Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
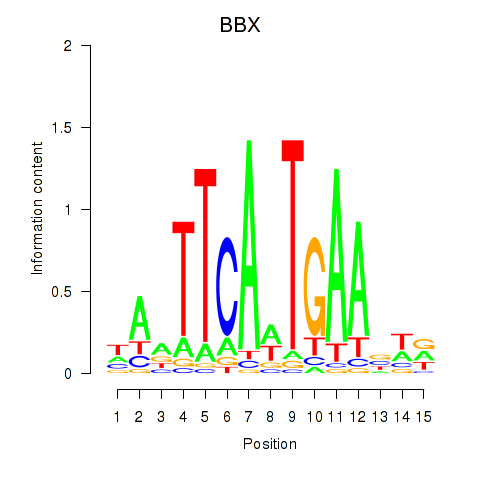
Results for BBX
Z-value: 0.66
Transcription factors associated with BBX
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
BBX
|
ENSG00000114439.14 | BBX |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| BBX | hg19_v2_chr3_+_107318157_107318172 | 0.75 | 3.2e-02 | Click! |
Activity profile of BBX motif
Sorted Z-values of BBX motif
Network of associatons between targets according to the STRING database.
First level regulatory network of BBX
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_159080806 | 0.49 |
ENST00000590648.1 |
FAM198B |
family with sequence similarity 198, member B |
| chr6_+_72922505 | 0.38 |
ENST00000401910.3 |
RIMS1 |
regulating synaptic membrane exocytosis 1 |
| chr6_+_72922590 | 0.36 |
ENST00000523963.1 |
RIMS1 |
regulating synaptic membrane exocytosis 1 |
| chr9_-_95166841 | 0.35 |
ENST00000262551.4 |
OGN |
osteoglycin |
| chr9_-_95166884 | 0.35 |
ENST00000375561.5 |
OGN |
osteoglycin |
| chr14_+_22458631 | 0.31 |
ENST00000390444.1 |
TRAV16 |
T cell receptor alpha variable 16 |
| chr9_-_95244781 | 0.30 |
ENST00000375544.3 ENST00000375543.1 ENST00000395538.3 ENST00000450139.2 |
ASPN |
asporin |
| chr4_-_186696425 | 0.30 |
ENST00000430503.1 ENST00000319454.6 ENST00000450341.1 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr4_-_186697044 | 0.26 |
ENST00000437304.2 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr8_-_108510224 | 0.22 |
ENST00000517746.1 ENST00000297450.3 |
ANGPT1 |
angiopoietin 1 |
| chr12_-_52887034 | 0.20 |
ENST00000330722.6 |
KRT6A |
keratin 6A |
| chr4_-_186733363 | 0.19 |
ENST00000393523.2 ENST00000393528.3 ENST00000449407.2 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr9_-_5833027 | 0.19 |
ENST00000339450.5 |
ERMP1 |
endoplasmic reticulum metallopeptidase 1 |
| chr10_+_111765562 | 0.17 |
ENST00000360162.3 |
ADD3 |
adducin 3 (gamma) |
| chr11_+_58390132 | 0.16 |
ENST00000361987.4 |
CNTF |
ciliary neurotrophic factor |
| chr19_+_9361606 | 0.15 |
ENST00000456448.1 |
OR7E24 |
olfactory receptor, family 7, subfamily E, member 24 |
| chr3_+_122399444 | 0.14 |
ENST00000474629.2 |
PARP14 |
poly (ADP-ribose) polymerase family, member 14 |
| chr2_+_110656005 | 0.14 |
ENST00000437679.2 |
LIMS3 |
LIM and senescent cell antigen-like domains 3 |
| chr6_-_29343068 | 0.13 |
ENST00000396806.3 |
OR12D3 |
olfactory receptor, family 12, subfamily D, member 3 |
| chr2_+_210517895 | 0.12 |
ENST00000447185.1 |
MAP2 |
microtubule-associated protein 2 |
| chr12_+_4382917 | 0.12 |
ENST00000261254.3 |
CCND2 |
cyclin D2 |
| chr12_-_8765446 | 0.12 |
ENST00000537228.1 ENST00000229335.6 |
AICDA |
activation-induced cytidine deaminase |
| chr7_-_16921601 | 0.11 |
ENST00000402239.3 ENST00000310398.2 ENST00000414935.1 |
AGR3 |
anterior gradient 3 |
| chr4_-_987164 | 0.11 |
ENST00000398520.2 |
SLC26A1 |
solute carrier family 26 (anion exchanger), member 1 |
| chr4_-_104021009 | 0.11 |
ENST00000509245.1 ENST00000296424.4 |
BDH2 |
3-hydroxybutyrate dehydrogenase, type 2 |
| chr4_-_987217 | 0.11 |
ENST00000361661.2 ENST00000398516.2 |
SLC26A1 |
solute carrier family 26 (anion exchanger), member 1 |
| chr4_+_174818390 | 0.10 |
ENST00000509968.1 ENST00000512263.1 |
RP11-161D15.1 |
RP11-161D15.1 |
| chr3_+_111260856 | 0.10 |
ENST00000352690.4 |
CD96 |
CD96 molecule |
| chr3_+_111260954 | 0.09 |
ENST00000283285.5 |
CD96 |
CD96 molecule |
| chr2_-_120124258 | 0.09 |
ENST00000409877.1 ENST00000409523.1 ENST00000409466.2 ENST00000414534.1 |
C2orf76 |
chromosome 2 open reading frame 76 |
| chrX_-_102531717 | 0.09 |
ENST00000372680.1 |
TCEAL5 |
transcription elongation factor A (SII)-like 5 |
| chr3_-_112218378 | 0.08 |
ENST00000334529.5 |
BTLA |
B and T lymphocyte associated |
| chr2_+_101437487 | 0.08 |
ENST00000427413.1 ENST00000542504.1 |
NPAS2 |
neuronal PAS domain protein 2 |
| chr1_-_206785789 | 0.08 |
ENST00000437518.1 ENST00000367114.3 |
EIF2D |
eukaryotic translation initiation factor 2D |
| chr4_+_155484155 | 0.08 |
ENST00000509493.1 |
FGB |
fibrinogen beta chain |
| chr1_+_229440129 | 0.08 |
ENST00000366688.3 |
SPHAR |
S-phase response (cyclin related) |
| chr3_+_138067666 | 0.08 |
ENST00000475711.1 ENST00000464896.1 |
MRAS |
muscle RAS oncogene homolog |
| chr22_+_32871224 | 0.08 |
ENST00000452138.1 ENST00000382058.3 ENST00000397426.1 |
FBXO7 |
F-box protein 7 |
| chr7_+_117120017 | 0.08 |
ENST00000003084.6 ENST00000454343.1 |
CFTR |
cystic fibrosis transmembrane conductance regulator (ATP-binding cassette sub-family C, member 7) |
| chr1_+_117963209 | 0.08 |
ENST00000449370.2 |
MAN1A2 |
mannosidase, alpha, class 1A, member 2 |
| chr1_+_152784447 | 0.08 |
ENST00000360090.3 |
LCE1B |
late cornified envelope 1B |
| chr19_+_6887571 | 0.07 |
ENST00000250572.8 ENST00000381407.5 ENST00000312053.4 ENST00000450315.3 ENST00000381404.4 |
EMR1 |
egf-like module containing, mucin-like, hormone receptor-like 1 |
| chr5_+_140514782 | 0.07 |
ENST00000231134.5 |
PCDHB5 |
protocadherin beta 5 |
| chr5_+_179921344 | 0.07 |
ENST00000261951.4 |
CNOT6 |
CCR4-NOT transcription complex, subunit 6 |
| chr12_-_4754339 | 0.07 |
ENST00000228850.1 |
AKAP3 |
A kinase (PRKA) anchor protein 3 |
| chr14_+_94492674 | 0.07 |
ENST00000203664.5 ENST00000553723.1 |
OTUB2 |
OTU domain, ubiquitin aldehyde binding 2 |
| chr19_+_7828035 | 0.07 |
ENST00000327325.5 ENST00000394122.2 ENST00000248228.4 ENST00000334806.5 ENST00000359059.5 ENST00000357361.2 ENST00000596363.1 ENST00000595751.1 ENST00000596707.1 ENST00000597522.1 ENST00000595496.1 |
CLEC4M |
C-type lectin domain family 4, member M |
| chr1_-_28527152 | 0.06 |
ENST00000321830.5 |
AL353354.1 |
Uncharacterized protein |
| chr1_-_198509804 | 0.06 |
ENST00000489986.1 ENST00000367382.1 |
ATP6V1G3 |
ATPase, H+ transporting, lysosomal 13kDa, V1 subunit G3 |
| chr15_+_54901540 | 0.06 |
ENST00000539562.2 |
UNC13C |
unc-13 homolog C (C. elegans) |
| chr6_-_107235331 | 0.06 |
ENST00000433965.1 ENST00000430094.1 |
RP1-60O19.1 |
RP1-60O19.1 |
| chr2_+_12246664 | 0.06 |
ENST00000449986.1 |
AC096559.1 |
AC096559.1 |
| chr11_-_18956556 | 0.06 |
ENST00000302797.3 |
MRGPRX1 |
MAS-related GPR, member X1 |
| chrX_+_54834004 | 0.05 |
ENST00000375068.1 |
MAGED2 |
melanoma antigen family D, 2 |
| chr10_+_24497704 | 0.05 |
ENST00000376456.4 ENST00000458595.1 |
KIAA1217 |
KIAA1217 |
| chr7_+_135611542 | 0.05 |
ENST00000416501.1 |
AC015987.2 |
AC015987.2 |
| chr5_-_78809950 | 0.05 |
ENST00000334082.6 |
HOMER1 |
homer homolog 1 (Drosophila) |
| chrX_+_54834159 | 0.05 |
ENST00000375053.2 ENST00000347546.4 ENST00000375062.4 |
MAGED2 |
melanoma antigen family D, 2 |
| chr22_-_39928823 | 0.05 |
ENST00000334678.3 |
RPS19BP1 |
ribosomal protein S19 binding protein 1 |
| chr3_-_112218205 | 0.05 |
ENST00000383680.4 |
BTLA |
B and T lymphocyte associated |
| chr11_-_105010320 | 0.05 |
ENST00000532895.1 ENST00000530950.1 |
CARD18 |
caspase recruitment domain family, member 18 |
| chr7_+_117120106 | 0.05 |
ENST00000426809.1 |
CFTR |
cystic fibrosis transmembrane conductance regulator (ATP-binding cassette sub-family C, member 7) |
| chrX_+_140677562 | 0.05 |
ENST00000370518.3 |
SPANXA2 |
SPANX family, member A2 |
| chr10_-_94003003 | 0.04 |
ENST00000412050.4 |
CPEB3 |
cytoplasmic polyadenylation element binding protein 3 |
| chrX_-_140786896 | 0.04 |
ENST00000370515.3 |
SPANXD |
SPANX family, member D |
| chr4_+_155484103 | 0.04 |
ENST00000302068.4 |
FGB |
fibrinogen beta chain |
| chr6_-_133084580 | 0.03 |
ENST00000525270.1 ENST00000530536.1 ENST00000524919.1 |
VNN2 |
vanin 2 |
| chr1_-_59043166 | 0.03 |
ENST00000371225.2 |
TACSTD2 |
tumor-associated calcium signal transducer 2 |
| chr4_-_46126093 | 0.03 |
ENST00000295452.4 |
GABRG1 |
gamma-aminobutyric acid (GABA) A receptor, gamma 1 |
| chr20_-_30310336 | 0.03 |
ENST00000434194.1 ENST00000376062.2 |
BCL2L1 |
BCL2-like 1 |
| chr5_+_150040403 | 0.03 |
ENST00000517768.1 ENST00000297130.4 |
MYOZ3 |
myozenin 3 |
| chr3_+_94657086 | 0.03 |
ENST00000463200.1 |
LINC00879 |
long intergenic non-protein coding RNA 879 |
| chr6_-_161085291 | 0.03 |
ENST00000316300.5 |
LPA |
lipoprotein, Lp(a) |
| chr11_+_61717336 | 0.03 |
ENST00000378042.3 |
BEST1 |
bestrophin 1 |
| chr5_+_140079919 | 0.03 |
ENST00000274712.3 |
ZMAT2 |
zinc finger, matrin-type 2 |
| chr11_+_61717279 | 0.03 |
ENST00000378043.4 |
BEST1 |
bestrophin 1 |
| chr1_-_153123345 | 0.03 |
ENST00000368748.4 |
SPRR2G |
small proline-rich protein 2G |
| chr3_+_94657118 | 0.03 |
ENST00000466089.1 ENST00000470465.1 |
LINC00879 |
long intergenic non-protein coding RNA 879 |
| chr8_-_6783588 | 0.03 |
ENST00000297436.2 |
DEFA6 |
defensin, alpha 6, Paneth cell-specific |
| chr20_+_23420885 | 0.03 |
ENST00000246020.2 |
CSTL1 |
cystatin-like 1 |
| chr5_+_161494521 | 0.03 |
ENST00000356592.3 |
GABRG2 |
gamma-aminobutyric acid (GABA) A receptor, gamma 2 |
| chr1_+_55464600 | 0.03 |
ENST00000371265.4 |
BSND |
Bartter syndrome, infantile, with sensorineural deafness (Barttin) |
| chr1_+_52082751 | 0.03 |
ENST00000447887.1 ENST00000435686.2 ENST00000428468.1 ENST00000453295.1 |
OSBPL9 |
oxysterol binding protein-like 9 |
| chr8_-_28347737 | 0.02 |
ENST00000517673.1 ENST00000518734.1 ENST00000346498.2 ENST00000380254.2 |
FBXO16 |
F-box protein 16 |
| chr17_-_17184605 | 0.02 |
ENST00000268717.5 |
COPS3 |
COP9 signalosome subunit 3 |
| chr17_-_8263538 | 0.02 |
ENST00000535173.1 |
AC135178.1 |
HCG1985372; Uncharacterized protein; cDNA FLJ37541 fis, clone BRCAN2026340 |
| chr11_+_22689648 | 0.02 |
ENST00000278187.3 |
GAS2 |
growth arrest-specific 2 |
| chr20_-_30310693 | 0.02 |
ENST00000307677.4 ENST00000420653.1 |
BCL2L1 |
BCL2-like 1 |
| chr1_+_192544857 | 0.02 |
ENST00000367459.3 ENST00000469578.2 |
RGS1 |
regulator of G-protein signaling 1 |
| chr2_-_231084617 | 0.02 |
ENST00000409815.2 |
SP110 |
SP110 nuclear body protein |
| chr2_+_102927962 | 0.02 |
ENST00000233954.1 ENST00000393393.3 ENST00000410040.1 |
IL1RL1 IL18R1 |
interleukin 1 receptor-like 1 interleukin 18 receptor 1 |
| chr20_-_30310656 | 0.02 |
ENST00000376055.4 |
BCL2L1 |
BCL2-like 1 |
| chr2_+_63816295 | 0.02 |
ENST00000539945.1 ENST00000544381.1 |
MDH1 |
malate dehydrogenase 1, NAD (soluble) |
| chr3_+_140981456 | 0.02 |
ENST00000504264.1 |
ACPL2 |
acid phosphatase-like 2 |
| chr11_+_18154059 | 0.02 |
ENST00000531264.1 |
MRGPRX3 |
MAS-related GPR, member X3 |
| chr5_+_125758813 | 0.02 |
ENST00000285689.3 ENST00000515200.1 |
GRAMD3 |
GRAM domain containing 3 |
| chr14_+_57671888 | 0.02 |
ENST00000391612.1 |
AL391152.1 |
AL391152.1 |
| chr20_-_7921090 | 0.02 |
ENST00000378789.3 |
HAO1 |
hydroxyacid oxidase (glycolate oxidase) 1 |
| chr7_+_5085452 | 0.02 |
ENST00000353796.3 ENST00000396912.1 ENST00000396904.2 |
RBAK RBAK-RBAKDN |
RB-associated KRAB zinc finger RBAK-RBAKDN readthrough |
| chr13_-_81801115 | 0.02 |
ENST00000567258.1 |
LINC00564 |
long intergenic non-protein coding RNA 564 |
| chr9_+_35161998 | 0.02 |
ENST00000396787.1 ENST00000378495.3 ENST00000378496.4 |
UNC13B |
unc-13 homolog B (C. elegans) |
| chr12_-_100378006 | 0.02 |
ENST00000547776.2 ENST00000329257.7 ENST00000547010.1 |
ANKS1B |
ankyrin repeat and sterile alpha motif domain containing 1B |
| chr4_+_108910870 | 0.02 |
ENST00000403312.1 ENST00000603302.1 ENST00000309522.3 |
HADH |
hydroxyacyl-CoA dehydrogenase |
| chr17_+_40950900 | 0.01 |
ENST00000588527.1 |
CNTD1 |
cyclin N-terminal domain containing 1 |
| chr7_-_130080977 | 0.01 |
ENST00000223208.5 |
CEP41 |
centrosomal protein 41kDa |
| chr19_+_49109990 | 0.01 |
ENST00000321762.1 |
SPACA4 |
sperm acrosome associated 4 |
| chr12_-_99548270 | 0.01 |
ENST00000546568.1 ENST00000332712.7 ENST00000546960.1 |
ANKS1B |
ankyrin repeat and sterile alpha motif domain containing 1B |
| chr2_+_113816215 | 0.01 |
ENST00000346807.3 |
IL36RN |
interleukin 36 receptor antagonist |
| chr14_+_23235886 | 0.01 |
ENST00000604262.1 ENST00000431881.2 ENST00000412791.1 ENST00000358043.5 |
OXA1L |
oxidase (cytochrome c) assembly 1-like |
| chr17_-_59668550 | 0.00 |
ENST00000521764.1 |
NACA2 |
nascent polypeptide-associated complex alpha subunit 2 |
| chr18_-_13915530 | 0.00 |
ENST00000327606.3 |
MC2R |
melanocortin 2 receptor (adrenocorticotropic hormone) |
| chr6_+_29424958 | 0.00 |
ENST00000377136.1 ENST00000377133.1 |
OR2H1 |
olfactory receptor, family 2, subfamily H, member 1 |
| chr16_+_56672571 | 0.00 |
ENST00000290705.8 |
MT1A |
metallothionein 1A |
| chr2_-_62081254 | 0.00 |
ENST00000405894.3 |
FAM161A |
family with sequence similarity 161, member A |
| chr7_-_16872932 | 0.00 |
ENST00000419572.2 ENST00000412973.1 |
AGR2 |
anterior gradient 2 |
| chr9_-_13175823 | 0.00 |
ENST00000545857.1 |
MPDZ |
multiple PDZ domain protein |
| chr13_+_77522632 | 0.00 |
ENST00000377462.1 |
IRG1 |
immunoresponsive 1 homolog (mouse) |
| chr6_+_54173227 | 0.00 |
ENST00000259782.4 ENST00000370864.3 |
TINAG |
tubulointerstitial nephritis antigen |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0070171 | negative regulation of tooth mineralization(GO:0070171) |
| 0.1 | 0.2 | GO:0002728 | negative regulation of natural killer cell cytokine production(GO:0002728) |
| 0.1 | 0.2 | GO:0051710 | cytolysis by symbiont of host cells(GO:0001897) regulation of cytolysis in other organism(GO:0051710) |
| 0.0 | 0.1 | GO:1902161 | positive regulation of cyclic nucleotide-gated ion channel activity(GO:1902161) |
| 0.0 | 0.8 | GO:0097151 | positive regulation of inhibitory postsynaptic potential(GO:0097151) modulation of inhibitory postsynaptic potential(GO:0098828) |
| 0.0 | 0.1 | GO:0090310 | negative regulation of methylation-dependent chromatin silencing(GO:0090310) |
| 0.0 | 0.6 | GO:0042340 | keratan sulfate catabolic process(GO:0042340) |
| 0.0 | 0.2 | GO:0046668 | regulation of retinal cell programmed cell death(GO:0046668) |
| 0.0 | 0.2 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.0 | 0.2 | GO:0050428 | purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.0 | 0.7 | GO:0061049 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.0 | 0.0 | GO:0007206 | phospholipase C-activating G-protein coupled glutamate receptor signaling pathway(GO:0007206) |
| 0.0 | 0.0 | GO:1900365 | positive regulation of mRNA polyadenylation(GO:1900365) |
| 0.0 | 0.1 | GO:0046968 | peptide antigen transport(GO:0046968) |
| 0.0 | 0.1 | GO:1904381 | Golgi apparatus mannose trimming(GO:1904381) |
| 0.0 | 0.1 | GO:0070966 | nuclear-transcribed mRNA catabolic process, no-go decay(GO:0070966) |
| 0.0 | 0.1 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.0 | 0.1 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0097058 | CRLF-CLCF1 complex(GO:0097058) |
| 0.0 | 0.1 | GO:0097442 | CA3 pyramidal cell dendrite(GO:0097442) |
| 0.0 | 0.1 | GO:0031205 | endoplasmic reticulum Sec complex(GO:0031205) |
| 0.0 | 0.1 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 0.0 | 0.8 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 0.1 | GO:1990037 | Lewy body core(GO:1990037) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0005260 | channel-conductance-controlling ATPase activity(GO:0005260) |
| 0.0 | 0.1 | GO:0003858 | 3-hydroxybutyrate dehydrogenase activity(GO:0003858) |
| 0.0 | 0.2 | GO:0005127 | ciliary neurotrophic factor receptor binding(GO:0005127) |
| 0.0 | 0.7 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.2 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.0 | 0.1 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.0 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 0.6 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |