Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
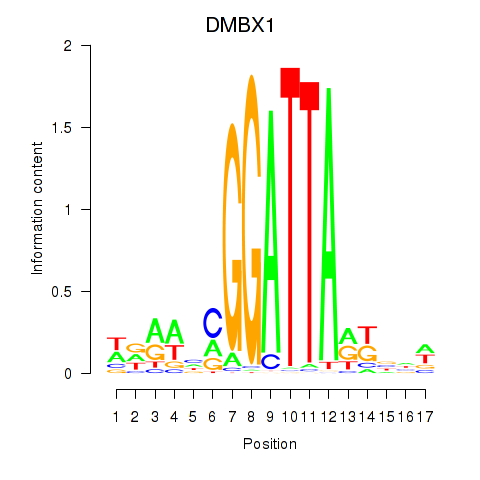
Results for DMBX1
Z-value: 0.48
Transcription factors associated with DMBX1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
DMBX1
|
ENSG00000197587.6 | DMBX1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| DMBX1 | hg19_v2_chr1_+_46972668_46972669 | 0.10 | 8.1e-01 | Click! |
Activity profile of DMBX1 motif
Sorted Z-values of DMBX1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of DMBX1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_+_118187379 | 0.51 |
ENST00000369230.3 |
PNLIPRP3 |
pancreatic lipase-related protein 3 |
| chr18_+_29027696 | 0.46 |
ENST00000257189.4 |
DSG3 |
desmoglein 3 |
| chrX_-_102565932 | 0.41 |
ENST00000372674.1 ENST00000372677.3 |
BEX2 |
brain expressed X-linked 2 |
| chr2_-_70780770 | 0.37 |
ENST00000444975.1 ENST00000445399.1 ENST00000418333.2 |
TGFA |
transforming growth factor, alpha |
| chr2_-_70780572 | 0.36 |
ENST00000450929.1 |
TGFA |
transforming growth factor, alpha |
| chr17_-_4643114 | 0.36 |
ENST00000293778.6 |
CXCL16 |
chemokine (C-X-C motif) ligand 16 |
| chr8_-_125577940 | 0.33 |
ENST00000519168.1 ENST00000395508.2 |
MTSS1 |
metastasis suppressor 1 |
| chrX_-_32173579 | 0.28 |
ENST00000359836.1 ENST00000343523.2 ENST00000378707.3 ENST00000541735.1 ENST00000474231.1 |
DMD |
dystrophin |
| chr12_+_7941989 | 0.15 |
ENST00000229307.4 |
NANOG |
Nanog homeobox |
| chr6_+_89791507 | 0.14 |
ENST00000354922.3 |
PNRC1 |
proline-rich nuclear receptor coactivator 1 |
| chr11_-_36619771 | 0.13 |
ENST00000311485.3 ENST00000527033.1 ENST00000532616.1 |
RAG2 |
recombination activating gene 2 |
| chr19_-_6459746 | 0.12 |
ENST00000301454.4 ENST00000334510.5 |
SLC25A23 |
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 23 |
| chr3_+_49058444 | 0.12 |
ENST00000326925.6 ENST00000395458.2 |
NDUFAF3 |
NADH dehydrogenase (ubiquinone) complex I, assembly factor 3 |
| chr6_+_130339710 | 0.12 |
ENST00000526087.1 ENST00000533560.1 ENST00000361794.2 |
L3MBTL3 |
l(3)mbt-like 3 (Drosophila) |
| chr7_+_1727755 | 0.12 |
ENST00000424383.2 |
ELFN1 |
extracellular leucine-rich repeat and fibronectin type III domain containing 1 |
| chr3_+_49057876 | 0.11 |
ENST00000326912.4 |
NDUFAF3 |
NADH dehydrogenase (ubiquinone) complex I, assembly factor 3 |
| chr1_-_94586651 | 0.10 |
ENST00000535735.1 ENST00000370225.3 |
ABCA4 |
ATP-binding cassette, sub-family A (ABC1), member 4 |
| chr4_+_140586922 | 0.10 |
ENST00000265498.1 ENST00000506797.1 |
MGST2 |
microsomal glutathione S-transferase 2 |
| chr11_+_61248583 | 0.10 |
ENST00000432063.2 ENST00000338608.2 |
PPP1R32 |
protein phosphatase 1, regulatory subunit 32 |
| chr6_-_111927062 | 0.09 |
ENST00000359831.4 |
TRAF3IP2 |
TRAF3 interacting protein 2 |
| chr20_+_31805131 | 0.08 |
ENST00000375454.3 ENST00000375452.3 |
BPIFA3 |
BPI fold containing family A, member 3 |
| chr3_+_188664988 | 0.08 |
ENST00000433971.1 |
TPRG1 |
tumor protein p63 regulated 1 |
| chr15_-_85201779 | 0.08 |
ENST00000360476.3 ENST00000394588.3 |
NMB |
neuromedin B |
| chr20_-_43729750 | 0.08 |
ENST00000537075.1 ENST00000306117.1 |
KCNS1 |
potassium voltage-gated channel, delayed-rectifier, subfamily S, member 1 |
| chr15_+_74165945 | 0.08 |
ENST00000535547.2 ENST00000300504.2 ENST00000562056.1 |
TBC1D21 |
TBC1 domain family, member 21 |
| chr10_+_74451883 | 0.07 |
ENST00000373053.3 ENST00000357157.6 |
MCU |
mitochondrial calcium uniporter |
| chr20_+_34556488 | 0.07 |
ENST00000373973.3 ENST00000349339.1 ENST00000538900.1 |
CNBD2 |
cyclic nucleotide binding domain containing 2 |
| chr3_-_11685345 | 0.07 |
ENST00000430365.2 |
VGLL4 |
vestigial like 4 (Drosophila) |
| chr17_+_4643300 | 0.07 |
ENST00000433935.1 |
ZMYND15 |
zinc finger, MYND-type containing 15 |
| chr9_-_21995249 | 0.07 |
ENST00000494262.1 |
CDKN2A |
cyclin-dependent kinase inhibitor 2A |
| chr17_+_4643337 | 0.07 |
ENST00000592813.1 |
ZMYND15 |
zinc finger, MYND-type containing 15 |
| chr1_+_120839005 | 0.06 |
ENST00000369390.3 ENST00000452190.1 |
FAM72B |
family with sequence similarity 72, member B |
| chr3_+_136649311 | 0.06 |
ENST00000469404.1 ENST00000467911.1 |
NCK1 |
NCK adaptor protein 1 |
| chr6_-_138893661 | 0.06 |
ENST00000427025.2 |
NHSL1 |
NHS-like 1 |
| chr1_-_143913143 | 0.06 |
ENST00000400889.1 |
FAM72D |
family with sequence similarity 72, member D |
| chr1_+_206138457 | 0.06 |
ENST00000367128.3 ENST00000431655.2 |
FAM72A |
family with sequence similarity 72, member A |
| chr9_-_21995300 | 0.06 |
ENST00000498628.2 |
CDKN2A |
cyclin-dependent kinase inhibitor 2A |
| chr2_-_165812028 | 0.06 |
ENST00000303735.4 |
SLC38A11 |
solute carrier family 38, member 11 |
| chr19_+_36157715 | 0.05 |
ENST00000379013.2 ENST00000222275.2 |
UPK1A |
uroplakin 1A |
| chr2_-_165811983 | 0.05 |
ENST00000409058.1 ENST00000409149.3 |
SLC38A11 |
solute carrier family 38, member 11 |
| chr12_+_115800817 | 0.05 |
ENST00000547948.1 |
RP11-116D17.1 |
HCG2038717; Uncharacterized protein |
| chr12_-_21487829 | 0.05 |
ENST00000445053.1 ENST00000452078.1 ENST00000458504.1 ENST00000422327.1 ENST00000421294.1 |
SLCO1A2 |
solute carrier organic anion transporter family, member 1A2 |
| chr9_-_21994344 | 0.05 |
ENST00000530628.2 ENST00000361570.3 |
CDKN2A |
cyclin-dependent kinase inhibitor 2A |
| chr7_-_135412925 | 0.04 |
ENST00000354042.4 |
SLC13A4 |
solute carrier family 13 (sodium/sulfate symporter), member 4 |
| chr12_-_118628350 | 0.04 |
ENST00000537952.1 ENST00000537822.1 |
TAOK3 |
TAO kinase 3 |
| chr19_+_46001697 | 0.04 |
ENST00000451287.2 ENST00000324688.4 |
PPM1N |
protein phosphatase, Mg2+/Mn2+ dependent, 1N (putative) |
| chr9_-_21994597 | 0.04 |
ENST00000579755.1 |
CDKN2A |
cyclin-dependent kinase inhibitor 2A |
| chr4_+_169552748 | 0.04 |
ENST00000504519.1 ENST00000512127.1 |
PALLD |
palladin, cytoskeletal associated protein |
| chr2_-_165811756 | 0.04 |
ENST00000409662.1 |
SLC38A11 |
solute carrier family 38, member 11 |
| chr6_-_26250835 | 0.03 |
ENST00000446824.2 |
HIST1H3F |
histone cluster 1, H3f |
| chr15_+_63413990 | 0.03 |
ENST00000261893.4 |
LACTB |
lactamase, beta |
| chr16_-_20338748 | 0.03 |
ENST00000575582.1 ENST00000341642.5 ENST00000381362.4 ENST00000572347.1 ENST00000572478.1 ENST00000302555.5 |
GP2 |
glycoprotein 2 (zymogen granule membrane) |
| chr17_-_38859996 | 0.03 |
ENST00000264651.2 |
KRT24 |
keratin 24 |
| chr19_-_17186229 | 0.03 |
ENST00000253669.5 ENST00000448593.2 |
HAUS8 |
HAUS augmin-like complex, subunit 8 |
| chr17_+_68100989 | 0.03 |
ENST00000585558.1 ENST00000392670.1 |
KCNJ16 |
potassium inwardly-rectifying channel, subfamily J, member 16 |
| chr19_+_17448348 | 0.03 |
ENST00000324894.8 ENST00000358792.7 ENST00000600625.1 |
GTPBP3 |
GTP binding protein 3 (mitochondrial) |
| chr2_+_73461410 | 0.03 |
ENST00000399032.2 ENST00000398422.2 ENST00000537131.1 ENST00000538797.1 |
CCT7 |
chaperonin containing TCP1, subunit 7 (eta) |
| chr2_+_73461364 | 0.03 |
ENST00000540468.1 ENST00000539919.1 ENST00000258091.5 |
CCT7 |
chaperonin containing TCP1, subunit 7 (eta) |
| chr17_+_60536002 | 0.03 |
ENST00000582809.1 |
TLK2 |
tousled-like kinase 2 |
| chr17_-_12921270 | 0.03 |
ENST00000578071.1 ENST00000426905.3 ENST00000395962.2 ENST00000583371.1 ENST00000338034.4 |
ELAC2 |
elaC ribonuclease Z 2 |
| chr1_-_28384598 | 0.03 |
ENST00000373864.1 |
EYA3 |
eyes absent homolog 3 (Drosophila) |
| chr12_-_56321649 | 0.02 |
ENST00000454792.2 ENST00000408946.2 |
WIBG |
within bgcn homolog (Drosophila) |
| chr11_+_124055923 | 0.02 |
ENST00000318666.6 |
OR10D3 |
olfactory receptor, family 10, subfamily D, member 3 (non-functional) |
| chr1_+_109756523 | 0.02 |
ENST00000234677.2 ENST00000369923.4 |
SARS |
seryl-tRNA synthetase |
| chr19_+_41284121 | 0.02 |
ENST00000594800.1 ENST00000357052.2 ENST00000602173.1 |
RAB4B |
RAB4B, member RAS oncogene family |
| chr2_-_14541060 | 0.02 |
ENST00000418420.1 ENST00000417751.1 |
LINC00276 |
long intergenic non-protein coding RNA 276 |
| chr4_+_619386 | 0.02 |
ENST00000496514.1 |
PDE6B |
phosphodiesterase 6B, cGMP-specific, rod, beta |
| chr11_-_45940343 | 0.02 |
ENST00000532681.1 |
PEX16 |
peroxisomal biogenesis factor 16 |
| chr12_-_56321397 | 0.02 |
ENST00000557259.1 ENST00000549939.1 |
WIBG |
within bgcn homolog (Drosophila) |
| chr14_+_22265444 | 0.02 |
ENST00000390430.2 |
TRAV8-1 |
T cell receptor alpha variable 8-1 |
| chr19_-_17185848 | 0.01 |
ENST00000593360.1 |
HAUS8 |
HAUS augmin-like complex, subunit 8 |
| chr21_+_18811205 | 0.01 |
ENST00000440664.1 |
C21orf37 |
chromosome 21 open reading frame 37 |
| chr10_-_35104185 | 0.01 |
ENST00000374789.3 ENST00000374788.3 ENST00000346874.4 ENST00000374794.3 ENST00000350537.4 ENST00000374790.3 ENST00000374776.1 ENST00000374773.1 ENST00000545693.1 ENST00000545260.1 ENST00000340077.5 |
PARD3 |
par-3 family cell polarity regulator |
| chrX_+_72062617 | 0.01 |
ENST00000440247.1 |
DMRTC1B |
DMRT-like family C1B |
| chr4_-_54232144 | 0.01 |
ENST00000388940.4 ENST00000503450.1 ENST00000401642.3 |
SCFD2 |
sec1 family domain containing 2 |
| chr3_-_149470229 | 0.01 |
ENST00000473414.1 |
COMMD2 |
COMM domain containing 2 |
| chr3_-_10362725 | 0.01 |
ENST00000397109.3 ENST00000428626.1 ENST00000445064.1 ENST00000431352.1 ENST00000397117.1 ENST00000337354.4 ENST00000383801.2 ENST00000432213.1 ENST00000350697.3 |
SEC13 |
SEC13 homolog (S. cerevisiae) |
| chrX_-_71792477 | 0.01 |
ENST00000421523.1 ENST00000415409.1 ENST00000373559.4 ENST00000373556.3 ENST00000373560.2 ENST00000373583.1 ENST00000429103.2 ENST00000373571.1 ENST00000373554.1 |
HDAC8 |
histone deacetylase 8 |
| chr1_-_13452656 | 0.01 |
ENST00000376132.3 |
PRAMEF13 |
PRAME family member 13 |
| chr17_-_39324424 | 0.01 |
ENST00000391356.2 |
KRTAP4-3 |
keratin associated protein 4-3 |
| chr12_+_54718904 | 0.00 |
ENST00000262061.2 ENST00000549043.1 ENST00000552218.1 ENST00000553231.1 ENST00000552362.1 ENST00000455864.2 ENST00000416254.2 ENST00000549116.1 ENST00000551779.1 |
COPZ1 |
coatomer protein complex, subunit zeta 1 |
| chr18_-_35145593 | 0.00 |
ENST00000334919.5 ENST00000591282.1 ENST00000588597.1 |
CELF4 |
CUGBP, Elav-like family member 4 |
| chr3_-_179692042 | 0.00 |
ENST00000468741.1 |
PEX5L |
peroxisomal biogenesis factor 5-like |
| chr15_+_63414760 | 0.00 |
ENST00000557972.1 |
LACTB |
lactamase, beta |
| chr12_-_110937351 | 0.00 |
ENST00000552130.2 |
VPS29 |
vacuolar protein sorting 29 homolog (S. cerevisiae) |
| chr17_+_48911942 | 0.00 |
ENST00000426127.1 |
WFIKKN2 |
WAP, follistatin/kazal, immunoglobulin, kunitz and netrin domain containing 2 |
| chr8_-_110656995 | 0.00 |
ENST00000276646.9 ENST00000533065.1 |
SYBU |
syntabulin (syntaxin-interacting) |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0030035 | microspike assembly(GO:0030035) |
| 0.0 | 0.7 | GO:0045741 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) |
| 0.0 | 0.1 | GO:0002331 | pre-B cell allelic exclusion(GO:0002331) |
| 0.0 | 0.3 | GO:0014809 | regulation of skeletal muscle contraction by regulation of release of sequestered calcium ion(GO:0014809) |
| 0.0 | 0.2 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 0.0 | 0.1 | GO:1903676 | regulation of cap-dependent translational initiation(GO:1903674) positive regulation of cap-dependent translational initiation(GO:1903676) |
| 0.0 | 0.2 | GO:2000111 | senescence-associated heterochromatin focus assembly(GO:0035986) positive regulation of macrophage apoptotic process(GO:2000111) |
| 0.0 | 0.2 | GO:0001714 | endodermal cell fate specification(GO:0001714) |
| 0.0 | 0.4 | GO:0010818 | T cell chemotaxis(GO:0010818) |
| 0.0 | 0.1 | GO:0031087 | deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | GO:0016013 | syntrophin complex(GO:0016013) |
| 0.0 | 0.2 | GO:0035985 | senescence-associated heterochromatin focus(GO:0035985) |
| 0.0 | 0.1 | GO:1990246 | uniplex complex(GO:1990246) |
| 0.0 | 0.5 | GO:0030057 | desmosome(GO:0030057) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0004464 | leukotriene-C4 synthase activity(GO:0004464) |
| 0.0 | 0.5 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.4 | GO:0005041 | low-density lipoprotein receptor activity(GO:0005041) |
| 0.0 | 0.1 | GO:0031705 | bombesin receptor binding(GO:0031705) |
| 0.0 | 0.1 | GO:0071074 | eukaryotic initiation factor eIF2 binding(GO:0071074) |
| 0.0 | 0.3 | GO:0050998 | nitric-oxide synthase binding(GO:0050998) |
| 0.0 | 0.7 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |