 | ferric-chelate reductase activity(GO:0000293) |
| 0.1 | 0.2 | GO:0072345 | NAADP-sensitive calcium-release channel activity(GO:0072345) |
| 0.1 | 0.6 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.1 | 0.2 | GO:0008532 | N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase activity(GO:0008532) |
| 0.1 | 0.2 | GO:0019959 | interleukin-8 binding(GO:0019959) |
| 0.1 | 0.2 | GO:0031493 | nucleosomal histone binding(GO:0031493) |
| 0.0 | 0.1 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 0.0 | 0.7 | GO:0031014 | troponin T binding(GO:0031014) |
| 0.0 | 0.1 | GO:0004827 | proline-tRNA ligase activity(GO:0004827) |
| 0.0 | 2.1 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.4 | GO:0005497 | androgen binding(GO:0005497) |
| 0.0 | 0.2 | GO:0005105 | type 1 fibroblast growth factor receptor binding(GO:0005105) |
| 0.0 | 0.4 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.0 | 1.2 | GO:0051428 | peptide hormone receptor binding(GO:0051428) |
| 0.0 | 0.1 | GO:0004031 | aldehyde oxidase activity(GO:0004031) |
| 0.0 | 0.3 | GO:0042608 | T cell receptor binding(GO:0042608) |
| 0.0 | 0.2 | GO:0030229 | very-low-density lipoprotein particle receptor activity(GO:0030229) |
| 0.0 | 1.1 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 0.2 | GO:0008273 | calcium, potassium:sodium antiporter activity(GO:0008273) |
| 0.0 | 0.3 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.0 | 0.6 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 1.8 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.0 | 3.1 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.1 | GO:0010348 | lithium:proton antiporter activity(GO:0010348) |
| 0.0 | 0.7 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.0 | 0.2 | GO:0004977 | melanocortin receptor activity(GO:0004977) |
| 0.0 | 0.1 | GO:0008665 | 2'-phosphotransferase activity(GO:0008665) |
| 0.0 | 0.8 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.1 | GO:0031780 | corticotropin hormone receptor binding(GO:0031780) type 5 melanocortin receptor binding(GO:0031783) |
| 0.0 | 0.2 | GO:0008097 | 5S rRNA binding(GO:0008097) |
| 0.0 | 0.1 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.0 | 0.2 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.0 | 0.4 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.0 | 0.1 | GO:0016160 | amylase activity(GO:0016160) |
| 0.0 | 0.0 | GO:0033754 | indoleamine 2,3-dioxygenase activity(GO:0033754) |
| 0.0 | 0.1 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.0 | 0.3 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.0 | 0.5 | GO:0001848 | complement binding(GO:0001848) |
| 0.0 | 0.2 | GO:0070679 | inositol 1,4,5 trisphosphate binding(GO:0070679) |
| 0.0 | 0.4 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.0 | 0.2 | GO:0001013 | RNA polymerase I regulatory region DNA binding(GO:0001013) RNA polymerase I regulatory region sequence-specific DNA binding(GO:0001163) RNA polymerase I CORE element sequence-specific DNA binding(GO:0001164) |
| 0.0 | 0.0 | GO:0052593 | tryptamine:oxygen oxidoreductase (deaminating) activity(GO:0052593) aminoacetone:oxygen oxidoreductase(deaminating) activity(GO:0052594) aliphatic-amine oxidase activity(GO:0052595) phenethylamine:oxygen oxidoreductase (deaminating) activity(GO:0052596) |
| 0.0 | 0.2 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.8 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
| 0.0 | 1.2 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.0 | 2.4 | PID INSULIN PATHWAY | Insulin Pathway |
| 0.0 | 1.2 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 0.9 | SIG PIP3 SIGNALING IN B LYMPHOCYTES | Genes related to PIP3 signaling in B lymphocytes |
| 0.0 | 0.6 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 2.4 | PID P73PATHWAY | p73 transcription factor network |
| 0.0 | 0.2 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.0 | 0.5 | PID IL8 CXCR1 PATHWAY | IL8- and CXCR1-mediated signaling events |
| 0.0 | 0.4 | PID AR NONGENOMIC PATHWAY | Nongenotropic Androgen signaling |
| 0.0 | 0.2 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.0 | 0.5 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.2 | REACTOME ACYL CHAIN REMODELLING OF PI | Genes involved in Acyl chain remodelling of PI |
| 0.1 | 1.8 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.0 | 1.2 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.0 | 0.7 | REACTOME SHC1 EVENTS IN ERBB4 SIGNALING | Genes involved in SHC1 events in ERBB4 signaling |
| 0.0 | 0.6 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.0 | 0.5 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.0 | 0.3 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 0.0 | 0.8 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.0 | 1.0 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.3 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 0.0 | 0.3 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.0 | 0.3 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.0 | 0.2 | REACTOME SIGNALING BY FGFR3 MUTANTS | Genes involved in Signaling by FGFR3 mutants |
| 0.0 | 0.2 | REACTOME ACTIVATION OF CHAPERONES BY ATF6 ALPHA | Genes involved in Activation of Chaperones by ATF6-alpha |