Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
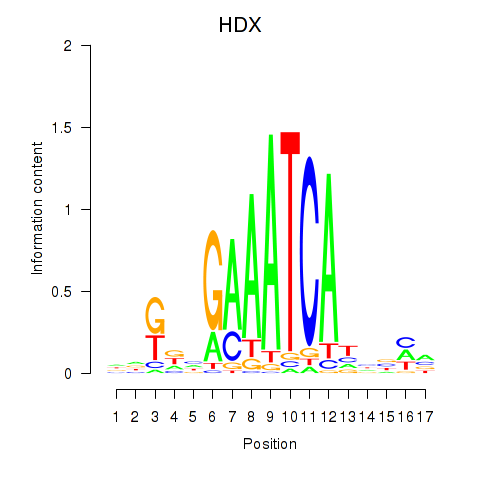
Results for HDX
Z-value: 0.46
Transcription factors associated with HDX
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HDX
|
ENSG00000165259.9 | HDX |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HDX | hg19_v2_chrX_-_83757399_83757487 | -0.76 | 2.8e-02 | Click! |
Activity profile of HDX motif
Sorted Z-values of HDX motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HDX
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr18_+_29027696 | 0.95 |
ENST00000257189.4 |
DSG3 |
desmoglein 3 |
| chr2_-_113594279 | 0.87 |
ENST00000416750.1 ENST00000418817.1 ENST00000263341.2 |
IL1B |
interleukin 1, beta |
| chr1_-_153363452 | 0.82 |
ENST00000368732.1 ENST00000368733.3 |
S100A8 |
S100 calcium binding protein A8 |
| chr1_-_209825674 | 0.54 |
ENST00000367030.3 ENST00000356082.4 |
LAMB3 |
laminin, beta 3 |
| chr5_+_140165876 | 0.49 |
ENST00000504120.2 ENST00000394633.3 ENST00000378133.3 |
PCDHA1 |
protocadherin alpha 1 |
| chr2_+_113735575 | 0.23 |
ENST00000376489.2 ENST00000259205.4 |
IL36G |
interleukin 36, gamma |
| chr7_-_121944491 | 0.23 |
ENST00000331178.4 ENST00000427185.2 ENST00000442488.2 |
FEZF1 |
FEZ family zinc finger 1 |
| chr1_+_84630367 | 0.20 |
ENST00000370680.1 |
PRKACB |
protein kinase, cAMP-dependent, catalytic, beta |
| chr5_-_35230434 | 0.20 |
ENST00000504500.1 |
PRLR |
prolactin receptor |
| chr1_-_11024258 | 0.19 |
ENST00000418570.2 |
C1orf127 |
chromosome 1 open reading frame 127 |
| chr1_+_117602925 | 0.19 |
ENST00000369466.4 |
TTF2 |
transcription termination factor, RNA polymerase II |
| chr6_-_116381918 | 0.18 |
ENST00000606080.1 |
FRK |
fyn-related kinase |
| chr12_-_39734783 | 0.18 |
ENST00000552961.1 |
KIF21A |
kinesin family member 21A |
| chr13_+_97928395 | 0.15 |
ENST00000445661.2 |
MBNL2 |
muscleblind-like splicing regulator 2 |
| chr2_+_128403439 | 0.11 |
ENST00000544369.1 |
GPR17 |
G protein-coupled receptor 17 |
| chr2_+_128403720 | 0.11 |
ENST00000272644.3 |
GPR17 |
G protein-coupled receptor 17 |
| chr15_-_54025300 | 0.10 |
ENST00000559418.1 |
WDR72 |
WD repeat domain 72 |
| chr19_-_41934635 | 0.10 |
ENST00000321702.2 |
B3GNT8 |
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 8 |
| chr7_-_135412925 | 0.09 |
ENST00000354042.4 |
SLC13A4 |
solute carrier family 13 (sodium/sulfate symporter), member 4 |
| chr1_-_169680745 | 0.08 |
ENST00000236147.4 |
SELL |
selectin L |
| chr6_+_42584847 | 0.07 |
ENST00000372883.3 |
UBR2 |
ubiquitin protein ligase E3 component n-recognin 2 |
| chr7_-_69062391 | 0.07 |
ENST00000436600.2 |
RP5-942I16.1 |
RP5-942I16.1 |
| chr1_+_197237352 | 0.06 |
ENST00000538660.1 ENST00000367400.3 ENST00000367399.2 |
CRB1 |
crumbs homolog 1 (Drosophila) |
| chr18_-_53804580 | 0.06 |
ENST00000590484.1 ENST00000589293.1 ENST00000587904.1 ENST00000591974.1 |
RP11-456O19.4 |
RP11-456O19.4 |
| chr5_+_60933634 | 0.05 |
ENST00000505642.1 |
C5orf64 |
chromosome 5 open reading frame 64 |
| chr5_-_135290705 | 0.04 |
ENST00000274507.1 |
LECT2 |
leukocyte cell-derived chemotaxin 2 |
| chr19_+_36602104 | 0.04 |
ENST00000585332.1 ENST00000262637.4 |
OVOL3 |
ovo-like zinc finger 3 |
| chr12_+_101869096 | 0.03 |
ENST00000551346.1 |
SPIC |
Spi-C transcription factor (Spi-1/PU.1 related) |
| chrX_+_55649833 | 0.03 |
ENST00000339140.3 |
FOXR2 |
forkhead box R2 |
| chr5_-_41213607 | 0.03 |
ENST00000337836.5 ENST00000433294.1 |
C6 |
complement component 6 |
| chr19_-_58485895 | 0.02 |
ENST00000314391.3 |
C19orf18 |
chromosome 19 open reading frame 18 |
| chr12_-_118797475 | 0.02 |
ENST00000541786.1 ENST00000419821.2 ENST00000541878.1 |
TAOK3 |
TAO kinase 3 |
| chr11_-_66675371 | 0.02 |
ENST00000393955.2 |
PC |
pyruvate carboxylase |
| chr3_+_107241882 | 0.02 |
ENST00000416476.2 |
BBX |
bobby sox homolog (Drosophila) |
| chr1_+_47489240 | 0.01 |
ENST00000371901.3 |
CYP4X1 |
cytochrome P450, family 4, subfamily X, polypeptide 1 |
| chr1_+_75600567 | 0.00 |
ENST00000356261.3 |
LHX8 |
LIM homeobox 8 |
| chr5_-_35230771 | 0.00 |
ENST00000342362.5 |
PRLR |
prolactin receptor |
| chr5_-_35230649 | 0.00 |
ENST00000382002.5 |
PRLR |
prolactin receptor |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.9 | GO:0060557 | positive regulation of vitamin metabolic process(GO:0046136) positive regulation of vitamin D biosynthetic process(GO:0060557) positive regulation of calcidiol 1-monooxygenase activity(GO:0060559) |
| 0.1 | 0.8 | GO:0032119 | sequestering of zinc ion(GO:0032119) |
| 0.0 | 0.2 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 0.0 | 0.2 | GO:0038161 | prolactin signaling pathway(GO:0038161) |
| 0.0 | 0.1 | GO:0030311 | poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 0.0 | 0.1 | GO:0097338 | response to clozapine(GO:0097338) |
| 0.0 | 0.5 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.0 | 0.2 | GO:0002862 | negative regulation of inflammatory response to antigenic stimulus(GO:0002862) |
| 0.0 | 0.1 | GO:0071596 | ubiquitin-dependent protein catabolic process via the N-end rule pathway(GO:0071596) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.0 | 0.9 | GO:0030057 | desmosome(GO:0030057) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.8 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 0.0 | 1.1 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 0.2 | GO:0004925 | prolactin receptor activity(GO:0004925) |
| 0.0 | 0.1 | GO:0016262 | protein N-acetylglucosaminyltransferase activity(GO:0016262) |
| 0.0 | 0.1 | GO:0070728 | leucine binding(GO:0070728) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.0 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.0 | 0.5 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.0 | 0.8 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.9 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.0 | 0.9 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 0.2 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.0 | 0.2 | REACTOME EICOSANOID LIGAND BINDING RECEPTORS | Genes involved in Eicosanoid ligand-binding receptors |