Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
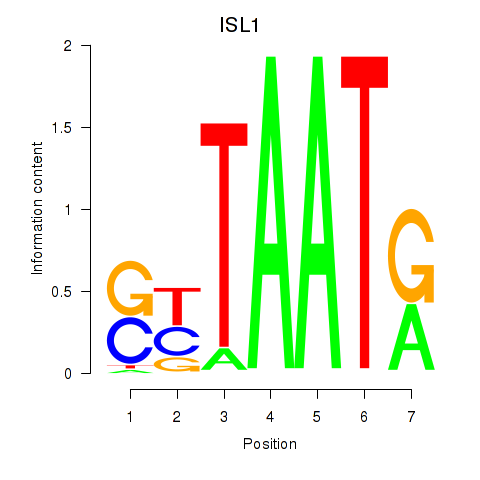
Results for ISL1
Z-value: 0.55
Transcription factors associated with ISL1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ISL1
|
ENSG00000016082.10 | ISL1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ISL1 | hg19_v2_chr5_+_50678921_50678921 | 0.13 | 7.6e-01 | Click! |
Activity profile of ISL1 motif
Sorted Z-values of ISL1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ISL1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_+_137203557 | 0.49 |
ENST00000515645.1 |
MYOT |
myotilin |
| chr5_+_137203541 | 0.36 |
ENST00000421631.2 |
MYOT |
myotilin |
| chr4_-_186877806 | 0.31 |
ENST00000355634.5 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr14_+_22458631 | 0.29 |
ENST00000390444.1 |
TRAV16 |
T cell receptor alpha variable 16 |
| chr1_-_108231101 | 0.23 |
ENST00000544443.1 ENST00000415432.2 |
VAV3 |
vav 3 guanine nucleotide exchange factor |
| chrX_+_135252050 | 0.22 |
ENST00000449474.1 ENST00000345434.3 |
FHL1 |
four and a half LIM domains 1 |
| chrX_+_135251835 | 0.21 |
ENST00000456445.1 |
FHL1 |
four and a half LIM domains 1 |
| chr13_-_70682590 | 0.21 |
ENST00000377844.4 |
KLHL1 |
kelch-like family member 1 |
| chr4_-_186877502 | 0.21 |
ENST00000431902.1 ENST00000284776.7 ENST00000415274.1 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr1_+_196621156 | 0.20 |
ENST00000359637.2 |
CFH |
complement factor H |
| chr12_-_50290839 | 0.19 |
ENST00000552863.1 |
FAIM2 |
Fas apoptotic inhibitory molecule 2 |
| chr2_-_175711133 | 0.19 |
ENST00000409597.1 ENST00000413882.1 |
CHN1 |
chimerin 1 |
| chr10_-_116444371 | 0.18 |
ENST00000533213.2 ENST00000369252.4 |
ABLIM1 |
actin binding LIM protein 1 |
| chrX_+_135251783 | 0.17 |
ENST00000394153.2 |
FHL1 |
four and a half LIM domains 1 |
| chr4_+_113970772 | 0.16 |
ENST00000504454.1 ENST00000394537.3 ENST00000357077.4 ENST00000264366.6 |
ANK2 |
ankyrin 2, neuronal |
| chr2_+_207804278 | 0.15 |
ENST00000272852.3 |
CPO |
carboxypeptidase O |
| chr17_+_66521936 | 0.15 |
ENST00000592800.1 |
PRKAR1A |
protein kinase, cAMP-dependent, regulatory, type I, alpha |
| chr5_+_32710736 | 0.14 |
ENST00000415685.2 |
NPR3 |
natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C) |
| chr14_-_21566731 | 0.13 |
ENST00000360947.3 |
ZNF219 |
zinc finger protein 219 |
| chr12_-_87232644 | 0.13 |
ENST00000549405.2 |
MGAT4C |
mannosyl (alpha-1,3-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase, isozyme C (putative) |
| chr19_-_50143452 | 0.13 |
ENST00000246792.3 |
RRAS |
related RAS viral (r-ras) oncogene homolog |
| chr6_+_35996859 | 0.12 |
ENST00000472333.1 |
MAPK14 |
mitogen-activated protein kinase 14 |
| chr4_-_87281196 | 0.12 |
ENST00000359221.3 |
MAPK10 |
mitogen-activated protein kinase 10 |
| chr3_-_21792838 | 0.12 |
ENST00000281523.2 |
ZNF385D |
zinc finger protein 385D |
| chr8_-_101571964 | 0.12 |
ENST00000520552.1 ENST00000521345.1 ENST00000523000.1 ENST00000335659.3 ENST00000358990.3 ENST00000519597.1 |
ANKRD46 |
ankyrin repeat domain 46 |
| chr5_+_32711419 | 0.11 |
ENST00000265074.8 |
NPR3 |
natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C) |
| chr2_+_202937972 | 0.11 |
ENST00000541917.1 ENST00000295844.3 |
AC079354.1 |
uncharacterized protein KIAA2012 |
| chr1_+_156024525 | 0.11 |
ENST00000368305.4 |
LAMTOR2 |
late endosomal/lysosomal adaptor, MAPK and MTOR activator 2 |
| chr1_+_156024552 | 0.11 |
ENST00000368304.5 ENST00000368302.3 |
LAMTOR2 |
late endosomal/lysosomal adaptor, MAPK and MTOR activator 2 |
| chr17_-_21156578 | 0.11 |
ENST00000399011.2 ENST00000468196.1 |
C17orf103 |
chromosome 17 open reading frame 103 |
| chr8_-_7220490 | 0.11 |
ENST00000400078.2 |
ZNF705G |
zinc finger protein 705G |
| chr2_+_219472637 | 0.10 |
ENST00000417849.1 |
PLCD4 |
phospholipase C, delta 4 |
| chr1_-_95391315 | 0.10 |
ENST00000545882.1 ENST00000415017.1 |
CNN3 |
calponin 3, acidic |
| chr1_+_82266053 | 0.10 |
ENST00000370715.1 ENST00000370713.1 ENST00000319517.6 ENST00000370717.2 ENST00000394879.1 ENST00000271029.4 ENST00000335786.5 |
LPHN2 |
latrophilin 2 |
| chr4_+_146539415 | 0.10 |
ENST00000281317.5 |
MMAA |
methylmalonic aciduria (cobalamin deficiency) cblA type |
| chr7_+_13141097 | 0.10 |
ENST00000411542.1 |
AC011288.2 |
AC011288.2 |
| chr10_+_114710425 | 0.10 |
ENST00000352065.5 ENST00000369395.1 |
TCF7L2 |
transcription factor 7-like 2 (T-cell specific, HMG-box) |
| chr1_+_217804661 | 0.10 |
ENST00000366933.4 |
SPATA17 |
spermatogenesis associated 17 |
| chr17_+_6918354 | 0.10 |
ENST00000552775.1 |
C17orf49 |
chromosome 17 open reading frame 49 |
| chr17_+_35732916 | 0.10 |
ENST00000586700.1 |
C17orf78 |
chromosome 17 open reading frame 78 |
| chr20_-_5426332 | 0.10 |
ENST00000420529.1 |
LINC00658 |
long intergenic non-protein coding RNA 658 |
| chr17_+_35732955 | 0.09 |
ENST00000300618.4 |
C17orf78 |
chromosome 17 open reading frame 78 |
| chr7_+_150811705 | 0.09 |
ENST00000335367.3 |
AGAP3 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 3 |
| chr14_-_21567009 | 0.09 |
ENST00000556174.1 ENST00000554478.1 ENST00000553980.1 ENST00000421093.2 |
ZNF219 |
zinc finger protein 219 |
| chr3_+_57875711 | 0.09 |
ENST00000442599.2 |
SLMAP |
sarcolemma associated protein |
| chr21_-_34185944 | 0.09 |
ENST00000479548.1 |
C21orf62 |
chromosome 21 open reading frame 62 |
| chr1_+_150980889 | 0.09 |
ENST00000450884.1 ENST00000271620.3 ENST00000271619.8 ENST00000368937.1 ENST00000431193.1 ENST00000368936.1 |
PRUNE |
prune exopolyphosphatase |
| chr8_-_101571933 | 0.09 |
ENST00000520311.1 |
ANKRD46 |
ankyrin repeat domain 46 |
| chr17_-_2996290 | 0.09 |
ENST00000331459.1 |
OR1D2 |
olfactory receptor, family 1, subfamily D, member 2 |
| chr8_+_26247878 | 0.09 |
ENST00000518611.1 |
BNIP3L |
BCL2/adenovirus E1B 19kDa interacting protein 3-like |
| chr8_-_101321584 | 0.09 |
ENST00000523167.1 |
RNF19A |
ring finger protein 19A, RBR E3 ubiquitin protein ligase |
| chr13_+_46276441 | 0.09 |
ENST00000310521.1 ENST00000533564.1 |
SPERT |
spermatid associated |
| chr1_-_79472365 | 0.09 |
ENST00000370742.3 |
ELTD1 |
EGF, latrophilin and seven transmembrane domain containing 1 |
| chr5_+_32788945 | 0.09 |
ENST00000326958.1 |
AC026703.1 |
AC026703.1 |
| chr10_-_27529486 | 0.09 |
ENST00000375888.1 |
ACBD5 |
acyl-CoA binding domain containing 5 |
| chr3_+_119501557 | 0.08 |
ENST00000337940.4 |
NR1I2 |
nuclear receptor subfamily 1, group I, member 2 |
| chr12_-_10251576 | 0.08 |
ENST00000315330.4 |
CLEC1A |
C-type lectin domain family 1, member A |
| chr11_-_59951889 | 0.08 |
ENST00000532169.1 ENST00000534596.1 |
MS4A6A |
membrane-spanning 4-domains, subfamily A, member 6A |
| chr21_+_17792672 | 0.08 |
ENST00000602620.1 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr2_-_227050079 | 0.08 |
ENST00000423838.1 |
AC068138.1 |
AC068138.1 |
| chr3_-_185538849 | 0.08 |
ENST00000421047.2 |
IGF2BP2 |
insulin-like growth factor 2 mRNA binding protein 2 |
| chr4_+_74347400 | 0.08 |
ENST00000226355.3 |
AFM |
afamin |
| chr1_-_119682812 | 0.08 |
ENST00000537870.1 |
WARS2 |
tryptophanyl tRNA synthetase 2, mitochondrial |
| chr14_-_45252031 | 0.08 |
ENST00000556405.1 |
RP11-398E10.1 |
RP11-398E10.1 |
| chr1_+_84873913 | 0.08 |
ENST00000370662.3 |
DNASE2B |
deoxyribonuclease II beta |
| chr2_+_211421262 | 0.08 |
ENST00000233072.5 |
CPS1 |
carbamoyl-phosphate synthase 1, mitochondrial |
| chr7_-_31380502 | 0.08 |
ENST00000297142.3 |
NEUROD6 |
neuronal differentiation 6 |
| chr12_-_10251603 | 0.08 |
ENST00000457018.2 |
CLEC1A |
C-type lectin domain family 1, member A |
| chr4_+_123300121 | 0.08 |
ENST00000446706.1 ENST00000296513.2 |
ADAD1 |
adenosine deaminase domain containing 1 (testis-specific) |
| chr12_-_53012343 | 0.07 |
ENST00000305748.3 |
KRT73 |
keratin 73 |
| chr16_-_4665023 | 0.07 |
ENST00000591897.1 |
UBALD1 |
UBA-like domain containing 1 |
| chr11_+_60995849 | 0.07 |
ENST00000537932.1 |
PGA4 |
pepsinogen 4, group I (pepsinogen A) |
| chr17_-_57229155 | 0.07 |
ENST00000584089.1 |
SKA2 |
spindle and kinetochore associated complex subunit 2 |
| chr11_-_40315640 | 0.07 |
ENST00000278198.2 |
LRRC4C |
leucine rich repeat containing 4C |
| chr1_-_160254913 | 0.07 |
ENST00000440949.3 ENST00000368072.5 ENST00000608310.1 ENST00000556710.1 |
PEX19 DCAF8 DCAF8 |
peroxisomal biogenesis factor 19 DDB1 and CUL4 associated factor 8 DDB1- and CUL4-associated factor 8 |
| chr11_+_61248583 | 0.07 |
ENST00000432063.2 ENST00000338608.2 |
PPP1R32 |
protein phosphatase 1, regulatory subunit 32 |
| chr9_+_2029019 | 0.07 |
ENST00000382194.1 |
SMARCA2 |
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr16_-_4664860 | 0.07 |
ENST00000587615.1 ENST00000587649.1 ENST00000590965.1 ENST00000591401.1 ENST00000283474.7 |
UBALD1 |
UBA-like domain containing 1 |
| chr15_-_72767490 | 0.07 |
ENST00000565181.1 |
RP11-1007O24.3 |
RP11-1007O24.3 |
| chr3_-_121740969 | 0.06 |
ENST00000393631.1 ENST00000273691.3 ENST00000344209.5 |
ILDR1 |
immunoglobulin-like domain containing receptor 1 |
| chr9_+_131084846 | 0.06 |
ENST00000608951.1 |
COQ4 |
coenzyme Q4 |
| chr17_+_37824217 | 0.06 |
ENST00000394246.1 |
PNMT |
phenylethanolamine N-methyltransferase |
| chr16_-_52119019 | 0.06 |
ENST00000561513.1 ENST00000565742.1 |
LINC00919 |
long intergenic non-protein coding RNA 919 |
| chr17_+_37824411 | 0.06 |
ENST00000269582.2 |
PNMT |
phenylethanolamine N-methyltransferase |
| chr1_-_197169672 | 0.06 |
ENST00000367405.4 |
ZBTB41 |
zinc finger and BTB domain containing 41 |
| chr14_-_61190754 | 0.06 |
ENST00000216513.4 |
SIX4 |
SIX homeobox 4 |
| chr6_-_111136513 | 0.06 |
ENST00000368911.3 |
CDK19 |
cyclin-dependent kinase 19 |
| chr9_-_134585221 | 0.06 |
ENST00000372190.3 ENST00000427994.1 |
RAPGEF1 |
Rap guanine nucleotide exchange factor (GEF) 1 |
| chr3_-_138312971 | 0.06 |
ENST00000485115.1 ENST00000484888.1 ENST00000468900.1 ENST00000542237.1 ENST00000481834.1 |
CEP70 |
centrosomal protein 70kDa |
| chrX_+_10031499 | 0.06 |
ENST00000454666.1 |
WWC3 |
WWC family member 3 |
| chr22_+_30279144 | 0.06 |
ENST00000401950.2 ENST00000333027.3 ENST00000445401.1 ENST00000323630.5 ENST00000351488.3 |
MTMR3 |
myotubularin related protein 3 |
| chr12_-_42631529 | 0.06 |
ENST00000548917.1 |
YAF2 |
YY1 associated factor 2 |
| chr2_+_171034646 | 0.06 |
ENST00000409044.3 ENST00000408978.4 |
MYO3B |
myosin IIIB |
| chr8_+_110098850 | 0.06 |
ENST00000518632.1 |
TRHR |
thyrotropin-releasing hormone receptor |
| chr6_-_11779014 | 0.05 |
ENST00000229583.5 |
ADTRP |
androgen-dependent TFPI-regulating protein |
| chr8_+_21911054 | 0.05 |
ENST00000519850.1 ENST00000381470.3 |
DMTN |
dematin actin binding protein |
| chr5_-_94417314 | 0.05 |
ENST00000505208.1 |
MCTP1 |
multiple C2 domains, transmembrane 1 |
| chr6_+_132455118 | 0.05 |
ENST00000458028.1 |
LINC01013 |
long intergenic non-protein coding RNA 1013 |
| chr11_+_128563652 | 0.05 |
ENST00000527786.2 |
FLI1 |
Fli-1 proto-oncogene, ETS transcription factor |
| chr5_-_94417339 | 0.05 |
ENST00000429576.2 ENST00000508509.1 ENST00000510732.1 |
MCTP1 |
multiple C2 domains, transmembrane 1 |
| chr12_+_122356488 | 0.05 |
ENST00000397454.2 |
WDR66 |
WD repeat domain 66 |
| chr2_+_28618532 | 0.05 |
ENST00000545753.1 |
FOSL2 |
FOS-like antigen 2 |
| chr2_+_234216454 | 0.05 |
ENST00000447536.1 ENST00000409110.1 |
SAG |
S-antigen; retina and pineal gland (arrestin) |
| chr2_-_74735707 | 0.05 |
ENST00000233630.6 |
PCGF1 |
polycomb group ring finger 1 |
| chr4_-_87281224 | 0.05 |
ENST00000395169.3 ENST00000395161.2 |
MAPK10 |
mitogen-activated protein kinase 10 |
| chrX_+_144908928 | 0.05 |
ENST00000408967.2 |
TMEM257 |
transmembrane protein 257 |
| chr8_-_19540086 | 0.05 |
ENST00000332246.6 |
CSGALNACT1 |
chondroitin sulfate N-acetylgalactosaminyltransferase 1 |
| chr4_+_169418255 | 0.05 |
ENST00000505667.1 ENST00000511948.1 |
PALLD |
palladin, cytoskeletal associated protein |
| chr21_-_34185989 | 0.05 |
ENST00000487113.1 ENST00000382373.4 |
C21orf62 |
chromosome 21 open reading frame 62 |
| chr19_-_42806842 | 0.05 |
ENST00000596265.1 |
PAFAH1B3 |
platelet-activating factor acetylhydrolase 1b, catalytic subunit 3 (29kDa) |
| chr8_-_19540266 | 0.05 |
ENST00000311540.4 |
CSGALNACT1 |
chondroitin sulfate N-acetylgalactosaminyltransferase 1 |
| chr6_+_108977520 | 0.05 |
ENST00000540898.1 |
FOXO3 |
forkhead box O3 |
| chr1_+_197170592 | 0.05 |
ENST00000535699.1 |
CRB1 |
crumbs homolog 1 (Drosophila) |
| chr12_+_54402790 | 0.05 |
ENST00000040584.4 |
HOXC8 |
homeobox C8 |
| chr17_-_39471947 | 0.05 |
ENST00000334202.3 |
KRTAP17-1 |
keratin associated protein 17-1 |
| chr16_-_4524543 | 0.05 |
ENST00000573571.1 ENST00000404295.3 ENST00000574425.1 |
NMRAL1 |
NmrA-like family domain containing 1 |
| chr12_-_5352315 | 0.04 |
ENST00000536518.1 |
RP11-319E16.1 |
RP11-319E16.1 |
| chr18_+_616711 | 0.04 |
ENST00000579494.1 |
CLUL1 |
clusterin-like 1 (retinal) |
| chr18_+_616672 | 0.04 |
ENST00000338387.7 |
CLUL1 |
clusterin-like 1 (retinal) |
| chr5_+_52856456 | 0.04 |
ENST00000296684.5 ENST00000506765.1 |
NDUFS4 |
NADH dehydrogenase (ubiquinone) Fe-S protein 4, 18kDa (NADH-coenzyme Q reductase) |
| chr17_-_47045949 | 0.04 |
ENST00000357424.2 |
GIP |
gastric inhibitory polypeptide |
| chr18_-_40857493 | 0.04 |
ENST00000255224.3 |
SYT4 |
synaptotagmin IV |
| chr3_+_142342228 | 0.04 |
ENST00000337777.3 |
PLS1 |
plastin 1 |
| chr1_-_154842741 | 0.04 |
ENST00000271915.4 |
KCNN3 |
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 3 |
| chr6_+_31515337 | 0.04 |
ENST00000376148.4 ENST00000376145.4 |
NFKBIL1 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor-like 1 |
| chr18_+_46065393 | 0.04 |
ENST00000256413.3 |
CTIF |
CBP80/20-dependent translation initiation factor |
| chr18_-_52989217 | 0.04 |
ENST00000570287.2 |
TCF4 |
transcription factor 4 |
| chr3_+_158787041 | 0.04 |
ENST00000471575.1 ENST00000476809.1 ENST00000485419.1 |
IQCJ-SCHIP1 |
IQCJ-SCHIP1 readthrough |
| chr3_+_35721106 | 0.04 |
ENST00000474696.1 ENST00000412048.1 ENST00000396482.2 ENST00000432682.1 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chrY_-_6740649 | 0.04 |
ENST00000383036.1 ENST00000383037.4 |
AMELY |
amelogenin, Y-linked |
| chr12_-_53045948 | 0.04 |
ENST00000309680.3 |
KRT2 |
keratin 2 |
| chr3_+_57875738 | 0.04 |
ENST00000417128.1 ENST00000438794.1 |
SLMAP |
sarcolemma associated protein |
| chr6_-_32498046 | 0.04 |
ENST00000374975.3 |
HLA-DRB5 |
major histocompatibility complex, class II, DR beta 5 |
| chr8_-_42234745 | 0.04 |
ENST00000220812.2 |
DKK4 |
dickkopf WNT signaling pathway inhibitor 4 |
| chr3_+_179322481 | 0.04 |
ENST00000259037.3 |
NDUFB5 |
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 5, 16kDa |
| chr4_+_74269956 | 0.04 |
ENST00000295897.4 ENST00000415165.2 ENST00000503124.1 ENST00000509063.1 ENST00000401494.3 |
ALB |
albumin |
| chr20_-_48732472 | 0.03 |
ENST00000340309.3 ENST00000415862.2 ENST00000371677.3 ENST00000420027.2 |
UBE2V1 |
ubiquitin-conjugating enzyme E2 variant 1 |
| chr13_+_78315295 | 0.03 |
ENST00000351546.3 |
SLAIN1 |
SLAIN motif family, member 1 |
| chr9_+_131084815 | 0.03 |
ENST00000300452.3 ENST00000609948.1 |
COQ4 |
coenzyme Q4 |
| chr16_+_20775358 | 0.03 |
ENST00000440284.2 |
ACSM3 |
acyl-CoA synthetase medium-chain family member 3 |
| chr12_-_57037284 | 0.03 |
ENST00000551570.1 |
ATP5B |
ATP synthase, H+ transporting, mitochondrial F1 complex, beta polypeptide |
| chr3_-_146262637 | 0.03 |
ENST00000472349.1 ENST00000342435.4 |
PLSCR1 |
phospholipid scramblase 1 |
| chr4_-_168155169 | 0.03 |
ENST00000534949.1 ENST00000535728.1 |
SPOCK3 |
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3 |
| chr1_+_221054584 | 0.03 |
ENST00000549319.1 |
HLX |
H2.0-like homeobox |
| chr4_+_169418195 | 0.03 |
ENST00000261509.6 ENST00000335742.7 |
PALLD |
palladin, cytoskeletal associated protein |
| chr17_-_7760457 | 0.03 |
ENST00000576384.1 |
LSMD1 |
LSM domain containing 1 |
| chr18_-_21977748 | 0.03 |
ENST00000399441.4 ENST00000319481.3 |
OSBPL1A |
oxysterol binding protein-like 1A |
| chr10_+_123923205 | 0.03 |
ENST00000369004.3 ENST00000260733.3 |
TACC2 |
transforming, acidic coiled-coil containing protein 2 |
| chr6_+_168418553 | 0.03 |
ENST00000354419.2 ENST00000351261.3 |
KIF25 |
kinesin family member 25 |
| chr1_-_156217822 | 0.03 |
ENST00000368270.1 |
PAQR6 |
progestin and adipoQ receptor family member VI |
| chr3_-_47023455 | 0.03 |
ENST00000446836.1 ENST00000425441.1 |
CCDC12 |
coiled-coil domain containing 12 |
| chr7_+_99724317 | 0.03 |
ENST00000398075.2 ENST00000421390.1 |
MBLAC1 |
metallo-beta-lactamase domain containing 1 |
| chr18_+_55018044 | 0.03 |
ENST00000324000.3 |
ST8SIA3 |
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 3 |
| chr1_-_156217875 | 0.03 |
ENST00000292291.5 |
PAQR6 |
progestin and adipoQ receptor family member VI |
| chr7_+_90338712 | 0.03 |
ENST00000265741.3 ENST00000406263.1 |
CDK14 |
cyclin-dependent kinase 14 |
| chr15_-_55541227 | 0.03 |
ENST00000566877.1 |
RAB27A |
RAB27A, member RAS oncogene family |
| chr14_+_37126765 | 0.03 |
ENST00000402703.2 |
PAX9 |
paired box 9 |
| chr8_-_70016408 | 0.03 |
ENST00000518540.1 |
RP11-600K15.1 |
RP11-600K15.1 |
| chr12_+_21525818 | 0.03 |
ENST00000240652.3 ENST00000542023.1 ENST00000537593.1 |
IAPP |
islet amyloid polypeptide |
| chrX_-_23926004 | 0.03 |
ENST00000379226.4 ENST00000379220.3 |
APOO |
apolipoprotein O |
| chr1_+_192544857 | 0.03 |
ENST00000367459.3 ENST00000469578.2 |
RGS1 |
regulator of G-protein signaling 1 |
| chr7_-_14026063 | 0.02 |
ENST00000443608.1 ENST00000438956.1 |
ETV1 |
ets variant 1 |
| chr1_+_15986364 | 0.02 |
ENST00000345034.1 |
RSC1A1 |
regulatory solute carrier protein, family 1, member 1 |
| chr22_+_40390930 | 0.02 |
ENST00000333407.6 |
FAM83F |
family with sequence similarity 83, member F |
| chr21_-_34186006 | 0.02 |
ENST00000490358.1 |
C21orf62 |
chromosome 21 open reading frame 62 |
| chr3_-_100712292 | 0.02 |
ENST00000495063.1 ENST00000530539.1 |
ABI3BP |
ABI family, member 3 (NESH) binding protein |
| chr12_+_81110684 | 0.02 |
ENST00000228644.3 |
MYF5 |
myogenic factor 5 |
| chr7_+_20687017 | 0.02 |
ENST00000258738.6 |
ABCB5 |
ATP-binding cassette, sub-family B (MDR/TAP), member 5 |
| chrX_-_100129128 | 0.02 |
ENST00000372960.4 ENST00000372964.1 ENST00000217885.5 |
NOX1 |
NADPH oxidase 1 |
| chr18_-_53177984 | 0.02 |
ENST00000543082.1 |
TCF4 |
transcription factor 4 |
| chr6_-_100912785 | 0.02 |
ENST00000369208.3 |
SIM1 |
single-minded family bHLH transcription factor 1 |
| chr6_-_123958141 | 0.02 |
ENST00000334268.4 |
TRDN |
triadin |
| chr2_+_166152283 | 0.02 |
ENST00000375427.2 |
SCN2A |
sodium channel, voltage-gated, type II, alpha subunit |
| chr17_-_41132010 | 0.02 |
ENST00000409103.1 ENST00000360221.4 |
PTGES3L-AARSD1 |
PTGES3L-AARSD1 readthrough |
| chr4_-_103998439 | 0.02 |
ENST00000503230.1 ENST00000503818.1 |
SLC9B2 |
solute carrier family 9, subfamily B (NHA2, cation proton antiporter 2), member 2 |
| chr1_+_197382957 | 0.02 |
ENST00000367397.1 |
CRB1 |
crumbs homolog 1 (Drosophila) |
| chr8_-_116680833 | 0.02 |
ENST00000220888.5 |
TRPS1 |
trichorhinophalangeal syndrome I |
| chr6_+_90272027 | 0.02 |
ENST00000522441.1 |
ANKRD6 |
ankyrin repeat domain 6 |
| chr3_-_143567262 | 0.02 |
ENST00000474151.1 ENST00000316549.6 |
SLC9A9 |
solute carrier family 9, subfamily A (NHE9, cation proton antiporter 9), member 9 |
| chr16_+_20775024 | 0.02 |
ENST00000289416.5 |
ACSM3 |
acyl-CoA synthetase medium-chain family member 3 |
| chr17_+_1933404 | 0.02 |
ENST00000263083.6 ENST00000571418.1 |
DPH1 |
diphthamide biosynthesis 1 |
| chrX_-_100129320 | 0.02 |
ENST00000372966.3 |
NOX1 |
NADPH oxidase 1 |
| chr13_-_99667960 | 0.02 |
ENST00000448493.2 |
DOCK9 |
dedicator of cytokinesis 9 |
| chr14_+_93357749 | 0.02 |
ENST00000557689.1 |
RP11-862G15.1 |
RP11-862G15.1 |
| chr6_-_117150198 | 0.02 |
ENST00000310357.3 ENST00000368549.3 ENST00000530250.1 |
GPRC6A |
G protein-coupled receptor, family C, group 6, member A |
| chr15_+_76352178 | 0.02 |
ENST00000388942.3 |
C15orf27 |
chromosome 15 open reading frame 27 |
| chr16_+_3493611 | 0.02 |
ENST00000407558.4 ENST00000572169.1 ENST00000572757.1 ENST00000573593.1 ENST00000570372.1 ENST00000424546.2 ENST00000575733.1 ENST00000573201.1 ENST00000574950.1 ENST00000573580.1 ENST00000608722.1 |
NAA60 NAA60 |
N(alpha)-acetyltransferase 60, NatF catalytic subunit N-alpha-acetyltransferase 60 |
| chrX_-_77914825 | 0.02 |
ENST00000321110.1 |
ZCCHC5 |
zinc finger, CCHC domain containing 5 |
| chr6_-_128222103 | 0.02 |
ENST00000434358.1 ENST00000543064.1 ENST00000368248.2 |
THEMIS |
thymocyte selection associated |
| chr21_-_47575481 | 0.02 |
ENST00000291670.5 ENST00000397748.1 ENST00000359679.2 ENST00000355384.2 ENST00000397746.3 ENST00000397743.1 |
FTCD |
formimidoyltransferase cyclodeaminase |
| chr4_+_71494461 | 0.02 |
ENST00000396073.3 |
ENAM |
enamelin |
| chr15_+_48498480 | 0.01 |
ENST00000380993.3 ENST00000396577.3 |
SLC12A1 |
solute carrier family 12 (sodium/potassium/chloride transporter), member 1 |
| chr19_-_42759300 | 0.01 |
ENST00000222329.4 |
ERF |
Ets2 repressor factor |
| chr12_-_102591604 | 0.01 |
ENST00000329406.4 |
PMCH |
pro-melanin-concentrating hormone |
| chr10_-_36813162 | 0.01 |
ENST00000440465.1 |
NAMPTL |
nicotinamide phosphoribosyltransferase-like |
| chr11_-_66496430 | 0.01 |
ENST00000533211.1 |
SPTBN2 |
spectrin, beta, non-erythrocytic 2 |
| chr18_-_13915530 | 0.01 |
ENST00000327606.3 |
MC2R |
melanocortin 2 receptor (adrenocorticotropic hormone) |
| chr3_-_49066811 | 0.01 |
ENST00000442157.1 ENST00000326739.4 |
IMPDH2 |
IMP (inosine 5'-monophosphate) dehydrogenase 2 |
| chr18_+_46065483 | 0.01 |
ENST00000382998.4 |
CTIF |
CBP80/20-dependent translation initiation factor |
| chr9_+_37667978 | 0.01 |
ENST00000539465.1 |
FRMPD1 |
FERM and PDZ domain containing 1 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0036371 | protein localization to T-tubule(GO:0036371) |
| 0.0 | 0.1 | GO:0042418 | epinephrine biosynthetic process(GO:0042418) |
| 0.0 | 0.1 | GO:0009236 | cobalamin biosynthetic process(GO:0009236) |
| 0.0 | 0.1 | GO:0014835 | myoblast differentiation involved in skeletal muscle regeneration(GO:0014835) |
| 0.0 | 0.1 | GO:0050653 | chondroitin sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0050653) |
| 0.0 | 0.2 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 0.0 | 0.1 | GO:1902725 | negative regulation of satellite cell differentiation(GO:1902725) |
| 0.0 | 0.1 | GO:0071400 | cellular response to oleic acid(GO:0071400) |
| 0.0 | 0.1 | GO:1904562 | phosphatidylinositol 5-phosphate metabolic process(GO:1904562) |
| 0.0 | 0.3 | GO:0002158 | osteoclast proliferation(GO:0002158) |
| 0.0 | 0.1 | GO:0006436 | tryptophanyl-tRNA aminoacylation(GO:0006436) |
| 0.0 | 0.0 | GO:0001544 | initiation of primordial ovarian follicle growth(GO:0001544) |
| 0.0 | 0.0 | GO:0042488 | positive regulation of odontogenesis of dentin-containing tooth(GO:0042488) |
| 0.0 | 0.1 | GO:0010909 | regulation of heparan sulfate proteoglycan biosynthetic process(GO:0010908) positive regulation of heparan sulfate proteoglycan biosynthetic process(GO:0010909) canonical Wnt signaling pathway involved in positive regulation of epithelial to mesenchymal transition(GO:0044334) positive regulation of proteoglycan biosynthetic process(GO:1902730) |
| 0.0 | 0.5 | GO:0003301 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.0 | 0.1 | GO:0035694 | mitochondrial protein catabolic process(GO:0035694) |
| 0.0 | 0.2 | GO:0021694 | cerebellar Purkinje cell layer formation(GO:0021694) cerebellar Purkinje cell differentiation(GO:0021702) |
| 0.0 | 0.1 | GO:0030242 | pexophagy(GO:0030242) |
| 0.0 | 0.0 | GO:0052331 | hemolysis by symbiont of host erythrocytes(GO:0019836) hemolysis in other organism(GO:0044179) hemolysis in other organism involved in symbiotic interaction(GO:0052331) |
| 0.0 | 0.2 | GO:0021680 | cerebellar Purkinje cell layer development(GO:0021680) |
| 0.0 | 0.0 | GO:0045726 | positive regulation of integrin biosynthetic process(GO:0045726) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | GO:0071986 | Ragulator complex(GO:0071986) |
| 0.0 | 0.1 | GO:0001534 | radial spoke(GO:0001534) |
| 0.0 | 0.1 | GO:0070369 | beta-catenin-TCF7L2 complex(GO:0070369) |
| 0.0 | 0.0 | GO:0032127 | dense core granule membrane(GO:0032127) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | GO:0016941 | natriuretic peptide receptor activity(GO:0016941) |
| 0.0 | 0.1 | GO:0004603 | phenylethanolamine N-methyltransferase activity(GO:0004603) |
| 0.0 | 0.1 | GO:0008955 | peptidoglycan glycosyltransferase activity(GO:0008955) |
| 0.0 | 0.2 | GO:0004705 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.0 | 0.1 | GO:0004531 | deoxyribonuclease II activity(GO:0004531) |
| 0.0 | 0.1 | GO:0004087 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 0.0 | 0.1 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.0 | 0.1 | GO:0070506 | high-density lipoprotein particle receptor activity(GO:0070506) |
| 0.0 | 0.2 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.0 | 0.1 | GO:0008431 | vitamin E binding(GO:0008431) |
| 0.0 | 0.5 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.9 | GO:0051393 | alpha-actinin binding(GO:0051393) |
| 0.0 | 0.1 | GO:0004830 | tryptophan-tRNA ligase activity(GO:0004830) |
| 0.0 | 0.0 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.0 | 0.1 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.0 | 0.0 | GO:0030171 | voltage-gated proton channel activity(GO:0030171) |
| 0.0 | 0.1 | GO:0002046 | opsin binding(GO:0002046) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |