Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
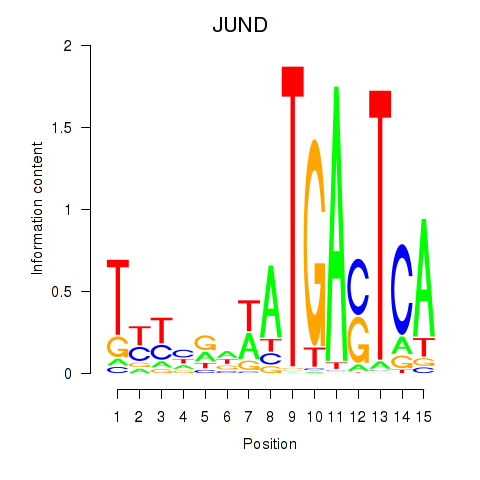
Results for JUND
Z-value: 0.40
Transcription factors associated with JUND
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
JUND
|
ENSG00000130522.4 | JUND |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| JUND | hg19_v2_chr19_-_18391708_18391739 | -0.32 | 4.4e-01 | Click! |
Activity profile of JUND motif
Sorted Z-values of JUND motif
Network of associatons between targets according to the STRING database.
First level regulatory network of JUND
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_+_147582348 | 0.32 |
ENST00000514389.1 |
SPINK6 |
serine peptidase inhibitor, Kazal type 6 |
| chr11_-_51412448 | 0.22 |
ENST00000319760.6 |
OR4A5 |
olfactory receptor, family 4, subfamily A, member 5 |
| chr5_-_60140009 | 0.19 |
ENST00000505959.1 |
ELOVL7 |
ELOVL fatty acid elongase 7 |
| chr5_-_60140089 | 0.19 |
ENST00000507047.1 ENST00000438340.1 ENST00000425382.1 ENST00000508821.1 |
ELOVL7 |
ELOVL fatty acid elongase 7 |
| chrX_+_99899180 | 0.19 |
ENST00000373004.3 |
SRPX2 |
sushi-repeat containing protein, X-linked 2 |
| chr5_-_140013275 | 0.16 |
ENST00000512545.1 ENST00000302014.6 ENST00000401743.2 |
CD14 |
CD14 molecule |
| chr2_+_102615416 | 0.16 |
ENST00000393414.2 |
IL1R2 |
interleukin 1 receptor, type II |
| chr1_+_160709076 | 0.15 |
ENST00000359331.4 ENST00000495334.1 |
SLAMF7 |
SLAM family member 7 |
| chr10_+_49892904 | 0.14 |
ENST00000360890.2 |
WDFY4 |
WDFY family member 4 |
| chr5_+_147582387 | 0.14 |
ENST00000325630.2 |
SPINK6 |
serine peptidase inhibitor, Kazal type 6 |
| chr5_+_140227048 | 0.14 |
ENST00000532602.1 |
PCDHA9 |
protocadherin alpha 9 |
| chr21_+_42733870 | 0.14 |
ENST00000330714.3 ENST00000436410.1 ENST00000435611.1 |
MX2 |
myxovirus (influenza virus) resistance 2 (mouse) |
| chr1_+_160709055 | 0.12 |
ENST00000368043.3 ENST00000368042.3 ENST00000458602.2 ENST00000458104.2 |
SLAMF7 |
SLAM family member 7 |
| chr4_-_123542224 | 0.11 |
ENST00000264497.3 |
IL21 |
interleukin 21 |
| chr8_-_135522425 | 0.11 |
ENST00000521673.1 |
ZFAT |
zinc finger and AT hook domain containing |
| chr18_-_19283649 | 0.11 |
ENST00000584464.1 ENST00000578270.1 |
ABHD3 |
abhydrolase domain containing 3 |
| chr9_-_127263265 | 0.10 |
ENST00000373587.3 |
NR5A1 |
nuclear receptor subfamily 5, group A, member 1 |
| chr17_+_28256874 | 0.10 |
ENST00000541045.1 ENST00000536908.2 |
EFCAB5 |
EF-hand calcium binding domain 5 |
| chr11_-_2162162 | 0.10 |
ENST00000381389.1 |
IGF2 |
insulin-like growth factor 2 (somatomedin A) |
| chr16_-_72206034 | 0.10 |
ENST00000537465.1 ENST00000237353.10 |
PMFBP1 |
polyamine modulated factor 1 binding protein 1 |
| chr11_-_2162468 | 0.09 |
ENST00000434045.2 |
IGF2 |
insulin-like growth factor 2 (somatomedin A) |
| chr1_-_6662919 | 0.09 |
ENST00000377658.4 ENST00000377663.3 |
KLHL21 |
kelch-like family member 21 |
| chr7_+_55177416 | 0.09 |
ENST00000450046.1 ENST00000454757.2 |
EGFR |
epidermal growth factor receptor |
| chr16_+_19467772 | 0.09 |
ENST00000219821.5 ENST00000561503.1 ENST00000564959.1 |
TMC5 |
transmembrane channel-like 5 |
| chr4_+_77356248 | 0.09 |
ENST00000296043.6 |
SHROOM3 |
shroom family member 3 |
| chr1_+_160709029 | 0.09 |
ENST00000444090.2 ENST00000441662.2 |
SLAMF7 |
SLAM family member 7 |
| chr9_-_39239171 | 0.08 |
ENST00000358144.2 |
CNTNAP3 |
contactin associated protein-like 3 |
| chr11_-_5207612 | 0.08 |
ENST00000380367.1 |
OR52A1 |
olfactory receptor, family 52, subfamily A, member 1 |
| chr14_+_22977587 | 0.08 |
ENST00000390504.1 |
TRAJ33 |
T cell receptor alpha joining 33 |
| chr1_-_223853348 | 0.08 |
ENST00000366872.5 |
CAPN8 |
calpain 8 |
| chr3_-_48601206 | 0.08 |
ENST00000273610.3 |
UCN2 |
urocortin 2 |
| chr3_-_155011483 | 0.07 |
ENST00000489090.1 |
RP11-451G4.2 |
RP11-451G4.2 |
| chr3_+_98699880 | 0.07 |
ENST00000473756.1 |
LINC00973 |
long intergenic non-protein coding RNA 973 |
| chr5_+_126988732 | 0.07 |
ENST00000395322.3 |
CTXN3 |
cortexin 3 |
| chr21_+_26934165 | 0.07 |
ENST00000456917.1 |
MIR155HG |
MIR155 host gene (non-protein coding) |
| chr1_-_153521714 | 0.07 |
ENST00000368713.3 |
S100A3 |
S100 calcium binding protein A3 |
| chr15_-_42565606 | 0.07 |
ENST00000307216.6 ENST00000448392.1 |
TMEM87A |
transmembrane protein 87A |
| chr18_+_55888767 | 0.06 |
ENST00000431212.2 ENST00000586268.1 ENST00000587190.1 |
NEDD4L |
neural precursor cell expressed, developmentally down-regulated 4-like, E3 ubiquitin protein ligase |
| chr1_-_153521597 | 0.06 |
ENST00000368712.1 |
S100A3 |
S100 calcium binding protein A3 |
| chr15_+_89181974 | 0.06 |
ENST00000306072.5 |
ISG20 |
interferon stimulated exonuclease gene 20kDa |
| chr6_+_44194762 | 0.06 |
ENST00000371708.1 |
SLC29A1 |
solute carrier family 29 (equilibrative nucleoside transporter), member 1 |
| chr5_+_145317356 | 0.06 |
ENST00000511217.1 |
SH3RF2 |
SH3 domain containing ring finger 2 |
| chr4_-_104119528 | 0.06 |
ENST00000380026.3 ENST00000503705.1 ENST00000265148.3 |
CENPE |
centromere protein E, 312kDa |
| chr12_-_95009837 | 0.06 |
ENST00000551457.1 |
TMCC3 |
transmembrane and coiled-coil domain family 3 |
| chr12_-_71182695 | 0.06 |
ENST00000342084.4 |
PTPRR |
protein tyrosine phosphatase, receptor type, R |
| chr10_+_120116527 | 0.06 |
ENST00000445161.1 |
LINC00867 |
long intergenic non-protein coding RNA 867 |
| chr3_-_98241760 | 0.06 |
ENST00000507874.1 ENST00000502299.1 ENST00000508659.1 ENST00000510545.1 ENST00000511667.1 ENST00000394185.2 ENST00000394181.2 ENST00000508902.1 ENST00000341181.6 ENST00000437922.1 ENST00000394180.2 |
CLDND1 |
claudin domain containing 1 |
| chrX_-_154563889 | 0.06 |
ENST00000369449.2 ENST00000321926.4 |
CLIC2 |
chloride intracellular channel 2 |
| chr8_+_120220561 | 0.05 |
ENST00000276681.6 |
MAL2 |
mal, T-cell differentiation protein 2 (gene/pseudogene) |
| chr7_+_79998864 | 0.05 |
ENST00000435819.1 |
CD36 |
CD36 molecule (thrombospondin receptor) |
| chr3_+_127771212 | 0.05 |
ENST00000243253.3 ENST00000481210.1 |
SEC61A1 |
Sec61 alpha 1 subunit (S. cerevisiae) |
| chr19_+_11546093 | 0.05 |
ENST00000591462.1 |
PRKCSH |
protein kinase C substrate 80K-H |
| chr3_+_127770455 | 0.05 |
ENST00000464451.1 |
SEC61A1 |
Sec61 alpha 1 subunit (S. cerevisiae) |
| chr10_-_14996017 | 0.05 |
ENST00000378241.1 ENST00000456122.1 ENST00000418843.1 ENST00000378249.1 ENST00000396817.2 ENST00000378255.1 ENST00000378254.1 ENST00000378278.2 ENST00000357717.2 |
DCLRE1C |
DNA cross-link repair 1C |
| chr1_-_152297679 | 0.05 |
ENST00000368799.1 |
FLG |
filaggrin |
| chr3_-_150996239 | 0.05 |
ENST00000309170.3 |
P2RY14 |
purinergic receptor P2Y, G-protein coupled, 14 |
| chr4_-_100356551 | 0.05 |
ENST00000209665.4 |
ADH7 |
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide |
| chr16_-_67281413 | 0.05 |
ENST00000258201.4 |
FHOD1 |
formin homology 2 domain containing 1 |
| chr10_-_74114714 | 0.05 |
ENST00000338820.3 ENST00000394903.2 ENST00000444643.2 |
DNAJB12 |
DnaJ (Hsp40) homolog, subfamily B, member 12 |
| chr14_+_94492674 | 0.05 |
ENST00000203664.5 ENST00000553723.1 |
OTUB2 |
OTU domain, ubiquitin aldehyde binding 2 |
| chr10_+_127661942 | 0.04 |
ENST00000417114.1 ENST00000445510.1 ENST00000368691.1 |
FANK1 |
fibronectin type III and ankyrin repeat domains 1 |
| chr10_-_128210005 | 0.04 |
ENST00000284694.7 ENST00000454341.1 ENST00000432642.1 ENST00000392694.1 |
C10orf90 |
chromosome 10 open reading frame 90 |
| chr7_+_100663353 | 0.04 |
ENST00000306151.4 |
MUC17 |
mucin 17, cell surface associated |
| chr12_-_95510743 | 0.04 |
ENST00000551521.1 |
FGD6 |
FYVE, RhoGEF and PH domain containing 6 |
| chr3_-_105588231 | 0.04 |
ENST00000545639.1 ENST00000394027.3 ENST00000438603.1 ENST00000447441.1 ENST00000443752.1 |
CBLB |
Cbl proto-oncogene B, E3 ubiquitin protein ligase |
| chr1_+_159796534 | 0.04 |
ENST00000289707.5 |
SLAMF8 |
SLAM family member 8 |
| chr21_-_43786634 | 0.04 |
ENST00000291527.2 |
TFF1 |
trefoil factor 1 |
| chr17_-_45899126 | 0.04 |
ENST00000007414.3 ENST00000392507.3 |
OSBPL7 |
oxysterol binding protein-like 7 |
| chrX_+_133371077 | 0.04 |
ENST00000517294.1 ENST00000370809.4 |
CCDC160 |
coiled-coil domain containing 160 |
| chr8_+_85097110 | 0.04 |
ENST00000517638.1 ENST00000522647.1 |
RALYL |
RALY RNA binding protein-like |
| chr5_-_138533973 | 0.04 |
ENST00000507002.1 ENST00000505830.1 ENST00000508639.1 ENST00000265195.5 |
SIL1 |
SIL1 nucleotide exchange factor |
| chr10_-_30024716 | 0.04 |
ENST00000375398.2 ENST00000375400.3 |
SVIL |
supervillin |
| chr12_-_8218997 | 0.04 |
ENST00000307637.4 |
C3AR1 |
complement component 3a receptor 1 |
| chr12_-_8765446 | 0.04 |
ENST00000537228.1 ENST00000229335.6 |
AICDA |
activation-induced cytidine deaminase |
| chr10_-_14996070 | 0.04 |
ENST00000378258.1 ENST00000453695.2 ENST00000378246.2 |
DCLRE1C |
DNA cross-link repair 1C |
| chr5_-_138534071 | 0.04 |
ENST00000394817.2 |
SIL1 |
SIL1 nucleotide exchange factor |
| chrX_-_153718953 | 0.04 |
ENST00000369649.4 ENST00000393586.1 |
SLC10A3 |
solute carrier family 10, member 3 |
| chr1_+_15256230 | 0.03 |
ENST00000376028.4 ENST00000400798.2 |
KAZN |
kazrin, periplakin interacting protein |
| chr11_-_102826434 | 0.03 |
ENST00000340273.4 ENST00000260302.3 |
MMP13 |
matrix metallopeptidase 13 (collagenase 3) |
| chr3_-_48632593 | 0.03 |
ENST00000454817.1 ENST00000328333.8 |
COL7A1 |
collagen, type VII, alpha 1 |
| chr4_-_100356291 | 0.03 |
ENST00000476959.1 ENST00000482593.1 |
ADH7 |
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide |
| chr3_+_184032283 | 0.03 |
ENST00000346169.2 ENST00000414031.1 |
EIF4G1 |
eukaryotic translation initiation factor 4 gamma, 1 |
| chr7_+_130126165 | 0.03 |
ENST00000427521.1 ENST00000416162.2 ENST00000378576.4 |
MEST |
mesoderm specific transcript |
| chr2_-_187713891 | 0.03 |
ENST00000295131.2 |
ZSWIM2 |
zinc finger, SWIM-type containing 2 |
| chr22_-_37545972 | 0.03 |
ENST00000216223.5 |
IL2RB |
interleukin 2 receptor, beta |
| chr17_-_25568687 | 0.03 |
ENST00000581944.1 |
RP11-663N22.1 |
RP11-663N22.1 |
| chr3_+_184032313 | 0.03 |
ENST00000392537.2 ENST00000444134.1 ENST00000450424.1 ENST00000421110.1 ENST00000382330.3 ENST00000426123.1 ENST00000350481.5 ENST00000455679.1 ENST00000440448.1 |
EIF4G1 |
eukaryotic translation initiation factor 4 gamma, 1 |
| chr12_-_7818474 | 0.03 |
ENST00000229304.4 |
APOBEC1 |
apolipoprotein B mRNA editing enzyme, catalytic polypeptide 1 |
| chr18_+_9885760 | 0.03 |
ENST00000536353.2 ENST00000584255.1 |
TXNDC2 |
thioredoxin domain containing 2 (spermatozoa) |
| chr12_+_62654119 | 0.03 |
ENST00000353364.3 ENST00000549523.1 ENST00000280377.5 |
USP15 |
ubiquitin specific peptidase 15 |
| chr6_-_44281043 | 0.03 |
ENST00000244571.4 |
AARS2 |
alanyl-tRNA synthetase 2, mitochondrial |
| chr16_-_80603558 | 0.03 |
ENST00000567317.1 |
RP11-18F14.1 |
RP11-18F14.1 |
| chr7_-_57207571 | 0.03 |
ENST00000331162.4 |
ZNF479 |
zinc finger protein 479 |
| chr16_-_75301886 | 0.03 |
ENST00000393422.2 |
BCAR1 |
breast cancer anti-estrogen resistance 1 |
| chr3_-_122512619 | 0.03 |
ENST00000383659.1 ENST00000306103.2 |
HSPBAP1 |
HSPB (heat shock 27kDa) associated protein 1 |
| chr1_+_117297007 | 0.03 |
ENST00000369478.3 ENST00000369477.1 |
CD2 |
CD2 molecule |
| chr18_+_9885961 | 0.03 |
ENST00000306084.6 |
TXNDC2 |
thioredoxin domain containing 2 (spermatozoa) |
| chr15_+_38226827 | 0.03 |
ENST00000559502.1 ENST00000558148.1 ENST00000558158.1 |
TMCO5A |
transmembrane and coiled-coil domains 5A |
| chr15_+_42566384 | 0.03 |
ENST00000440615.2 ENST00000318010.8 |
GANC |
glucosidase, alpha; neutral C |
| chr1_+_213224572 | 0.03 |
ENST00000543470.1 ENST00000366960.3 ENST00000366959.3 ENST00000543354.1 |
RPS6KC1 |
ribosomal protein S6 kinase, 52kDa, polypeptide 1 |
| chr15_+_59903975 | 0.03 |
ENST00000560585.1 ENST00000396065.1 |
GCNT3 |
glucosaminyl (N-acetyl) transferase 3, mucin type |
| chr20_+_48429233 | 0.02 |
ENST00000417961.1 |
SLC9A8 |
solute carrier family 9, subfamily A (NHE8, cation proton antiporter 8), member 8 |
| chr15_+_89182178 | 0.02 |
ENST00000559876.1 |
ISG20 |
interferon stimulated exonuclease gene 20kDa |
| chr7_-_33080506 | 0.02 |
ENST00000381626.2 ENST00000409467.1 ENST00000449201.1 |
NT5C3A |
5'-nucleotidase, cytosolic IIIA |
| chr8_-_19102999 | 0.02 |
ENST00000517949.1 |
RP11-1080G15.1 |
RP11-1080G15.1 |
| chr8_-_82359662 | 0.02 |
ENST00000519260.1 ENST00000256103.2 |
PMP2 |
peripheral myelin protein 2 |
| chr1_-_117753540 | 0.02 |
ENST00000328189.3 ENST00000369458.3 |
VTCN1 |
V-set domain containing T cell activation inhibitor 1 |
| chr16_+_28996114 | 0.02 |
ENST00000395461.3 |
LAT |
linker for activation of T cells |
| chr2_+_169926047 | 0.02 |
ENST00000428522.1 ENST00000450153.1 ENST00000421653.1 |
DHRS9 |
dehydrogenase/reductase (SDR family) member 9 |
| chr3_+_156807663 | 0.02 |
ENST00000467995.1 ENST00000474477.1 ENST00000471719.1 |
LINC00881 |
long intergenic non-protein coding RNA 881 |
| chr10_+_5005598 | 0.02 |
ENST00000442997.1 |
AKR1C1 |
aldo-keto reductase family 1, member C1 |
| chr3_-_157251383 | 0.02 |
ENST00000487753.1 ENST00000489602.1 ENST00000461299.1 ENST00000479987.1 |
VEPH1 |
ventricular zone expressed PH domain-containing 1 |
| chr2_-_145275211 | 0.02 |
ENST00000462355.1 |
ZEB2 |
zinc finger E-box binding homeobox 2 |
| chr17_-_28257080 | 0.02 |
ENST00000579954.1 ENST00000540801.1 ENST00000269033.3 ENST00000590153.1 ENST00000582084.1 |
SSH2 |
slingshot protein phosphatase 2 |
| chr20_+_48429356 | 0.02 |
ENST00000361573.2 ENST00000541138.1 ENST00000539601.1 |
SLC9A8 |
solute carrier family 9, subfamily A (NHE8, cation proton antiporter 8), member 8 |
| chr16_+_31366536 | 0.02 |
ENST00000562522.1 |
ITGAX |
integrin, alpha X (complement component 3 receptor 4 subunit) |
| chr1_+_223889310 | 0.02 |
ENST00000434648.1 |
CAPN2 |
calpain 2, (m/II) large subunit |
| chr14_+_73525229 | 0.02 |
ENST00000527432.1 ENST00000531500.1 ENST00000525321.1 ENST00000526754.1 |
RBM25 |
RNA binding motif protein 25 |
| chr3_-_11762202 | 0.02 |
ENST00000445411.1 ENST00000404339.1 ENST00000273038.3 |
VGLL4 |
vestigial like 4 (Drosophila) |
| chr10_-_50396407 | 0.02 |
ENST00000374153.2 ENST00000374151.3 |
C10orf128 |
chromosome 10 open reading frame 128 |
| chr3_-_37216055 | 0.02 |
ENST00000336686.4 |
LRRFIP2 |
leucine rich repeat (in FLII) interacting protein 2 |
| chr1_+_85527987 | 0.02 |
ENST00000326813.8 ENST00000294664.6 ENST00000528899.1 |
WDR63 |
WD repeat domain 63 |
| chr6_+_150070857 | 0.02 |
ENST00000544496.1 |
PCMT1 |
protein-L-isoaspartate (D-aspartate) O-methyltransferase |
| chr16_-_20681177 | 0.02 |
ENST00000524149.1 |
ACSM1 |
acyl-CoA synthetase medium-chain family member 1 |
| chr12_+_9142131 | 0.02 |
ENST00000356986.3 ENST00000266551.4 |
KLRG1 |
killer cell lectin-like receptor subfamily G, member 1 |
| chr9_-_95432536 | 0.02 |
ENST00000287996.3 |
IPPK |
inositol 1,3,4,5,6-pentakisphosphate 2-kinase |
| chr1_+_196912902 | 0.02 |
ENST00000476712.2 ENST00000367415.5 |
CFHR2 |
complement factor H-related 2 |
| chr16_+_31366455 | 0.01 |
ENST00000268296.4 |
ITGAX |
integrin, alpha X (complement component 3 receptor 4 subunit) |
| chr2_+_201980827 | 0.01 |
ENST00000309955.3 ENST00000443227.1 ENST00000341222.6 ENST00000355558.4 ENST00000340870.5 ENST00000341582.6 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr10_-_14996321 | 0.01 |
ENST00000378289.4 |
DCLRE1C |
DNA cross-link repair 1C |
| chr5_+_145583107 | 0.01 |
ENST00000506502.1 |
RBM27 |
RNA binding motif protein 27 |
| chr2_+_36923830 | 0.01 |
ENST00000379242.3 ENST00000389975.3 |
VIT |
vitrin |
| chr4_+_95128996 | 0.01 |
ENST00000457823.2 |
SMARCAD1 |
SWI/SNF-related, matrix-associated actin-dependent regulator of chromatin, subfamily a, containing DEAD/H box 1 |
| chr17_+_30771279 | 0.01 |
ENST00000261712.3 ENST00000578213.1 ENST00000457654.2 ENST00000579451.1 |
PSMD11 |
proteasome (prosome, macropain) 26S subunit, non-ATPase, 11 |
| chr2_-_152830479 | 0.01 |
ENST00000360283.6 |
CACNB4 |
calcium channel, voltage-dependent, beta 4 subunit |
| chr15_+_89182156 | 0.01 |
ENST00000379224.5 |
ISG20 |
interferon stimulated exonuclease gene 20kDa |
| chr12_+_62654155 | 0.01 |
ENST00000312635.6 ENST00000393654.3 ENST00000549237.1 |
USP15 |
ubiquitin specific peptidase 15 |
| chr12_-_53343602 | 0.01 |
ENST00000546897.1 ENST00000552551.1 |
KRT8 |
keratin 8 |
| chr6_-_32145861 | 0.01 |
ENST00000336984.6 |
AGPAT1 |
1-acylglycerol-3-phosphate O-acyltransferase 1 |
| chr14_+_73525265 | 0.01 |
ENST00000525161.1 |
RBM25 |
RNA binding motif protein 25 |
| chr18_+_32290218 | 0.01 |
ENST00000348997.5 ENST00000588949.1 ENST00000597599.1 |
DTNA |
dystrobrevin, alpha |
| chr1_+_236687881 | 0.01 |
ENST00000526634.1 |
LGALS8 |
lectin, galactoside-binding, soluble, 8 |
| chr4_+_159727272 | 0.01 |
ENST00000379346.3 |
FNIP2 |
folliculin interacting protein 2 |
| chr12_+_52695617 | 0.01 |
ENST00000293525.5 |
KRT86 |
keratin 86 |
| chr15_-_74504597 | 0.01 |
ENST00000416286.3 |
STRA6 |
stimulated by retinoic acid 6 |
| chr1_+_207262881 | 0.01 |
ENST00000451804.2 |
C4BPB |
complement component 4 binding protein, beta |
| chr3_+_11178779 | 0.01 |
ENST00000438284.2 |
HRH1 |
histamine receptor H1 |
| chr16_+_23847339 | 0.01 |
ENST00000303531.7 |
PRKCB |
protein kinase C, beta |
| chr3_-_98241713 | 0.01 |
ENST00000502288.1 ENST00000512147.1 ENST00000510541.1 ENST00000503621.1 ENST00000511081.1 |
CLDND1 |
claudin domain containing 1 |
| chrX_+_51942963 | 0.01 |
ENST00000375625.3 |
RP11-363G10.2 |
RP11-363G10.2 |
| chr4_-_123377880 | 0.01 |
ENST00000226730.4 |
IL2 |
interleukin 2 |
| chr19_-_46285736 | 0.00 |
ENST00000291270.4 ENST00000447742.2 ENST00000354227.5 |
DMPK |
dystrophia myotonica-protein kinase |
| chr10_-_81708854 | 0.00 |
ENST00000372292.3 |
SFTPD |
surfactant protein D |
| chr8_-_17752912 | 0.00 |
ENST00000398054.1 ENST00000381840.2 |
FGL1 |
fibrinogen-like 1 |
| chr14_+_58765103 | 0.00 |
ENST00000355431.3 ENST00000348476.3 ENST00000395168.3 |
ARID4A |
AT rich interactive domain 4A (RBP1-like) |
| chr15_-_74504560 | 0.00 |
ENST00000449139.2 |
STRA6 |
stimulated by retinoic acid 6 |
| chr8_-_17752996 | 0.00 |
ENST00000381841.2 ENST00000427924.1 |
FGL1 |
fibrinogen-like 1 |
| chr10_+_17686124 | 0.00 |
ENST00000377524.3 |
STAM |
signal transducing adaptor molecule (SH3 domain and ITAM motif) 1 |
| chr1_-_205744574 | 0.00 |
ENST00000367139.3 ENST00000235932.4 ENST00000437324.2 ENST00000414729.1 |
RAB7L1 |
RAB7, member RAS oncogene family-like 1 |
| chr22_+_22749343 | 0.00 |
ENST00000390298.2 |
IGLV7-43 |
immunoglobulin lambda variable 7-43 |
| chr15_+_67420441 | 0.00 |
ENST00000558894.1 |
SMAD3 |
SMAD family member 3 |
| chrX_-_51797351 | 0.00 |
ENST00000375644.3 |
RP11-114H20.1 |
RP11-114H20.1 |
| chr3_-_170587974 | 0.00 |
ENST00000463836.1 |
RPL22L1 |
ribosomal protein L22-like 1 |
| chr19_-_11545920 | 0.00 |
ENST00000356392.4 ENST00000591179.1 |
CCDC151 |
coiled-coil domain containing 151 |
| chr3_+_30647994 | 0.00 |
ENST00000295754.5 |
TGFBR2 |
transforming growth factor, beta receptor II (70/80kDa) |
| chr3_-_46608010 | 0.00 |
ENST00000395905.3 |
LRRC2 |
leucine rich repeat containing 2 |
| chr19_-_46285646 | 0.00 |
ENST00000458663.2 |
DMPK |
dystrophia myotonica-protein kinase |
| chr17_+_76311791 | 0.00 |
ENST00000586321.1 |
AC061992.2 |
AC061992.2 |
| chr13_+_98605902 | 0.00 |
ENST00000460070.1 ENST00000481455.1 ENST00000261574.5 ENST00000493281.1 ENST00000463157.1 ENST00000471898.1 ENST00000489058.1 ENST00000481689.1 |
IPO5 |
importin 5 |
| chr18_-_5419797 | 0.00 |
ENST00000542146.1 ENST00000427684.2 |
EPB41L3 |
erythrocyte membrane protein band 4.1-like 3 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | GO:0019368 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.0 | 0.2 | GO:0038123 | toll-like receptor TLR1:TLR2 signaling pathway(GO:0038123) response to triacyl bacterial lipopeptide(GO:0071725) cellular response to triacyl bacterial lipopeptide(GO:0071727) |
| 0.0 | 0.2 | GO:0050712 | negative regulation of interleukin-1 alpha production(GO:0032690) negative regulation of interleukin-1 alpha secretion(GO:0050712) |
| 0.0 | 0.1 | GO:0039019 | pronephric nephron development(GO:0039019) |
| 0.0 | 0.1 | GO:0010430 | fatty acid omega-oxidation(GO:0010430) |
| 0.0 | 0.1 | GO:0007538 | primary sex determination(GO:0007538) |
| 0.0 | 0.1 | GO:0043006 | activation of phospholipase A2 activity by calcium-mediated signaling(GO:0043006) |
| 0.0 | 0.1 | GO:0090310 | negative regulation of methylation-dependent chromatin silencing(GO:0090310) |
| 0.0 | 0.2 | GO:0038028 | insulin receptor signaling pathway via phosphatidylinositol 3-kinase(GO:0038028) |
| 0.0 | 0.1 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.0 | 0.2 | GO:0090050 | positive regulation of cell migration involved in sprouting angiogenesis(GO:0090050) |
| 0.0 | 0.1 | GO:2001288 | positive regulation of caveolin-mediated endocytosis(GO:2001288) |
| 0.0 | 0.1 | GO:0003373 | dynamin polymerization involved in membrane fission(GO:0003373) dynamin polymerization involved in mitochondrial fission(GO:0003374) |
| 0.0 | 0.1 | GO:0015862 | uridine transport(GO:0015862) |
| 0.0 | 0.0 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 0.0 | 0.1 | GO:0097489 | multivesicular body, internal vesicle lumen(GO:0097489) |
| 0.0 | 0.1 | GO:0005827 | polar microtubule(GO:0005827) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | GO:0009922 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.0 | 0.2 | GO:0016019 | peptidoglycan receptor activity(GO:0016019) |
| 0.0 | 0.1 | GO:0008859 | exoribonuclease II activity(GO:0008859) |
| 0.0 | 0.2 | GO:0004910 | interleukin-1, Type II, blocking receptor activity(GO:0004910) |
| 0.0 | 0.1 | GO:0052740 | 1-acyl-2-lysophosphatidylserine acylhydrolase activity(GO:0052740) |
| 0.0 | 0.1 | GO:0004031 | aldehyde oxidase activity(GO:0004031) |
| 0.0 | 0.1 | GO:0051431 | corticotropin-releasing hormone receptor 2 binding(GO:0051431) |
| 0.0 | 0.1 | GO:0005006 | epidermal growth factor-activated receptor activity(GO:0005006) |
| 0.0 | 0.1 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 0.0 | 0.0 | GO:0019976 | interleukin-2 receptor activity(GO:0004911) interleukin-2 binding(GO:0019976) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 0.2 | REACTOME IKK COMPLEX RECRUITMENT MEDIATED BY RIP1 | Genes involved in IKK complex recruitment mediated by RIP1 |