Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
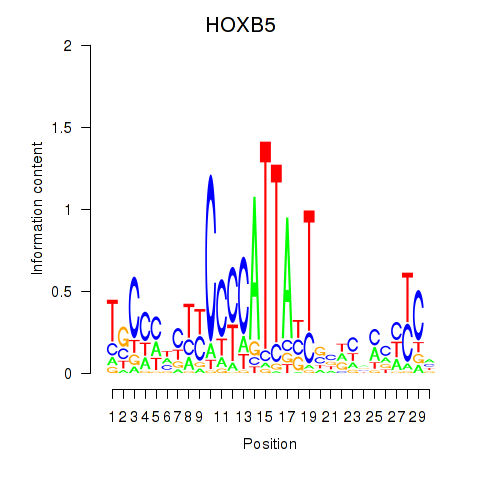
Results for HOXB5
Z-value: 0.28
Transcription factors associated with HOXB5
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HOXB5
|
ENSG00000120075.5 | HOXB5 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HOXB5 | hg19_v2_chr17_-_46671323_46671323 | -0.40 | 3.2e-01 | Click! |
Activity profile of HOXB5 motif
Sorted Z-values of HOXB5 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HOXB5
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_-_16045220 | 0.16 |
ENST00000326742.8 |
CYP4F11 |
cytochrome P450, family 4, subfamily F, polypeptide 11 |
| chr5_-_35048047 | 0.12 |
ENST00000231420.6 |
AGXT2 |
alanine--glyoxylate aminotransferase 2 |
| chr6_+_121756809 | 0.09 |
ENST00000282561.3 |
GJA1 |
gap junction protein, alpha 1, 43kDa |
| chrX_+_100333709 | 0.09 |
ENST00000372930.4 |
TMEM35 |
transmembrane protein 35 |
| chr11_-_125648690 | 0.08 |
ENST00000436890.2 ENST00000358524.3 |
PATE2 |
prostate and testis expressed 2 |
| chr8_-_133097902 | 0.07 |
ENST00000262283.5 |
OC90 |
Otoconin-90 |
| chr15_+_74218787 | 0.07 |
ENST00000261921.7 |
LOXL1 |
lysyl oxidase-like 1 |
| chr8_-_93107443 | 0.06 |
ENST00000360348.2 ENST00000520428.1 ENST00000518992.1 ENST00000520556.1 ENST00000518317.1 ENST00000521319.1 ENST00000521375.1 ENST00000518449.1 |
RUNX1T1 |
runt-related transcription factor 1; translocated to, 1 (cyclin D-related) |
| chr1_+_4714792 | 0.06 |
ENST00000378190.3 |
AJAP1 |
adherens junctions associated protein 1 |
| chr2_+_17721920 | 0.06 |
ENST00000295156.4 |
VSNL1 |
visinin-like 1 |
| chr6_-_97285336 | 0.05 |
ENST00000229955.3 ENST00000417980.1 |
GPR63 |
G protein-coupled receptor 63 |
| chr8_-_93107827 | 0.05 |
ENST00000520724.1 ENST00000518844.1 |
RUNX1T1 |
runt-related transcription factor 1; translocated to, 1 (cyclin D-related) |
| chr8_-_93107696 | 0.05 |
ENST00000436581.2 ENST00000520583.1 ENST00000519061.1 |
RUNX1T1 |
runt-related transcription factor 1; translocated to, 1 (cyclin D-related) |
| chr5_+_175298573 | 0.05 |
ENST00000512824.1 |
CPLX2 |
complexin 2 |
| chr10_-_103603568 | 0.04 |
ENST00000356640.2 |
KCNIP2 |
Kv channel interacting protein 2 |
| chr12_-_53614155 | 0.04 |
ENST00000543726.1 |
RARG |
retinoic acid receptor, gamma |
| chr12_-_53614043 | 0.04 |
ENST00000338561.5 |
RARG |
retinoic acid receptor, gamma |
| chr6_-_133119668 | 0.04 |
ENST00000275227.4 ENST00000538764.1 |
SLC18B1 |
solute carrier family 18, subfamily B, member 1 |
| chr16_+_31539197 | 0.04 |
ENST00000564707.1 |
AHSP |
alpha hemoglobin stabilizing protein |
| chr3_+_45986511 | 0.03 |
ENST00000458629.1 ENST00000457814.1 |
CXCR6 |
chemokine (C-X-C motif) receptor 6 |
| chr2_-_220173685 | 0.03 |
ENST00000423636.2 ENST00000442029.1 ENST00000412847.1 |
PTPRN |
protein tyrosine phosphatase, receptor type, N |
| chr16_-_84651647 | 0.03 |
ENST00000564057.1 |
COTL1 |
coactosin-like 1 (Dictyostelium) |
| chr13_-_54706954 | 0.02 |
ENST00000606706.1 ENST00000607494.1 ENST00000427299.2 ENST00000423442.2 ENST00000451744.1 |
LINC00458 |
long intergenic non-protein coding RNA 458 |
| chr17_+_6679528 | 0.02 |
ENST00000321535.4 |
FBXO39 |
F-box protein 39 |
| chrX_+_135230712 | 0.02 |
ENST00000535737.1 |
FHL1 |
four and a half LIM domains 1 |
| chr16_-_30102547 | 0.02 |
ENST00000279386.2 |
TBX6 |
T-box 6 |
| chr14_+_21498666 | 0.02 |
ENST00000481535.1 |
TPPP2 |
tubulin polymerization-promoting protein family member 2 |
| chr1_+_235492300 | 0.01 |
ENST00000476121.1 ENST00000497327.1 |
GGPS1 |
geranylgeranyl diphosphate synthase 1 |
| chr19_+_34745442 | 0.01 |
ENST00000299505.6 ENST00000588470.1 ENST00000589583.1 ENST00000588338.2 |
KIAA0355 |
KIAA0355 |
| chr18_+_32558208 | 0.01 |
ENST00000436190.2 |
MAPRE2 |
microtubule-associated protein, RP/EB family, member 2 |
| chr14_+_21498360 | 0.01 |
ENST00000321760.6 ENST00000460647.2 ENST00000530140.2 ENST00000472458.1 |
TPPP2 |
tubulin polymerization-promoting protein family member 2 |
| chr1_+_41157671 | 0.01 |
ENST00000534399.1 ENST00000372653.1 |
NFYC |
nuclear transcription factor Y, gamma |
| chr12_+_54955235 | 0.01 |
ENST00000550620.1 |
PDE1B |
phosphodiesterase 1B, calmodulin-dependent |
| chr2_+_219283815 | 0.01 |
ENST00000248444.5 ENST00000454069.1 ENST00000392114.2 |
VIL1 |
villin 1 |
| chr11_-_57089671 | 0.01 |
ENST00000532437.1 |
TNKS1BP1 |
tankyrase 1 binding protein 1, 182kDa |
| chr12_-_75601778 | 0.01 |
ENST00000550433.1 ENST00000548513.1 |
KCNC2 |
potassium voltage-gated channel, Shaw-related subfamily, member 2 |
| chr1_+_41157421 | 0.01 |
ENST00000372654.1 |
NFYC |
nuclear transcription factor Y, gamma |
| chr1_+_116915855 | 0.01 |
ENST00000295598.5 |
ATP1A1 |
ATPase, Na+/K+ transporting, alpha 1 polypeptide |
| chr6_-_31509714 | 0.01 |
ENST00000456662.1 ENST00000431908.1 ENST00000456976.1 ENST00000428450.1 ENST00000453105.2 ENST00000418897.1 ENST00000415382.2 ENST00000449074.2 ENST00000419020.1 ENST00000428098.1 |
DDX39B |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 39B |
| chr7_+_155090271 | 0.01 |
ENST00000476756.1 |
INSIG1 |
insulin induced gene 1 |
| chr3_-_195538760 | 0.01 |
ENST00000475231.1 |
MUC4 |
mucin 4, cell surface associated |
| chr15_-_72523454 | 0.01 |
ENST00000565154.1 ENST00000565184.1 ENST00000389093.3 ENST00000449901.2 ENST00000335181.5 ENST00000319622.6 |
PKM |
pyruvate kinase, muscle |
| chr1_+_235491714 | 0.01 |
ENST00000471812.1 ENST00000358966.2 ENST00000282841.5 ENST00000391855.2 |
GGPS1 |
geranylgeranyl diphosphate synthase 1 |
| chr6_-_31510181 | 0.01 |
ENST00000458640.1 ENST00000396172.1 ENST00000417556.2 |
DDX39B |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 39B |
| chr19_-_42724261 | 0.01 |
ENST00000595337.1 |
DEDD2 |
death effector domain containing 2 |
| chr17_-_42143963 | 0.01 |
ENST00000585388.1 ENST00000293406.3 |
LSM12 |
LSM12 homolog (S. cerevisiae) |
| chr12_+_56915776 | 0.01 |
ENST00000550726.1 ENST00000542360.1 |
RBMS2 |
RNA binding motif, single stranded interacting protein 2 |
| chr1_-_119530428 | 0.01 |
ENST00000369429.3 |
TBX15 |
T-box 15 |
| chr18_-_33077556 | 0.01 |
ENST00000589273.1 ENST00000586489.1 |
INO80C |
INO80 complex subunit C |
| chr8_+_118147498 | 0.01 |
ENST00000519688.1 ENST00000456015.2 |
SLC30A8 |
solute carrier family 30 (zinc transporter), member 8 |
| chr19_+_7571968 | 0.01 |
ENST00000599312.1 |
CTD-2207O23.12 |
Uncharacterized protein |
| chr3_-_195538728 | 0.01 |
ENST00000349607.4 ENST00000346145.4 |
MUC4 |
mucin 4, cell surface associated |
| chr1_+_41157361 | 0.01 |
ENST00000427410.2 ENST00000447388.3 ENST00000425457.2 ENST00000453631.1 ENST00000456393.2 |
NFYC |
nuclear transcription factor Y, gamma |
| chr7_-_102252038 | 0.01 |
ENST00000461209.1 |
RASA4 |
RAS p21 protein activator 4 |
| chr12_-_6798616 | 0.01 |
ENST00000355772.4 ENST00000417772.3 ENST00000396801.3 ENST00000396799.2 |
ZNF384 |
zinc finger protein 384 |
| chr7_+_130020932 | 0.00 |
ENST00000484324.1 |
CPA1 |
carboxypeptidase A1 (pancreatic) |
| chr1_+_145576007 | 0.00 |
ENST00000369298.1 |
PIAS3 |
protein inhibitor of activated STAT, 3 |
| chr10_+_114135952 | 0.00 |
ENST00000356116.1 ENST00000433418.1 ENST00000354273.4 |
ACSL5 |
acyl-CoA synthetase long-chain family member 5 |
| chr16_-_1821721 | 0.00 |
ENST00000219302.3 |
NME3 |
NME/NM23 nucleoside diphosphate kinase 3 |
| chr1_+_145575980 | 0.00 |
ENST00000393045.2 |
PIAS3 |
protein inhibitor of activated STAT, 3 |
| chr12_-_6798410 | 0.00 |
ENST00000361959.3 ENST00000436774.2 ENST00000544482.1 |
ZNF384 |
zinc finger protein 384 |
| chr12_-_6798523 | 0.00 |
ENST00000319770.3 |
ZNF384 |
zinc finger protein 384 |
| chr1_+_155658849 | 0.00 |
ENST00000368336.5 ENST00000343043.3 ENST00000421487.2 ENST00000535183.1 ENST00000465375.1 ENST00000470830.1 |
DAP3 |
death associated protein 3 |
| chr15_-_49103235 | 0.00 |
ENST00000380950.2 |
CEP152 |
centrosomal protein 152kDa |
| chr3_-_128186091 | 0.00 |
ENST00000319153.3 |
DNAJB8 |
DnaJ (Hsp40) homolog, subfamily B, member 8 |
| chr3_+_130613226 | 0.00 |
ENST00000509662.1 ENST00000328560.8 ENST00000428331.2 ENST00000359644.3 ENST00000422190.2 |
ATP2C1 |
ATPase, Ca++ transporting, type 2C, member 1 |
| chr16_-_50402836 | 0.00 |
ENST00000394688.3 |
BRD7 |
bromodomain containing 7 |
| chr6_-_32820529 | 0.00 |
ENST00000425148.2 |
TAP1 |
transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) |
| chr6_-_31869769 | 0.00 |
ENST00000375527.2 |
ZBTB12 |
zinc finger and BTB domain containing 12 |
| chr16_-_58004992 | 0.00 |
ENST00000564448.1 ENST00000251102.8 ENST00000311183.4 |
CNGB1 |
cyclic nucleotide gated channel beta 1 |
| chr6_+_167525277 | 0.00 |
ENST00000400926.2 |
CCR6 |
chemokine (C-C motif) receptor 6 |
| chr19_-_12267524 | 0.00 |
ENST00000455799.1 ENST00000355738.1 ENST00000439556.2 ENST00000542938.1 |
ZNF625 |
zinc finger protein 625 |
| chr16_-_30381580 | 0.00 |
ENST00000409939.3 |
TBC1D10B |
TBC1 domain family, member 10B |
| chr1_+_162351503 | 0.00 |
ENST00000458626.2 |
C1orf226 |
chromosome 1 open reading frame 226 |
| chr11_+_4664650 | 0.00 |
ENST00000396952.5 |
OR51E1 |
olfactory receptor, family 51, subfamily E, member 1 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | GO:0042377 | menaquinone catabolic process(GO:0042361) vitamin K catabolic process(GO:0042377) |
| 0.0 | 0.1 | GO:0009436 | glyoxylate catabolic process(GO:0009436) |
| 0.0 | 0.1 | GO:0010645 | regulation of cell communication by chemical coupling(GO:0010645) positive regulation of cell communication by chemical coupling(GO:0010652) |
| 0.0 | 0.1 | GO:0030860 | regulation of polarized epithelial cell differentiation(GO:0030860) |
| 0.0 | 0.1 | GO:0003430 | growth plate cartilage chondrocyte growth(GO:0003430) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0044214 | spanning component of plasma membrane(GO:0044214) spanning component of membrane(GO:0089717) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0008453 | alanine-glyoxylate transaminase activity(GO:0008453) |
| 0.0 | 0.1 | GO:0086075 | gap junction channel activity involved in cardiac conduction electrical coupling(GO:0086075) |