Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
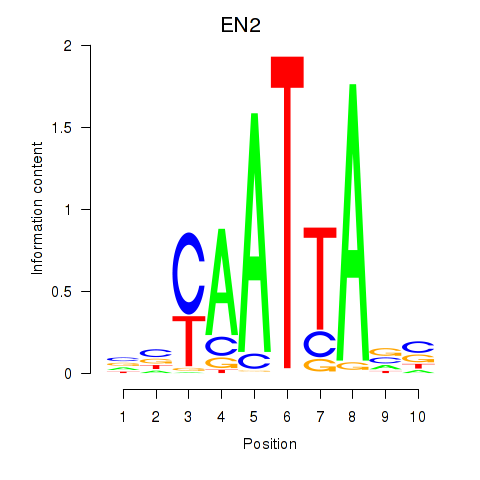
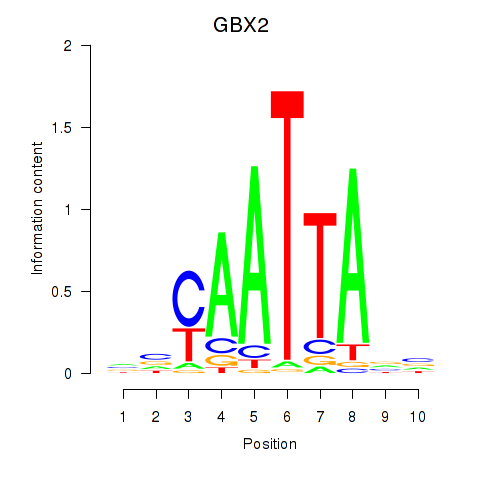
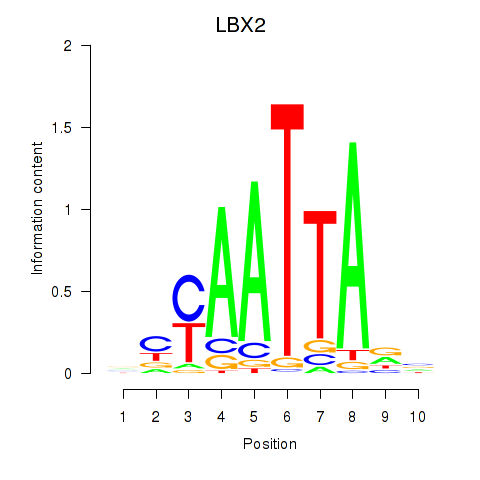
Results for EN2_GBX2_LBX2
Z-value: 0.34
Transcription factors associated with EN2_GBX2_LBX2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
EN2
|
ENSG00000164778.4 | EN2 |
|
GBX2
|
ENSG00000168505.6 | GBX2 |
|
LBX2
|
ENSG00000179528.11 | LBX2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| GBX2 | hg19_v2_chr2_-_237076992_237077012 | 0.64 | 8.6e-02 | Click! |
| EN2 | hg19_v2_chr7_+_155250824_155250824 | -0.06 | 8.8e-01 | Click! |
Activity profile of EN2_GBX2_LBX2 motif
Sorted Z-values of EN2_GBX2_LBX2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of EN2_GBX2_LBX2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr16_+_53133070 | 0.31 |
ENST00000565832.1 |
CHD9 |
chromodomain helicase DNA binding protein 9 |
| chr17_+_18086392 | 0.28 |
ENST00000541285.1 |
ALKBH5 |
alkB, alkylation repair homolog 5 (E. coli) |
| chr17_+_59489112 | 0.24 |
ENST00000335108.2 |
C17orf82 |
chromosome 17 open reading frame 82 |
| chr1_+_40713573 | 0.22 |
ENST00000372766.3 |
TMCO2 |
transmembrane and coiled-coil domains 2 |
| chr6_-_31107127 | 0.18 |
ENST00000259845.4 |
PSORS1C2 |
psoriasis susceptibility 1 candidate 2 |
| chr6_-_32157947 | 0.17 |
ENST00000375050.4 |
PBX2 |
pre-B-cell leukemia homeobox 2 |
| chr7_+_77428066 | 0.16 |
ENST00000422959.2 ENST00000307305.8 ENST00000424760.1 |
PHTF2 |
putative homeodomain transcription factor 2 |
| chr18_-_71959159 | 0.15 |
ENST00000494131.2 ENST00000397914.4 ENST00000340533.4 |
CYB5A |
cytochrome b5 type A (microsomal) |
| chr14_-_51027838 | 0.13 |
ENST00000555216.1 |
MAP4K5 |
mitogen-activated protein kinase kinase kinase kinase 5 |
| chr11_+_67250490 | 0.13 |
ENST00000528641.2 ENST00000279146.3 |
AIP |
aryl hydrocarbon receptor interacting protein |
| chr11_-_40315640 | 0.13 |
ENST00000278198.2 |
LRRC4C |
leucine rich repeat containing 4C |
| chr1_+_101003687 | 0.13 |
ENST00000315033.4 |
GPR88 |
G protein-coupled receptor 88 |
| chr12_+_81110684 | 0.12 |
ENST00000228644.3 |
MYF5 |
myogenic factor 5 |
| chr9_+_2159850 | 0.12 |
ENST00000416751.1 |
SMARCA2 |
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr2_-_73460334 | 0.11 |
ENST00000258083.2 |
PRADC1 |
protease-associated domain containing 1 |
| chr5_-_147286065 | 0.10 |
ENST00000318315.4 ENST00000515291.1 |
C5orf46 |
chromosome 5 open reading frame 46 |
| chr4_+_71458012 | 0.09 |
ENST00000449493.2 |
AMBN |
ameloblastin (enamel matrix protein) |
| chr13_-_36788718 | 0.09 |
ENST00000317764.6 ENST00000379881.3 |
SOHLH2 |
spermatogenesis and oogenesis specific basic helix-loop-helix 2 |
| chrX_-_46187069 | 0.09 |
ENST00000446884.1 |
RP1-30G7.2 |
RP1-30G7.2 |
| chr9_+_12693336 | 0.09 |
ENST00000381137.2 ENST00000388918.5 |
TYRP1 |
tyrosinase-related protein 1 |
| chr1_-_151762943 | 0.08 |
ENST00000368825.3 ENST00000368823.1 ENST00000458431.2 ENST00000368827.6 ENST00000368824.3 |
TDRKH |
tudor and KH domain containing |
| chr1_-_232651312 | 0.08 |
ENST00000262861.4 |
SIPA1L2 |
signal-induced proliferation-associated 1 like 2 |
| chr12_-_53171128 | 0.08 |
ENST00000332411.2 |
KRT76 |
keratin 76 |
| chr12_+_7013897 | 0.08 |
ENST00000007969.8 ENST00000323702.5 |
LRRC23 |
leucine rich repeat containing 23 |
| chr7_+_77428149 | 0.08 |
ENST00000415251.2 ENST00000275575.7 |
PHTF2 |
putative homeodomain transcription factor 2 |
| chr12_+_7014064 | 0.08 |
ENST00000443597.2 |
LRRC23 |
leucine rich repeat containing 23 |
| chr4_-_57547454 | 0.08 |
ENST00000556376.2 |
HOPX |
HOP homeobox |
| chr15_-_55562582 | 0.08 |
ENST00000396307.2 |
RAB27A |
RAB27A, member RAS oncogene family |
| chr4_-_57547870 | 0.08 |
ENST00000381260.3 ENST00000420433.1 ENST00000554144.1 ENST00000557328.1 |
HOPX |
HOP homeobox |
| chr18_-_44181442 | 0.08 |
ENST00000398722.4 |
LOXHD1 |
lipoxygenase homology domains 1 |
| chr9_+_136501478 | 0.07 |
ENST00000393056.2 ENST00000263611.2 |
DBH |
dopamine beta-hydroxylase (dopamine beta-monooxygenase) |
| chr6_+_43968306 | 0.07 |
ENST00000442114.2 ENST00000336600.5 ENST00000439969.2 |
C6orf223 |
chromosome 6 open reading frame 223 |
| chr6_+_34204642 | 0.07 |
ENST00000347617.6 ENST00000401473.3 ENST00000311487.5 ENST00000447654.1 ENST00000395004.3 |
HMGA1 |
high mobility group AT-hook 1 |
| chr6_+_111408698 | 0.07 |
ENST00000368851.5 |
SLC16A10 |
solute carrier family 16 (aromatic amino acid transporter), member 10 |
| chr5_-_95297534 | 0.07 |
ENST00000513343.1 ENST00000431061.2 |
ELL2 |
elongation factor, RNA polymerase II, 2 |
| chr16_+_69345243 | 0.07 |
ENST00000254950.11 |
VPS4A |
vacuolar protein sorting 4 homolog A (S. cerevisiae) |
| chr8_+_22424551 | 0.07 |
ENST00000523348.1 |
SORBS3 |
sorbin and SH3 domain containing 3 |
| chr19_-_14064114 | 0.06 |
ENST00000585607.1 ENST00000538517.2 ENST00000587458.1 ENST00000538371.2 |
PODNL1 |
podocan-like 1 |
| chr11_-_31531121 | 0.06 |
ENST00000532287.1 ENST00000526776.1 ENST00000534812.1 ENST00000529749.1 ENST00000278200.1 ENST00000530023.1 ENST00000533642.1 |
IMMP1L |
IMP1 inner mitochondrial membrane peptidase-like (S. cerevisiae) |
| chr6_-_167040731 | 0.06 |
ENST00000265678.4 |
RPS6KA2 |
ribosomal protein S6 kinase, 90kDa, polypeptide 2 |
| chr6_+_39760129 | 0.06 |
ENST00000274867.4 |
DAAM2 |
dishevelled associated activator of morphogenesis 2 |
| chr20_-_50722183 | 0.06 |
ENST00000371523.4 |
ZFP64 |
ZFP64 zinc finger protein |
| chr14_+_32798547 | 0.06 |
ENST00000557354.1 ENST00000557102.1 ENST00000557272.1 |
AKAP6 |
A kinase (PRKA) anchor protein 6 |
| chr2_-_227050079 | 0.06 |
ENST00000423838.1 |
AC068138.1 |
AC068138.1 |
| chr1_+_153190053 | 0.06 |
ENST00000368744.3 |
PRR9 |
proline rich 9 |
| chr6_+_26199737 | 0.06 |
ENST00000359985.1 |
HIST1H2BF |
histone cluster 1, H2bf |
| chr14_+_32798462 | 0.06 |
ENST00000280979.4 |
AKAP6 |
A kinase (PRKA) anchor protein 6 |
| chr19_-_7040190 | 0.06 |
ENST00000381394.4 |
MBD3L4 |
methyl-CpG binding domain protein 3-like 4 |
| chr15_-_55563072 | 0.06 |
ENST00000567380.1 ENST00000565972.1 ENST00000569493.1 |
RAB27A |
RAB27A, member RAS oncogene family |
| chr19_+_7030589 | 0.05 |
ENST00000329753.5 |
MBD3L5 |
methyl-CpG binding domain protein 3-like 5 |
| chr2_-_61697862 | 0.05 |
ENST00000398571.2 |
USP34 |
ubiquitin specific peptidase 34 |
| chr15_-_55562479 | 0.05 |
ENST00000564609.1 |
RAB27A |
RAB27A, member RAS oncogene family |
| chr5_-_35938674 | 0.04 |
ENST00000397366.1 ENST00000513623.1 ENST00000514524.1 ENST00000397367.2 |
CAPSL |
calcyphosine-like |
| chr12_-_56694142 | 0.04 |
ENST00000550655.1 ENST00000548567.1 ENST00000551430.2 ENST00000351328.3 |
CS |
citrate synthase |
| chr15_-_37393406 | 0.04 |
ENST00000338564.5 ENST00000558313.1 ENST00000340545.5 |
MEIS2 |
Meis homeobox 2 |
| chr3_-_195538728 | 0.04 |
ENST00000349607.4 ENST00000346145.4 |
MUC4 |
mucin 4, cell surface associated |
| chr14_-_74551096 | 0.04 |
ENST00000350259.4 |
ALDH6A1 |
aldehyde dehydrogenase 6 family, member A1 |
| chr1_+_145507587 | 0.03 |
ENST00000330165.8 ENST00000369307.3 |
RBM8A |
RNA binding motif protein 8A |
| chr6_-_26199499 | 0.03 |
ENST00000377831.5 |
HIST1H3D |
histone cluster 1, H3d |
| chr12_-_57328187 | 0.03 |
ENST00000293502.1 |
SDR9C7 |
short chain dehydrogenase/reductase family 9C, member 7 |
| chr3_-_195538760 | 0.03 |
ENST00000475231.1 |
MUC4 |
mucin 4, cell surface associated |
| chr13_-_41593425 | 0.03 |
ENST00000239882.3 |
ELF1 |
E74-like factor 1 (ets domain transcription factor) |
| chr9_+_80912059 | 0.03 |
ENST00000347159.2 ENST00000376588.3 |
PSAT1 |
phosphoserine aminotransferase 1 |
| chr7_-_99717463 | 0.03 |
ENST00000437822.2 |
TAF6 |
TAF6 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 80kDa |
| chr3_+_115342349 | 0.03 |
ENST00000393780.3 |
GAP43 |
growth associated protein 43 |
| chr8_-_42234745 | 0.03 |
ENST00000220812.2 |
DKK4 |
dickkopf WNT signaling pathway inhibitor 4 |
| chrX_-_70326455 | 0.03 |
ENST00000374251.5 |
CXorf65 |
chromosome X open reading frame 65 |
| chr18_+_32173276 | 0.03 |
ENST00000591816.1 ENST00000588125.1 ENST00000598334.1 ENST00000588684.1 ENST00000554864.3 ENST00000399121.5 ENST00000595022.1 ENST00000269190.7 ENST00000399097.3 |
DTNA |
dystrobrevin, alpha |
| chr4_-_89205705 | 0.03 |
ENST00000295908.7 ENST00000510548.2 ENST00000508256.1 |
PPM1K |
protein phosphatase, Mg2+/Mn2+ dependent, 1K |
| chr4_-_89205879 | 0.03 |
ENST00000608933.1 ENST00000315194.4 ENST00000514204.1 |
PPM1K |
protein phosphatase, Mg2+/Mn2+ dependent, 1K |
| chrX_+_130192318 | 0.03 |
ENST00000370922.1 |
ARHGAP36 |
Rho GTPase activating protein 36 |
| chr1_+_152943122 | 0.03 |
ENST00000328051.2 |
SPRR4 |
small proline-rich protein 4 |
| chr3_+_173116225 | 0.03 |
ENST00000457714.1 |
NLGN1 |
neuroligin 1 |
| chr21_+_31768348 | 0.03 |
ENST00000355459.2 |
KRTAP13-1 |
keratin associated protein 13-1 |
| chr5_-_10761206 | 0.03 |
ENST00000432074.2 ENST00000230895.6 |
DAP |
death-associated protein |
| chr3_-_112360116 | 0.03 |
ENST00000206423.3 ENST00000439685.2 |
CCDC80 |
coiled-coil domain containing 80 |
| chr3_+_172468505 | 0.03 |
ENST00000427830.1 ENST00000417960.1 ENST00000428567.1 ENST00000366090.2 ENST00000426894.1 |
ECT2 |
epithelial cell transforming sequence 2 oncogene |
| chr11_-_121986923 | 0.03 |
ENST00000560104.1 |
BLID |
BH3-like motif containing, cell death inducer |
| chr2_-_224467093 | 0.02 |
ENST00000305409.2 |
SCG2 |
secretogranin II |
| chr17_-_9694614 | 0.02 |
ENST00000330255.5 ENST00000571134.1 |
DHRS7C |
dehydrogenase/reductase (SDR family) member 7C |
| chr5_+_176811431 | 0.02 |
ENST00000512593.1 ENST00000324417.5 |
SLC34A1 |
solute carrier family 34 (type II sodium/phosphate contransporter), member 1 |
| chrX_+_129473916 | 0.02 |
ENST00000545805.1 ENST00000543953.1 ENST00000218197.5 |
SLC25A14 |
solute carrier family 25 (mitochondrial carrier, brain), member 14 |
| chr14_-_74551172 | 0.02 |
ENST00000553458.1 |
ALDH6A1 |
aldehyde dehydrogenase 6 family, member A1 |
| chr11_-_124981475 | 0.02 |
ENST00000532156.1 ENST00000532407.1 ENST00000279968.4 ENST00000527766.1 ENST00000529583.1 ENST00000524373.1 ENST00000527271.1 ENST00000526175.1 ENST00000529609.1 ENST00000533273.1 ENST00000531909.1 ENST00000529530.1 |
TMEM218 |
transmembrane protein 218 |
| chr14_-_104181771 | 0.02 |
ENST00000554913.1 ENST00000554974.1 ENST00000553361.1 ENST00000555055.1 ENST00000555964.1 ENST00000556682.1 ENST00000445556.1 ENST00000553332.1 ENST00000352127.7 |
XRCC3 |
X-ray repair complementing defective repair in Chinese hamster cells 3 |
| chr8_+_92261516 | 0.02 |
ENST00000276609.3 ENST00000309536.2 |
SLC26A7 |
solute carrier family 26 (anion exchanger), member 7 |
| chr7_-_99716952 | 0.02 |
ENST00000523306.1 ENST00000344095.4 ENST00000417349.1 ENST00000493322.1 ENST00000520135.1 ENST00000418432.2 ENST00000460673.2 ENST00000452041.1 ENST00000452438.2 ENST00000451699.1 ENST00000453269.2 |
TAF6 |
TAF6 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 80kDa |
| chr19_-_40931891 | 0.02 |
ENST00000357949.4 |
SERTAD1 |
SERTA domain containing 1 |
| chr8_-_18744528 | 0.02 |
ENST00000523619.1 |
PSD3 |
pleckstrin and Sec7 domain containing 3 |
| chr8_-_7243080 | 0.02 |
ENST00000400156.4 |
ZNF705G |
zinc finger protein 705G |
| chr6_+_131571535 | 0.02 |
ENST00000474850.2 |
AKAP7 |
A kinase (PRKA) anchor protein 7 |
| chr1_-_68698197 | 0.02 |
ENST00000370973.2 ENST00000370971.1 |
WLS |
wntless Wnt ligand secretion mediator |
| chr3_-_141747950 | 0.02 |
ENST00000497579.1 |
TFDP2 |
transcription factor Dp-2 (E2F dimerization partner 2) |
| chr11_-_71823796 | 0.02 |
ENST00000545680.1 ENST00000543587.1 ENST00000538393.1 ENST00000535234.1 ENST00000227618.4 ENST00000535503.1 |
ANAPC15 |
anaphase promoting complex subunit 15 |
| chr2_-_163008903 | 0.02 |
ENST00000418842.2 ENST00000375497.3 |
GCG |
glucagon |
| chr5_+_148737562 | 0.02 |
ENST00000274569.4 |
PCYOX1L |
prenylcysteine oxidase 1 like |
| chr11_-_71823715 | 0.02 |
ENST00000545944.1 ENST00000502597.2 |
ANAPC15 |
anaphase promoting complex subunit 15 |
| chr12_-_16759711 | 0.02 |
ENST00000447609.1 |
LMO3 |
LIM domain only 3 (rhombotin-like 2) |
| chr3_+_172468472 | 0.02 |
ENST00000232458.5 ENST00000392692.3 |
ECT2 |
epithelial cell transforming sequence 2 oncogene |
| chr2_-_77749474 | 0.02 |
ENST00000409093.1 ENST00000409088.3 |
LRRTM4 |
leucine rich repeat transmembrane neuronal 4 |
| chr1_+_214161272 | 0.02 |
ENST00000498508.2 ENST00000366958.4 |
PROX1 |
prospero homeobox 1 |
| chr20_+_18488137 | 0.02 |
ENST00000450074.1 ENST00000262544.2 ENST00000336714.3 ENST00000377475.3 |
SEC23B |
Sec23 homolog B (S. cerevisiae) |
| chr19_+_48949030 | 0.02 |
ENST00000253237.5 |
GRWD1 |
glutamate-rich WD repeat containing 1 |
| chr14_+_104182061 | 0.02 |
ENST00000216602.6 |
ZFYVE21 |
zinc finger, FYVE domain containing 21 |
| chr20_+_18447771 | 0.02 |
ENST00000377603.4 |
POLR3F |
polymerase (RNA) III (DNA directed) polypeptide F, 39 kDa |
| chr1_-_157746909 | 0.02 |
ENST00000392274.3 ENST00000361516.3 ENST00000368181.4 |
FCRL2 |
Fc receptor-like 2 |
| chr7_-_25268104 | 0.02 |
ENST00000222674.2 |
NPVF |
neuropeptide VF precursor |
| chr11_-_118023490 | 0.02 |
ENST00000324727.4 |
SCN4B |
sodium channel, voltage-gated, type IV, beta subunit |
| chr7_+_99717230 | 0.02 |
ENST00000262932.3 |
CNPY4 |
canopy FGF signaling regulator 4 |
| chr17_-_7307358 | 0.02 |
ENST00000576017.1 ENST00000302422.3 ENST00000535512.1 |
TMEM256 TMEM256-PLSCR3 |
transmembrane protein 256 TMEM256-PLSCR3 readthrough (NMD candidate) |
| chr4_-_26492076 | 0.02 |
ENST00000295589.3 |
CCKAR |
cholecystokinin A receptor |
| chr5_-_95297678 | 0.02 |
ENST00000237853.4 |
ELL2 |
elongation factor, RNA polymerase II, 2 |
| chr12_+_7014126 | 0.02 |
ENST00000415834.1 ENST00000436789.1 |
LRRC23 |
leucine rich repeat containing 23 |
| chr4_-_174256276 | 0.02 |
ENST00000296503.5 |
HMGB2 |
high mobility group box 2 |
| chr11_-_71823266 | 0.02 |
ENST00000538919.1 ENST00000539395.1 ENST00000542531.1 |
ANAPC15 |
anaphase promoting complex subunit 15 |
| chr12_+_52695617 | 0.02 |
ENST00000293525.5 |
KRT86 |
keratin 86 |
| chr1_+_210501589 | 0.01 |
ENST00000413764.2 ENST00000541565.1 |
HHAT |
hedgehog acyltransferase |
| chr19_-_7698599 | 0.01 |
ENST00000311069.5 |
PCP2 |
Purkinje cell protein 2 |
| chr10_-_48416849 | 0.01 |
ENST00000249598.1 |
GDF2 |
growth differentiation factor 2 |
| chr14_+_104182105 | 0.01 |
ENST00000311141.2 |
ZFYVE21 |
zinc finger, FYVE domain containing 21 |
| chr10_-_99447024 | 0.01 |
ENST00000370626.3 |
AVPI1 |
arginine vasopressin-induced 1 |
| chr9_-_136933615 | 0.01 |
ENST00000371834.2 |
BRD3 |
bromodomain containing 3 |
| chr3_-_157217328 | 0.01 |
ENST00000392832.2 ENST00000543418.1 |
VEPH1 |
ventricular zone expressed PH domain-containing 1 |
| chr11_+_75526212 | 0.01 |
ENST00000356136.3 |
UVRAG |
UV radiation resistance associated |
| chr6_+_140175987 | 0.01 |
ENST00000414038.1 ENST00000431609.1 |
RP5-899B16.1 |
RP5-899B16.1 |
| chr15_+_80351910 | 0.01 |
ENST00000261749.6 ENST00000561060.1 |
ZFAND6 |
zinc finger, AN1-type domain 6 |
| chr6_+_130339710 | 0.01 |
ENST00000526087.1 ENST00000533560.1 ENST00000361794.2 |
L3MBTL3 |
l(3)mbt-like 3 (Drosophila) |
| chr7_+_74072288 | 0.01 |
ENST00000443166.1 |
GTF2I |
general transcription factor IIi |
| chr1_+_42846443 | 0.01 |
ENST00000410070.2 ENST00000431473.3 |
RIMKLA |
ribosomal modification protein rimK-like family member A |
| chr17_+_7792101 | 0.01 |
ENST00000358181.4 ENST00000330494.7 |
CHD3 |
chromodomain helicase DNA binding protein 3 |
| chr3_+_152879985 | 0.01 |
ENST00000323534.2 |
RAP2B |
RAP2B, member of RAS oncogene family |
| chrX_-_53461305 | 0.01 |
ENST00000168216.6 |
HSD17B10 |
hydroxysteroid (17-beta) dehydrogenase 10 |
| chr3_+_35722424 | 0.01 |
ENST00000396481.2 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chr4_+_71587669 | 0.01 |
ENST00000381006.3 ENST00000226328.4 |
RUFY3 |
RUN and FYVE domain containing 3 |
| chr3_-_27764190 | 0.01 |
ENST00000537516.1 |
EOMES |
eomesodermin |
| chr3_+_133292759 | 0.01 |
ENST00000431519.2 |
CDV3 |
CDV3 homolog (mouse) |
| chr19_-_41870026 | 0.01 |
ENST00000243578.3 |
B9D2 |
B9 protein domain 2 |
| chr1_-_17766198 | 0.01 |
ENST00000375436.4 |
RCC2 |
regulator of chromosome condensation 2 |
| chr10_-_99205607 | 0.01 |
ENST00000477692.2 ENST00000485122.2 ENST00000370886.5 ENST00000370885.4 ENST00000370902.3 ENST00000370884.5 |
EXOSC1 |
exosome component 1 |
| chr17_-_8093471 | 0.01 |
ENST00000389017.4 |
C17orf59 |
chromosome 17 open reading frame 59 |
| chr2_+_105654441 | 0.01 |
ENST00000258455.3 |
MRPS9 |
mitochondrial ribosomal protein S9 |
| chr1_+_179923873 | 0.01 |
ENST00000367607.3 ENST00000491495.2 |
CEP350 |
centrosomal protein 350kDa |
| chr19_+_49990811 | 0.01 |
ENST00000391857.4 ENST00000467825.2 |
RPL13A |
ribosomal protein L13a |
| chr16_+_103816 | 0.01 |
ENST00000383018.3 ENST00000417493.1 |
SNRNP25 |
small nuclear ribonucleoprotein 25kDa (U11/U12) |
| chrX_+_95939711 | 0.01 |
ENST00000373049.4 ENST00000324765.8 |
DIAPH2 |
diaphanous-related formin 2 |
| chr17_-_48785216 | 0.01 |
ENST00000285243.6 |
ANKRD40 |
ankyrin repeat domain 40 |
| chr7_-_27169801 | 0.01 |
ENST00000511914.1 |
HOXA4 |
homeobox A4 |
| chr3_+_156799587 | 0.01 |
ENST00000469196.1 |
RP11-6F2.5 |
RP11-6F2.5 |
| chr1_+_203463245 | 0.01 |
ENST00000367222.2 |
OPTC |
opticin |
| chr21_-_43816052 | 0.01 |
ENST00000398405.1 |
TMPRSS3 |
transmembrane protease, serine 3 |
| chr6_-_49681235 | 0.01 |
ENST00000339139.4 |
CRISP2 |
cysteine-rich secretory protein 2 |
| chr3_+_35722487 | 0.01 |
ENST00000441454.1 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chr5_+_125758813 | 0.01 |
ENST00000285689.3 ENST00000515200.1 |
GRAMD3 |
GRAM domain containing 3 |
| chr5_+_125758865 | 0.01 |
ENST00000542322.1 ENST00000544396.1 |
GRAMD3 |
GRAM domain containing 3 |
| chr12_-_6233828 | 0.01 |
ENST00000572068.1 ENST00000261405.5 |
VWF |
von Willebrand factor |
| chr7_-_137028534 | 0.01 |
ENST00000348225.2 |
PTN |
pleiotrophin |
| chr4_-_69111401 | 0.01 |
ENST00000332644.5 |
TMPRSS11B |
transmembrane protease, serine 11B |
| chr7_-_137028498 | 0.01 |
ENST00000393083.2 |
PTN |
pleiotrophin |
| chr4_-_174255536 | 0.01 |
ENST00000446922.2 |
HMGB2 |
high mobility group box 2 |
| chr9_-_136933134 | 0.01 |
ENST00000303407.7 |
BRD3 |
bromodomain containing 3 |
| chr4_-_54232144 | 0.01 |
ENST00000388940.4 ENST00000503450.1 ENST00000401642.3 |
SCFD2 |
sec1 family domain containing 2 |
| chr8_+_9413410 | 0.01 |
ENST00000520408.1 ENST00000310430.6 ENST00000522110.1 |
TNKS |
tankyrase, TRF1-interacting ankyrin-related ADP-ribose polymerase |
| chr7_-_92777606 | 0.01 |
ENST00000437805.1 ENST00000446959.1 ENST00000439952.1 ENST00000414791.1 ENST00000446033.1 ENST00000411955.1 ENST00000318238.4 |
SAMD9L |
sterile alpha motif domain containing 9-like |
| chr4_+_89206076 | 0.01 |
ENST00000500009.2 |
RP11-10L7.1 |
RP11-10L7.1 |
| chr2_+_242089833 | 0.01 |
ENST00000404405.3 ENST00000439916.1 ENST00000406106.3 ENST00000401987.1 |
PPP1R7 |
protein phosphatase 1, regulatory subunit 7 |
| chr6_-_26250835 | 0.01 |
ENST00000446824.2 |
HIST1H3F |
histone cluster 1, H3f |
| chr15_+_40733387 | 0.01 |
ENST00000416165.1 |
BAHD1 |
bromo adjacent homology domain containing 1 |
| chr9_-_116840728 | 0.01 |
ENST00000265132.3 |
AMBP |
alpha-1-microglobulin/bikunin precursor |
| chr10_-_56561022 | 0.01 |
ENST00000373965.2 ENST00000414778.1 ENST00000395438.1 ENST00000409834.1 ENST00000395445.1 ENST00000395446.1 ENST00000395442.1 ENST00000395440.1 ENST00000395432.2 ENST00000361849.3 ENST00000395433.1 ENST00000320301.6 ENST00000395430.1 ENST00000437009.1 |
PCDH15 |
protocadherin-related 15 |
| chr14_+_96000930 | 0.00 |
ENST00000331334.4 |
GLRX5 |
glutaredoxin 5 |
| chr5_+_179159813 | 0.00 |
ENST00000292599.3 |
MAML1 |
mastermind-like 1 (Drosophila) |
| chr17_-_74163159 | 0.00 |
ENST00000591615.1 |
RNF157 |
ring finger protein 157 |
| chr1_+_160370344 | 0.00 |
ENST00000368061.2 |
VANGL2 |
VANGL planar cell polarity protein 2 |
| chr2_+_65216462 | 0.00 |
ENST00000234256.3 |
SLC1A4 |
solute carrier family 1 (glutamate/neutral amino acid transporter), member 4 |
| chr6_-_100912785 | 0.00 |
ENST00000369208.3 |
SIM1 |
single-minded family bHLH transcription factor 1 |
| chr7_-_122339162 | 0.00 |
ENST00000340112.2 |
RNF133 |
ring finger protein 133 |
| chr12_+_28410128 | 0.00 |
ENST00000381259.1 ENST00000381256.1 |
CCDC91 |
coiled-coil domain containing 91 |
| chr8_+_50824233 | 0.00 |
ENST00000522124.1 |
SNTG1 |
syntrophin, gamma 1 |
| chr1_-_156399184 | 0.00 |
ENST00000368243.1 ENST00000357975.4 ENST00000310027.5 ENST00000400991.2 |
C1orf61 |
chromosome 1 open reading frame 61 |
| chr2_-_157189180 | 0.00 |
ENST00000539077.1 ENST00000424077.1 ENST00000426264.1 ENST00000339562.4 ENST00000421709.1 |
NR4A2 |
nuclear receptor subfamily 4, group A, member 2 |
| chrX_-_153602991 | 0.00 |
ENST00000369850.3 ENST00000422373.1 |
FLNA |
filamin A, alpha |
| chr1_+_62901968 | 0.00 |
ENST00000452143.1 ENST00000442679.1 ENST00000371146.1 |
USP1 |
ubiquitin specific peptidase 1 |
| chr19_-_7697857 | 0.00 |
ENST00000598935.1 |
PCP2 |
Purkinje cell protein 2 |
| chr2_+_47630255 | 0.00 |
ENST00000406134.1 |
MSH2 |
mutS homolog 2 |
| chr2_+_47630108 | 0.00 |
ENST00000233146.2 ENST00000454849.1 ENST00000543555.1 |
MSH2 |
mutS homolog 2 |
| chr12_+_53662110 | 0.00 |
ENST00000552462.1 |
ESPL1 |
extra spindle pole bodies homolog 1 (S. cerevisiae) |
| chr17_-_9929581 | 0.00 |
ENST00000437099.2 ENST00000396115.2 |
GAS7 |
growth arrest-specific 7 |
| chr5_+_53751445 | 0.00 |
ENST00000302005.1 |
HSPB3 |
heat shock 27kDa protein 3 |
| chr1_-_2461684 | 0.00 |
ENST00000378453.3 |
HES5 |
hes family bHLH transcription factor 5 |
| chrX_-_53461288 | 0.00 |
ENST00000375298.4 ENST00000375304.5 |
HSD17B10 |
hydroxysteroid (17-beta) dehydrogenase 10 |
| chr18_-_31803169 | 0.00 |
ENST00000590712.1 |
NOL4 |
nucleolar protein 4 |
| chr15_+_34638066 | 0.00 |
ENST00000333756.4 |
NUTM1 |
NUT midline carcinoma, family member 1 |
| chr5_+_31193847 | 0.00 |
ENST00000514738.1 ENST00000265071.2 |
CDH6 |
cadherin 6, type 2, K-cadherin (fetal kidney) |
| chr8_+_7783859 | 0.00 |
ENST00000400120.3 |
ZNF705B |
zinc finger protein 705B |
| chr4_+_169013666 | 0.00 |
ENST00000359299.3 |
ANXA10 |
annexin A10 |
| chr17_+_43238438 | 0.00 |
ENST00000593138.1 ENST00000586681.1 |
HEXIM2 |
hexamethylene bis-acetamide inducible 2 |
| chr8_+_24298531 | 0.00 |
ENST00000175238.6 |
ADAM7 |
ADAM metallopeptidase domain 7 |
| chr12_-_101604185 | 0.00 |
ENST00000536262.2 |
SLC5A8 |
solute carrier family 5 (sodium/monocarboxylate cotransporter), member 8 |
| chr8_+_24298597 | 0.00 |
ENST00000380789.1 |
ADAM7 |
ADAM metallopeptidase domain 7 |
| chr5_+_141303373 | 0.00 |
ENST00000432126.2 ENST00000194118.4 |
KIAA0141 |
KIAA0141 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0035553 | oxidative RNA demethylation(GO:0035513) oxidative single-stranded RNA demethylation(GO:0035553) |
| 0.0 | 0.1 | GO:0061743 | motor learning(GO:0061743) |
| 0.0 | 0.2 | GO:1903435 | positive regulation of constitutive secretory pathway(GO:1903435) |
| 0.0 | 0.1 | GO:0051344 | regulation of cyclic-nucleotide phosphodiesterase activity(GO:0051342) negative regulation of cyclic-nucleotide phosphodiesterase activity(GO:0051344) |
| 0.0 | 0.1 | GO:1902261 | positive regulation of delayed rectifier potassium channel activity(GO:1902261) |
| 0.0 | 0.1 | GO:0019859 | pyrimidine nucleobase catabolic process(GO:0006208) thymine catabolic process(GO:0006210) thymine metabolic process(GO:0019859) |
| 0.0 | 0.1 | GO:0090402 | oncogene-induced cell senescence(GO:0090402) |
| 0.0 | 0.1 | GO:0090611 | ubiquitin-independent protein catabolic process via the multivesicular body sorting pathway(GO:0090611) |
| 0.0 | 0.1 | GO:0042421 | norepinephrine biosynthetic process(GO:0042421) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0034751 | aryl hydrocarbon receptor complex(GO:0034751) |
| 0.0 | 0.1 | GO:0042720 | mitochondrial inner membrane peptidase complex(GO:0042720) |
| 0.0 | 0.1 | GO:0071547 | piP-body(GO:0071547) |
| 0.0 | 0.2 | GO:0033093 | Weibel-Palade body(GO:0033093) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0035515 | oxidative RNA demethylase activity(GO:0035515) |
| 0.0 | 0.1 | GO:0015173 | aromatic amino acid transmembrane transporter activity(GO:0015173) |
| 0.0 | 0.2 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.0 | 0.1 | GO:0030021 | extracellular matrix structural constituent conferring compression resistance(GO:0030021) structural constituent of tooth enamel(GO:0030345) |
| 0.0 | 0.1 | GO:0030197 | extracellular matrix constituent, lubricant activity(GO:0030197) |
| 0.0 | 0.1 | GO:0016715 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced ascorbate as one donor, and incorporation of one atom of oxygen(GO:0016715) |