Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
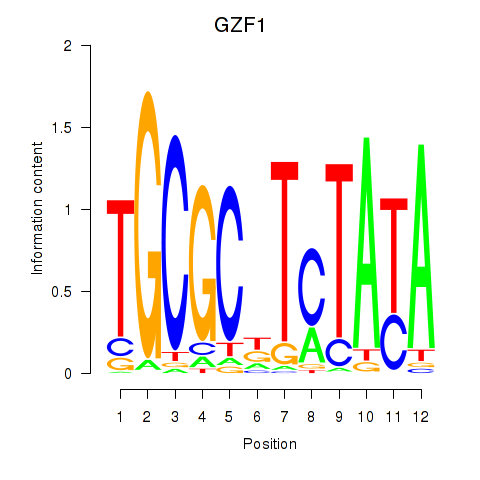
Results for GZF1
Z-value: 0.77
Transcription factors associated with GZF1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
GZF1
|
ENSG00000125812.11 | GZF1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| GZF1 | hg19_v2_chr20_+_23342783_23342957 | 0.59 | 1.3e-01 | Click! |
Activity profile of GZF1 motif
Sorted Z-values of GZF1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of GZF1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr14_-_92413353 | 1.79 |
ENST00000556154.1 |
FBLN5 |
fibulin 5 |
| chr14_-_92413727 | 1.55 |
ENST00000267620.10 |
FBLN5 |
fibulin 5 |
| chr5_+_92919043 | 1.07 |
ENST00000327111.3 |
NR2F1 |
nuclear receptor subfamily 2, group F, member 1 |
| chr14_-_52535712 | 0.92 |
ENST00000216286.5 ENST00000541773.1 |
NID2 |
nidogen 2 (osteonidogen) |
| chr14_+_101292445 | 0.72 |
ENST00000429159.2 ENST00000520714.1 ENST00000522771.2 ENST00000424076.3 ENST00000423456.1 ENST00000521404.1 ENST00000556736.1 ENST00000451743.2 ENST00000398518.2 ENST00000554639.1 ENST00000452120.2 ENST00000519709.1 ENST00000412736.2 |
MEG3 |
maternally expressed 3 (non-protein coding) |
| chr1_+_79115503 | 0.64 |
ENST00000370747.4 ENST00000438486.1 ENST00000545124.1 |
IFI44 |
interferon-induced protein 44 |
| chr3_-_114035026 | 0.62 |
ENST00000570269.1 |
RP11-553L6.5 |
RP11-553L6.5 |
| chr1_-_144994909 | 0.61 |
ENST00000369347.4 ENST00000369354.3 |
PDE4DIP |
phosphodiesterase 4D interacting protein |
| chrX_+_54835493 | 0.60 |
ENST00000396224.1 |
MAGED2 |
melanoma antigen family D, 2 |
| chr4_-_89619386 | 0.60 |
ENST00000323061.5 |
NAP1L5 |
nucleosome assembly protein 1-like 5 |
| chr1_-_144995074 | 0.55 |
ENST00000534536.1 |
PDE4DIP |
phosphodiesterase 4D interacting protein |
| chr7_-_27183263 | 0.52 |
ENST00000222726.3 |
HOXA5 |
homeobox A5 |
| chr1_+_26606608 | 0.45 |
ENST00000319041.6 |
SH3BGRL3 |
SH3 domain binding glutamic acid-rich protein like 3 |
| chr1_-_144995002 | 0.42 |
ENST00000369356.4 |
PDE4DIP |
phosphodiesterase 4D interacting protein |
| chr19_-_44384291 | 0.42 |
ENST00000324394.6 |
ZNF404 |
zinc finger protein 404 |
| chrX_+_102883620 | 0.41 |
ENST00000372626.3 |
TCEAL1 |
transcription elongation factor A (SII)-like 1 |
| chr6_+_30524663 | 0.37 |
ENST00000376560.3 |
PRR3 |
proline rich 3 |
| chr6_+_30525051 | 0.34 |
ENST00000376557.3 |
PRR3 |
proline rich 3 |
| chr3_-_87040233 | 0.33 |
ENST00000398399.2 |
VGLL3 |
vestigial like 3 (Drosophila) |
| chr16_+_86612112 | 0.31 |
ENST00000320241.3 |
FOXL1 |
forkhead box L1 |
| chr12_+_107168418 | 0.30 |
ENST00000392839.2 ENST00000548914.1 ENST00000355478.2 ENST00000552619.1 ENST00000549643.1 |
RIC8B |
RIC8 guanine nucleotide exchange factor B |
| chr20_-_31331212 | 0.29 |
ENST00000474815.2 ENST00000446419.2 |
COMMD7 |
COMM domain containing 7 |
| chr1_-_144994840 | 0.26 |
ENST00000369351.3 ENST00000369349.3 |
PDE4DIP |
phosphodiesterase 4D interacting protein |
| chr20_+_42086525 | 0.25 |
ENST00000244020.3 |
SRSF6 |
serine/arginine-rich splicing factor 6 |
| chr7_-_26904317 | 0.25 |
ENST00000345317.2 |
SKAP2 |
src kinase associated phosphoprotein 2 |
| chr19_-_36545128 | 0.22 |
ENST00000538849.1 |
THAP8 |
THAP domain containing 8 |
| chr9_+_131133598 | 0.18 |
ENST00000372853.4 ENST00000452446.1 ENST00000372850.1 ENST00000372847.1 |
URM1 |
ubiquitin related modifier 1 |
| chr13_+_98086445 | 0.18 |
ENST00000245304.4 |
RAP2A |
RAP2A, member of RAS oncogene family |
| chr1_-_42384343 | 0.16 |
ENST00000372584.1 |
HIVEP3 |
human immunodeficiency virus type I enhancer binding protein 3 |
| chr15_+_44580955 | 0.15 |
ENST00000345795.2 ENST00000360824.3 |
CASC4 |
cancer susceptibility candidate 4 |
| chr9_-_14308004 | 0.14 |
ENST00000493697.1 |
NFIB |
nuclear factor I/B |
| chr12_+_104458235 | 0.14 |
ENST00000229330.4 |
HCFC2 |
host cell factor C2 |
| chr2_-_88427568 | 0.14 |
ENST00000393750.3 ENST00000295834.3 |
FABP1 |
fatty acid binding protein 1, liver |
| chr8_+_23430157 | 0.12 |
ENST00000399967.3 |
FP15737 |
FP15737 |
| chr17_-_29648761 | 0.12 |
ENST00000247270.3 ENST00000462804.2 |
EVI2A |
ecotropic viral integration site 2A |
| chr16_-_30582888 | 0.12 |
ENST00000563707.1 ENST00000567855.1 |
ZNF688 |
zinc finger protein 688 |
| chr11_-_17498348 | 0.11 |
ENST00000389817.3 ENST00000302539.4 |
ABCC8 |
ATP-binding cassette, sub-family C (CFTR/MRP), member 8 |
| chr1_-_226374373 | 0.11 |
ENST00000366812.5 |
ACBD3 |
acyl-CoA binding domain containing 3 |
| chr6_-_30524951 | 0.11 |
ENST00000376621.3 |
GNL1 |
guanine nucleotide binding protein-like 1 |
| chr12_-_76478446 | 0.10 |
ENST00000393263.3 ENST00000548044.1 ENST00000547704.1 ENST00000431879.3 ENST00000549596.1 ENST00000550934.1 ENST00000551600.1 ENST00000547479.1 ENST00000547773.1 ENST00000544816.1 ENST00000542344.1 ENST00000548273.1 |
NAP1L1 |
nucleosome assembly protein 1-like 1 |
| chr14_-_53619816 | 0.10 |
ENST00000323669.5 ENST00000395606.1 ENST00000357758.3 |
DDHD1 |
DDHD domain containing 1 |
| chr12_-_76478417 | 0.10 |
ENST00000552342.1 |
NAP1L1 |
nucleosome assembly protein 1-like 1 |
| chr12_-_76478386 | 0.09 |
ENST00000535020.2 |
NAP1L1 |
nucleosome assembly protein 1-like 1 |
| chr19_-_59084922 | 0.09 |
ENST00000215057.2 ENST00000599369.1 |
MZF1 |
myeloid zinc finger 1 |
| chr14_-_59950724 | 0.08 |
ENST00000481608.1 |
L3HYPDH |
L-3-hydroxyproline dehydratase (trans-) |
| chr2_-_27603582 | 0.07 |
ENST00000323703.6 ENST00000436006.1 |
ZNF513 |
zinc finger protein 513 |
| chr19_-_59084647 | 0.06 |
ENST00000594234.1 ENST00000596039.1 |
MZF1 |
myeloid zinc finger 1 |
| chr22_-_43355858 | 0.05 |
ENST00000402229.1 ENST00000407585.1 ENST00000453079.1 |
PACSIN2 |
protein kinase C and casein kinase substrate in neurons 2 |
| chr11_+_101918153 | 0.05 |
ENST00000434758.2 ENST00000526781.1 ENST00000534360.1 |
C11orf70 |
chromosome 11 open reading frame 70 |
| chrX_+_48367338 | 0.04 |
ENST00000359882.4 ENST00000537758.1 ENST00000367574.4 ENST00000355961.4 ENST00000489940.1 ENST00000361988.3 |
PORCN |
porcupine homolog (Drosophila) |
| chr3_+_30647994 | 0.03 |
ENST00000295754.5 |
TGFBR2 |
transforming growth factor, beta receptor II (70/80kDa) |
| chr10_+_60028818 | 0.03 |
ENST00000333926.5 |
CISD1 |
CDGSH iron sulfur domain 1 |
| chr9_+_96338647 | 0.02 |
ENST00000359246.4 |
PHF2 |
PHD finger protein 2 |
| chr7_-_105029812 | 0.02 |
ENST00000482897.1 |
SRPK2 |
SRSF protein kinase 2 |
| chr16_-_70285797 | 0.01 |
ENST00000435634.1 |
EXOSC6 |
exosome component 6 |
| chr16_-_46797149 | 0.01 |
ENST00000536476.1 |
MYLK3 |
myosin light chain kinase 3 |
| chr3_+_30648066 | 0.00 |
ENST00000359013.4 |
TGFBR2 |
transforming growth factor, beta receptor II (70/80kDa) |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 3.3 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.2 | 0.5 | GO:0060435 | bronchiole development(GO:0060435) |
| 0.1 | 0.6 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.1 | 0.3 | GO:0061146 | Peyer's patch morphogenesis(GO:0061146) |
| 0.0 | 0.2 | GO:0034227 | tRNA thio-modification(GO:0034227) |
| 0.0 | 0.1 | GO:1901079 | positive regulation of relaxation of muscle(GO:1901079) |
| 0.0 | 0.2 | GO:0060501 | positive regulation of epithelial cell proliferation involved in lung morphogenesis(GO:0060501) |
| 0.0 | 0.9 | GO:0071711 | basement membrane organization(GO:0071711) |
| 0.0 | 0.1 | GO:2000795 | negative regulation of epithelial cell proliferation involved in lung morphogenesis(GO:2000795) |
| 0.0 | 0.2 | GO:0045198 | establishment of epithelial cell apical/basal polarity(GO:0045198) |
| 0.0 | 0.1 | GO:0032000 | positive regulation of fatty acid beta-oxidation(GO:0032000) |
| 0.0 | 0.0 | GO:0002514 | B cell tolerance induction(GO:0002514) regulation of B cell tolerance induction(GO:0002661) positive regulation of B cell tolerance induction(GO:0002663) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 3.3 | GO:0071953 | elastic fiber(GO:0071953) |
| 0.0 | 0.1 | GO:0008282 | ATP-sensitive potassium channel complex(GO:0008282) |
| 0.0 | 0.1 | GO:0045179 | apical cortex(GO:0045179) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.1 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 0.0 | 0.1 | GO:0008281 | sulfonylurea receptor activity(GO:0008281) |
| 0.0 | 0.4 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 0.1 | GO:0047661 | racemase and epimerase activity, acting on amino acids and derivatives(GO:0016855) racemase activity, acting on amino acids and derivatives(GO:0036361) amino-acid racemase activity(GO:0047661) |
| 0.0 | 0.1 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.0 | 3.3 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.2 | GO:0016783 | sulfurtransferase activity(GO:0016783) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.9 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 3.3 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.1 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |