Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
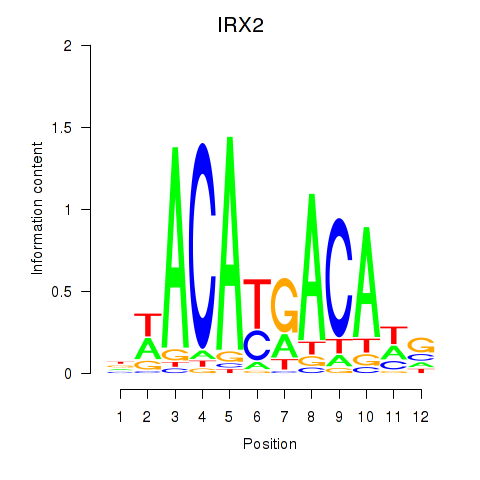
Results for IRX2
Z-value: 0.76
Transcription factors associated with IRX2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
IRX2
|
ENSG00000170561.8 | IRX2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| IRX2 | hg19_v2_chr5_-_2751762_2751784 | 0.21 | 6.2e-01 | Click! |
Activity profile of IRX2 motif
Sorted Z-values of IRX2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of IRX2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_-_100027943 | 1.37 |
ENST00000260702.3 |
LOXL4 |
lysyl oxidase-like 4 |
| chr5_+_36608422 | 0.83 |
ENST00000381918.3 |
SLC1A3 |
solute carrier family 1 (glial high affinity glutamate transporter), member 3 |
| chr16_+_68678892 | 0.70 |
ENST00000429102.2 |
CDH3 |
cadherin 3, type 1, P-cadherin (placental) |
| chr3_-_112565703 | 0.63 |
ENST00000488794.1 |
CD200R1L |
CD200 receptor 1-like |
| chr11_+_62648336 | 0.55 |
ENST00000338663.7 |
SLC3A2 |
solute carrier family 3 (amino acid transporter heavy chain), member 2 |
| chr3_+_122044084 | 0.48 |
ENST00000264474.3 ENST00000479204.1 |
CSTA |
cystatin A (stefin A) |
| chrX_-_117119243 | 0.42 |
ENST00000539496.1 ENST00000469946.1 |
KLHL13 |
kelch-like family member 13 |
| chr6_-_136847099 | 0.42 |
ENST00000438100.2 |
MAP7 |
microtubule-associated protein 7 |
| chr19_-_51466681 | 0.40 |
ENST00000456750.2 |
KLK6 |
kallikrein-related peptidase 6 |
| chr4_-_22444733 | 0.37 |
ENST00000508133.1 |
GPR125 |
G protein-coupled receptor 125 |
| chr9_-_47314222 | 0.36 |
ENST00000420228.1 ENST00000438517.1 ENST00000414020.1 |
AL953854.2 |
AL953854.2 |
| chr6_-_136847610 | 0.34 |
ENST00000454590.1 ENST00000432797.2 |
MAP7 |
microtubule-associated protein 7 |
| chr17_-_26127525 | 0.33 |
ENST00000313735.6 |
NOS2 |
nitric oxide synthase 2, inducible |
| chr16_+_82068830 | 0.31 |
ENST00000199936.4 |
HSD17B2 |
hydroxysteroid (17-beta) dehydrogenase 2 |
| chr4_+_88896819 | 0.30 |
ENST00000237623.7 ENST00000395080.3 ENST00000508233.1 ENST00000360804.4 |
SPP1 |
secreted phosphoprotein 1 |
| chr1_+_31883048 | 0.28 |
ENST00000536859.1 |
SERINC2 |
serine incorporator 2 |
| chr18_-_53070913 | 0.23 |
ENST00000568186.1 ENST00000564228.1 |
TCF4 |
transcription factor 4 |
| chr8_-_124428569 | 0.23 |
ENST00000521903.1 |
ATAD2 |
ATPase family, AAA domain containing 2 |
| chr8_-_86253888 | 0.22 |
ENST00000522389.1 ENST00000432364.2 ENST00000517618.1 |
CA1 |
carbonic anhydrase I |
| chr18_-_33702078 | 0.22 |
ENST00000586829.1 |
SLC39A6 |
solute carrier family 39 (zinc transporter), member 6 |
| chrX_+_135618258 | 0.22 |
ENST00000440515.1 ENST00000456412.1 |
VGLL1 |
vestigial like 1 (Drosophila) |
| chr3_+_97887544 | 0.20 |
ENST00000356526.2 |
OR5H15 |
olfactory receptor, family 5, subfamily H, member 15 |
| chr5_+_147258266 | 0.20 |
ENST00000296694.4 |
SCGB3A2 |
secretoglobin, family 3A, member 2 |
| chr12_+_9144626 | 0.19 |
ENST00000543895.1 |
KLRG1 |
killer cell lectin-like receptor subfamily G, member 1 |
| chr16_+_69373323 | 0.19 |
ENST00000254940.5 |
NIP7 |
NIP7, nucleolar pre-rRNA processing protein |
| chr17_-_2614927 | 0.18 |
ENST00000435359.1 |
CLUH |
clustered mitochondria (cluA/CLU1) homolog |
| chr8_+_40018977 | 0.18 |
ENST00000520487.1 |
RP11-470M17.2 |
RP11-470M17.2 |
| chr6_-_9933500 | 0.17 |
ENST00000492169.1 |
OFCC1 |
orofacial cleft 1 candidate 1 |
| chr12_+_66218903 | 0.17 |
ENST00000393577.3 |
HMGA2 |
high mobility group AT-hook 2 |
| chr16_+_56899114 | 0.16 |
ENST00000566786.1 ENST00000438926.2 ENST00000563236.1 ENST00000262502.5 |
SLC12A3 |
solute carrier family 12 (sodium/chloride transporter), member 3 |
| chr4_-_80247162 | 0.16 |
ENST00000286794.4 |
NAA11 |
N(alpha)-acetyltransferase 11, NatA catalytic subunit |
| chr10_-_61122220 | 0.15 |
ENST00000422313.2 ENST00000435852.2 ENST00000442566.3 ENST00000373868.2 ENST00000277705.6 ENST00000373867.3 ENST00000419214.2 |
FAM13C |
family with sequence similarity 13, member C |
| chr7_-_105332084 | 0.15 |
ENST00000472195.1 |
ATXN7L1 |
ataxin 7-like 1 |
| chr2_+_159651821 | 0.15 |
ENST00000309950.3 ENST00000409042.1 |
DAPL1 |
death associated protein-like 1 |
| chr15_-_44486632 | 0.14 |
ENST00000484674.1 |
FRMD5 |
FERM domain containing 5 |
| chr6_+_24775153 | 0.14 |
ENST00000356509.3 ENST00000230056.3 |
GMNN |
geminin, DNA replication inhibitor |
| chr11_+_111126707 | 0.13 |
ENST00000280325.4 |
C11orf53 |
chromosome 11 open reading frame 53 |
| chr4_+_114214125 | 0.13 |
ENST00000509550.1 |
ANK2 |
ankyrin 2, neuronal |
| chr15_+_38226827 | 0.13 |
ENST00000559502.1 ENST00000558148.1 ENST00000558158.1 |
TMCO5A |
transmembrane and coiled-coil domains 5A |
| chr2_-_166060552 | 0.13 |
ENST00000283254.7 ENST00000453007.1 |
SCN3A |
sodium channel, voltage-gated, type III, alpha subunit |
| chr6_-_53013620 | 0.13 |
ENST00000259803.7 |
GCM1 |
glial cells missing homolog 1 (Drosophila) |
| chr3_+_31574189 | 0.13 |
ENST00000295770.2 |
STT3B |
STT3B, subunit of the oligosaccharyltransferase complex (catalytic) |
| chr15_-_80263506 | 0.13 |
ENST00000335661.6 |
BCL2A1 |
BCL2-related protein A1 |
| chr17_-_60885700 | 0.13 |
ENST00000583600.1 |
MARCH10 |
membrane-associated ring finger (C3HC4) 10, E3 ubiquitin protein ligase |
| chr8_+_94767072 | 0.13 |
ENST00000452276.1 ENST00000453321.3 ENST00000498673.1 ENST00000518319.1 |
TMEM67 |
transmembrane protein 67 |
| chr5_-_94417339 | 0.12 |
ENST00000429576.2 ENST00000508509.1 ENST00000510732.1 |
MCTP1 |
multiple C2 domains, transmembrane 1 |
| chr11_-_59383617 | 0.12 |
ENST00000263847.1 |
OSBP |
oxysterol binding protein |
| chr17_-_3337135 | 0.12 |
ENST00000248384.1 |
OR1E2 |
olfactory receptor, family 1, subfamily E, member 2 |
| chr6_-_31620403 | 0.12 |
ENST00000451898.1 ENST00000439687.2 ENST00000362049.6 ENST00000424480.1 |
BAG6 |
BCL2-associated athanogene 6 |
| chr12_+_66218212 | 0.12 |
ENST00000393578.3 ENST00000425208.2 ENST00000536545.1 ENST00000354636.3 |
HMGA2 |
high mobility group AT-hook 2 |
| chr1_-_111506562 | 0.12 |
ENST00000485275.2 ENST00000369763.4 |
LRIF1 |
ligand dependent nuclear receptor interacting factor 1 |
| chr4_-_14889791 | 0.12 |
ENST00000509654.1 ENST00000515031.1 ENST00000505089.2 |
LINC00504 |
long intergenic non-protein coding RNA 504 |
| chr10_+_127661942 | 0.12 |
ENST00000417114.1 ENST00000445510.1 ENST00000368691.1 |
FANK1 |
fibronectin type III and ankyrin repeat domains 1 |
| chr11_+_33037652 | 0.12 |
ENST00000311388.3 |
DEPDC7 |
DEP domain containing 7 |
| chr11_-_118305921 | 0.12 |
ENST00000532619.1 |
RP11-770J1.4 |
RP11-770J1.4 |
| chr2_-_166060571 | 0.11 |
ENST00000360093.3 |
SCN3A |
sodium channel, voltage-gated, type III, alpha subunit |
| chr17_-_60885659 | 0.11 |
ENST00000311269.5 |
MARCH10 |
membrane-associated ring finger (C3HC4) 10, E3 ubiquitin protein ligase |
| chr1_+_113009163 | 0.11 |
ENST00000256640.5 |
WNT2B |
wingless-type MMTV integration site family, member 2B |
| chr17_-_5322786 | 0.11 |
ENST00000225696.4 |
NUP88 |
nucleoporin 88kDa |
| chr17_-_27054952 | 0.11 |
ENST00000580518.1 |
TLCD1 |
TLC domain containing 1 |
| chr5_-_35048047 | 0.10 |
ENST00000231420.6 |
AGXT2 |
alanine--glyoxylate aminotransferase 2 |
| chrX_-_118739835 | 0.10 |
ENST00000542113.1 ENST00000304449.5 |
NKRF |
NFKB repressing factor |
| chr13_-_24007815 | 0.10 |
ENST00000382298.3 |
SACS |
spastic ataxia of Charlevoix-Saguenay (sacsin) |
| chrX_-_70331298 | 0.10 |
ENST00000456850.2 ENST00000473378.1 ENST00000487883.1 ENST00000374202.2 |
IL2RG |
interleukin 2 receptor, gamma |
| chr9_+_35792151 | 0.10 |
ENST00000342694.2 |
NPR2 |
natriuretic peptide receptor B/guanylate cyclase B (atrionatriuretic peptide receptor B) |
| chr1_+_204839959 | 0.10 |
ENST00000404076.1 |
NFASC |
neurofascin |
| chr3_+_132843652 | 0.10 |
ENST00000508711.1 |
TMEM108 |
transmembrane protein 108 |
| chr5_-_94417314 | 0.10 |
ENST00000505208.1 |
MCTP1 |
multiple C2 domains, transmembrane 1 |
| chr2_+_196313239 | 0.10 |
ENST00000413290.1 |
AC064834.1 |
AC064834.1 |
| chr2_+_133874577 | 0.10 |
ENST00000596384.1 |
AC011755.1 |
HCG2006742; Protein LOC100996685 |
| chr4_-_153332886 | 0.09 |
ENST00000603841.1 |
FBXW7 |
F-box and WD repeat domain containing 7, E3 ubiquitin protein ligase |
| chr9_+_79792410 | 0.09 |
ENST00000357409.5 |
VPS13A |
vacuolar protein sorting 13 homolog A (S. cerevisiae) |
| chrX_+_30261847 | 0.09 |
ENST00000378981.3 ENST00000397550.1 |
MAGEB1 |
melanoma antigen family B, 1 |
| chr8_+_98900132 | 0.09 |
ENST00000520016.1 |
MATN2 |
matrilin 2 |
| chr1_-_207095324 | 0.08 |
ENST00000530505.1 ENST00000367091.3 ENST00000442471.2 |
FAIM3 |
Fas apoptotic inhibitory molecule 3 |
| chr19_+_13134772 | 0.08 |
ENST00000587760.1 ENST00000585575.1 |
NFIX |
nuclear factor I/X (CCAAT-binding transcription factor) |
| chr17_+_7761013 | 0.08 |
ENST00000571846.1 |
CYB5D1 |
cytochrome b5 domain containing 1 |
| chr20_+_56136136 | 0.08 |
ENST00000319441.4 ENST00000543666.1 |
PCK1 |
phosphoenolpyruvate carboxykinase 1 (soluble) |
| chr16_-_4401258 | 0.08 |
ENST00000577031.1 |
PAM16 |
presequence translocase-associated motor 16 homolog (S. cerevisiae) |
| chr8_+_81397846 | 0.08 |
ENST00000379091.4 |
ZBTB10 |
zinc finger and BTB domain containing 10 |
| chr1_-_156698181 | 0.08 |
ENST00000313146.6 |
ISG20L2 |
interferon stimulated exonuclease gene 20kDa-like 2 |
| chr6_-_31620455 | 0.08 |
ENST00000437771.1 ENST00000404765.2 ENST00000375964.6 ENST00000211379.5 |
BAG6 |
BCL2-associated athanogene 6 |
| chr21_-_31538971 | 0.08 |
ENST00000286808.3 |
CLDN17 |
claudin 17 |
| chr4_-_38806404 | 0.08 |
ENST00000308979.2 ENST00000505940.1 ENST00000515861.1 |
TLR1 |
toll-like receptor 1 |
| chr8_+_30244580 | 0.08 |
ENST00000523115.1 ENST00000519647.1 |
RBPMS |
RNA binding protein with multiple splicing |
| chr3_-_194072019 | 0.08 |
ENST00000429275.1 ENST00000323830.3 |
CPN2 |
carboxypeptidase N, polypeptide 2 |
| chr2_+_113479063 | 0.07 |
ENST00000327581.4 |
NT5DC4 |
5'-nucleotidase domain containing 4 |
| chr11_-_33795893 | 0.07 |
ENST00000526785.1 ENST00000534136.1 ENST00000265651.3 ENST00000530401.1 ENST00000448981.2 |
FBXO3 |
F-box protein 3 |
| chr15_-_45670924 | 0.07 |
ENST00000396659.3 |
GATM |
glycine amidinotransferase (L-arginine:glycine amidinotransferase) |
| chr1_+_166958346 | 0.07 |
ENST00000367872.4 |
MAEL |
maelstrom spermatogenic transposon silencer |
| chr12_-_102513843 | 0.07 |
ENST00000551744.2 ENST00000552283.1 |
NUP37 |
nucleoporin 37kDa |
| chr14_+_75536280 | 0.07 |
ENST00000238686.8 |
ZC2HC1C |
zinc finger, C2HC-type containing 1C |
| chr13_-_22033392 | 0.07 |
ENST00000320220.9 ENST00000415724.1 ENST00000422251.1 ENST00000382466.3 ENST00000542645.1 ENST00000400590.3 |
ZDHHC20 |
zinc finger, DHHC-type containing 20 |
| chr12_+_66218598 | 0.07 |
ENST00000541363.1 |
HMGA2 |
high mobility group AT-hook 2 |
| chr1_+_166958497 | 0.07 |
ENST00000367870.2 |
MAEL |
maelstrom spermatogenic transposon silencer |
| chr3_-_122102065 | 0.06 |
ENST00000479899.1 ENST00000291458.5 ENST00000497726.1 |
CCDC58 |
coiled-coil domain containing 58 |
| chr7_+_95115210 | 0.06 |
ENST00000428113.1 ENST00000325885.5 |
ASB4 |
ankyrin repeat and SOCS box containing 4 |
| chr17_+_7761301 | 0.06 |
ENST00000332439.4 ENST00000570446.1 |
CYB5D1 |
cytochrome b5 domain containing 1 |
| chr11_+_6866883 | 0.06 |
ENST00000299454.4 ENST00000379831.2 |
OR10A5 |
olfactory receptor, family 10, subfamily A, member 5 |
| chr6_+_27925019 | 0.06 |
ENST00000244623.1 |
OR2B6 |
olfactory receptor, family 2, subfamily B, member 6 |
| chr21_+_43823983 | 0.06 |
ENST00000291535.6 ENST00000450356.1 ENST00000319294.6 ENST00000398367.1 |
UBASH3A |
ubiquitin associated and SH3 domain containing A |
| chr1_+_215747118 | 0.06 |
ENST00000448333.1 |
KCTD3 |
potassium channel tetramerization domain containing 3 |
| chr7_-_150020578 | 0.06 |
ENST00000478393.1 |
ACTR3C |
ARP3 actin-related protein 3 homolog C (yeast) |
| chr1_-_157811588 | 0.06 |
ENST00000368174.4 |
CD5L |
CD5 molecule-like |
| chr4_+_159131630 | 0.06 |
ENST00000504569.1 ENST00000509278.1 ENST00000514558.1 ENST00000503200.1 |
TMEM144 |
transmembrane protein 144 |
| chr6_-_159466136 | 0.06 |
ENST00000367066.3 ENST00000326965.6 |
TAGAP |
T-cell activation RhoGTPase activating protein |
| chr1_-_173572181 | 0.06 |
ENST00000536496.1 ENST00000367714.3 |
SLC9C2 |
solute carrier family 9, member C2 (putative) |
| chrX_+_148855726 | 0.05 |
ENST00000370416.4 |
HSFX1 |
heat shock transcription factor family, X linked 1 |
| chr8_-_82359662 | 0.05 |
ENST00000519260.1 ENST00000256103.2 |
PMP2 |
peripheral myelin protein 2 |
| chr12_+_128399917 | 0.05 |
ENST00000544645.1 |
LINC00507 |
long intergenic non-protein coding RNA 507 |
| chr10_+_96522361 | 0.05 |
ENST00000371321.3 |
CYP2C19 |
cytochrome P450, family 2, subfamily C, polypeptide 19 |
| chrX_-_100129320 | 0.05 |
ENST00000372966.3 |
NOX1 |
NADPH oxidase 1 |
| chr8_+_24151553 | 0.05 |
ENST00000265769.4 ENST00000540823.1 ENST00000397649.3 |
ADAM28 |
ADAM metallopeptidase domain 28 |
| chr12_+_112856690 | 0.05 |
ENST00000392597.1 ENST00000351677.2 |
PTPN11 |
protein tyrosine phosphatase, non-receptor type 11 |
| chr2_+_48844937 | 0.05 |
ENST00000448460.1 ENST00000437125.1 ENST00000430487.2 |
GTF2A1L |
general transcription factor IIA, 1-like |
| chr16_-_5083917 | 0.05 |
ENST00000312251.3 ENST00000381955.3 |
NAGPA |
N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase |
| chr1_+_67673297 | 0.05 |
ENST00000425614.1 ENST00000395227.1 |
IL23R |
interleukin 23 receptor |
| chr14_+_62462541 | 0.05 |
ENST00000430451.2 |
SYT16 |
synaptotagmin XVI |
| chr1_-_173793458 | 0.05 |
ENST00000356198.2 |
CENPL |
centromere protein L |
| chr2_+_234637754 | 0.05 |
ENST00000482026.1 ENST00000609767.1 |
UGT1A3 UGT1A1 |
UDP glucuronosyltransferase 1 family, polypeptide A3 UDP glucuronosyltransferase 1 family, polypeptide A8 |
| chr12_-_16758304 | 0.05 |
ENST00000320122.6 |
LMO3 |
LIM domain only 3 (rhombotin-like 2) |
| chr12_-_112856623 | 0.05 |
ENST00000551291.2 |
RPL6 |
ribosomal protein L6 |
| chr14_+_75536335 | 0.05 |
ENST00000554763.1 ENST00000439583.2 ENST00000526130.1 ENST00000525046.1 |
ZC2HC1C |
zinc finger, C2HC-type containing 1C |
| chr12_-_10007448 | 0.04 |
ENST00000538152.1 |
CLEC2B |
C-type lectin domain family 2, member B |
| chr2_+_48844973 | 0.04 |
ENST00000403751.3 |
GTF2A1L |
general transcription factor IIA, 1-like |
| chr16_+_58426296 | 0.04 |
ENST00000426538.2 ENST00000328514.7 ENST00000318129.5 |
GINS3 |
GINS complex subunit 3 (Psf3 homolog) |
| chrX_-_148676974 | 0.04 |
ENST00000524178.1 |
HSFX2 |
heat shock transcription factor family, X linked 2 |
| chr17_-_8055747 | 0.04 |
ENST00000317276.4 ENST00000581703.1 |
PER1 |
period circadian clock 1 |
| chrX_-_100129128 | 0.04 |
ENST00000372960.4 ENST00000372964.1 ENST00000217885.5 |
NOX1 |
NADPH oxidase 1 |
| chr4_+_187187337 | 0.04 |
ENST00000492972.2 |
F11 |
coagulation factor XI |
| chr4_-_168155417 | 0.04 |
ENST00000511269.1 ENST00000506697.1 ENST00000512042.1 |
SPOCK3 |
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3 |
| chr2_+_168675182 | 0.04 |
ENST00000305861.1 |
B3GALT1 |
UDP-Gal:betaGlcNAc beta 1,3-galactosyltransferase, polypeptide 1 |
| chr12_-_16758059 | 0.04 |
ENST00000261169.6 |
LMO3 |
LIM domain only 3 (rhombotin-like 2) |
| chr6_+_88117683 | 0.04 |
ENST00000369562.4 |
C6ORF165 |
UPF0704 protein C6orf165 |
| chr4_-_71705060 | 0.03 |
ENST00000514161.1 |
GRSF1 |
G-rich RNA sequence binding factor 1 |
| chr1_-_151148442 | 0.03 |
ENST00000441701.1 ENST00000416280.2 |
TMOD4 |
tropomodulin 4 (muscle) |
| chr4_+_158141899 | 0.03 |
ENST00000264426.9 ENST00000506284.1 |
GRIA2 |
glutamate receptor, ionotropic, AMPA 2 |
| chr7_-_130598059 | 0.03 |
ENST00000432045.2 |
MIR29B1 |
microRNA 29a |
| chr5_-_180242534 | 0.03 |
ENST00000333055.3 ENST00000513431.1 |
MGAT1 |
mannosyl (alpha-1,3-)-glycoprotein beta-1,2-N-acetylglucosaminyltransferase |
| chr4_-_71705027 | 0.03 |
ENST00000545193.1 |
GRSF1 |
G-rich RNA sequence binding factor 1 |
| chr9_-_35812236 | 0.03 |
ENST00000340291.2 |
SPAG8 |
sperm associated antigen 8 |
| chr14_-_65289812 | 0.03 |
ENST00000389720.3 ENST00000389721.5 ENST00000389722.3 |
SPTB |
spectrin, beta, erythrocytic |
| chr1_-_151148492 | 0.03 |
ENST00000295314.4 |
TMOD4 |
tropomodulin 4 (muscle) |
| chr12_+_128399965 | 0.03 |
ENST00000540882.1 ENST00000542089.1 |
LINC00507 |
long intergenic non-protein coding RNA 507 |
| chr1_-_109399682 | 0.03 |
ENST00000369995.3 ENST00000370001.3 |
AKNAD1 |
AKNA domain containing 1 |
| chrX_-_119077695 | 0.03 |
ENST00000371410.3 |
NKAP |
NFKB activating protein |
| chr14_+_23790655 | 0.02 |
ENST00000397276.2 |
PABPN1 |
poly(A) binding protein, nuclear 1 |
| chr3_-_167371740 | 0.02 |
ENST00000466760.1 ENST00000479765.1 |
WDR49 |
WD repeat domain 49 |
| chr9_-_113100088 | 0.02 |
ENST00000374510.4 ENST00000423740.2 ENST00000374511.3 ENST00000374507.4 |
TXNDC8 |
thioredoxin domain containing 8 (spermatozoa) |
| chr11_-_110561721 | 0.02 |
ENST00000357139.3 |
ARHGAP20 |
Rho GTPase activating protein 20 |
| chrX_-_49042778 | 0.02 |
ENST00000538114.1 ENST00000376310.3 ENST00000376317.3 ENST00000417014.1 |
PRICKLE3 |
prickle homolog 3 (Drosophila) |
| chr10_+_102672712 | 0.02 |
ENST00000370271.3 ENST00000370269.3 ENST00000609386.1 |
FAM178A |
family with sequence similarity 178, member A |
| chr12_+_40787194 | 0.02 |
ENST00000425730.2 ENST00000454784.4 |
MUC19 |
mucin 19, oligomeric |
| chr1_+_29138654 | 0.02 |
ENST00000234961.2 |
OPRD1 |
opioid receptor, delta 1 |
| chr4_+_109571740 | 0.02 |
ENST00000361564.4 |
OSTC |
oligosaccharyltransferase complex subunit (non-catalytic) |
| chr4_+_83821835 | 0.02 |
ENST00000302236.5 |
THAP9 |
THAP domain containing 9 |
| chr6_+_13272904 | 0.02 |
ENST00000379335.3 ENST00000379329.1 |
PHACTR1 |
phosphatase and actin regulator 1 |
| chrX_+_155227246 | 0.02 |
ENST00000244174.5 ENST00000424344.3 |
IL9R |
interleukin 9 receptor |
| chr2_+_27851863 | 0.02 |
ENST00000264718.3 ENST00000610189.1 |
GPN1 |
GPN-loop GTPase 1 |
| chr1_-_190446759 | 0.01 |
ENST00000367462.3 |
BRINP3 |
bone morphogenetic protein/retinoic acid inducible neural-specific 3 |
| chr1_-_207095212 | 0.01 |
ENST00000420007.2 |
FAIM3 |
Fas apoptotic inhibitory molecule 3 |
| chr20_-_29978286 | 0.01 |
ENST00000376315.2 |
DEFB119 |
defensin, beta 119 |
| chr4_+_109571800 | 0.01 |
ENST00000512478.2 |
OSTC |
oligosaccharyltransferase complex subunit (non-catalytic) |
| chr14_-_106312010 | 0.01 |
ENST00000390556.2 |
IGHD |
immunoglobulin heavy constant delta |
| chr1_+_16083154 | 0.01 |
ENST00000375771.1 |
FBLIM1 |
filamin binding LIM protein 1 |
| chr10_+_11047259 | 0.01 |
ENST00000379261.4 ENST00000416382.2 |
CELF2 |
CUGBP, Elav-like family member 2 |
| chr19_-_29704448 | 0.01 |
ENST00000304863.4 |
UQCRFS1 |
ubiquinol-cytochrome c reductase, Rieske iron-sulfur polypeptide 1 |
| chrX_-_132095419 | 0.01 |
ENST00000370836.2 ENST00000521489.1 |
HS6ST2 |
heparan sulfate 6-O-sulfotransferase 2 |
| chr11_+_59824127 | 0.01 |
ENST00000278865.3 |
MS4A3 |
membrane-spanning 4-domains, subfamily A, member 3 (hematopoietic cell-specific) |
| chr20_+_43849941 | 0.01 |
ENST00000372769.3 |
SEMG2 |
semenogelin II |
| chrX_+_155227371 | 0.01 |
ENST00000369423.2 ENST00000540897.1 |
IL9R |
interleukin 9 receptor |
| chr22_+_23046750 | 0.01 |
ENST00000390307.2 |
IGLV3-22 |
immunoglobulin lambda variable 3-22 (gene/pseudogene) |
| chr18_+_32455201 | 0.01 |
ENST00000590831.2 |
DTNA |
dystrobrevin, alpha |
| chr19_-_40336969 | 0.01 |
ENST00000599134.1 ENST00000597634.1 ENST00000598417.1 ENST00000601274.1 ENST00000594309.1 ENST00000221801.3 |
FBL |
fibrillarin |
| chr19_+_16308711 | 0.00 |
ENST00000429941.2 ENST00000444449.2 ENST00000589822.1 |
AP1M1 |
adaptor-related protein complex 1, mu 1 subunit |
| chr3_-_126327398 | 0.00 |
ENST00000383572.2 |
TXNRD3NB |
thioredoxin reductase 3 neighbor |
| chr17_+_74075263 | 0.00 |
ENST00000334586.5 ENST00000392503.2 |
ZACN |
zinc activated ligand-gated ion channel |
| chr5_-_39364586 | 0.00 |
ENST00000263408.4 |
C9 |
complement component 9 |
| chr22_-_36013368 | 0.00 |
ENST00000442617.1 ENST00000397326.2 ENST00000397328.1 ENST00000451685.1 |
MB |
myoglobin |
| chr10_-_4285923 | 0.00 |
ENST00000418372.1 ENST00000608792.1 |
LINC00702 |
long intergenic non-protein coding RNA 702 |
| chrX_+_19373700 | 0.00 |
ENST00000379804.1 |
PDHA1 |
pyruvate dehydrogenase (lipoamide) alpha 1 |
| chr8_+_67104323 | 0.00 |
ENST00000518412.1 ENST00000518035.1 ENST00000517725.1 |
LINC00967 |
long intergenic non-protein coding RNA 967 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0051796 | regulation of monophenol monooxygenase activity(GO:0032771) positive regulation of monophenol monooxygenase activity(GO:0032773) negative regulation of catagen(GO:0051796) regulation of hair cycle by canonical Wnt signaling pathway(GO:0060901) positive regulation of melanosome transport(GO:1902910) |
| 0.1 | 0.8 | GO:0006537 | glutamate biosynthetic process(GO:0006537) gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 0.1 | 0.5 | GO:0060356 | leucine import(GO:0060356) |
| 0.1 | 0.3 | GO:2000866 | positive regulation of estrogen secretion(GO:2000863) positive regulation of estradiol secretion(GO:2000866) |
| 0.1 | 0.4 | GO:2000685 | mesodermal-endodermal cell signaling(GO:0003131) programmed DNA elimination(GO:0031049) chromosome breakage(GO:0031052) histone H2A-S139 phosphorylation(GO:0035978) positive regulation of cellular response to X-ray(GO:2000685) |
| 0.1 | 0.3 | GO:1904219 | regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904217) positive regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904219) positive regulation of serine C-palmitoyltransferase activity(GO:1904222) |
| 0.0 | 0.2 | GO:1904379 | protein localization to cytosolic proteasome complex(GO:1904327) protein localization to cytosolic proteasome complex involved in ERAD pathway(GO:1904379) |
| 0.0 | 0.1 | GO:0036371 | protein localization to T-tubule(GO:0036371) |
| 0.0 | 0.1 | GO:0043686 | co-translational protein modification(GO:0043686) |
| 0.0 | 0.2 | GO:0030920 | N-terminal peptidyl-serine acetylation(GO:0017198) N-terminal peptidyl-glutamic acid acetylation(GO:0018002) peptidyl-serine acetylation(GO:0030920) |
| 0.0 | 0.1 | GO:0009436 | glyoxylate catabolic process(GO:0009436) |
| 0.0 | 0.1 | GO:0060018 | astrocyte fate commitment(GO:0060018) |
| 0.0 | 0.1 | GO:0042495 | detection of triacyl bacterial lipopeptide(GO:0042495) detection of bacterial lipopeptide(GO:0070340) |
| 0.0 | 0.1 | GO:1900194 | negative regulation of oocyte maturation(GO:1900194) |
| 0.0 | 0.1 | GO:0071163 | DNA replication preinitiation complex assembly(GO:0071163) |
| 0.0 | 0.3 | GO:0031284 | positive regulation of guanylate cyclase activity(GO:0031284) positive regulation of killing of cells of other organism(GO:0051712) |
| 0.0 | 0.1 | GO:0045726 | positive regulation of integrin biosynthetic process(GO:0045726) |
| 0.0 | 0.3 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.0 | 0.1 | GO:0006114 | glycerol biosynthetic process(GO:0006114) |
| 0.0 | 0.1 | GO:2000638 | regulation of SREBP signaling pathway(GO:2000638) negative regulation of SREBP signaling pathway(GO:2000639) |
| 0.0 | 0.2 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.0 | 0.1 | GO:0061582 | intestinal epithelial cell migration(GO:0061582) |
| 0.0 | 0.1 | GO:0038110 | interleukin-2-mediated signaling pathway(GO:0038110) |
| 0.0 | 0.4 | GO:0016540 | protein autoprocessing(GO:0016540) |
| 0.0 | 0.1 | GO:0060492 | foregut regionalization(GO:0060423) lung field specification(GO:0060424) lung induction(GO:0060492) |
| 0.0 | 0.1 | GO:2000324 | positive regulation of glucocorticoid receptor signaling pathway(GO:2000324) |
| 0.0 | 0.1 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.0 | 0.2 | GO:0071578 | zinc II ion transmembrane import(GO:0071578) |
| 0.0 | 0.1 | GO:0000055 | ribosomal large subunit export from nucleus(GO:0000055) ribosomal small subunit export from nucleus(GO:0000056) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0030849 | autosome(GO:0030849) |
| 0.0 | 0.2 | GO:0071818 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.0 | 0.4 | GO:0035985 | senescence-associated heterochromatin focus(GO:0035985) |
| 0.0 | 0.1 | GO:0035354 | Toll-like receptor 1-Toll-like receptor 2 protein complex(GO:0035354) |
| 0.0 | 0.1 | GO:0097454 | Schwann cell microvillus(GO:0097454) |
| 0.0 | 0.2 | GO:0031415 | NatA complex(GO:0031415) |
| 0.0 | 0.1 | GO:0001405 | presequence translocase-associated import motor(GO:0001405) |
| 0.0 | 0.2 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.0 | 0.1 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
| 0.0 | 0.1 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 0.0 | 0.1 | GO:0070852 | cell body fiber(GO:0070852) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.8 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.1 | 0.4 | GO:0035501 | MH1 domain binding(GO:0035501) |
| 0.1 | 0.3 | GO:0004517 | nitric-oxide synthase activity(GO:0004517) |
| 0.1 | 0.3 | GO:0047006 | 17-alpha,20-alpha-dihydroxypregn-4-en-3-one dehydrogenase activity(GO:0047006) |
| 0.1 | 1.4 | GO:0016641 | oxidoreductase activity, acting on the CH-NH2 group of donors, oxygen as acceptor(GO:0016641) |
| 0.0 | 0.2 | GO:1990190 | peptide-glutamate-N-acetyltransferase activity(GO:1990190) |
| 0.0 | 0.5 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
| 0.0 | 0.1 | GO:0004911 | interleukin-2 receptor activity(GO:0004911) interleukin-2 binding(GO:0019976) |
| 0.0 | 0.2 | GO:0004064 | arylesterase activity(GO:0004064) |
| 0.0 | 0.1 | GO:0004613 | phosphoenolpyruvate carboxykinase activity(GO:0004611) phosphoenolpyruvate carboxykinase (GTP) activity(GO:0004613) |
| 0.0 | 0.1 | GO:0030171 | voltage-gated proton channel activity(GO:0030171) |
| 0.0 | 0.1 | GO:0008453 | alanine-glyoxylate transaminase activity(GO:0008453) |
| 0.0 | 0.1 | GO:0035663 | Toll-like receptor 2 binding(GO:0035663) |
| 0.0 | 0.3 | GO:0022889 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.0 | 0.2 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.0 | 0.1 | GO:0016941 | natriuretic peptide receptor activity(GO:0016941) |
| 0.0 | 0.1 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.0 | 0.3 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.0 | 0.1 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.0 | 0.2 | GO:0005549 | odorant binding(GO:0005549) |
| 0.0 | 0.0 | GO:0047275 | glucosaminylgalactosylglucosylceramide beta-galactosyltransferase activity(GO:0047275) |
| 0.0 | 0.1 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.0 | 0.1 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 0.0 | 0.2 | GO:0004579 | dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.0 | 0.2 | GO:0015377 | cation:chloride symporter activity(GO:0015377) |
| 0.0 | 0.1 | GO:0035614 | snRNA stem-loop binding(GO:0035614) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 1.4 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.0 | 0.2 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |