Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
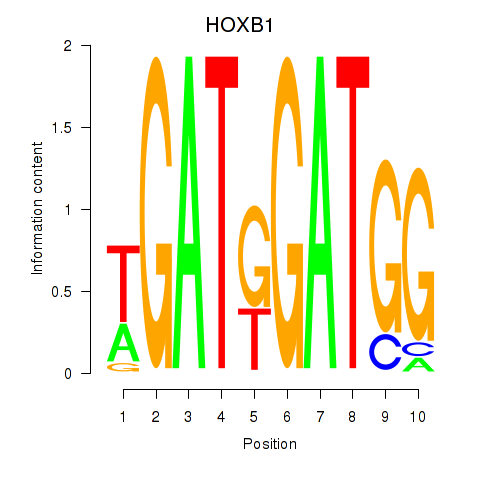
Results for HOXA2_HOXB1
Z-value: 0.47
Transcription factors associated with HOXA2_HOXB1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HOXA2
|
ENSG00000105996.5 | HOXA2 |
|
HOXB1
|
ENSG00000120094.6 | HOXB1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HOXA2 | hg19_v2_chr7_-_27142290_27142430 | 0.86 | 5.9e-03 | Click! |
| HOXB1 | hg19_v2_chr17_-_46608272_46608385 | -0.42 | 3.0e-01 | Click! |
Activity profile of HOXA2_HOXB1 motif
Sorted Z-values of HOXA2_HOXB1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HOXA2_HOXB1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_-_93616340 | 0.76 |
ENST00000557420.1 ENST00000542321.2 |
RGMA |
repulsive guidance molecule family member a |
| chr15_-_93616892 | 0.52 |
ENST00000556658.1 ENST00000538818.1 ENST00000425933.2 |
RGMA |
repulsive guidance molecule family member a |
| chr2_-_190044480 | 0.49 |
ENST00000374866.3 |
COL5A2 |
collagen, type V, alpha 2 |
| chr12_-_15038779 | 0.49 |
ENST00000228938.5 ENST00000539261.1 |
MGP |
matrix Gla protein |
| chr1_-_182641367 | 0.31 |
ENST00000508450.1 |
RGS8 |
regulator of G-protein signaling 8 |
| chr11_-_2162162 | 0.28 |
ENST00000381389.1 |
IGF2 |
insulin-like growth factor 2 (somatomedin A) |
| chr1_-_108507631 | 0.26 |
ENST00000527011.1 ENST00000370056.4 |
VAV3 |
vav 3 guanine nucleotide exchange factor |
| chr12_+_21679220 | 0.25 |
ENST00000256969.2 |
C12orf39 |
chromosome 12 open reading frame 39 |
| chr11_-_111782484 | 0.25 |
ENST00000533971.1 |
CRYAB |
crystallin, alpha B |
| chr5_+_147582348 | 0.25 |
ENST00000514389.1 |
SPINK6 |
serine peptidase inhibitor, Kazal type 6 |
| chr11_-_111782696 | 0.24 |
ENST00000227251.3 ENST00000526180.1 |
CRYAB |
crystallin, alpha B |
| chr1_-_92351769 | 0.22 |
ENST00000212355.4 |
TGFBR3 |
transforming growth factor, beta receptor III |
| chr2_-_45236540 | 0.20 |
ENST00000303077.6 |
SIX2 |
SIX homeobox 2 |
| chr6_+_151561085 | 0.18 |
ENST00000402676.2 |
AKAP12 |
A kinase (PRKA) anchor protein 12 |
| chr1_-_182642017 | 0.16 |
ENST00000367557.4 ENST00000258302.4 |
RGS8 |
regulator of G-protein signaling 8 |
| chr10_+_95848824 | 0.16 |
ENST00000371385.3 ENST00000371375.1 |
PLCE1 |
phospholipase C, epsilon 1 |
| chr16_+_15596123 | 0.16 |
ENST00000452191.2 |
C16orf45 |
chromosome 16 open reading frame 45 |
| chr6_+_151561506 | 0.15 |
ENST00000253332.1 |
AKAP12 |
A kinase (PRKA) anchor protein 12 |
| chr2_+_12857043 | 0.14 |
ENST00000381465.2 |
TRIB2 |
tribbles pseudokinase 2 |
| chr2_-_145278475 | 0.14 |
ENST00000558170.2 |
ZEB2 |
zinc finger E-box binding homeobox 2 |
| chr17_+_39975455 | 0.14 |
ENST00000455106.1 |
FKBP10 |
FK506 binding protein 10, 65 kDa |
| chr2_+_12857015 | 0.14 |
ENST00000155926.4 |
TRIB2 |
tribbles pseudokinase 2 |
| chr17_+_39975544 | 0.13 |
ENST00000544340.1 |
FKBP10 |
FK506 binding protein 10, 65 kDa |
| chr7_-_150864635 | 0.13 |
ENST00000297537.4 |
GBX1 |
gastrulation brain homeobox 1 |
| chr10_-_79397202 | 0.13 |
ENST00000372437.1 ENST00000372408.2 ENST00000372403.4 |
KCNMA1 |
potassium large conductance calcium-activated channel, subfamily M, alpha member 1 |
| chr20_+_34802295 | 0.12 |
ENST00000432603.1 |
EPB41L1 |
erythrocyte membrane protein band 4.1-like 1 |
| chr10_-_79397316 | 0.12 |
ENST00000372421.5 ENST00000457953.1 |
KCNMA1 |
potassium large conductance calcium-activated channel, subfamily M, alpha member 1 |
| chr17_+_58755184 | 0.11 |
ENST00000589222.1 ENST00000407086.3 ENST00000390652.5 |
BCAS3 |
breast carcinoma amplified sequence 3 |
| chr5_-_10761206 | 0.10 |
ENST00000432074.2 ENST00000230895.6 |
DAP |
death-associated protein |
| chr5_+_162864575 | 0.10 |
ENST00000512163.1 ENST00000393929.1 ENST00000340828.2 ENST00000511683.2 ENST00000510097.1 ENST00000511490.2 ENST00000510664.1 |
CCNG1 |
cyclin G1 |
| chr7_-_27142290 | 0.10 |
ENST00000222718.5 |
HOXA2 |
homeobox A2 |
| chr19_-_45735138 | 0.10 |
ENST00000252482.3 |
EXOC3L2 |
exocyst complex component 3-like 2 |
| chr10_-_79397479 | 0.09 |
ENST00000404771.3 |
KCNMA1 |
potassium large conductance calcium-activated channel, subfamily M, alpha member 1 |
| chr10_-_98945677 | 0.09 |
ENST00000266058.4 ENST00000371041.3 |
SLIT1 |
slit homolog 1 (Drosophila) |
| chr16_+_85645007 | 0.09 |
ENST00000405402.2 |
GSE1 |
Gse1 coiled-coil protein |
| chr4_-_90757364 | 0.09 |
ENST00000508895.1 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr10_-_75457554 | 0.09 |
ENST00000374094.4 ENST00000443782.2 |
AGAP5 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 5 |
| chr16_-_49890016 | 0.09 |
ENST00000563137.2 |
ZNF423 |
zinc finger protein 423 |
| chrX_+_99899180 | 0.09 |
ENST00000373004.3 |
SRPX2 |
sushi-repeat containing protein, X-linked 2 |
| chr19_+_35849723 | 0.09 |
ENST00000594310.1 |
FFAR3 |
free fatty acid receptor 3 |
| chr4_-_90756769 | 0.09 |
ENST00000345009.4 ENST00000505199.1 ENST00000502987.1 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr2_-_154335300 | 0.08 |
ENST00000325926.3 |
RPRM |
reprimo, TP53 dependent G2 arrest mediator candidate |
| chrX_-_13835147 | 0.08 |
ENST00000493677.1 ENST00000355135.2 |
GPM6B |
glycoprotein M6B |
| chr9_-_73029540 | 0.08 |
ENST00000377126.2 |
KLF9 |
Kruppel-like factor 9 |
| chr4_+_129732467 | 0.07 |
ENST00000413543.2 |
PHF17 |
jade family PHD finger 1 |
| chr17_-_42277203 | 0.07 |
ENST00000587097.1 |
ATXN7L3 |
ataxin 7-like 3 |
| chr2_-_200323414 | 0.07 |
ENST00000443023.1 |
SATB2 |
SATB homeobox 2 |
| chr17_-_42276574 | 0.07 |
ENST00000589805.1 |
ATXN7L3 |
ataxin 7-like 3 |
| chr2_+_26624775 | 0.07 |
ENST00000288710.2 |
DRC1 |
dynein regulatory complex subunit 1 homolog (Chlamydomonas) |
| chr2_-_200322723 | 0.06 |
ENST00000417098.1 |
SATB2 |
SATB homeobox 2 |
| chr4_-_113437328 | 0.06 |
ENST00000313341.3 |
NEUROG2 |
neurogenin 2 |
| chr5_-_125930929 | 0.06 |
ENST00000553117.1 ENST00000447989.2 ENST00000409134.3 |
ALDH7A1 |
aldehyde dehydrogenase 7 family, member A1 |
| chr7_-_14029283 | 0.06 |
ENST00000433547.1 ENST00000405192.2 |
ETV1 |
ets variant 1 |
| chr1_+_178062855 | 0.06 |
ENST00000448150.3 |
RASAL2 |
RAS protein activator like 2 |
| chrX_-_13835461 | 0.06 |
ENST00000316715.4 ENST00000356942.5 |
GPM6B |
glycoprotein M6B |
| chr20_+_34742650 | 0.06 |
ENST00000373945.1 ENST00000338074.2 |
EPB41L1 |
erythrocyte membrane protein band 4.1-like 1 |
| chr2_-_145277882 | 0.06 |
ENST00000465070.1 ENST00000444559.1 |
ZEB2 |
zinc finger E-box binding homeobox 2 |
| chr7_-_144107320 | 0.06 |
ENST00000483238.1 ENST00000467773.1 |
NOBOX |
NOBOX oogenesis homeobox |
| chr1_-_67390474 | 0.06 |
ENST00000371023.3 ENST00000371022.3 ENST00000371026.3 ENST00000431318.1 |
WDR78 |
WD repeat domain 78 |
| chr11_+_57365150 | 0.05 |
ENST00000457869.1 ENST00000340687.6 ENST00000378323.4 ENST00000378324.2 ENST00000403558.1 |
SERPING1 |
serpin peptidase inhibitor, clade G (C1 inhibitor), member 1 |
| chr7_-_27170352 | 0.05 |
ENST00000428284.2 ENST00000360046.5 |
HOXA4 |
homeobox A4 |
| chr7_-_14029515 | 0.05 |
ENST00000430479.1 ENST00000405218.2 ENST00000343495.5 |
ETV1 |
ets variant 1 |
| chr7_+_26591441 | 0.05 |
ENST00000420912.1 ENST00000457000.1 ENST00000430426.1 |
AC004947.2 |
AC004947.2 |
| chr7_-_100881041 | 0.05 |
ENST00000412417.1 ENST00000414035.1 |
CLDN15 |
claudin 15 |
| chr6_-_112575687 | 0.05 |
ENST00000521398.1 ENST00000424408.2 ENST00000243219.3 |
LAMA4 |
laminin, alpha 4 |
| chr19_+_44645700 | 0.05 |
ENST00000592437.1 |
ZNF234 |
zinc finger protein 234 |
| chr9_-_73477826 | 0.05 |
ENST00000396285.1 ENST00000396292.4 ENST00000396280.5 ENST00000358082.3 ENST00000408909.2 ENST00000377097.3 |
TRPM3 |
transient receptor potential cation channel, subfamily M, member 3 |
| chr6_+_32006042 | 0.05 |
ENST00000418967.2 |
CYP21A2 |
cytochrome P450, family 21, subfamily A, polypeptide 2 |
| chr5_+_133861790 | 0.05 |
ENST00000395003.1 |
PHF15 |
jade family PHD finger 2 |
| chr10_+_102891048 | 0.04 |
ENST00000467928.2 |
TLX1 |
T-cell leukemia homeobox 1 |
| chr11_-_126870655 | 0.04 |
ENST00000525144.2 |
KIRREL3 |
kin of IRRE like 3 (Drosophila) |
| chr6_+_32006159 | 0.04 |
ENST00000478281.1 ENST00000471671.1 ENST00000435122.2 |
CYP21A2 |
cytochrome P450, family 21, subfamily A, polypeptide 2 |
| chr2_-_197226875 | 0.04 |
ENST00000409111.1 |
HECW2 |
HECT, C2 and WW domain containing E3 ubiquitin protein ligase 2 |
| chr9_+_44867571 | 0.04 |
ENST00000377548.2 |
RP11-160N1.10 |
RP11-160N1.10 |
| chr10_-_65028817 | 0.04 |
ENST00000542921.1 |
JMJD1C |
jumonji domain containing 1C |
| chr3_+_191046810 | 0.03 |
ENST00000392455.3 ENST00000392456.3 |
CCDC50 |
coiled-coil domain containing 50 |
| chr8_-_143999259 | 0.03 |
ENST00000323110.2 |
CYP11B2 |
cytochrome P450, family 11, subfamily B, polypeptide 2 |
| chr10_-_65028938 | 0.03 |
ENST00000402544.1 |
JMJD1C |
jumonji domain containing 1C |
| chr10_-_98945515 | 0.03 |
ENST00000371070.4 |
SLIT1 |
slit homolog 1 (Drosophila) |
| chr12_+_57610562 | 0.03 |
ENST00000349394.5 |
NXPH4 |
neurexophilin 4 |
| chr18_+_56887381 | 0.03 |
ENST00000256857.2 ENST00000529320.2 ENST00000420468.2 |
GRP |
gastrin-releasing peptide |
| chr12_-_49393092 | 0.03 |
ENST00000421952.2 |
DDN |
dendrin |
| chr10_-_46342921 | 0.03 |
ENST00000448048.2 |
AGAP4 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 4 |
| chrX_+_48455866 | 0.03 |
ENST00000376729.5 ENST00000218056.5 |
WDR13 |
WD repeat domain 13 |
| chrX_+_95939711 | 0.03 |
ENST00000373049.4 ENST00000324765.8 |
DIAPH2 |
diaphanous-related formin 2 |
| chr12_-_58135903 | 0.03 |
ENST00000257897.3 |
AGAP2 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 2 |
| chrX_+_95939638 | 0.03 |
ENST00000373061.3 ENST00000373054.4 ENST00000355827.4 |
DIAPH2 |
diaphanous-related formin 2 |
| chr2_+_242089833 | 0.03 |
ENST00000404405.3 ENST00000439916.1 ENST00000406106.3 ENST00000401987.1 |
PPP1R7 |
protein phosphatase 1, regulatory subunit 7 |
| chr19_+_44645731 | 0.02 |
ENST00000426739.2 |
ZNF234 |
zinc finger protein 234 |
| chr9_+_44868935 | 0.02 |
ENST00000448436.2 |
RP11-160N1.10 |
RP11-160N1.10 |
| chr6_-_112575758 | 0.02 |
ENST00000431543.2 ENST00000453937.2 ENST00000368638.4 ENST00000389463.4 |
LAMA4 |
laminin, alpha 4 |
| chr13_+_52586517 | 0.02 |
ENST00000523764.1 ENST00000521508.1 |
ALG11 |
ALG11, alpha-1,2-mannosyltransferase |
| chr21_-_43430440 | 0.02 |
ENST00000398505.3 ENST00000310826.5 ENST00000449949.1 ENST00000398499.1 ENST00000398497.2 ENST00000398511.3 |
ZBTB21 |
zinc finger and BTB domain containing 21 |
| chr17_+_43239231 | 0.02 |
ENST00000591576.1 ENST00000591070.1 ENST00000592695.1 |
HEXIM2 |
hexamethylene bis-acetamide inducible 2 |
| chr21_+_19617140 | 0.02 |
ENST00000299295.2 ENST00000338326.3 |
CHODL |
chondrolectin |
| chr6_-_112575838 | 0.02 |
ENST00000455073.1 |
LAMA4 |
laminin, alpha 4 |
| chr1_+_67390578 | 0.02 |
ENST00000371018.3 ENST00000355977.6 ENST00000357692.2 ENST00000401041.1 ENST00000371016.1 ENST00000371014.1 ENST00000371012.2 |
MIER1 |
mesoderm induction early response 1, transcriptional regulator |
| chr3_-_156877997 | 0.02 |
ENST00000295926.3 |
CCNL1 |
cyclin L1 |
| chr11_-_128712362 | 0.02 |
ENST00000392664.2 |
KCNJ1 |
potassium inwardly-rectifying channel, subfamily J, member 1 |
| chr10_-_100995603 | 0.02 |
ENST00000370552.3 ENST00000370549.1 |
HPSE2 |
heparanase 2 |
| chr6_-_112575912 | 0.02 |
ENST00000522006.1 ENST00000230538.7 ENST00000519932.1 |
LAMA4 |
laminin, alpha 4 |
| chr13_+_36050881 | 0.02 |
ENST00000537702.1 |
NBEA |
neurobeachin |
| chr10_-_100995540 | 0.02 |
ENST00000370546.1 ENST00000404542.1 |
HPSE2 |
heparanase 2 |
| chr8_-_67874805 | 0.02 |
ENST00000563496.1 |
TCF24 |
transcription factor 24 |
| chr20_+_30467600 | 0.02 |
ENST00000375934.4 ENST00000375922.4 |
TTLL9 |
tubulin tyrosine ligase-like family, member 9 |
| chr3_+_189507460 | 0.01 |
ENST00000434928.1 |
TP63 |
tumor protein p63 |
| chr19_+_52074502 | 0.01 |
ENST00000545217.1 ENST00000262259.2 ENST00000596504.1 |
ZNF175 |
zinc finger protein 175 |
| chr6_+_29426230 | 0.01 |
ENST00000442615.1 |
OR2H1 |
olfactory receptor, family 2, subfamily H, member 1 |
| chr2_-_44588679 | 0.01 |
ENST00000409411.1 |
PREPL |
prolyl endopeptidase-like |
| chr6_-_159065741 | 0.01 |
ENST00000367085.3 ENST00000367089.3 |
DYNLT1 |
dynein, light chain, Tctex-type 1 |
| chr12_+_56414795 | 0.01 |
ENST00000431367.2 |
IKZF4 |
IKAROS family zinc finger 4 (Eos) |
| chr19_+_47421933 | 0.01 |
ENST00000404338.3 |
ARHGAP35 |
Rho GTPase activating protein 35 |
| chr12_+_57482877 | 0.01 |
ENST00000342556.6 ENST00000357680.4 |
NAB2 |
NGFI-A binding protein 2 (EGR1 binding protein 2) |
| chr3_-_150264272 | 0.01 |
ENST00000491660.1 ENST00000487153.1 ENST00000239944.2 |
SERP1 |
stress-associated endoplasmic reticulum protein 1 |
| chr11_+_47430133 | 0.01 |
ENST00000531974.1 ENST00000531419.1 ENST00000531865.1 ENST00000362021.4 ENST00000354884.4 |
SLC39A13 |
solute carrier family 39 (zinc transporter), member 13 |
| chr2_-_44588694 | 0.01 |
ENST00000409957.1 |
PREPL |
prolyl endopeptidase-like |
| chr15_-_64673630 | 0.01 |
ENST00000558008.1 ENST00000559519.1 ENST00000380258.2 |
KIAA0101 |
KIAA0101 |
| chr3_-_187388173 | 0.01 |
ENST00000287641.3 |
SST |
somatostatin |
| chr15_+_75335604 | 0.01 |
ENST00000563393.1 |
PPCDC |
phosphopantothenoylcysteine decarboxylase |
| chr15_-_95870310 | 0.01 |
ENST00000508732.2 |
CTD-2536I1.1 |
CTD-2536I1.1 |
| chr18_-_53070913 | 0.01 |
ENST00000568186.1 ENST00000564228.1 |
TCF4 |
transcription factor 4 |
| chr6_+_33387868 | 0.01 |
ENST00000418600.2 |
SYNGAP1 |
synaptic Ras GTPase activating protein 1 |
| chr8_+_143530791 | 0.00 |
ENST00000517894.1 |
BAI1 |
brain-specific angiogenesis inhibitor 1 |
| chr13_-_110438914 | 0.00 |
ENST00000375856.3 |
IRS2 |
insulin receptor substrate 2 |
| chr1_+_52082751 | 0.00 |
ENST00000447887.1 ENST00000435686.2 ENST00000428468.1 ENST00000453295.1 |
OSBPL9 |
oxysterol binding protein-like 9 |
| chr8_-_79717750 | 0.00 |
ENST00000263851.4 ENST00000379113.2 |
IL7 |
interleukin 7 |
| chr8_-_143961236 | 0.00 |
ENST00000377675.3 ENST00000517471.1 ENST00000292427.4 |
CYP11B1 |
cytochrome P450, family 11, subfamily B, polypeptide 1 |
| chr17_+_37824411 | 0.00 |
ENST00000269582.2 |
PNMT |
phenylethanolamine N-methyltransferase |
| chr8_+_23386305 | 0.00 |
ENST00000519973.1 |
SLC25A37 |
solute carrier family 25 (mitochondrial iron transporter), member 37 |
| chr1_-_109584716 | 0.00 |
ENST00000531337.1 ENST00000529074.1 ENST00000369965.4 |
WDR47 |
WD repeat domain 47 |
| chr1_+_12524965 | 0.00 |
ENST00000471923.1 |
VPS13D |
vacuolar protein sorting 13 homolog D (S. cerevisiae) |
| chr1_-_109584768 | 0.00 |
ENST00000357672.3 |
WDR47 |
WD repeat domain 47 |
| chr1_-_109584608 | 0.00 |
ENST00000400794.3 ENST00000528747.1 ENST00000369962.3 ENST00000361054.3 |
WDR47 |
WD repeat domain 47 |
| chrX_+_125953746 | 0.00 |
ENST00000371125.3 |
CXorf64 |
chromosome X open reading frame 64 |
| chr7_-_120497178 | 0.00 |
ENST00000441017.1 ENST00000424710.1 ENST00000433758.1 |
TSPAN12 |
tetraspanin 12 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.1 | 0.3 | GO:0000412 | histone peptidyl-prolyl isomerization(GO:0000412) |
| 0.1 | 1.3 | GO:0048681 | negative regulation of axon regeneration(GO:0048681) |
| 0.1 | 0.2 | GO:0007181 | transforming growth factor beta receptor complex assembly(GO:0007181) |
| 0.1 | 0.3 | GO:0042091 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 0.1 | 0.2 | GO:0072137 | condensed mesenchymal cell proliferation(GO:0072137) |
| 0.0 | 0.2 | GO:1903976 | negative regulation of glial cell migration(GO:1903976) |
| 0.0 | 0.2 | GO:0051585 | negative regulation of dopamine uptake involved in synaptic transmission(GO:0051585) norepinephrine uptake(GO:0051620) regulation of norepinephrine uptake(GO:0051621) negative regulation of norepinephrine uptake(GO:0051622) negative regulation of catecholamine uptake involved in synaptic transmission(GO:0051945) regulation of glutathione peroxidase activity(GO:1903282) positive regulation of glutathione peroxidase activity(GO:1903284) positive regulation of hydrogen peroxide catabolic process(GO:1903285) positive regulation of peroxidase activity(GO:2000470) |
| 0.0 | 0.1 | GO:0035283 | central nervous system segmentation(GO:0035283) brain segmentation(GO:0035284) |
| 0.0 | 0.2 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.0 | 0.1 | GO:0033563 | dorsal/ventral axon guidance(GO:0033563) |
| 0.0 | 0.3 | GO:0038028 | insulin receptor signaling pathway via phosphatidylinositol 3-kinase(GO:0038028) |
| 0.0 | 0.3 | GO:0034465 | response to carbon monoxide(GO:0034465) |
| 0.0 | 0.1 | GO:0051581 | negative regulation of neurotransmitter uptake(GO:0051581) serotonin uptake(GO:0051610) regulation of serotonin uptake(GO:0051611) negative regulation of serotonin uptake(GO:0051612) |
| 0.0 | 0.5 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.0 | 0.2 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 0.0 | 0.1 | GO:0019230 | proprioception(GO:0019230) |
| 0.0 | 0.5 | GO:0060159 | regulation of dopamine receptor signaling pathway(GO:0060159) |
| 0.0 | 0.1 | GO:0002879 | positive regulation of acute inflammatory response to non-antigenic stimulus(GO:0002879) |
| 0.0 | 0.3 | GO:0010739 | positive regulation of protein kinase A signaling(GO:0010739) |
| 0.0 | 0.1 | GO:0031456 | glycine betaine biosynthetic process from choline(GO:0019285) glycine betaine metabolic process(GO:0031455) glycine betaine biosynthetic process(GO:0031456) |
| 0.0 | 0.5 | GO:0001502 | cartilage condensation(GO:0001502) |
| 0.0 | 0.1 | GO:0021902 | commitment of neuronal cell to specific neuron type in forebrain(GO:0021902) |
| 0.0 | 0.1 | GO:0051414 | response to cortisol(GO:0051414) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.1 | 0.2 | GO:0034673 | inhibin-betaglycan-ActRII complex(GO:0034673) |
| 0.0 | 0.5 | GO:0097512 | cardiac myofibril(GO:0097512) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.3 | GO:1990459 | transferrin receptor binding(GO:1990459) |
| 0.0 | 0.3 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 0.0 | 0.1 | GO:0010698 | acetyltransferase activator activity(GO:0010698) |
| 0.0 | 0.2 | GO:0060961 | phospholipase D inhibitor activity(GO:0060961) |
| 0.0 | 0.2 | GO:0005114 | type II transforming growth factor beta receptor binding(GO:0005114) |
| 0.0 | 0.3 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.0 | 0.5 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 0.3 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.0 | 0.1 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.0 | 0.0 | GO:0004507 | steroid 11-beta-monooxygenase activity(GO:0004507) corticosterone 18-monooxygenase activity(GO:0047783) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.2 | PID BMP PATHWAY | BMP receptor signaling |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.3 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.3 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |