Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
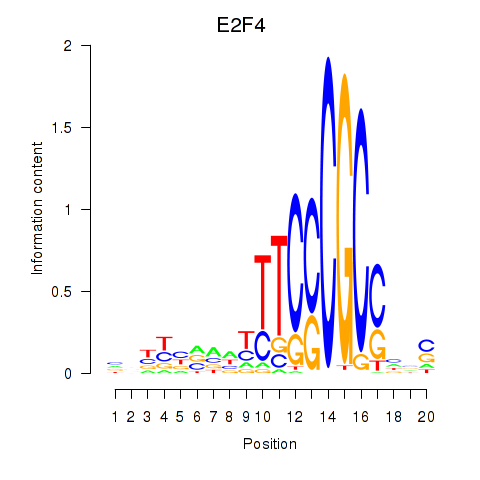
Results for E2F4
Z-value: 1.13
Transcription factors associated with E2F4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
E2F4
|
ENSG00000205250.4 | E2F4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| E2F4 | hg19_v2_chr16_+_67226019_67226127 | 0.29 | 4.8e-01 | Click! |
Activity profile of E2F4 motif
Sorted Z-values of E2F4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of E2F4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_+_39687596 | 0.59 |
ENST00000339852.4 |
NCCRP1 |
non-specific cytotoxic cell receptor protein 1 homolog (zebrafish) |
| chr9_+_131314859 | 0.46 |
ENST00000358161.5 ENST00000372731.4 ENST00000372739.3 |
SPTAN1 |
spectrin, alpha, non-erythrocytic 1 |
| chr15_+_91260552 | 0.44 |
ENST00000355112.3 ENST00000560509.1 |
BLM |
Bloom syndrome, RecQ helicase-like |
| chrX_+_30671476 | 0.42 |
ENST00000378946.3 ENST00000378943.3 ENST00000378945.3 ENST00000427190.1 ENST00000378941.3 |
GK |
glycerol kinase |
| chr10_-_70231639 | 0.38 |
ENST00000551118.2 ENST00000358410.3 ENST00000399180.2 ENST00000399179.2 |
DNA2 |
DNA replication helicase/nuclease 2 |
| chr13_+_32889605 | 0.38 |
ENST00000380152.3 ENST00000544455.1 ENST00000530893.2 |
BRCA2 |
breast cancer 2, early onset |
| chr1_-_197115818 | 0.36 |
ENST00000367409.4 ENST00000294732.7 |
ASPM |
asp (abnormal spindle) homolog, microcephaly associated (Drosophila) |
| chr16_+_838614 | 0.36 |
ENST00000262315.9 ENST00000455171.2 |
CHTF18 |
CTF18, chromosome transmission fidelity factor 18 homolog (S. cerevisiae) |
| chr3_+_10068095 | 0.35 |
ENST00000287647.3 ENST00000383807.1 ENST00000383806.1 ENST00000419585.1 |
FANCD2 |
Fanconi anemia, complementation group D2 |
| chr1_-_36235529 | 0.35 |
ENST00000318121.3 ENST00000373220.3 ENST00000520551.1 |
CLSPN |
claspin |
| chr2_+_157292859 | 0.34 |
ENST00000438166.2 |
GPD2 |
glycerol-3-phosphate dehydrogenase 2 (mitochondrial) |
| chr8_-_95907423 | 0.32 |
ENST00000396133.3 ENST00000308108.4 |
CCNE2 |
cyclin E2 |
| chr10_+_92631709 | 0.29 |
ENST00000413330.1 ENST00000277882.3 |
RPP30 |
ribonuclease P/MRP 30kDa subunit |
| chr4_-_113558014 | 0.29 |
ENST00000503172.1 ENST00000505019.1 ENST00000309071.5 |
C4orf21 |
chromosome 4 open reading frame 21 |
| chr5_-_43313574 | 0.28 |
ENST00000325110.6 ENST00000433297.2 |
HMGCS1 |
3-hydroxy-3-methylglutaryl-CoA synthase 1 (soluble) |
| chr16_+_2510081 | 0.28 |
ENST00000361837.4 ENST00000569496.1 ENST00000567489.1 ENST00000563531.1 ENST00000483320.1 |
C16orf59 |
chromosome 16 open reading frame 59 |
| chr4_+_178230985 | 0.28 |
ENST00000264596.3 |
NEIL3 |
nei endonuclease VIII-like 3 (E. coli) |
| chr19_-_6424783 | 0.27 |
ENST00000398148.3 |
KHSRP |
KH-type splicing regulatory protein |
| chr14_-_55658323 | 0.26 |
ENST00000554067.1 ENST00000247191.2 |
DLGAP5 |
discs, large (Drosophila) homolog-associated protein 5 |
| chr6_+_31865552 | 0.25 |
ENST00000469372.1 ENST00000497706.1 |
C2 |
complement component 2 |
| chr1_-_36235559 | 0.25 |
ENST00000251195.5 |
CLSPN |
claspin |
| chr12_-_57472522 | 0.25 |
ENST00000379391.3 ENST00000300128.4 |
TMEM194A |
transmembrane protein 194A |
| chr17_-_41277317 | 0.25 |
ENST00000497488.1 ENST00000489037.1 ENST00000470026.1 ENST00000586385.1 ENST00000591534.1 ENST00000591849.1 |
BRCA1 |
breast cancer 1, early onset |
| chr1_+_173837488 | 0.25 |
ENST00000427304.1 ENST00000432989.1 ENST00000367702.1 |
ZBTB37 |
zinc finger and BTB domain containing 37 |
| chr22_+_19467261 | 0.23 |
ENST00000455750.1 ENST00000437685.2 ENST00000263201.1 ENST00000404724.3 |
CDC45 |
cell division cycle 45 |
| chr1_-_26185844 | 0.23 |
ENST00000538789.1 ENST00000374298.3 |
AUNIP |
aurora kinase A and ninein interacting protein |
| chr17_-_41277370 | 0.23 |
ENST00000476777.1 ENST00000491747.2 ENST00000478531.1 ENST00000477152.1 ENST00000357654.3 ENST00000493795.1 ENST00000493919.1 |
BRCA1 |
breast cancer 1, early onset |
| chr6_+_151561506 | 0.22 |
ENST00000253332.1 |
AKAP12 |
A kinase (PRKA) anchor protein 12 |
| chr2_+_157292933 | 0.22 |
ENST00000540309.1 |
GPD2 |
glycerol-3-phosphate dehydrogenase 2 (mitochondrial) |
| chr17_-_41277467 | 0.22 |
ENST00000494123.1 ENST00000346315.3 ENST00000309486.4 ENST00000468300.1 ENST00000354071.3 ENST00000352993.3 ENST00000471181.2 |
BRCA1 |
breast cancer 1, early onset |
| chr7_-_129845188 | 0.22 |
ENST00000462753.1 ENST00000471077.1 ENST00000473456.1 ENST00000336804.8 |
TMEM209 |
transmembrane protein 209 |
| chr8_-_124408652 | 0.22 |
ENST00000287394.5 |
ATAD2 |
ATPase family, AAA domain containing 2 |
| chr4_-_113558079 | 0.22 |
ENST00000445203.2 |
C4orf21 |
chromosome 4 open reading frame 21 |
| chr6_+_83073952 | 0.21 |
ENST00000543496.1 |
TPBG |
trophoblast glycoprotein |
| chr1_+_63833261 | 0.21 |
ENST00000371108.4 |
ALG6 |
ALG6, alpha-1,3-glucosyltransferase |
| chr9_-_110251836 | 0.21 |
ENST00000374672.4 |
KLF4 |
Kruppel-like factor 4 (gut) |
| chr15_+_89631647 | 0.21 |
ENST00000569550.1 ENST00000565066.1 ENST00000565973.1 |
ABHD2 |
abhydrolase domain containing 2 |
| chr7_+_100199800 | 0.20 |
ENST00000223061.5 |
PCOLCE |
procollagen C-endopeptidase enhancer |
| chr12_-_77459306 | 0.20 |
ENST00000547316.1 ENST00000416496.2 ENST00000550669.1 ENST00000322886.7 |
E2F7 |
E2F transcription factor 7 |
| chr6_-_71666732 | 0.20 |
ENST00000230053.6 |
B3GAT2 |
beta-1,3-glucuronyltransferase 2 (glucuronosyltransferase S) |
| chr1_-_186344802 | 0.20 |
ENST00000451586.1 |
TPR |
translocated promoter region, nuclear basket protein |
| chr1_+_171454639 | 0.20 |
ENST00000392078.3 ENST00000426496.2 |
PRRC2C |
proline-rich coiled-coil 2C |
| chr6_+_31707725 | 0.20 |
ENST00000375755.3 ENST00000375742.3 ENST00000375750.3 ENST00000425703.1 ENST00000534153.4 ENST00000375703.3 ENST00000375740.3 |
MSH5 |
mutS homolog 5 |
| chr13_+_34392200 | 0.20 |
ENST00000434425.1 |
RFC3 |
replication factor C (activator 1) 3, 38kDa |
| chr7_-_150780487 | 0.20 |
ENST00000482202.1 |
TMUB1 |
transmembrane and ubiquitin-like domain containing 1 |
| chr17_-_34890759 | 0.19 |
ENST00000431794.3 |
MYO19 |
myosin XIX |
| chr2_+_173292390 | 0.19 |
ENST00000442250.1 ENST00000458358.1 ENST00000409080.1 |
ITGA6 |
integrin, alpha 6 |
| chr8_+_124084899 | 0.19 |
ENST00000287380.1 ENST00000309336.3 ENST00000519418.1 ENST00000327098.5 ENST00000522420.1 ENST00000521676.1 ENST00000378080.2 |
TBC1D31 |
TBC1 domain family, member 31 |
| chrX_-_14891150 | 0.19 |
ENST00000452869.1 ENST00000398334.1 ENST00000324138.3 |
FANCB |
Fanconi anemia, complementation group B |
| chr19_+_55795493 | 0.19 |
ENST00000309383.1 |
BRSK1 |
BR serine/threonine kinase 1 |
| chr1_+_11333245 | 0.19 |
ENST00000376810.5 |
UBIAD1 |
UbiA prenyltransferase domain containing 1 |
| chr7_-_129845313 | 0.19 |
ENST00000397622.2 |
TMEM209 |
transmembrane protein 209 |
| chr4_+_56815102 | 0.19 |
ENST00000257287.4 |
CEP135 |
centrosomal protein 135kDa |
| chr19_-_5622991 | 0.18 |
ENST00000252542.4 |
SAFB2 |
scaffold attachment factor B2 |
| chr12_-_53574671 | 0.18 |
ENST00000444623.1 |
CSAD |
cysteine sulfinic acid decarboxylase |
| chr14_+_36003248 | 0.18 |
ENST00000307169.3 |
INSM2 |
insulinoma-associated 2 |
| chr16_-_2264779 | 0.18 |
ENST00000333503.7 |
PGP |
phosphoglycolate phosphatase |
| chr18_+_33767473 | 0.18 |
ENST00000261326.5 |
MOCOS |
molybdenum cofactor sulfurase |
| chr15_+_51973680 | 0.18 |
ENST00000542355.2 |
SCG3 |
secretogranin III |
| chr2_+_10262857 | 0.18 |
ENST00000304567.5 |
RRM2 |
ribonucleotide reductase M2 |
| chr7_+_138145145 | 0.17 |
ENST00000415680.2 |
TRIM24 |
tripartite motif containing 24 |
| chr15_+_51973550 | 0.17 |
ENST00000220478.3 |
SCG3 |
secretogranin III |
| chr2_-_11484710 | 0.17 |
ENST00000315872.6 |
ROCK2 |
Rho-associated, coiled-coil containing protein kinase 2 |
| chr9_-_131709858 | 0.17 |
ENST00000372586.3 |
DOLK |
dolichol kinase |
| chr2_+_27440229 | 0.17 |
ENST00000264705.4 ENST00000403525.1 |
CAD |
carbamoyl-phosphate synthetase 2, aspartate transcarbamylase, and dihydroorotase |
| chr8_-_101157680 | 0.17 |
ENST00000428847.2 |
FBXO43 |
F-box protein 43 |
| chr12_+_110338063 | 0.16 |
ENST00000405876.4 |
TCHP |
trichoplein, keratin filament binding |
| chr1_+_173837214 | 0.16 |
ENST00000367704.1 |
ZBTB37 |
zinc finger and BTB domain containing 37 |
| chr15_+_40987327 | 0.16 |
ENST00000423169.2 ENST00000267868.3 ENST00000557850.1 ENST00000532743.1 ENST00000382643.3 |
RAD51 |
RAD51 recombinase |
| chr19_-_48673552 | 0.16 |
ENST00000536218.1 ENST00000596549.1 |
LIG1 |
ligase I, DNA, ATP-dependent |
| chr1_+_171454659 | 0.16 |
ENST00000367742.3 ENST00000338920.4 |
PRRC2C |
proline-rich coiled-coil 2C |
| chr7_-_99698338 | 0.16 |
ENST00000354230.3 ENST00000425308.1 |
MCM7 |
minichromosome maintenance complex component 7 |
| chr4_-_157892498 | 0.16 |
ENST00000502773.1 |
PDGFC |
platelet derived growth factor C |
| chr5_-_127873659 | 0.15 |
ENST00000262464.4 |
FBN2 |
fibrillin 2 |
| chr3_+_133502877 | 0.15 |
ENST00000466490.2 |
SRPRB |
signal recognition particle receptor, B subunit |
| chr4_-_157892055 | 0.15 |
ENST00000422544.2 |
PDGFC |
platelet derived growth factor C |
| chr6_-_20212630 | 0.15 |
ENST00000324607.7 ENST00000541730.1 ENST00000536798.1 |
MBOAT1 |
membrane bound O-acyltransferase domain containing 1 |
| chr1_+_179262905 | 0.15 |
ENST00000539888.1 ENST00000540564.1 ENST00000535686.1 ENST00000367619.3 |
SOAT1 |
sterol O-acyltransferase 1 |
| chr19_+_7953384 | 0.15 |
ENST00000306708.6 |
LRRC8E |
leucine rich repeat containing 8 family, member E |
| chr1_-_182922505 | 0.15 |
ENST00000367547.3 |
SHCBP1L |
SHC SH2-domain binding protein 1-like |
| chr1_+_28844648 | 0.15 |
ENST00000373832.1 ENST00000373831.3 |
RCC1 |
regulator of chromosome condensation 1 |
| chr5_+_109025067 | 0.15 |
ENST00000261483.4 |
MAN2A1 |
mannosidase, alpha, class 2A, member 1 |
| chr16_+_699319 | 0.15 |
ENST00000549091.1 ENST00000293879.4 |
WDR90 |
WD repeat domain 90 |
| chr19_-_47616992 | 0.15 |
ENST00000253048.5 |
ZC3H4 |
zinc finger CCCH-type containing 4 |
| chr19_-_48673580 | 0.15 |
ENST00000427526.2 |
LIG1 |
ligase I, DNA, ATP-dependent |
| chr7_+_138145076 | 0.14 |
ENST00000343526.4 |
TRIM24 |
tripartite motif containing 24 |
| chr12_+_110338323 | 0.14 |
ENST00000312777.5 ENST00000536408.2 |
TCHP |
trichoplein, keratin filament binding |
| chr21_+_47743995 | 0.14 |
ENST00000359568.5 |
PCNT |
pericentrin |
| chr4_-_157892167 | 0.14 |
ENST00000541126.1 |
PDGFC |
platelet derived growth factor C |
| chr6_+_43739697 | 0.14 |
ENST00000230480.6 |
VEGFA |
vascular endothelial growth factor A |
| chr6_-_139308777 | 0.14 |
ENST00000529597.1 ENST00000415951.2 ENST00000367663.4 ENST00000409812.2 |
REPS1 |
RALBP1 associated Eps domain containing 1 |
| chr11_-_118972575 | 0.14 |
ENST00000432443.2 |
DPAGT1 |
dolichyl-phosphate (UDP-N-acetylglucosamine) N-acetylglucosaminephosphotransferase 1 (GlcNAc-1-P transferase) |
| chr10_-_70166946 | 0.14 |
ENST00000388768.2 |
RUFY2 |
RUN and FYVE domain containing 2 |
| chr8_-_38325219 | 0.14 |
ENST00000533668.1 ENST00000413133.2 ENST00000397108.4 ENST00000526742.1 ENST00000525001.1 ENST00000425967.3 ENST00000529552.1 ENST00000397113.2 |
FGFR1 |
fibroblast growth factor receptor 1 |
| chr19_+_5623186 | 0.14 |
ENST00000538656.1 |
SAFB |
scaffold attachment factor B |
| chr7_-_152373216 | 0.14 |
ENST00000359321.1 |
XRCC2 |
X-ray repair complementing defective repair in Chinese hamster cells 2 |
| chr12_-_53574376 | 0.14 |
ENST00000267085.4 ENST00000379850.3 ENST00000379846.1 ENST00000424990.1 |
CSAD |
cysteine sulfinic acid decarboxylase |
| chr1_+_114471809 | 0.14 |
ENST00000426820.2 |
HIPK1 |
homeodomain interacting protein kinase 1 |
| chr13_+_34392185 | 0.14 |
ENST00000380071.3 |
RFC3 |
replication factor C (activator 1) 3, 38kDa |
| chrX_+_100353153 | 0.14 |
ENST00000423383.1 ENST00000218507.5 ENST00000403304.2 ENST00000435570.1 |
CENPI |
centromere protein I |
| chr2_+_17935383 | 0.14 |
ENST00000524465.1 ENST00000381254.2 ENST00000532257.1 |
GEN1 |
GEN1 Holliday junction 5' flap endonuclease |
| chr6_-_139309378 | 0.14 |
ENST00000450536.2 ENST00000258062.5 |
REPS1 |
RALBP1 associated Eps domain containing 1 |
| chr1_-_43833628 | 0.14 |
ENST00000413844.2 ENST00000372458.3 |
ELOVL1 |
ELOVL fatty acid elongase 1 |
| chr5_+_139781393 | 0.13 |
ENST00000360839.2 ENST00000297183.6 ENST00000421134.1 ENST00000394723.3 ENST00000511151.1 |
ANKHD1 |
ankyrin repeat and KH domain containing 1 |
| chr4_-_185655278 | 0.13 |
ENST00000281453.5 |
MLF1IP |
centromere protein U |
| chr1_-_33283754 | 0.13 |
ENST00000373477.4 |
YARS |
tyrosyl-tRNA synthetase |
| chr2_+_17935119 | 0.13 |
ENST00000317402.7 |
GEN1 |
GEN1 Holliday junction 5' flap endonuclease |
| chr1_-_47779762 | 0.13 |
ENST00000371877.3 ENST00000360380.3 ENST00000337817.5 ENST00000447475.2 |
STIL |
SCL/TAL1 interrupting locus |
| chr10_+_60936347 | 0.13 |
ENST00000373880.4 |
PHYHIPL |
phytanoyl-CoA 2-hydroxylase interacting protein-like |
| chr3_-_10028366 | 0.13 |
ENST00000429759.1 |
EMC3 |
ER membrane protein complex subunit 3 |
| chr21_-_44496441 | 0.13 |
ENST00000359624.3 ENST00000352178.5 |
CBS |
cystathionine-beta-synthase |
| chr9_-_35732362 | 0.13 |
ENST00000314888.9 ENST00000540444.1 |
TLN1 |
talin 1 |
| chrX_-_99987088 | 0.13 |
ENST00000372981.1 ENST00000263033.5 |
SYTL4 |
synaptotagmin-like 4 |
| chr4_+_128802016 | 0.13 |
ENST00000270861.5 ENST00000515069.1 ENST00000513090.1 ENST00000507249.1 |
PLK4 |
polo-like kinase 4 |
| chr11_+_82612740 | 0.13 |
ENST00000524921.1 ENST00000528759.1 ENST00000525361.1 ENST00000430323.2 ENST00000533655.1 ENST00000532764.1 ENST00000532589.1 ENST00000525388.1 |
C11orf82 |
chromosome 11 open reading frame 82 |
| chr1_+_173793777 | 0.13 |
ENST00000239457.5 |
DARS2 |
aspartyl-tRNA synthetase 2, mitochondrial |
| chr19_+_5623083 | 0.13 |
ENST00000292123.5 ENST00000592224.1 ENST00000454510.1 ENST00000433404.1 ENST00000588852.1 |
SAFB |
scaffold attachment factor B |
| chr1_+_114472481 | 0.13 |
ENST00000369555.2 |
HIPK1 |
homeodomain interacting protein kinase 1 |
| chr3_-_27525826 | 0.13 |
ENST00000454389.1 ENST00000440156.1 ENST00000437179.1 ENST00000446700.1 ENST00000455077.1 ENST00000435667.2 ENST00000388777.4 ENST00000425128.2 |
SLC4A7 |
solute carrier family 4, sodium bicarbonate cotransporter, member 7 |
| chr11_+_62623621 | 0.12 |
ENST00000535296.1 |
SLC3A2 |
solute carrier family 3 (amino acid transporter heavy chain), member 2 |
| chr5_+_139781445 | 0.12 |
ENST00000532219.1 ENST00000394722.3 |
ANKHD1-EIF4EBP3 ANKHD1 |
ANKHD1-EIF4EBP3 readthrough ankyrin repeat and KH domain containing 1 |
| chr3_+_160117418 | 0.12 |
ENST00000465903.1 ENST00000485645.1 ENST00000360111.2 ENST00000472991.1 ENST00000467468.1 ENST00000469762.1 ENST00000489573.1 ENST00000462787.1 ENST00000490207.1 ENST00000485867.1 |
SMC4 |
structural maintenance of chromosomes 4 |
| chr1_+_173793641 | 0.12 |
ENST00000361951.4 |
DARS2 |
aspartyl-tRNA synthetase 2, mitochondrial |
| chr19_+_14017116 | 0.12 |
ENST00000589606.1 |
CC2D1A |
coiled-coil and C2 domain containing 1A |
| chr5_+_156887027 | 0.12 |
ENST00000435489.2 ENST00000311946.7 |
NIPAL4 |
NIPA-like domain containing 4 |
| chrX_-_122866874 | 0.12 |
ENST00000245838.8 ENST00000355725.4 |
THOC2 |
THO complex 2 |
| chr17_-_43339453 | 0.12 |
ENST00000543122.1 |
SPATA32 |
spermatogenesis associated 32 |
| chr8_-_67525473 | 0.12 |
ENST00000522677.3 |
MYBL1 |
v-myb avian myeloblastosis viral oncogene homolog-like 1 |
| chr6_+_97372596 | 0.12 |
ENST00000369261.4 |
KLHL32 |
kelch-like family member 32 |
| chr11_+_62623512 | 0.12 |
ENST00000377892.1 |
SLC3A2 |
solute carrier family 3 (amino acid transporter heavy chain), member 2 |
| chr3_+_32280159 | 0.12 |
ENST00000458535.2 ENST00000307526.3 |
CMTM8 |
CKLF-like MARVEL transmembrane domain containing 8 |
| chrX_+_129473859 | 0.12 |
ENST00000424447.1 |
SLC25A14 |
solute carrier family 25 (mitochondrial carrier, brain), member 14 |
| chr5_+_892745 | 0.12 |
ENST00000166345.3 |
TRIP13 |
thyroid hormone receptor interactor 13 |
| chr11_+_62623544 | 0.12 |
ENST00000377890.2 ENST00000377891.2 ENST00000377889.2 |
SLC3A2 |
solute carrier family 3 (amino acid transporter heavy chain), member 2 |
| chr21_+_34915337 | 0.12 |
ENST00000290239.6 ENST00000381692.2 ENST00000300278.4 ENST00000381679.4 |
SON |
SON DNA binding protein |
| chr7_-_102985288 | 0.12 |
ENST00000379263.3 |
DNAJC2 |
DnaJ (Hsp40) homolog, subfamily C, member 2 |
| chr1_+_28844778 | 0.12 |
ENST00000411533.1 |
RCC1 |
regulator of chromosome condensation 1 |
| chr18_+_77623668 | 0.12 |
ENST00000316249.3 |
KCNG2 |
potassium voltage-gated channel, subfamily G, member 2 |
| chr3_-_33759541 | 0.12 |
ENST00000468888.2 |
CLASP2 |
cytoplasmic linker associated protein 2 |
| chr6_+_167412835 | 0.11 |
ENST00000349556.4 |
FGFR1OP |
FGFR1 oncogene partner |
| chr9_-_35080013 | 0.11 |
ENST00000378643.3 |
FANCG |
Fanconi anemia, complementation group G |
| chr7_+_77428149 | 0.11 |
ENST00000415251.2 ENST00000275575.7 |
PHTF2 |
putative homeodomain transcription factor 2 |
| chr20_-_32274179 | 0.11 |
ENST00000343380.5 |
E2F1 |
E2F transcription factor 1 |
| chr1_-_229761717 | 0.11 |
ENST00000366675.3 ENST00000258281.2 ENST00000366674.1 |
TAF5L |
TAF5-like RNA polymerase II, p300/CBP-associated factor (PCAF)-associated factor, 65kDa |
| chr5_-_36152031 | 0.11 |
ENST00000296603.4 |
LMBRD2 |
LMBR1 domain containing 2 |
| chr10_-_33623310 | 0.11 |
ENST00000395995.1 ENST00000374823.5 ENST00000374821.5 ENST00000374816.3 |
NRP1 |
neuropilin 1 |
| chr12_-_15942503 | 0.11 |
ENST00000281172.5 |
EPS8 |
epidermal growth factor receptor pathway substrate 8 |
| chr7_+_77428066 | 0.11 |
ENST00000422959.2 ENST00000307305.8 ENST00000424760.1 |
PHTF2 |
putative homeodomain transcription factor 2 |
| chr8_+_48873479 | 0.11 |
ENST00000262105.2 |
MCM4 |
minichromosome maintenance complex component 4 |
| chr19_+_14017003 | 0.11 |
ENST00000318003.7 |
CC2D1A |
coiled-coil and C2 domain containing 1A |
| chr19_-_53238260 | 0.11 |
ENST00000453741.2 ENST00000602162.1 ENST00000601643.1 ENST00000596702.1 ENST00000600943.1 ENST00000543227.1 ENST00000540744.1 |
ZNF611 |
zinc finger protein 611 |
| chr6_-_139695757 | 0.11 |
ENST00000367651.2 |
CITED2 |
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 |
| chr2_-_97405775 | 0.11 |
ENST00000264963.4 ENST00000537039.1 ENST00000377079.4 ENST00000426463.2 ENST00000534882.1 |
LMAN2L |
lectin, mannose-binding 2-like |
| chr15_-_50647274 | 0.11 |
ENST00000543881.1 |
GABPB1 |
GA binding protein transcription factor, beta subunit 1 |
| chr3_-_33759699 | 0.11 |
ENST00000399362.4 ENST00000359576.5 ENST00000307312.7 |
CLASP2 |
cytoplasmic linker associated protein 2 |
| chr3_+_14989186 | 0.11 |
ENST00000435454.1 ENST00000323373.6 |
NR2C2 |
nuclear receptor subfamily 2, group C, member 2 |
| chr20_-_35724388 | 0.11 |
ENST00000344359.3 ENST00000373664.3 |
RBL1 |
retinoblastoma-like 1 (p107) |
| chr12_+_102513950 | 0.11 |
ENST00000378128.3 ENST00000327680.2 ENST00000541394.1 ENST00000543784.1 |
PARPBP |
PARP1 binding protein |
| chr11_-_64851496 | 0.11 |
ENST00000404147.3 ENST00000275517.3 |
CDCA5 |
cell division cycle associated 5 |
| chr17_-_5389477 | 0.11 |
ENST00000572834.1 ENST00000570848.1 ENST00000571971.1 ENST00000158771.4 |
DERL2 |
derlin 2 |
| chr12_+_8850471 | 0.11 |
ENST00000535829.1 ENST00000357529.3 |
RIMKLB |
ribosomal modification protein rimK-like family member B |
| chr1_-_156647189 | 0.11 |
ENST00000368223.3 |
NES |
nestin |
| chr11_+_32914579 | 0.11 |
ENST00000399302.2 |
QSER1 |
glutamine and serine rich 1 |
| chr3_-_27525865 | 0.10 |
ENST00000445684.1 |
SLC4A7 |
solute carrier family 4, sodium bicarbonate cotransporter, member 7 |
| chr12_-_15942309 | 0.10 |
ENST00000544064.1 ENST00000543523.1 ENST00000536793.1 |
EPS8 |
epidermal growth factor receptor pathway substrate 8 |
| chr3_-_136471204 | 0.10 |
ENST00000480733.1 ENST00000383202.2 ENST00000236698.5 ENST00000434713.2 |
STAG1 |
stromal antigen 1 |
| chr11_+_117198801 | 0.10 |
ENST00000527609.1 ENST00000533570.1 |
CEP164 |
centrosomal protein 164kDa |
| chr1_+_114472222 | 0.10 |
ENST00000369558.1 ENST00000369561.4 |
HIPK1 |
homeodomain interacting protein kinase 1 |
| chr12_-_53574418 | 0.10 |
ENST00000379843.3 ENST00000453446.2 ENST00000437073.1 |
CSAD |
cysteine sulfinic acid decarboxylase |
| chr15_-_50647370 | 0.10 |
ENST00000558970.1 ENST00000396464.3 ENST00000560825.1 |
GABPB1 |
GA binding protein transcription factor, beta subunit 1 |
| chr2_+_71357434 | 0.10 |
ENST00000244230.2 |
MPHOSPH10 |
M-phase phosphoprotein 10 (U3 small nucleolar ribonucleoprotein) |
| chr5_+_149737202 | 0.10 |
ENST00000451292.1 ENST00000377797.3 ENST00000445265.2 ENST00000323668.7 ENST00000439160.2 ENST00000394269.3 ENST00000427724.2 ENST00000504761.2 ENST00000513346.1 ENST00000515516.1 |
TCOF1 |
Treacher Collins-Franceschetti syndrome 1 |
| chr2_-_24583583 | 0.10 |
ENST00000355123.4 |
ITSN2 |
intersectin 2 |
| chr19_-_10613421 | 0.10 |
ENST00000393623.2 |
KEAP1 |
kelch-like ECH-associated protein 1 |
| chr9_+_131709966 | 0.10 |
ENST00000372577.2 |
NUP188 |
nucleoporin 188kDa |
| chr20_+_47662805 | 0.10 |
ENST00000262982.2 ENST00000542325.1 |
CSE1L |
CSE1 chromosome segregation 1-like (yeast) |
| chr6_-_41863098 | 0.10 |
ENST00000373006.1 |
USP49 |
ubiquitin specific peptidase 49 |
| chr8_+_48873453 | 0.10 |
ENST00000523944.1 |
MCM4 |
minichromosome maintenance complex component 4 |
| chr7_+_100026406 | 0.10 |
ENST00000414441.1 |
MEPCE |
methylphosphate capping enzyme |
| chr5_+_65222299 | 0.10 |
ENST00000284037.5 |
ERBB2IP |
erbb2 interacting protein |
| chr11_+_58910201 | 0.10 |
ENST00000528737.1 |
FAM111A |
family with sequence similarity 111, member A |
| chr16_+_24741013 | 0.10 |
ENST00000315183.7 ENST00000395799.3 |
TNRC6A |
trinucleotide repeat containing 6A |
| chr7_-_149158187 | 0.10 |
ENST00000247930.4 |
ZNF777 |
zinc finger protein 777 |
| chr1_+_64936428 | 0.10 |
ENST00000371073.2 ENST00000290039.5 |
CACHD1 |
cache domain containing 1 |
| chr3_+_184097905 | 0.10 |
ENST00000450923.1 |
CHRD |
chordin |
| chr16_-_66835480 | 0.10 |
ENST00000559050.1 ENST00000558713.2 ENST00000433154.1 ENST00000432602.1 ENST00000433574.1 ENST00000415744.1 |
CCDC79 |
coiled-coil domain containing 79 |
| chr2_-_203736334 | 0.10 |
ENST00000392237.2 ENST00000416760.1 ENST00000412210.1 |
ICA1L |
islet cell autoantigen 1,69kDa-like |
| chr19_+_50706866 | 0.09 |
ENST00000440075.2 ENST00000376970.2 ENST00000425460.1 ENST00000599920.1 ENST00000601313.1 |
MYH14 |
myosin, heavy chain 14, non-muscle |
| chr17_-_12921270 | 0.09 |
ENST00000578071.1 ENST00000426905.3 ENST00000395962.2 ENST00000583371.1 ENST00000338034.4 |
ELAC2 |
elaC ribonuclease Z 2 |
| chr12_+_56521840 | 0.09 |
ENST00000394048.5 |
ESYT1 |
extended synaptotagmin-like protein 1 |
| chr19_+_50887585 | 0.09 |
ENST00000440232.2 ENST00000601098.1 ENST00000599857.1 ENST00000593887.1 |
POLD1 |
polymerase (DNA directed), delta 1, catalytic subunit |
| chr10_+_105036909 | 0.09 |
ENST00000369849.4 |
INA |
internexin neuronal intermediate filament protein, alpha |
| chr22_-_31741757 | 0.09 |
ENST00000215919.3 |
PATZ1 |
POZ (BTB) and AT hook containing zinc finger 1 |
| chr5_-_79950371 | 0.09 |
ENST00000511032.1 ENST00000504396.1 ENST00000505337.1 |
DHFR |
dihydrofolate reductase |
| chr20_-_35402123 | 0.09 |
ENST00000373740.3 ENST00000426836.1 ENST00000373745.3 ENST00000448110.2 ENST00000438549.1 ENST00000447406.1 ENST00000373750.4 ENST00000373734.4 |
DSN1 |
DSN1, MIS12 kinetochore complex component |
| chr10_-_30024716 | 0.09 |
ENST00000375398.2 ENST00000375400.3 |
SVIL |
supervillin |
| chr8_-_67525524 | 0.09 |
ENST00000517885.1 |
MYBL1 |
v-myb avian myeloblastosis viral oncogene homolog-like 1 |
| chr5_+_79950463 | 0.09 |
ENST00000265081.6 |
MSH3 |
mutS homolog 3 |
| chr14_-_50154921 | 0.09 |
ENST00000553805.2 ENST00000554396.1 ENST00000216367.5 ENST00000539565.2 |
POLE2 |
polymerase (DNA directed), epsilon 2, accessory subunit |
| chr9_+_17134980 | 0.09 |
ENST00000380647.3 |
CNTLN |
centlein, centrosomal protein |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0070510 | regulation of histone H4-K20 methylation(GO:0070510) positive regulation of histone H4-K20 methylation(GO:0070512) |
| 0.1 | 0.4 | GO:0046167 | glycerol-3-phosphate biosynthetic process(GO:0046167) |
| 0.1 | 0.7 | GO:0033567 | DNA replication, Okazaki fragment processing(GO:0033567) |
| 0.1 | 0.4 | GO:0009786 | regulation of asymmetric cell division(GO:0009786) |
| 0.1 | 0.6 | GO:0006127 | glycerophosphate shuttle(GO:0006127) |
| 0.1 | 0.4 | GO:0042412 | taurine biosynthetic process(GO:0042412) |
| 0.1 | 0.3 | GO:0071930 | negative regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0071930) |
| 0.1 | 0.7 | GO:1900262 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 0.1 | 0.8 | GO:0048478 | replication fork protection(GO:0048478) |
| 0.1 | 0.3 | GO:0046832 | negative regulation of nucleobase-containing compound transport(GO:0032240) negative regulation of RNA export from nucleus(GO:0046832) |
| 0.1 | 0.3 | GO:0071140 | resolution of recombination intermediates(GO:0071139) resolution of mitotic recombination intermediates(GO:0071140) |
| 0.1 | 0.3 | GO:0006421 | asparaginyl-tRNA aminoacylation(GO:0006421) |
| 0.1 | 0.2 | GO:0036388 | pre-replicative complex assembly involved in nuclear cell cycle DNA replication(GO:0006267) pre-replicative complex assembly(GO:0036388) pre-replicative complex assembly involved in cell cycle DNA replication(GO:1902299) |
| 0.1 | 0.4 | GO:0060356 | leucine import(GO:0060356) |
| 0.1 | 0.2 | GO:0014740 | negative regulation of muscle hyperplasia(GO:0014740) |
| 0.1 | 0.2 | GO:0009107 | lipoate biosynthetic process(GO:0009107) |
| 0.1 | 0.2 | GO:0044205 | 'de novo' UMP biosynthetic process(GO:0044205) |
| 0.1 | 0.6 | GO:0033314 | mitotic DNA replication checkpoint(GO:0033314) |
| 0.1 | 0.2 | GO:1990414 | replication-born double-strand break repair via sister chromatid exchange(GO:1990414) |
| 0.1 | 0.1 | GO:0034085 | establishment of sister chromatid cohesion(GO:0034085) cohesin loading(GO:0071921) regulation of cohesin loading(GO:0071922) |
| 0.0 | 0.3 | GO:2000348 | regulation of CD40 signaling pathway(GO:2000348) |
| 0.0 | 0.1 | GO:1903570 | regulation of protein kinase D signaling(GO:1903570) positive regulation of protein kinase D signaling(GO:1903572) |
| 0.0 | 0.4 | GO:0060235 | lens induction in camera-type eye(GO:0060235) |
| 0.0 | 0.1 | GO:2000830 | vacuolar phosphate transport(GO:0007037) positive regulation of mitotic cell cycle DNA replication(GO:1903465) positive regulation of parathyroid hormone secretion(GO:2000830) |
| 0.0 | 0.3 | GO:0070562 | regulation of vitamin D receptor signaling pathway(GO:0070562) |
| 0.0 | 0.2 | GO:0051083 | 'de novo' cotranslational protein folding(GO:0051083) |
| 0.0 | 0.2 | GO:0006114 | glycerol biosynthetic process(GO:0006114) |
| 0.0 | 0.1 | GO:0006535 | cysteine biosynthetic process from serine(GO:0006535) |
| 0.0 | 0.2 | GO:0070358 | actin polymerization-dependent cell motility(GO:0070358) |
| 0.0 | 0.2 | GO:1904978 | regulation of endosome organization(GO:1904978) |
| 0.0 | 0.3 | GO:0007079 | mitotic chromosome movement towards spindle pole(GO:0007079) |
| 0.0 | 0.1 | GO:1900085 | negative regulation of peptidyl-tyrosine autophosphorylation(GO:1900085) negative regulation of inward rectifier potassium channel activity(GO:1903609) |
| 0.0 | 0.2 | GO:0035509 | negative regulation of myosin-light-chain-phosphatase activity(GO:0035509) negative regulation of bicellular tight junction assembly(GO:1903347) |
| 0.0 | 0.2 | GO:0042986 | positive regulation of amyloid precursor protein biosynthetic process(GO:0042986) |
| 0.0 | 0.2 | GO:0009233 | menaquinone metabolic process(GO:0009233) |
| 0.0 | 0.3 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.0 | 0.3 | GO:0006489 | dolichyl diphosphate biosynthetic process(GO:0006489) dolichyl diphosphate metabolic process(GO:0046465) |
| 0.0 | 0.1 | GO:0021919 | BMP signaling pathway involved in spinal cord dorsal/ventral patterning(GO:0021919) |
| 0.0 | 0.1 | GO:1990180 | mitochondrial tRNA 3'-end processing(GO:1990180) |
| 0.0 | 0.1 | GO:1902534 | single-organism membrane invagination(GO:1902534) |
| 0.0 | 0.3 | GO:0051096 | positive regulation of helicase activity(GO:0051096) |
| 0.0 | 0.1 | GO:0061428 | negative regulation of transcription from RNA polymerase II promoter in response to hypoxia(GO:0061428) |
| 0.0 | 0.3 | GO:2000628 | regulation of miRNA metabolic process(GO:2000628) |
| 0.0 | 0.1 | GO:0033504 | floor plate development(GO:0033504) |
| 0.0 | 0.1 | GO:0046601 | positive regulation of centriole replication(GO:0046601) |
| 0.0 | 0.1 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
| 0.0 | 0.1 | GO:0061441 | renal artery morphogenesis(GO:0061441) |
| 0.0 | 0.1 | GO:0046452 | dihydrofolate metabolic process(GO:0046452) |
| 0.0 | 0.2 | GO:0031936 | negative regulation of chromatin silencing(GO:0031936) |
| 0.0 | 0.1 | GO:0000727 | double-strand break repair via break-induced replication(GO:0000727) |
| 0.0 | 0.2 | GO:0019720 | Mo-molybdopterin cofactor biosynthetic process(GO:0006777) Mo-molybdopterin cofactor metabolic process(GO:0019720) |
| 0.0 | 0.2 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination(GO:2000042) |
| 0.0 | 0.1 | GO:0021526 | medial motor column neuron differentiation(GO:0021526) |
| 0.0 | 0.1 | GO:0006013 | mannose metabolic process(GO:0006013) |
| 0.0 | 0.3 | GO:2000427 | positive regulation of apoptotic cell clearance(GO:2000427) |
| 0.0 | 0.1 | GO:0034970 | histone H3-R2 methylation(GO:0034970) |
| 0.0 | 0.1 | GO:0050976 | detection of mechanical stimulus involved in sensory perception of touch(GO:0050976) |
| 0.0 | 0.3 | GO:0071372 | cellular response to follicle-stimulating hormone stimulus(GO:0071372) |
| 0.0 | 0.1 | GO:0000707 | meiotic DNA recombinase assembly(GO:0000707) |
| 0.0 | 0.2 | GO:0090091 | positive regulation of extracellular matrix disassembly(GO:0090091) |
| 0.0 | 0.3 | GO:1902018 | negative regulation of cilium assembly(GO:1902018) |
| 0.0 | 0.1 | GO:0060380 | regulation of single-stranded telomeric DNA binding(GO:0060380) positive regulation of single-stranded telomeric DNA binding(GO:0060381) |
| 0.0 | 0.5 | GO:0007171 | activation of transmembrane receptor protein tyrosine kinase activity(GO:0007171) |
| 0.0 | 0.1 | GO:0090521 | glomerular visceral epithelial cell migration(GO:0090521) |
| 0.0 | 0.0 | GO:0006106 | fumarate metabolic process(GO:0006106) |
| 0.0 | 0.2 | GO:0010739 | positive regulation of protein kinase A signaling(GO:0010739) |
| 0.0 | 0.3 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.0 | 0.0 | GO:0006424 | glutamyl-tRNA aminoacylation(GO:0006424) |
| 0.0 | 0.1 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 0.0 | 0.2 | GO:0051026 | chiasma assembly(GO:0051026) |
| 0.0 | 0.1 | GO:0006287 | base-excision repair, gap-filling(GO:0006287) |
| 0.0 | 0.2 | GO:0035583 | sequestering of TGFbeta in extracellular matrix(GO:0035583) |
| 0.0 | 0.1 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.0 | 0.2 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.0 | 0.0 | GO:0051758 | homologous chromosome movement towards spindle pole involved in homologous chromosome segregation(GO:0051758) |
| 0.0 | 0.1 | GO:0019368 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.0 | 0.1 | GO:0090526 | regulation of gluconeogenesis involved in cellular glucose homeostasis(GO:0090526) |
| 0.0 | 0.1 | GO:0002051 | osteoblast fate commitment(GO:0002051) |
| 0.0 | 0.0 | GO:0035494 | SNARE complex disassembly(GO:0035494) |
| 0.0 | 0.4 | GO:0051290 | protein heterotetramerization(GO:0051290) |
| 0.0 | 0.2 | GO:0010668 | ectodermal cell differentiation(GO:0010668) |
| 0.0 | 0.1 | GO:0072719 | cellular response to cisplatin(GO:0072719) |
| 0.0 | 0.1 | GO:0019236 | response to pheromone(GO:0019236) |
| 0.0 | 0.1 | GO:0072718 | response to cisplatin(GO:0072718) |
| 0.0 | 0.2 | GO:0010457 | centriole-centriole cohesion(GO:0010457) |
| 0.0 | 0.1 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 0.0 | 0.0 | GO:0046104 | thymidine metabolic process(GO:0046104) pyrimidine deoxyribonucleoside metabolic process(GO:0046125) |
| 0.0 | 0.5 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.0 | 0.1 | GO:0061055 | myotome development(GO:0061055) |
| 0.0 | 0.2 | GO:0045835 | negative regulation of meiotic nuclear division(GO:0045835) |
| 0.0 | 0.1 | GO:0045144 | meiotic sister chromatid segregation(GO:0045144) |
| 0.0 | 0.0 | GO:0031118 | rRNA pseudouridine synthesis(GO:0031118) |
| 0.0 | 0.1 | GO:0040031 | snRNA modification(GO:0040031) |
| 0.0 | 0.1 | GO:0007144 | female meiosis I(GO:0007144) |
| 0.0 | 0.1 | GO:0035811 | negative regulation of urine volume(GO:0035811) |
| 0.0 | 0.1 | GO:0010499 | proteasomal ubiquitin-independent protein catabolic process(GO:0010499) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0031436 | BRCA1-BARD1 complex(GO:0031436) |
| 0.1 | 0.6 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.1 | 0.1 | GO:0032302 | MutSbeta complex(GO:0032302) |
| 0.1 | 0.7 | GO:0031390 | Ctf18 RFC-like complex(GO:0031390) |
| 0.1 | 0.3 | GO:0097135 | cyclin E2-CDK2 complex(GO:0097135) |
| 0.1 | 0.2 | GO:0005656 | nuclear pre-replicative complex(GO:0005656) pre-replicative complex(GO:0036387) |
| 0.1 | 0.2 | GO:0034676 | integrin alpha6-beta4 complex(GO:0034676) |
| 0.1 | 0.3 | GO:0005726 | perichromatin fibrils(GO:0005726) |
| 0.1 | 1.0 | GO:0000800 | lateral element(GO:0000800) |
| 0.1 | 0.2 | GO:0032301 | MutSalpha complex(GO:0032301) |
| 0.1 | 0.5 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.0 | 0.3 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.0 | 0.1 | GO:0035189 | Rb-E2F complex(GO:0035189) |
| 0.0 | 0.2 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 0.0 | 0.3 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.0 | 0.2 | GO:0034457 | Mpp10 complex(GO:0034457) |
| 0.0 | 0.1 | GO:0000939 | condensed chromosome inner kinetochore(GO:0000939) |
| 0.0 | 0.6 | GO:0042575 | DNA polymerase complex(GO:0042575) |
| 0.0 | 0.5 | GO:0008091 | spectrin(GO:0008091) |
| 0.0 | 0.1 | GO:0017102 | methionyl glutamyl tRNA synthetase complex(GO:0017102) |
| 0.0 | 0.4 | GO:0042555 | MCM complex(GO:0042555) |
| 0.0 | 0.3 | GO:0005785 | signal recognition particle receptor complex(GO:0005785) |
| 0.0 | 0.1 | GO:0098536 | deuterosome(GO:0098536) |
| 0.0 | 0.1 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 0.0 | 0.1 | GO:0005797 | Golgi medial cisterna(GO:0005797) |
| 0.0 | 0.4 | GO:0097539 | ciliary transition fiber(GO:0097539) |
| 0.0 | 0.1 | GO:0008278 | cohesin complex(GO:0008278) |
| 0.0 | 0.1 | GO:0097513 | myosin II filament(GO:0097513) |
| 0.0 | 0.3 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.0 | 0.3 | GO:0042405 | nuclear inclusion body(GO:0042405) |
| 0.0 | 0.2 | GO:0032300 | mismatch repair complex(GO:0032300) |
| 0.0 | 0.1 | GO:0044609 | DBIRD complex(GO:0044609) |
| 0.0 | 0.1 | GO:0035061 | interchromatin granule(GO:0035061) |
| 0.0 | 0.1 | GO:0033063 | Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.0 | 0.2 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.0 | 0.2 | GO:0005828 | kinetochore microtubule(GO:0005828) |
| 0.0 | 0.2 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 0.0 | 0.2 | GO:0005883 | neurofilament(GO:0005883) |
| 0.0 | 0.0 | GO:0031523 | Myb complex(GO:0031523) |
| 0.0 | 0.1 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.0 | 0.1 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.0 | 0.2 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 0.0 | GO:0031510 | SUMO activating enzyme complex(GO:0031510) |
| 0.0 | 0.1 | GO:0000796 | condensin complex(GO:0000796) |
| 0.0 | 0.1 | GO:0000347 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.0 | 0.1 | GO:0071204 | histone pre-mRNA 3'end processing complex(GO:0071204) |
| 0.0 | 0.1 | GO:1990604 | IRE1-TRAF2-ASK1 complex(GO:1990604) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0052591 | sn-glycerol-3-phosphate:ubiquinone oxidoreductase activity(GO:0052590) sn-glycerol-3-phosphate:ubiquinone-8 oxidoreductase activity(GO:0052591) |
| 0.1 | 0.4 | GO:0004370 | glycerol kinase activity(GO:0004370) |
| 0.1 | 0.4 | GO:0004782 | sulfinoalanine decarboxylase activity(GO:0004782) |
| 0.1 | 1.3 | GO:0043142 | single-stranded DNA-dependent ATPase activity(GO:0043142) |
| 0.1 | 0.7 | GO:0000405 | bubble DNA binding(GO:0000405) |
| 0.1 | 0.4 | GO:0010484 | H3 histone acetyltransferase activity(GO:0010484) |
| 0.1 | 0.6 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) |
| 0.1 | 0.2 | GO:0016992 | lipoate synthase activity(GO:0016992) radical SAM enzyme activity(GO:0070283) |
| 0.1 | 0.2 | GO:0030292 | protein tyrosine kinase inhibitor activity(GO:0030292) |
| 0.1 | 0.3 | GO:0032142 | single guanine insertion binding(GO:0032142) |
| 0.1 | 0.2 | GO:0004088 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 0.0 | 0.2 | GO:0004815 | aspartate-tRNA ligase activity(GO:0004815) |
| 0.0 | 0.2 | GO:0015018 | galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase activity(GO:0015018) |
| 0.0 | 0.3 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.0 | 0.2 | GO:0043812 | phosphatidylinositol-4-phosphate phosphatase activity(GO:0043812) |
| 0.0 | 0.1 | GO:0098809 | cystathionine beta-synthase activity(GO:0004122) oxidoreductase activity, acting on other nitrogenous compounds as donors, cytochrome as acceptor(GO:0016662) nitrite reductase (NO-forming) activity(GO:0050421) carbon monoxide binding(GO:0070025) nitrite reductase activity(GO:0098809) |
| 0.0 | 0.2 | GO:0001010 | transcription factor activity, sequence-specific DNA binding transcription factor recruiting(GO:0001010) |
| 0.0 | 0.1 | GO:0070320 | inward rectifier potassium channel inhibitor activity(GO:0070320) |
| 0.0 | 0.2 | GO:0098519 | nucleotide phosphatase activity, acting on free nucleotides(GO:0098519) |
| 0.0 | 0.2 | GO:0030151 | molybdenum ion binding(GO:0030151) |
| 0.0 | 0.1 | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity(GO:0003976) |
| 0.0 | 0.2 | GO:0004748 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 0.0 | 0.3 | GO:0005087 | Ran guanyl-nucleotide exchange factor activity(GO:0005087) |
| 0.0 | 0.1 | GO:0071207 | histone pre-mRNA stem-loop binding(GO:0071207) |
| 0.0 | 0.3 | GO:0003910 | DNA ligase (ATP) activity(GO:0003910) |
| 0.0 | 0.3 | GO:0034056 | estrogen response element binding(GO:0034056) |
| 0.0 | 0.3 | GO:0008821 | crossover junction endodeoxyribonuclease activity(GO:0008821) |
| 0.0 | 0.1 | GO:0032093 | SAM domain binding(GO:0032093) |
| 0.0 | 0.2 | GO:0061649 | ubiquitinated histone binding(GO:0061649) |
| 0.0 | 0.2 | GO:0005047 | signal recognition particle binding(GO:0005047) |
| 0.0 | 0.1 | GO:0043183 | vascular endothelial growth factor receptor 1 binding(GO:0043183) |
| 0.0 | 0.3 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 0.4 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
| 0.0 | 0.2 | GO:0060002 | plus-end directed microfilament motor activity(GO:0060002) |
| 0.0 | 0.1 | GO:0003896 | DNA primase activity(GO:0003896) |
| 0.0 | 0.5 | GO:0005161 | platelet-derived growth factor receptor binding(GO:0005161) |
| 0.0 | 0.3 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.0 | 0.2 | GO:0030023 | extracellular matrix constituent conferring elasticity(GO:0030023) |
| 0.0 | 0.2 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.0 | 0.1 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.0 | 0.2 | GO:0043138 | 3'-5' DNA helicase activity(GO:0043138) |
| 0.0 | 0.0 | GO:0004797 | thymidine kinase activity(GO:0004797) |
| 0.0 | 0.2 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.0 | 0.1 | GO:0038085 | vascular endothelial growth factor binding(GO:0038085) |
| 0.0 | 0.0 | GO:0004818 | glutamate-tRNA ligase activity(GO:0004818) |
| 0.0 | 0.1 | GO:0051870 | methotrexate binding(GO:0051870) |
| 0.0 | 0.1 | GO:0102337 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.0 | 0.1 | GO:0016262 | protein N-acetylglucosaminyltransferase activity(GO:0016262) protein O-GlcNAc transferase activity(GO:0097363) |
| 0.0 | 0.0 | GO:0008940 | nitrate reductase activity(GO:0008940) molybdopterin adenylyltransferase activity(GO:0061598) molybdopterin molybdotransferase activity(GO:0061599) |
| 0.0 | 0.3 | GO:0005225 | volume-sensitive anion channel activity(GO:0005225) |
| 0.0 | 0.1 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.0 | 0.4 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.0 | 0.1 | GO:0004549 | tRNA-specific ribonuclease activity(GO:0004549) |
| 0.0 | 0.2 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.0 | 0.2 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.0 | 0.1 | GO:0016427 | tRNA (cytosine) methyltransferase activity(GO:0016427) |
| 0.0 | 0.3 | GO:0043495 | protein anchor(GO:0043495) |
| 0.0 | 0.2 | GO:0046527 | glucosyltransferase activity(GO:0046527) |
| 0.0 | 0.1 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.0 | 0.2 | GO:0047144 | 2-acylglycerol-3-phosphate O-acyltransferase activity(GO:0047144) |
| 0.0 | 0.1 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.0 | 0.1 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 0.0 | 0.2 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.0 | 0.2 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.0 | 0.1 | GO:0015924 | mannosyl-oligosaccharide mannosidase activity(GO:0015924) |
| 0.0 | 0.2 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.0 | 0.8 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.0 | 0.4 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 2.3 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 0.1 | PID PI3KCI AKT PATHWAY | Class I PI3K signaling events mediated by Akt |
| 0.0 | 1.1 | PID ATR PATHWAY | ATR signaling pathway |
| 0.0 | 0.3 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.0 | 0.2 | ST JAK STAT PATHWAY | Jak-STAT Pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.8 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.0 | 1.1 | REACTOME LAGGING STRAND SYNTHESIS | Genes involved in Lagging Strand Synthesis |
| 0.0 | 0.1 | REACTOME REPAIR SYNTHESIS FOR GAP FILLING BY DNA POL IN TC NER | Genes involved in Repair synthesis for gap-filling by DNA polymerase in TC-NER |
| 0.0 | 0.7 | REACTOME DNA STRAND ELONGATION | Genes involved in DNA strand elongation |
| 0.0 | 0.7 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 0.7 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.0 | 0.3 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.0 | 0.5 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 0.2 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.0 | 0.6 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.0 | 0.5 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 0.5 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.0 | 0.5 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.0 | 0.3 | REACTOME INITIAL TRIGGERING OF COMPLEMENT | Genes involved in Initial triggering of complement |
| 0.0 | 0.1 | REACTOME ACTIVATION OF THE PRE REPLICATIVE COMPLEX | Genes involved in Activation of the pre-replicative complex |
| 0.0 | 0.3 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |